

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
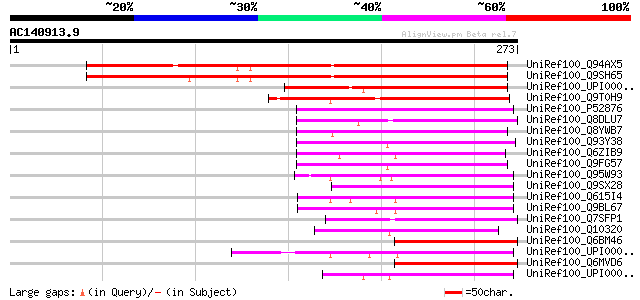
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94AX5 At1g64150/F22C12_10 [Arabidopsis thaliana] 246 6e-64
UniRef100_Q9SH65 F22C12.9 [Arabidopsis thaliana] 242 7e-63
UniRef100_UPI00002E160A UPI00002E160A UniRef100 entry 98 3e-19
UniRef100_Q9T0H9 Hypothetical protein T6G15.140 [Arabidopsis tha... 94 5e-18
UniRef100_P52876 Hypothetical UPF0016 protein sll0615 [Synechocy... 78 3e-13
UniRef100_Q8DLU7 Tlr0379 protein [Synechococcus elongatus] 77 4e-13
UniRef100_Q8YWB7 Alr1698 protein [Anabaena sp.] 76 8e-13
UniRef100_Q93Y38 Transmembrane protein FT27/PFT27-like [Arabidop... 72 2e-11
UniRef100_Q6ZIB9 Putative transmembrane protein [Oryza sativa] 72 2e-11
UniRef100_Q9FG57 Transmembrane protein FT27/PFT27-like [Arabidop... 71 3e-11
UniRef100_Q95W93 Putative membrane protein [Crithidia fasciculata] 65 2e-09
UniRef100_Q9SX28 F24J5.11 protein [Arabidopsis thaliana] 63 9e-09
UniRef100_Q615I4 Hypothetical protein CBG15669 [Caenorhabditis b... 62 1e-08
UniRef100_Q9BL67 Hypothetical protein Y54F10AL.1 [Caenorhabditis... 62 2e-08
UniRef100_Q7SFP1 Hypothetical protein [Neurospora crassa] 62 2e-08
UniRef100_Q10320 Hypothetical UPF0016 protein C17G8.08c in chrom... 62 2e-08
UniRef100_Q6BM46 Similar to ca|CA2327|IPF4782 Candida albicans I... 61 4e-08
UniRef100_UPI00003AD943 UPI00003AD943 UniRef100 entry 60 5e-08
UniRef100_Q6MVD6 Hypothetical protein B13D15.080 [Neurospora cra... 60 5e-08
UniRef100_UPI0000365A69 UPI0000365A69 UniRef100 entry 60 6e-08
>UniRef100_Q94AX5 At1g64150/F22C12_10 [Arabidopsis thaliana]
Length = 370
Score = 246 bits (627), Expect = 6e-64
Identities = 148/237 (62%), Positives = 168/237 (70%), Gaps = 13/237 (5%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRHFC*YFSLN 100
VL F+ S AL PA AAS S + FGD+GD+S+GFAS +FS
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFAS--AFLLIFFSEL 171
Query: 101 WETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHYVDELLPFRFGETDLPI 151
+ F +F +LG + L RTFHYVDE+LPFRFG TDLPI
Sbjct: 172 GDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHYVDEVLPFRFGGTDLPI 231
Query: 152 DDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTF 211
DDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG+GAGI+AAA+TI+STF
Sbjct: 232 DDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSGNGAGIVAAANTIISTF 290
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AVLGGSLLG FLSEK
Sbjct: 291 ALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAVLGGSLLGNFLSEK 347
>UniRef100_Q9SH65 F22C12.9 [Arabidopsis thaliana]
Length = 388
Score = 242 bits (618), Expect = 7e-63
Identities = 150/253 (59%), Positives = 170/253 (66%), Gaps = 27/253 (10%)
Query: 42 VLKFMLFSAFFALQDAFPAVAASDFATGLNS-IPIFGDVGDLSTGFASYHRH-FC*---- 95
VL F+ S AL PA AAS S + FGD+GD+S+GFAS FC
Sbjct: 114 VLMFLAVSGSVALLGTDPAFAASSIPNVTQSLVTSFGDLGDISSGFASVRESSFCTEPGT 173
Query: 96 -----------YFSLNWETRLFSLQHC*QLEIQPVLF-----SLGHLAH----LRRTFHY 135
+FS + F +F +LG + L RTFHY
Sbjct: 174 ISSSIPAFLLIFFSELGDKTFFIAALLAARNSAATVFVGTFGALGIMTIISVVLGRTFHY 233
Query: 136 VDELLPFRFGETDLPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSG 195
VDE+LPFRFG TDLPIDDIAAVCLLVYFGVSTLLDA S D K+D+EQKEAELAVS+ SG
Sbjct: 234 VDEVLPFRFGGTDLPIDDIAAVCLLVYFGVSTLLDAVS-DEGKADEEQKEAELAVSELSG 292
Query: 196 DGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
+GAGI+AAA+TI+STF LVFVAEWGDKSFFSTIALAAASSPLGVIAG+LAGHG ATL+AV
Sbjct: 293 NGAGIVAAANTIISTFALVFVAEWGDKSFFSTIALAAASSPLGVIAGALAGHGAATLLAV 352
Query: 256 LGGSLLGTFLSEK 268
LGGSLLG FLSEK
Sbjct: 353 LGGSLLGNFLSEK 365
>UniRef100_UPI00002E160A UPI00002E160A UniRef100 entry
Length = 157
Score = 97.8 bits (242), Expect = 3e-19
Identities = 56/124 (45%), Positives = 77/124 (61%), Gaps = 5/124 (4%)
Query: 149 LPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELA----VSDFSGDGAGILAAA 204
LPI + AAV +L++FGV TL DA + D + E+ ELA V S
Sbjct: 14 LPIGEYAAVAMLMFFGVKTLKDALAMDPSGATPEE-HGELAEATEVVCKSTSAQKRSPGV 72
Query: 205 STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTF 264
+ ++ TF L+F+AEWGD+S +TIAL AA +P+GV G+ GH +ATLIAV+GGSLL
Sbjct: 73 AALIETFTLIFIAEWGDRSMLATIALGAAQNPVGVAFGATLGHFIATLIAVVGGSLLSKR 132
Query: 265 LSEK 268
+SE+
Sbjct: 133 ISER 136
>UniRef100_Q9T0H9 Hypothetical protein T6G15.140 [Arabidopsis thaliana]
Length = 273
Score = 93.6 bits (231), Expect = 5e-18
Identities = 53/139 (38%), Positives = 85/139 (61%), Gaps = 12/139 (8%)
Query: 140 LPFRFGETDLPIDDIAAVCLLVYFGVSTLLDA---------SSSDSQKSDDEQKEAELAV 190
+P +F +T LPI + AA+ LL++FG+ ++ DA + ++ E EAE V
Sbjct: 137 VPAQF-QTTLPIGEYAAIALLMFFGLKSIKDAWDLPPVEAKNGEETGIELGEYSEAEELV 195
Query: 191 SDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVA 250
+ + + + +F LVF AEWGD+S +T+AL AA SPLGV +G++AGH VA
Sbjct: 196 KEKASKK--LTNPLEILWKSFSLVFFAEWGDRSMLATVALGAAQSPLGVASGAIAGHLVA 253
Query: 251 TLIAVLGGSLLGTFLSEKV 269
T++A++GG+ L ++SEK+
Sbjct: 254 TVLAIMGGAFLANYISEKL 272
Score = 39.3 bits (90), Expect = 0.11
Identities = 23/68 (33%), Positives = 37/68 (53%), Gaps = 2/68 (2%)
Query: 195 GDGAGILAAA--STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
G + +LAA S + F L+FV+E GDK+FF LA V+ GS+ + T+
Sbjct: 66 GGPSSVLAAVAKSGFTAAFSLIFVSEIGDKTFFIAALLAMQYEKTLVLLGSMGALSLMTI 125
Query: 253 IAVLGGSL 260
++V+ G +
Sbjct: 126 LSVVIGKI 133
>UniRef100_P52876 Hypothetical UPF0016 protein sll0615 [Synechocystis sp.]
Length = 206
Score = 77.8 bits (190), Expect = 3e-13
Identities = 42/117 (35%), Positives = 67/117 (56%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLV 214
A V L + FG L DA + + +E ++AE A++ + +V +F L
Sbjct: 70 AEVALFLIFGTKLLWDARRIKATANLEEMEDAEKAIASGEKKLKIVPRGWGIVVESFALT 129
Query: 215 FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
FVAEWGD++ +TIALAA+++ GV AG++ GH + +IAV+GG + +SEK +
Sbjct: 130 FVAEWGDRTQIATIALAASNNAWGVSAGAILGHTICAVIAVMGGKFVAGRISEKTVT 186
Score = 35.8 bits (81), Expect = 1.2
Identities = 21/54 (38%), Positives = 31/54 (56%), Gaps = 1/54 (1%)
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFL 265
LL+ V+E GDK+FF + LA V+ G + G T+++VL G + TFL
Sbjct: 10 LLITVSELGDKTFFIAMILAMRYPRRWVLVGVVGGLAAMTILSVLMGQIF-TFL 62
>UniRef100_Q8DLU7 Tlr0379 protein [Synechococcus elongatus]
Length = 211
Score = 77.4 bits (189), Expect = 4e-13
Identities = 50/124 (40%), Positives = 71/124 (56%), Gaps = 7/124 (5%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEA----ELAVSDFSGDGAGILAAASTIVST 210
AA+ L FG+ L+ ++ +DE + A E A ++ S G+ AA V
Sbjct: 70 AAILLFTIFGLRMLIQGWRMGNKPCEDECEAAVETVEKAEANLSRWGSNPAWAA--FVEA 127
Query: 211 FLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK-V 269
F L +AEWGD++ +TI LAAAS GV G++AGHG+ T IAVLGG L+ +SE+ +
Sbjct: 128 FSLTLMAEWGDRTQIATITLAAASQAFGVALGAIAGHGICTAIAVLGGGLIAGRISERTL 187
Query: 270 TSSG 273
T SG
Sbjct: 188 TLSG 191
>UniRef100_Q8YWB7 Alr1698 protein [Anabaena sp.]
Length = 209
Score = 76.3 bits (186), Expect = 8e-13
Identities = 42/118 (35%), Positives = 68/118 (57%), Gaps = 4/118 (3%)
Query: 155 AAVCLLVYFGVSTLLDAS----SSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVST 210
A + L + FG+ L DAS ++D + +E +EA+ AV S I+
Sbjct: 70 AEITLFIAFGLKLLYDASKMSAAADKAEVMEEMEEAKAAVEKADLQLPKQKTPLSIILEA 129
Query: 211 FLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
F+L F+AEWGD++ +TIALAA ++ +GV G++ GH + IAV+GG ++ +SE+
Sbjct: 130 FVLTFMAEWGDRTQIATIALAAGNNIIGVTIGAILGHAICAAIAVIGGKMIAGKISER 187
>UniRef100_Q93Y38 Transmembrane protein FT27/PFT27-like [Arabidopsis thaliana]
Length = 293
Score = 72.0 bits (175), Expect = 2e-11
Identities = 42/124 (33%), Positives = 67/124 (53%), Gaps = 6/124 (4%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS+ +++ E+ SG G +
Sbjct: 151 AATVLYAFFGLRLLYIAWRSTDSKSNQKKEMEEVEEKLESGQGKTPFRRLFSRFCTPIFL 210
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH V T +AV+GGS+L + +S++
Sbjct: 211 ESFILTFLAEWGDRSQIATIALATHKNAIGVAIGASIGHTVCTSLAVVGGSMLASRISQR 270
Query: 269 VTSS 272
++
Sbjct: 271 TVAT 274
Score = 34.3 bits (77), Expect = 3.6
Identities = 18/66 (27%), Positives = 36/66 (54%)
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
+ S+F ++ V E GD++F +A V++G+L+ V T+++ G ++ +S
Sbjct: 85 LFSSFSMILVTEIGDETFIIAALMAMRHPKATVLSGALSALFVMTILSTGLGRIVPNLIS 144
Query: 267 EKVTSS 272
K T+S
Sbjct: 145 RKHTNS 150
>UniRef100_Q6ZIB9 Putative transmembrane protein [Oryza sativa]
Length = 282
Score = 71.6 bits (174), Expect = 2e-11
Identities = 44/118 (37%), Positives = 65/118 (54%), Gaps = 5/118 (4%)
Query: 155 AAVCLLVYFGVSTLLDASSSDS---QKSDDEQKEAELAVSDFSGDGAGILAAAST--IVS 209
AA L +FG+ L A SDS QK + E+ E +L I + T +
Sbjct: 141 AATVLYAFFGLRLLYIAWRSDSKASQKKEIEEVEEKLEAGQGKSTFRRIFSRFCTPIFLE 200
Query: 210 TFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+F+L F+AEWGD+S +TIALA + +GV G+ GH + T AV+GGS+L + +S+
Sbjct: 201 SFVLTFLAEWGDRSQIATIALATHKNAVGVAVGATLGHTICTSFAVVGGSMLASKISQ 258
>UniRef100_Q9FG57 Transmembrane protein FT27/PFT27-like [Arabidopsis thaliana]
Length = 325
Score = 71.2 bits (173), Expect = 3e-11
Identities = 42/120 (35%), Positives = 65/120 (54%), Gaps = 6/120 (5%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILA------AASTIV 208
AA L +FG+ L A S KS+ +++ E+ SG G +
Sbjct: 151 AATVLYAFFGLRLLYIAWRSTDSKSNQKKEMEEVEEKLESGQGKTPFRRLFSRFCTPIFL 210
Query: 209 STFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+F+L F+AEWGD+S +TIALA + +GV G+ GH V T +AV+GGS+L + +S++
Sbjct: 211 ESFILTFLAEWGDRSQIATIALATHKNAIGVAIGASIGHTVCTSLAVVGGSMLASRISQR 270
Score = 34.3 bits (77), Expect = 3.6
Identities = 18/66 (27%), Positives = 36/66 (54%)
Query: 207 IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
+ S+F ++ V E GD++F +A V++G+L+ V T+++ G ++ +S
Sbjct: 85 LFSSFSMILVTEIGDETFIIAALMAMRHPKATVLSGALSALFVMTILSTGLGRIVPNLIS 144
Query: 267 EKVTSS 272
K T+S
Sbjct: 145 RKHTNS 150
>UniRef100_Q95W93 Putative membrane protein [Crithidia fasciculata]
Length = 259
Score = 65.1 bits (157), Expect = 2e-09
Identities = 49/139 (35%), Positives = 71/139 (50%), Gaps = 22/139 (15%)
Query: 154 IAAVCLLVYFGVSTLLDA---SSSDSQKSDDEQKEAELAVSDFSGDGA---GILAAA--- 204
+AAV LV FG L D + ++ ++S+DE EA A+ + A G +A++
Sbjct: 79 LAAVLFLV-FGGKILFDELVRNKAEDEESEDEMAEAAAALRRRDPNDAVETGSVASSVYT 137
Query: 205 ------------STIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
+V F L FVAEWGD+S +TIALAAA +P V G + GH + T
Sbjct: 138 SAPARRWRRLLNPVMVEAFTLTFVAEWGDRSQLATIALAAAKNPYAVTVGGVLGHALCTG 197
Query: 253 IAVLGGSLLGTFLSEKVTS 271
AVL G+L+ +S K +
Sbjct: 198 GAVLCGNLIAQRVSMKTVN 216
Score = 34.7 bits (78), Expect = 2.7
Identities = 18/63 (28%), Positives = 35/63 (54%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+S+ ++ V+E GDK+FF +A L V G+++ T+++ L G ++ + LS
Sbjct: 14 LSSLSMILVSEIGDKTFFIACLMAMRHPRLTVYLGAISALAAMTVLSALMGVIVPSLLSV 73
Query: 268 KVT 270
+T
Sbjct: 74 YLT 76
>UniRef100_Q9SX28 F24J5.11 protein [Arabidopsis thaliana]
Length = 228
Score = 62.8 bits (151), Expect = 9e-09
Identities = 35/99 (35%), Positives = 55/99 (55%), Gaps = 1/99 (1%)
Query: 174 SDSQKSDDEQKEAELAVSDFSGDGAGILAAASTI-VSTFLLVFVAEWGDKSFFSTIALAA 232
SD +K++D+ K +++ + A S I + F + F EWGDKS +TI LAA
Sbjct: 109 SDLKKTNDQSKNSKIEDEQKKQKRPFLTAFFSPIFLKAFSINFFGEWGDKSQLATIGLAA 168
Query: 233 ASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
+PLGV+ G + + T AVLGG L + +SE++ +
Sbjct: 169 DENPLGVVLGGIVAQTLCTTAAVLGGKSLASQISERIVA 207
>UniRef100_Q615I4 Hypothetical protein CBG15669 [Caenorhabditis briggsae]
Length = 268
Score = 62.4 bits (150), Expect = 1e-08
Identities = 42/136 (30%), Positives = 66/136 (47%), Gaps = 20/136 (14%)
Query: 156 AVCLLVYFGVSTLLDA---SSSDSQKSDDE------QKEAELAVSDFSG-DGAGILAAAS 205
+ L FG+ L + S ++ Q+ +E ++E EL S F +G G+ +
Sbjct: 107 STALFALFGLKMLHEGWTMSPNEGQEGFEEAQAEVAKREGELDASKFEMLEGGGVAPQSE 166
Query: 206 T----------IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
T + F L FVAEWGD+S +TI L A + GVI G + GH + T IAV
Sbjct: 167 TKKIFLFTSRIFIEAFTLTFVAEWGDRSQLTTIILGARENIAGVIGGGVLGHALCTGIAV 226
Query: 256 LGGSLLGTFLSEKVTS 271
+GG ++ +S + +
Sbjct: 227 IGGKIVAQRISVRTVT 242
>UniRef100_Q9BL67 Hypothetical protein Y54F10AL.1 [Caenorhabditis elegans]
Length = 297
Score = 62.0 bits (149), Expect = 2e-08
Identities = 38/137 (27%), Positives = 63/137 (45%), Gaps = 21/137 (15%)
Query: 156 AVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGD-----------GAGILAAA 204
+ L FG+ L + + + + +EA+ V+ G+ G G+ + +
Sbjct: 135 STALFALFGLKMLHEGWTMSPNEGQEGYEEAQAEVAKREGELDAGKFEMLEGGGGVASQS 194
Query: 205 ST----------IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIA 254
T + F L FVAEWGD+S +TI L A + GVI G + GH + T IA
Sbjct: 195 ETRKIFLFTSRIFIEAFSLTFVAEWGDRSQLTTIILGARENIAGVIGGGILGHALCTGIA 254
Query: 255 VLGGSLLGTFLSEKVTS 271
V+GG ++ +S + +
Sbjct: 255 VIGGKIVAQRISVRTVT 271
>UniRef100_Q7SFP1 Hypothetical protein [Neurospora crassa]
Length = 504
Score = 61.6 bits (148), Expect = 2e-08
Identities = 34/103 (33%), Positives = 59/103 (57%), Gaps = 2/103 (1%)
Query: 171 ASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIAL 230
+SS D++ + + ++ F+ + +L+ A V TF++ F+ EWGD+S +TIA+
Sbjct: 385 SSSPDARNASPRGRNLTECLAGFNNLVSLLLSPAW--VQTFVMTFLGEWGDRSQIATIAM 442
Query: 231 AAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTSSG 273
AA V G++ GHG+ T +AV+GG + +S KV + G
Sbjct: 443 AAGQDYWWVTLGAVVGHGICTSVAVIGGKAIAGKVSLKVITVG 485
Score = 36.2 bits (82), Expect = 0.94
Identities = 20/79 (25%), Positives = 40/79 (50%), Gaps = 1/79 (1%)
Query: 194 SGDGA-GILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
SG G G++ + +F ++ +E GDK+F +A L V +G+ A T+
Sbjct: 224 SGKGTEGVITPFHSFFLSFTMILFSEIGDKTFLVAALMAMKHDRLVVFSGAFAALITMTI 283
Query: 253 IAVLGGSLLGTFLSEKVTS 271
++ + G + T + +K+T+
Sbjct: 284 LSAVLGHAVPTLIPKKITN 302
>UniRef100_Q10320 Hypothetical UPF0016 protein C17G8.08c in chromosome I
[Schizosaccharomyces pombe]
Length = 287
Score = 61.6 bits (148), Expect = 2e-08
Identities = 30/101 (29%), Positives = 51/101 (49%), Gaps = 2/101 (1%)
Query: 165 VSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAA--ASTIVSTFLLVFVAEWGDK 222
+ LL+ ++ S + + +S G ++A + + F L FV+EWGD+
Sbjct: 159 IDQLLEEGAAPSHYTGHRSRSGHTLMSQLKSKGRNVMATLFSPLFIKAFALTFVSEWGDR 218
Query: 223 SFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGT 263
S +TIA+AA+ + GV G+ GH T +AV+ G + T
Sbjct: 219 SQIATIAMAASDNVYGVFMGANVGHACCTALAVISGKYIST 259
Score = 36.2 bits (82), Expect = 0.94
Identities = 21/65 (32%), Positives = 33/65 (50%)
Query: 206 TIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFL 265
+++ + ++F E GDK+F LA +S L V AGS + + TL+ VL G
Sbjct: 49 SLIFSISMIFGCEIGDKTFIVAALLAFENSRLTVFAGSYSALFIMTLLGVLLGHAAPLLF 108
Query: 266 SEKVT 270
K+T
Sbjct: 109 PRKLT 113
>UniRef100_Q6BM46 Similar to ca|CA2327|IPF4782 Candida albicans IPF4782 probable
membrane protein [Debaryomyces hansenii]
Length = 335
Score = 60.8 bits (146), Expect = 4e-08
Identities = 29/66 (43%), Positives = 42/66 (62%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
+ F++ F+ EWGD+S +TIA+AA S VI G++ GHGV T A +GG LL T +S
Sbjct: 248 IQVFVMTFLGEWGDRSQIATIAMAAGSDYWYVILGAIIGHGVCTAAACVGGKLLATRISM 307
Query: 268 KVTSSG 273
+ + G
Sbjct: 308 RNVTLG 313
>UniRef100_UPI00003AD943 UPI00003AD943 UniRef100 entry
Length = 258
Score = 60.5 bits (145), Expect = 5e-08
Identities = 47/175 (26%), Positives = 75/175 (42%), Gaps = 30/175 (17%)
Query: 120 LFSLGHLAHLRRTFHYVDELLPFRFGETDLPIDDIAAVCLLVYFGVSTLLDA--SSSDSQ 177
+ +LG + L F Y ++P + + L FG+ L + S D
Sbjct: 68 MLALGLMTCLSVLFGYATTVIPRVYTY-------YVSTALFAIFGIRMLREGLKMSPDEG 120
Query: 178 KSDDEQKEAELAVSD--------FSGDGAGILAAASTI-------------VSTFLLVFV 216
+ + E+ +AE+ D +G G + +TI V F L F+
Sbjct: 121 QEELEEVQAEIKKKDEELQRTKLLNGPGDVETGSTATIPQKKWLHFISPIFVQAFTLTFL 180
Query: 217 AEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
AEWGD+S +TI LAA P GV G GH + T +AV+GG ++ +S + +
Sbjct: 181 AEWGDRSQLTTIVLAAREDPYGVAVGGTVGHCLCTGLAVIGGRMIAQKISVRTVT 235
Score = 36.6 bits (83), Expect = 0.72
Identities = 18/51 (35%), Positives = 31/51 (60%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGG 258
V+ ++ V+E GDK+FF +A + L V+AG++ G+ T ++VL G
Sbjct: 32 VAAISVIIVSELGDKTFFIAAIMAMRYNRLTVLAGAMLALGLMTCLSVLFG 82
>UniRef100_Q6MVD6 Hypothetical protein B13D15.080 [Neurospora crassa]
Length = 481
Score = 60.5 bits (145), Expect = 5e-08
Identities = 28/66 (42%), Positives = 42/66 (63%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSE 267
V TF++ F+ EWGD+S +TIA+AA V G++ GHG+ T +AV+GG + +S
Sbjct: 397 VQTFVMTFLGEWGDRSQIATIAMAAGQDYWWVTLGAVVGHGICTSVAVIGGKAIAGKVSL 456
Query: 268 KVTSSG 273
KV + G
Sbjct: 457 KVITVG 462
Score = 36.2 bits (82), Expect = 0.94
Identities = 20/79 (25%), Positives = 40/79 (50%), Gaps = 1/79 (1%)
Query: 194 SGDGA-GILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATL 252
SG G G++ + +F ++ +E GDK+F +A L V +G+ A T+
Sbjct: 224 SGKGTEGVITPFHSFFLSFTMILFSEIGDKTFLVAALMAMKHDRLVVFSGAFAALITMTI 283
Query: 253 IAVLGGSLLGTFLSEKVTS 271
++ + G + T + +K+T+
Sbjct: 284 LSAVLGHAVPTLIPKKITN 302
>UniRef100_UPI0000365A69 UPI0000365A69 UniRef100 entry
Length = 259
Score = 60.1 bits (144), Expect = 6e-08
Identities = 36/112 (32%), Positives = 59/112 (52%), Gaps = 9/112 (8%)
Query: 169 LDASSSDSQKSDDEQKEAELA--VSDF-SGDGAGILAA------ASTIVSTFLLVFVAEW 219
L+ ++ +K D+E + ++LA D +G GA + + + F L F+AEW
Sbjct: 125 LEEVQAEIKKKDEELQRSKLANGTPDIEAGSGATVPQTKWYSFISPIFIQAFTLTFLAEW 184
Query: 220 GDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
GD+S +TI LAA P GV G GH + T +AV+GG ++ +S + +
Sbjct: 185 GDRSQLTTIILAAREDPFGVAVGGTLGHCLCTGLAVIGGRMVAQKISVRTVT 236
Score = 37.4 bits (85), Expect = 0.42
Identities = 27/86 (31%), Positives = 43/86 (49%), Gaps = 4/86 (4%)
Query: 174 SDSQKSDDEQKEAELAVSDFSGDG-AGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAA 232
S QK+ Q + D S G +G + A V+ ++ V+E GDK+FF +A
Sbjct: 1 SSLQKASPPQPVNPVFSEDESSKGNSGFIHA---FVAAISVIIVSELGDKTFFIAAIMAM 57
Query: 233 ASSPLGVIAGSLAGHGVATLIAVLGG 258
+ L V+ G++ GV T ++VL G
Sbjct: 58 RYNRLVVLTGAMLALGVMTCLSVLFG 83
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 408,310,663
Number of Sequences: 2790947
Number of extensions: 15554603
Number of successful extensions: 71006
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 71
Number of HSP's successfully gapped in prelim test: 66
Number of HSP's that attempted gapping in prelim test: 70856
Number of HSP's gapped (non-prelim): 193
length of query: 273
length of database: 848,049,833
effective HSP length: 125
effective length of query: 148
effective length of database: 499,181,458
effective search space: 73878855784
effective search space used: 73878855784
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC140913.9


