

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
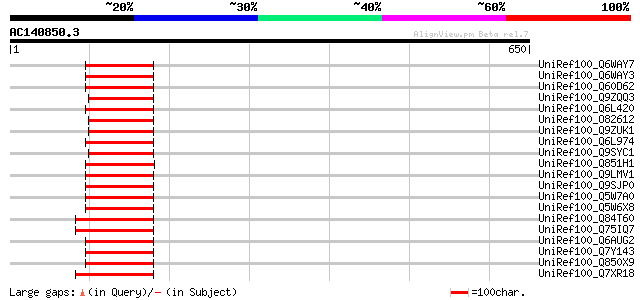
Query= AC140850.3 + phase: 0 /pseudo
(650 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum] 157 9e-37
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 156 2e-36
UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solan... 121 7e-26
UniRef100_Q9ZQQ3 Putative retroelement pol polyprotein [Arabidop... 121 7e-26
UniRef100_Q6L420 Putative polyprotein [Solanum demissum] 119 4e-25
UniRef100_O82612 T5H22.5 protein [Arabidopsis thaliana] 117 8e-25
UniRef100_Q9ZUK1 Putative retroelement pol polyprotein [Arabidop... 116 2e-24
UniRef100_Q6L974 GAG-POL [Vitis vinifera] 114 1e-23
UniRef100_Q9SYC1 F11M15.5 protein [Arabidopsis thaliana] 113 2e-23
UniRef100_Q851H1 Putative gag-pol polyprotein [Oryza sativa] 113 2e-23
UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana] 113 2e-23
UniRef100_Q9SJP0 Putative retroelement pol polyprotein [Arabidop... 112 3e-23
UniRef100_Q5W7A0 Putative polyprotein [Oryza sativa] 112 3e-23
UniRef100_Q5W6X8 Putative polyprotein [Oryza sativa] 112 3e-23
UniRef100_Q84T60 Putative GAG-POL [Oryza sativa] 112 3e-23
UniRef100_Q75IQ7 Putative retrotransposon protein [Oryza sativa] 112 3e-23
UniRef100_Q6AUG2 Putative polyprotein [Oryza sativa] 112 3e-23
UniRef100_Q7Y143 Putative gag-pol, 3'-partial [Oryza sativa] 112 4e-23
UniRef100_Q850X9 Putative gag-pol [Oryza sativa] 112 4e-23
UniRef100_Q7XR18 OSJNBb0022F23.12 protein [Oryza sativa] 111 6e-23
>UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 157 bits (397), Expect = 9e-37
Identities = 73/86 (84%), Positives = 78/86 (89%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
FL YP WVANIVPVPKKDGKVRMCVD+RDLN+ASPKD+FPLPHIDVLVDNTAQS VF
Sbjct: 1257 FLAVTSYPPWVANIVPVPKKDGKVRMCVDYRDLNRASPKDDFPLPHIDVLVDNTAQSSVF 1316
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
SFMDGFSGYN IKM+PED EKT+FIT
Sbjct: 1317 SFMDGFSGYNQIKMAPEDMEKTTFIT 1342
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 156 bits (394), Expect = 2e-36
Identities = 72/86 (83%), Positives = 78/86 (89%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
FL YP W+ANIVPVPKKDGKVRMCVD+RDLN+ASPKD+FPLPHIDVLVDNTAQS VF
Sbjct: 1255 FLAVTSYPPWMANIVPVPKKDGKVRMCVDYRDLNRASPKDDFPLPHIDVLVDNTAQSSVF 1314
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
SFMDGFSGYN IKM+PED EKT+FIT
Sbjct: 1315 SFMDGFSGYNQIKMAPEDMEKTTFIT 1340
>UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solanum demissum]
Length = 1784
Score = 121 bits (303), Expect = 7e-26
Identities = 55/85 (64%), Positives = 70/85 (81%)
Query: 96 LMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFS 155
+ +YP W+ANIVPVPKKDGKVRMCVD+RDLNKASPKD+F LP+I +L+DN A+ ++ S
Sbjct: 1634 IRVAQYPSWLANIVPVPKKDGKVRMCVDYRDLNKASPKDDFLLPNIHILLDNCAKHEIAS 1693
Query: 156 FMDGFSGYNLIKMSPEDREKTSFIT 180
F+D ++GY+ I M ED EKTSFIT
Sbjct: 1694 FVDCYAGYHQIIMDDEDAEKTSFIT 1718
>UniRef100_Q9ZQQ3 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1466
Score = 121 bits (303), Expect = 7e-26
Identities = 55/82 (67%), Positives = 69/82 (84%)
Query: 99 VEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMD 158
V+YP+W+AN V V KK+GK R+C+DF DLNKA PKD+FPLPHID LV++TA +++ SFMD
Sbjct: 496 VQYPDWLANTVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLSFMD 555
Query: 159 GFSGYNLIKMSPEDREKTSFIT 180
FSGYN I M+PED+EKT FIT
Sbjct: 556 AFSGYNQIMMNPEDQEKTLFIT 577
>UniRef100_Q6L420 Putative polyprotein [Solanum demissum]
Length = 1483
Score = 119 bits (297), Expect = 4e-25
Identities = 54/85 (63%), Positives = 68/85 (79%)
Query: 96 LMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFS 155
+ +YP W+ANIVPVPKKDGKVRMCVD RDLNKASPKD+FPLP+ +L+DN A+ ++ S
Sbjct: 588 IRVAQYPSWLANIVPVPKKDGKVRMCVDCRDLNKASPKDDFPLPNSHILLDNCAEHEIAS 647
Query: 156 FMDGFSGYNLIKMSPEDREKTSFIT 180
F+D ++GY+ I M ED EK SFIT
Sbjct: 648 FVDCYAGYHQIIMDDEDAEKMSFIT 672
>UniRef100_O82612 T5H22.5 protein [Arabidopsis thaliana]
Length = 637
Score = 117 bits (294), Expect = 8e-25
Identities = 54/82 (65%), Positives = 68/82 (82%)
Query: 99 VEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMD 158
V+YP+WVA V V KK+GK R+C+DF DLNKA PKD+FPLPHID LV++TA +++ +FMD
Sbjct: 373 VQYPDWVAITVVVKKKNGKDRVCIDFTDLNKACPKDSFPLPHIDRLVESTAGNELLTFMD 432
Query: 159 GFSGYNLIKMSPEDREKTSFIT 180
F GYN I M+PED+EKTSFIT
Sbjct: 433 AFLGYNQIMMNPEDQEKTSFIT 454
>UniRef100_Q9ZUK1 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1329
Score = 116 bits (291), Expect = 2e-24
Identities = 56/82 (68%), Positives = 66/82 (80%)
Query: 99 VEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMD 158
V+YPEW+AN V V KK+GK R+CVDF DLNKA KD FPLPHID LV++T ++ SFMD
Sbjct: 464 VKYPEWLANPVVVKKKNGKWRVCVDFTDLNKACSKDFFPLPHIDRLVESTTGHEMLSFMD 523
Query: 159 GFSGYNLIKMSPEDREKTSFIT 180
FSGYN I M+PED+EKTSFIT
Sbjct: 524 AFSGYNQILMNPEDQEKTSFIT 545
>UniRef100_Q6L974 GAG-POL [Vitis vinifera]
Length = 1027
Score = 114 bits (284), Expect = 1e-23
Identities = 50/86 (58%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V YP+W+AN+V VPKK+GK R+CVD+ +LN A PKD+FPLP ID +VD+T+ +
Sbjct: 49 FIREVSYPDWLANVVVVPKKEGKWRVCVDYTNLNNACPKDSFPLPRIDQIVDSTSGQGML 108
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
SF+D FSGY+ I MSP+D EK +FIT
Sbjct: 109 SFLDAFSGYHQIPMSPDDEEKIAFIT 134
>UniRef100_Q9SYC1 F11M15.5 protein [Arabidopsis thaliana]
Length = 1295
Score = 113 bits (283), Expect = 2e-23
Identities = 54/82 (65%), Positives = 65/82 (78%)
Query: 99 VEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMD 158
V+YPEW+AN V V KK+GK R+CVDF DLNKA PKD FPLPHID V++T ++ SFMD
Sbjct: 650 VKYPEWLANPVVVKKKNGKWRVCVDFTDLNKACPKDCFPLPHIDRHVESTTGHEMLSFMD 709
Query: 159 GFSGYNLIKMSPEDREKTSFIT 180
FS YN I M+P+D+EKTSFIT
Sbjct: 710 AFSAYNQILMNPDDQEKTSFIT 731
>UniRef100_Q851H1 Putative gag-pol polyprotein [Oryza sativa]
Length = 761
Score = 113 bits (283), Expect = 2e-23
Identities = 51/87 (58%), Positives = 68/87 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V VPK +GK+RMC+D+ +LNKA PKD FPLP ID +VD+TA +
Sbjct: 503 FIREVIHPEWLANPVVVPKANGKLRMCIDYTNLNKACPKDPFPLPRIDQIVDSTAGCDLL 562
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFITH 181
SF+D +SGY+ I+M+ ED EKT+FITH
Sbjct: 563 SFLDAYSGYHQIRMAREDEEKTAFITH 589
>UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana]
Length = 1838
Score = 113 bits (282), Expect = 2e-23
Identities = 52/85 (61%), Positives = 68/85 (79%)
Query: 96 LMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFS 155
++ V+YPEW+AN V V KK+GK R+CVD+ DLNKA PKD++PLPHID LV+ T+ + + S
Sbjct: 853 IIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLS 912
Query: 156 FMDGFSGYNLIKMSPEDREKTSFIT 180
FMD FSGYN I M +D+EKTSF+T
Sbjct: 913 FMDAFSGYNQILMHKDDQEKTSFVT 937
>UniRef100_Q9SJP0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1787
Score = 112 bits (281), Expect = 3e-23
Identities = 53/85 (62%), Positives = 65/85 (76%)
Query: 96 LMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFS 155
++ V YP+W+ N V V KK+GK R+C+DF DLNKA PKD+FPLPHID LV+ TA +++ S
Sbjct: 811 IVEVRYPDWLRNPVVVKKKNGKWRVCIDFTDLNKACPKDSFPLPHIDRLVEATAGNELLS 870
Query: 156 FMDGFSGYNLIKMSPEDREKTSFIT 180
FMD FSGYN I M DREKT FIT
Sbjct: 871 FMDAFSGYNQILMHQNDREKTVFIT 895
>UniRef100_Q5W7A0 Putative polyprotein [Oryza sativa]
Length = 1207
Score = 112 bits (280), Expect = 3e-23
Identities = 50/86 (58%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V VPK +GK+RMC+D+ DLNKA PKD +PLPHID +VD+TA +
Sbjct: 49 FIREVIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPHIDQIVDSTAGCDLL 108
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
F+D +SGY+ I+M+ ED EKT+FIT
Sbjct: 109 CFLDAYSGYHQIRMAREDEEKTAFIT 134
>UniRef100_Q5W6X8 Putative polyprotein [Oryza sativa]
Length = 1688
Score = 112 bits (280), Expect = 3e-23
Identities = 50/86 (58%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V VPK +GK+RMC+D+ DLNKA PKD +PLPHID +VD+TA +
Sbjct: 627 FIREVIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPHIDQIVDSTAGCDLL 686
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
F+D +SGY+ I+M+ ED EKT+FIT
Sbjct: 687 CFLDAYSGYHQIRMAREDEEKTAFIT 712
>UniRef100_Q84T60 Putative GAG-POL [Oryza sativa]
Length = 1380
Score = 112 bits (280), Expect = 3e-23
Identities = 56/98 (57%), Positives = 69/98 (70%)
Query: 83 LRVRFKSRLMRVFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHID 142
+R L F+ V +PEW+AN V V K +GK RMCVDF DLNKA PKD+FPLP ID
Sbjct: 840 IREELDKLLKAGFIKEVLHPEWLANPVMVRKANGKWRMCVDFTDLNKACPKDHFPLPRID 899
Query: 143 VLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSFIT 180
LVD+TA ++ SF+D +SGY+ I M+ ED EKTSFIT
Sbjct: 900 QLVDSTAGCELLSFIDAYSGYHQISMAKEDEEKTSFIT 937
>UniRef100_Q75IQ7 Putative retrotransposon protein [Oryza sativa]
Length = 1403
Score = 112 bits (280), Expect = 3e-23
Identities = 56/98 (57%), Positives = 69/98 (70%)
Query: 83 LRVRFKSRLMRVFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHID 142
+R L F+ V +PEW+AN V V K +GK RMCVDF DLNKA PKD+FPLP ID
Sbjct: 863 IREELDKLLKAGFIKEVLHPEWLANPVMVRKANGKWRMCVDFTDLNKACPKDHFPLPRID 922
Query: 143 VLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSFIT 180
LVD+TA ++ SF+D +SGY+ I M+ ED EKTSFIT
Sbjct: 923 QLVDSTAGCELLSFIDAYSGYHQISMAKEDEEKTSFIT 960
>UniRef100_Q6AUG2 Putative polyprotein [Oryza sativa]
Length = 1194
Score = 112 bits (280), Expect = 3e-23
Identities = 53/86 (61%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V V K +GK RMCVDF DLNKA PKD+FPLP ID LVD+TA ++
Sbjct: 1028 FIREVFHPEWLANPVMVRKSNGKWRMCVDFTDLNKACPKDHFPLPRIDQLVDSTAGCELL 1087
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
SF+D +SGY+ I M+ ED+EKT+FIT
Sbjct: 1088 SFLDAYSGYHQISMAKEDKEKTAFIT 1113
>UniRef100_Q7Y143 Putative gag-pol, 3'-partial [Oryza sativa]
Length = 948
Score = 112 bits (279), Expect = 4e-23
Identities = 50/86 (58%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V VPK +GK+RMC+D+ DLNKA PKD FPLPHID +VD+TA +
Sbjct: 670 FIREVIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPFPLPHIDQIVDSTAGCDLL 729
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
F+D +SGY+ I+M+ +D EKT+FIT
Sbjct: 730 CFLDAYSGYHQIRMARKDEEKTAFIT 755
>UniRef100_Q850X9 Putative gag-pol [Oryza sativa]
Length = 1635
Score = 112 bits (279), Expect = 4e-23
Identities = 50/86 (58%), Positives = 67/86 (77%)
Query: 95 FLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
F+ V +PEW+AN V VPK +GK+RMC+D+ DLNKA PKD FPLPHID +VD+TA +
Sbjct: 670 FIREVIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPFPLPHIDQIVDSTAGCDLL 729
Query: 155 SFMDGFSGYNLIKMSPEDREKTSFIT 180
F+D +SGY+ I+M+ +D EKT+FIT
Sbjct: 730 CFLDAYSGYHQIRMARKDEEKTAFIT 755
>UniRef100_Q7XR18 OSJNBb0022F23.12 protein [Oryza sativa]
Length = 1213
Score = 111 bits (278), Expect = 6e-23
Identities = 55/98 (56%), Positives = 69/98 (70%)
Query: 83 LRVRFKSRLMRVFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHID 142
+R L F+ V +PEW+AN V V K +GK RMCVDF DLNKA PKD+FPLP ID
Sbjct: 769 IREELDKLLKAGFIREVLHPEWLANPVMVRKANGKWRMCVDFTDLNKACPKDHFPLPRID 828
Query: 143 VLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSFIT 180
LVD+TA ++ SF+D +SGY+ I M+ ED EKT+FIT
Sbjct: 829 QLVDSTAGCELLSFLDAYSGYHQISMAKEDEEKTTFIT 866
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.347 0.151 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 909,495,975
Number of Sequences: 2790947
Number of extensions: 32858002
Number of successful extensions: 130128
Number of sequences better than 10.0: 1805
Number of HSP's better than 10.0 without gapping: 1637
Number of HSP's successfully gapped in prelim test: 168
Number of HSP's that attempted gapping in prelim test: 128131
Number of HSP's gapped (non-prelim): 2014
length of query: 650
length of database: 848,049,833
effective HSP length: 134
effective length of query: 516
effective length of database: 474,062,935
effective search space: 244616474460
effective search space used: 244616474460
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC140850.3


