

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140721.10 + phase: 0 /pseudo
(329 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
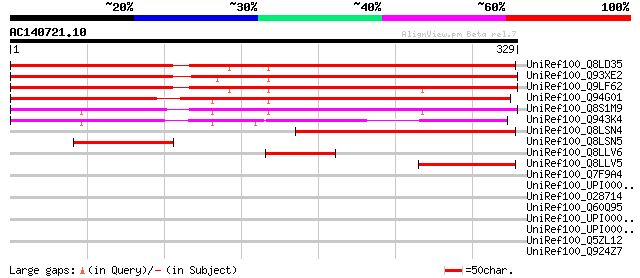
Sequences producing significant alignments: (bits) Value
UniRef100_Q8LD35 Nuclear matrix protein 1 [Arabidopsis thaliana] 405 e-111
UniRef100_Q93XE2 Nuclear matrix protein 1 [Lycopersicon esculentum] 393 e-108
UniRef100_Q9LF62 Hypothetical protein K10A8_100 [Arabidopsis tha... 380 e-104
UniRef100_Q94G01 Nuclear matrix protein 1 [Zea mays] 335 1e-90
UniRef100_Q8S1M9 Putative nuclear matrix protein [Oryza sativa] 316 6e-85
UniRef100_Q943K4 P0039A07.2 protein [Oryza sativa] 241 3e-62
UniRef100_Q8LSN4 Nuclear matrix protein 1 [Triticum aestivum] 204 3e-51
UniRef100_Q8LSN5 Nuclear matrix protein 1 [Triticum aestivum] 86 2e-15
UniRef100_Q8LLV6 Nuclear matrix protein 1 [Phaseolus lunatus] 79 2e-13
UniRef100_Q8LLV5 Nuclear matrix protein 1 [Hordeum vulgare] 74 7e-12
UniRef100_Q7F9A4 OSJNBa0065J03.22 protein [Oryza sativa] 40 0.067
UniRef100_UPI000026702D UPI000026702D UniRef100 entry 36 1.7
UniRef100_O28714 Chromosome segregation protein [Archaeoglobus f... 35 2.8
UniRef100_Q60Q95 Hypothetical protein CBG21911 [Caenorhabditis b... 35 3.7
UniRef100_UPI00003ACB9F UPI00003ACB9F UniRef100 entry 33 8.2
UniRef100_UPI000022072E UPI000022072E UniRef100 entry 33 8.2
UniRef100_Q5ZL12 Hypothetical protein [Gallus gallus] 33 8.2
UniRef100_Q924Z7 UCH37-interacting protein 1, isoform 1 [Mus mus... 33 8.2
>UniRef100_Q8LD35 Nuclear matrix protein 1 [Arabidopsis thaliana]
Length = 329
Score = 405 bits (1040), Expect = e-111
Identities = 223/339 (65%), Positives = 258/339 (75%), Gaps = 21/339 (6%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MA+KQME IQKKLG+L+YPRANA AQSLLFAGMERYAL EWLFF+LLGDKSPFSQQNLQG
Sbjct: 1 MAAKQMEEIQKKLGLLSYPRANAPAQSLLFAGMERYALLEWLFFKLLGDKSPFSQQNLQG 60
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
DA RDEET RIQYLAEIAKFLGIT TVD +AIQ H +D L N+V
Sbjct: 61 DAGVRDEETVRIQYLAEIAKFLGITPTVDIEAIQGHGTYEDRMEML----------RNIV 110
Query: 121 CIFVSIVLTSR*PRTYT**IL--------LQKNRHKYFLKNVNCFPQMFRFSPY--IP*V 170
+ + + + + + + + + F + FP + +P V
Sbjct: 111 DLVEASLFSDNQEWSIDEQVAKDIQLIDAIAERQSLIFSEECKLFPADVQIQSIYPLPDV 170
Query: 171 AELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMI 230
+ELE+KL+EQ+KIL NLQQKVDDLA+KHAYNPDEEYTEVESQLRA LESFLETAR FN I
Sbjct: 171 SELETKLSEQAKILSNLQQKVDDLAAKHAYNPDEEYTEVESQLRARLESFLETARAFNTI 230
Query: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSET-S 289
YTKEIRPWTHMMEVPQLHGFGPAANRLLEAY MLLKFLGNL+NLRDSHAAL+ GSS T +
Sbjct: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYNMLLKFLGNLKNLRDSHAALSIGSSGTVA 290
Query: 290 GGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKM 328
G PSSV+RI+S+CE+ +TV+NRDLGILSASIARE GE++
Sbjct: 291 GEPSSVTRIVSDCEAALTVLNRDLGILSASIAREQGERL 329
>UniRef100_Q93XE2 Nuclear matrix protein 1 [Lycopersicon esculentum]
Length = 331
Score = 393 bits (1010), Expect = e-108
Identities = 216/341 (63%), Positives = 254/341 (74%), Gaps = 23/341 (6%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MA+KQME IQKKL LNYPRANA AQSLLFAGMERYAL EWLFF+LLGDKSPFSQQNLQG
Sbjct: 1 MAAKQMEEIQKKLATLNYPRANAPAQSLLFAGMERYALLEWLFFKLLGDKSPFSQQNLQG 60
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
DA+DRDEET+RIQYLAEIAKFLGITTTVD +AIQ +D L L+
Sbjct: 61 DAVDRDEETSRIQYLAEIAKFLGITTTVDPEAIQGRGSYEDRMEML-----------RLI 109
Query: 121 CIFVSIVLTSR*P---------RTYT**ILLQKNRHKYFLKNVNCFPQMFRFSPY--IP* 169
V + + P + + + + + F + FP + +P
Sbjct: 110 VDLVEASMYADNPEWSVDEQVAKDIQLIDAIAEKQSQIFSEECKLFPADVQIQSIYPLPD 169
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNM 229
+++LE +L++QS LL+LQ+ VDDLASKH YNPDEEY +VE++LR HLESFL+TARTFN
Sbjct: 170 ISDLEKQLSDQSNRLLSLQEMVDDLASKHPYNPDEEYVDVEAKLRGHLESFLDTARTFNT 229
Query: 230 IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSET- 288
IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKML KFLGNL+NLRDSHAA+A GSSET
Sbjct: 230 IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLWKFLGNLKNLRDSHAAVAAGSSETV 289
Query: 289 SGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS 329
+G PSSV+RIISECE+ +T++NRDL ILSASIARE GE +S
Sbjct: 290 AGEPSSVTRIISECETALTLLNRDLAILSASIARERGEDIS 330
>UniRef100_Q9LF62 Hypothetical protein K10A8_100 [Arabidopsis thaliana]
Length = 383
Score = 380 bits (977), Expect = e-104
Identities = 222/374 (59%), Positives = 255/374 (67%), Gaps = 55/374 (14%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MA+KQME IQKKL +L+YPRANA AQSLLFAGMERYAL EWLFF+LLGDKSPFSQQNLQG
Sbjct: 1 MAAKQMEEIQKKLRLLSYPRANAPAQSLLFAGMERYALLEWLFFKLLGDKSPFSQQNLQG 60
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
DA RDEET RIQYLAEIAKFLGIT TVD +AIQ H +D L N+V
Sbjct: 61 DAGVRDEETVRIQYLAEIAKFLGITPTVDIEAIQGHGTYEDRMEML----------RNIV 110
Query: 121 CIFVSIVLTSR*PRTYT**IL--------LQKNRHKYFLKNVNCFPQMFRFSPY--IP*V 170
+ + + + + + + + + F + FP + +P V
Sbjct: 111 DLVEASLFSDNQEWSIDEQVAKDIQLIDAIAERQSLIFSEECKLFPADVQIQSIYPLPDV 170
Query: 171 AELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMI 230
+ELE+KL+EQ+KIL NLQQKVDDLA+KHAYNPDEEYTEVESQLRA LESFLETAR FN I
Sbjct: 171 SELETKLSEQAKILSNLQQKVDDLAAKHAYNPDEEYTEVESQLRARLESFLETARAFNTI 230
Query: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLK------------------------ 266
YTKEIRPWTHMMEVPQLHGFGPAANRLLEAY MLLK
Sbjct: 231 YTKEIRPWTHMMEVPQLHGFGPAANRLLEAYNMLLKVPHCLLLSVYLVVSTILSPYNLYF 290
Query: 267 ----------FLGNLQNLRDSHAALAFGSSET-SGGPSSVSRIISECESEMTVINRDLGI 315
FLGNL+NLRDSHAAL+ GSS T +G PSSV+RI+S+CE+ +TV+NRDLGI
Sbjct: 291 DSDTQIVWQQFLGNLKNLRDSHAALSIGSSGTVAGEPSSVTRIVSDCEAALTVLNRDLGI 350
Query: 316 LSASIAREHGEKMS 329
LSASIARE K S
Sbjct: 351 LSASIAREQATKDS 364
>UniRef100_Q94G01 Nuclear matrix protein 1 [Zea mays]
Length = 328
Score = 335 bits (859), Expect = 1e-90
Identities = 185/339 (54%), Positives = 230/339 (67%), Gaps = 28/339 (8%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MASKQME IQ+KL +L YPRANA AQSLLFAG+ERY L EWLFFRLLGD+SPF+QQN QG
Sbjct: 1 MASKQMEEIQRKLSLLEYPRANAPAQSLLFAGVERYRLLEWLFFRLLGDRSPFTQQNWQG 60
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
D+LDRDEE RIQ+LAEIA FLGIT + DT+AIQ Y + L+
Sbjct: 61 DSLDRDEENNRIQHLAEIANFLGITPSADTEAIQGR--------------GSYEERVELL 106
Query: 121 CIFVSIVLTS------------R*PRTYT**ILLQKNRHKYFLKNVNCFPQMFRFSPY-- 166
+ V +V S + + + + + + F + FP +
Sbjct: 107 HLIVDLVEASCYADNPEWSVDKQLEKDVQLVDSIAEKQAQIFSEECKLFPADVQIQSIYP 166
Query: 167 IP*VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETART 226
+P +AELE KL+E +K + NLQQ V +LASK+ YNP+E+Y E E +LR +L+SFLET ++
Sbjct: 167 LPDIAELELKLSEYTKKMSNLQQMVQELASKYDYNPNEDYAETELKLREYLQSFLETVKS 226
Query: 227 FNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSS 286
FN IYTKEI PWTHMMEVPQLHGFGPAANRLLEAY LLKFLGNL++LRDS+ A+A GS
Sbjct: 227 FNTIYTKEIHPWTHMMEVPQLHGFGPAANRLLEAYNTLLKFLGNLRSLRDSYTAMAAGSL 286
Query: 287 ETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHG 325
S PSSV++IIS+CES +T +N L ILS S+ARE G
Sbjct: 287 SASNEPSSVTKIISDCESALTFLNHSLSILSTSVAREQG 325
>UniRef100_Q8S1M9 Putative nuclear matrix protein [Oryza sativa]
Length = 383
Score = 316 bits (809), Expect = 6e-85
Identities = 188/395 (47%), Positives = 236/395 (59%), Gaps = 80/395 (20%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFR--------------- 45
MASKQME IQ+KL +L YPRANA AQSLLFAG+ERY L EWLFFR
Sbjct: 1 MASKQMEEIQRKLAVLAYPRANAPAQSLLFAGVERYRLLEWLFFRYAATHATFPISASQS 60
Query: 46 -----------------------LLGDKSPFSQQNLQGDALDRDEETARIQYLAEIAKFL 82
LLGD+SPF+QQN QGD+LDRDEE +RIQ+LAEIA FL
Sbjct: 61 ARLAAQLRSQLEIEGFVFSLLRRLLGDRSPFTQQNWQGDSLDRDEENSRIQHLAEIANFL 120
Query: 83 GITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLVCIFVSIVLTS------------ 130
GIT +VDT+AIQ D+ + L+C+ V +V S
Sbjct: 121 GITPSVDTEAIQGRGSYDER--------------VELLCLIVDLVEASCYADNPEWSVDE 166
Query: 131 R*PRTYT**ILLQKNRHKYFLKNVNCFPQMFRFSPY--IP*VAELESKLTEQSKILLNLQ 188
+ + + + + + F + FP + +P + ELE KL+E +K + NLQ
Sbjct: 167 QLAKDVLLVDSIAEKQAQIFSEECKLFPADVQIQSIYPLPDITELELKLSEYTKKMSNLQ 226
Query: 189 QKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIYTKEIRPWTHMMEVPQLH 248
V +LASK+ YNP+E+Y E E +LR HL+SFLET ++FNMIYTKEI PWTHMMEVPQLH
Sbjct: 227 LMVQELASKYDYNPNEDYAETELKLREHLQSFLETVKSFNMIYTKEIHPWTHMMEVPQLH 286
Query: 249 GFGPAANRLLEAYKMLLK--------------FLGNLQNLRDSHAALAFGSSETSGGPSS 294
GFGPAANRLLEAY LLK FL NL++LRDS+AA+A GS S PSS
Sbjct: 287 GFGPAANRLLEAYNTLLKLCAVFLLVSTVFCLFLSNLRSLRDSYAAMAAGSLSASNEPSS 346
Query: 295 VSRIISECESEMTVINRDLGILSASIAREHGEKMS 329
V++IIS+CES +T +N L ILS S+ARE GE ++
Sbjct: 347 VTKIISDCESALTFLNNSLSILSTSVAREQGETLN 381
>UniRef100_Q943K4 P0039A07.2 protein [Oryza sativa]
Length = 360
Score = 241 bits (614), Expect = 3e-62
Identities = 158/376 (42%), Positives = 204/376 (54%), Gaps = 101/376 (26%)
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFR--------------- 45
MASKQME IQ+KL +L YPRANA AQSLLFAG+ERY L EWLFFR
Sbjct: 1 MASKQMEEIQRKLAVLAYPRANAPAQSLLFAGVERYRLLEWLFFRYAATHATFPISASQS 60
Query: 46 -----------------------LLGDKSPFSQQNLQGDALDRDEETARIQYLAEIAKFL 82
LLGD+SPF+QQN QGD+LDRDEE +RIQ+LAEIA FL
Sbjct: 61 ARLAAQLRSQLEIEGFVFSLLRRLLGDRSPFTQQNWQGDSLDRDEENSRIQHLAEIANFL 120
Query: 83 GITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLVCIFVSIVLTS------------ 130
GIT +VDT+AIQ D+ + L+C+ V +V S
Sbjct: 121 GITPSVDTEAIQGRGSYDE--------------RVELLCLIVDLVEASCYADNPEWSVDE 166
Query: 131 R*PRTYT**ILLQKNRHKYFLKNVNCFP---QMFRFSPYIP*VAELESKLTEQSKILLNL 187
+ + + + + + F + FP Q+ P +P + ELE KL+E +K + NL
Sbjct: 167 QLAKDVLLVDSIAEKQAQIFSEECKLFPADVQIQSIYP-LPDITELELKLSEYTKKMSNL 225
Query: 188 QQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIYTKEIRPWTHMMEVPQL 247
Q V +LASK+ YNP+E+Y E E +LR HL+SFLET ++FNMIYT
Sbjct: 226 QLMVQELASKYDYNPNEDYAETELKLREHLQSFLETVKSFNMIYT--------------- 270
Query: 248 HGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESEMT 307
KFL NL++LRDS+AA+A GS S PSSV++IIS+CES +T
Sbjct: 271 ------------------KFLSNLRSLRDSYAAMAAGSLSASNEPSSVTKIISDCESALT 312
Query: 308 VINRDLGILSASIARE 323
+N L ILS S+ARE
Sbjct: 313 FLNNSLSILSTSVARE 328
>UniRef100_Q8LSN4 Nuclear matrix protein 1 [Triticum aestivum]
Length = 146
Score = 204 bits (519), Expect = 3e-51
Identities = 98/143 (68%), Positives = 117/143 (81%)
Query: 186 NLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIYTKEIRPWTHMMEVP 245
NLQQ V +LASK+ YNP+E+Y E E +LR HL+SFLET ++FNMIYTKEI PWTHMMEVP
Sbjct: 4 NLQQMVQELASKYDYNPNEDYAETELKLREHLQSFLETVKSFNMIYTKEIHPWTHMMEVP 63
Query: 246 QLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESE 305
QLHGFGPAANRLLEAY LLKFLGNL++LRDS+ A+A GS S PSSV++IIS+CES
Sbjct: 64 QLHGFGPAANRLLEAYNTLLKFLGNLRSLRDSYTAMAAGSLSNSSEPSSVTKIISDCESA 123
Query: 306 MTVINRDLGILSASIAREHGEKM 328
+T +N L ILS S+ARE GE +
Sbjct: 124 LTFLNNSLAILSTSVAREQGETL 146
>UniRef100_Q8LSN5 Nuclear matrix protein 1 [Triticum aestivum]
Length = 122
Score = 85.5 bits (210), Expect = 2e-15
Identities = 44/65 (67%), Positives = 52/65 (79%)
Query: 42 LFFRLLGDKSPFSQQNLQGDALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDD 101
LFFRLLGD+SPF+QQN Q D+LDRDEE +RIQ+LAEIA FLGIT +VDT+AIQ D+
Sbjct: 1 LFFRLLGDRSPFTQQNWQVDSLDRDEENSRIQHLAEIANFLGITPSVDTEAIQGRGSYDE 60
Query: 102 LFFFL 106
FL
Sbjct: 61 RVEFL 65
>UniRef100_Q8LLV6 Nuclear matrix protein 1 [Phaseolus lunatus]
Length = 123
Score = 79.0 bits (193), Expect = 2e-13
Identities = 38/45 (84%), Positives = 43/45 (95%)
Query: 167 IP*VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVES 211
+P V+ELESK +EQSKILLNLQQKVDDLASKHAY+PDEEYTEVE+
Sbjct: 78 LPDVSELESKFSEQSKILLNLQQKVDDLASKHAYHPDEEYTEVEA 122
>UniRef100_Q8LLV5 Nuclear matrix protein 1 [Hordeum vulgare]
Length = 66
Score = 73.6 bits (179), Expect = 7e-12
Identities = 34/63 (53%), Positives = 47/63 (73%)
Query: 266 KFLGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHG 325
KFLGNL++LRDS+ +A GS S PSS+++IIS+CES +T +N L ILS S+AR+ G
Sbjct: 4 KFLGNLRSLRDSYTTMAAGSLSNSSEPSSITKIISDCESVLTFLNNSLAILSTSVARDQG 63
Query: 326 EKM 328
E +
Sbjct: 64 ETL 66
>UniRef100_Q7F9A4 OSJNBa0065J03.22 protein [Oryza sativa]
Length = 274
Score = 40.4 bits (93), Expect = 0.067
Identities = 27/106 (25%), Positives = 54/106 (50%), Gaps = 5/106 (4%)
Query: 150 FLKNVNCFPQMFRFSPYIP*VAELESKLTEQSKI-LLNLQQKVDDLASKHAYNPDEEYTE 208
F++ +N F +M S ++P V L+ + ++ +L L + +++Y+ E +
Sbjct: 94 FIEVINLFGRML-ISSFVPSVNMLQGLVISYRELGVLRLGKATHGYMIRNSYDAQSENSA 152
Query: 209 VES---QLRAHLESFLETARTFNMIYTKEIRPWTHMMEVPQLHGFG 251
+E+ +L A S R F+ I+ K+I W+ M+E +HG+G
Sbjct: 153 LETSIVKLYASCGSINSAQRCFDSIHQKDIVAWSSMIEAYTIHGYG 198
>UniRef100_UPI000026702D UPI000026702D UniRef100 entry
Length = 229
Score = 35.8 bits (81), Expect = 1.7
Identities = 21/79 (26%), Positives = 36/79 (44%), Gaps = 3/79 (3%)
Query: 246 QLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESE 305
Q+ GFG + + ++ KF+ NLQ +HAA GS +T GP + + E
Sbjct: 139 QMTGFGGVLSFSIHKESLIQKFISNLQF---AHAAAHLGSVDTIVGPPKTTSHVENTSEE 195
Query: 306 MTVINRDLGILSASIAREH 324
+ + ++ S+ EH
Sbjct: 196 RSKLGIPENLIRCSVGVEH 214
>UniRef100_O28714 Chromosome segregation protein [Archaeoglobus fulgidus]
Length = 1156
Score = 35.0 bits (79), Expect = 2.8
Identities = 21/65 (32%), Positives = 36/65 (55%), Gaps = 1/65 (1%)
Query: 172 ELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIY 231
EL SK+ E + + L +K+++LA+K + DE E++S++ L S LE+ R Y
Sbjct: 248 ELTSKIPEINARIAELNEKINELAAKISELGDERSAEIQSRI-LELSSELESLRRAERFY 306
Query: 232 TKEIR 236
E +
Sbjct: 307 LDEAK 311
>UniRef100_Q60Q95 Hypothetical protein CBG21911 [Caenorhabditis briggsae]
Length = 1938
Score = 34.7 bits (78), Expect = 3.7
Identities = 40/143 (27%), Positives = 67/143 (45%), Gaps = 11/143 (7%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASK-HAYNPDEEYTEVES-QLRAHLESF---LETA 224
+A LE K KI+ + ++KVDD+ + A D T E +LR+ +++ +ET
Sbjct: 1446 IASLEKKQRAFDKIVDDWKRKVDDIQKEIDATTRDSRNTSTEVFKLRSSMDNLSEQIETL 1505
Query: 225 RTFNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQN-LRDSHAALAF 283
R N IY++EIR Q+ G + ++ + L + LQ+ L ++ AAL
Sbjct: 1506 RRENKIYSQEIRDINE-----QITQGGRTYQEVHKSVRRLEQEKDELQHALDEAEAALEA 1560
Query: 284 GSSETSGGPSSVSRIISECESEM 306
S+ V +I SE E +
Sbjct: 1561 EESKVLRLQIEVQQIRSEIEKRI 1583
>UniRef100_UPI00003ACB9F UPI00003ACB9F UniRef100 entry
Length = 748
Score = 33.5 bits (75), Expect = 8.2
Identities = 21/77 (27%), Positives = 39/77 (50%), Gaps = 4/77 (5%)
Query: 171 AELESKLTEQSKILLN--LQQKVDDLASKHA--YNPDEEYTEVESQLRAHLESFLETART 226
AE S+L+++ K + N LQ+K+++ H + DE++ +E++LR LE A
Sbjct: 93 AESSSQLSQEQKSVFNEDLQKKIEENERLHILYFEADEQHKRLEAELRTKLEVLESDAAQ 152
Query: 227 FNMIYTKEIRPWTHMME 243
+ R +T +E
Sbjct: 153 HQAVVDSLTRKYTDTIE 169
>UniRef100_UPI000022072E UPI000022072E UniRef100 entry
Length = 743
Score = 33.5 bits (75), Expect = 8.2
Identities = 31/157 (19%), Positives = 64/157 (40%), Gaps = 5/157 (3%)
Query: 172 ELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNMIY 231
E+ES+L E L+ ++ ++ +L S + ++ E EVES+L +E ++
Sbjct: 107 EVESELMEVESELMEVESELMELKSFYIFS---ELMEVESELMEVFSELMEVESELMEVF 163
Query: 232 TKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGG 291
++ + + +MEV + L+E L++ L + + E
Sbjct: 164 SELMEVESELMEV--FSELMEVESELMEVESELMEVFSELMEVESELMEVFSELMEVESE 221
Query: 292 PSSVSRIISECESEMTVINRDLGILSASIAREHGEKM 328
V + E ESE+ + +L + + + E M
Sbjct: 222 LMEVFSELMEVESELMEVESELMEVESELMEVESELM 258
>UniRef100_Q5ZL12 Hypothetical protein [Gallus gallus]
Length = 779
Score = 33.5 bits (75), Expect = 8.2
Identities = 21/77 (27%), Positives = 39/77 (50%), Gaps = 4/77 (5%)
Query: 171 AELESKLTEQSKILLN--LQQKVDDLASKHA--YNPDEEYTEVESQLRAHLESFLETART 226
AE S+L+++ K + N LQ+K+++ H + DE++ +E++LR LE A
Sbjct: 95 AESSSQLSQEQKSVFNEDLQKKIEENERLHILYFEADEQHKRLEAELRTKLEVLESDAAQ 154
Query: 227 FNMIYTKEIRPWTHMME 243
+ R +T +E
Sbjct: 155 HQAVVDSLTRKYTDTIE 171
>UniRef100_Q924Z7 UCH37-interacting protein 1, isoform 1 [Mus musculus]
Length = 364
Score = 33.5 bits (75), Expect = 8.2
Identities = 28/124 (22%), Positives = 51/124 (40%), Gaps = 6/124 (4%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLESFLETARTFNM 229
VA+L +L E + L L+ D ++H D + ++ +LR + F + F
Sbjct: 213 VADLARQLQESATKLQTLR---DQCFAQHKAGTDT--STIDQKLRLVISDFYQLILAFLQ 267
Query: 230 IYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETS 289
+Y E+R VP LH GP + + + L + + D+ A + +
Sbjct: 268 VYDDELRECCQR-PVPSLHPSGPIIQAVYQTLASCGQLLRAVMEIADTSAEAMKAARKQE 326
Query: 290 GGPS 293
G P+
Sbjct: 327 GEPN 330
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.332 0.143 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 471,934,554
Number of Sequences: 2790947
Number of extensions: 17371748
Number of successful extensions: 54610
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 54580
Number of HSP's gapped (non-prelim): 29
length of query: 329
length of database: 848,049,833
effective HSP length: 127
effective length of query: 202
effective length of database: 493,599,564
effective search space: 99707111928
effective search space used: 99707111928
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC140721.10


