

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140720.4 + phase: 0
(653 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
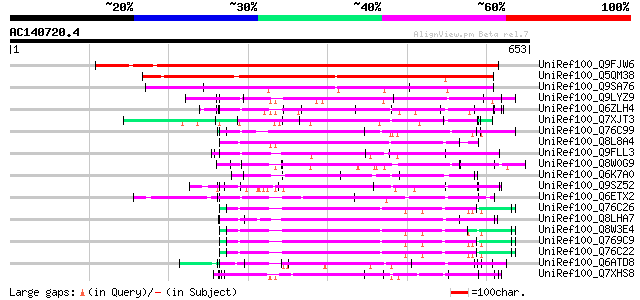
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJW6 Gb|AAD25748.1 [Arabidopsis thaliana] 626 e-178
UniRef100_Q5QM38 Pentatricopeptide (PPR) repeat-containing prote... 385 e-105
UniRef100_Q9SA76 T5I8.6 [Arabidopsis thaliana] 369 e-100
UniRef100_Q9LYZ9 Hypothetical protein F9G14_170 [Arabidopsis tha... 124 7e-27
UniRef100_Q6ZLH4 Putative pentatricopeptide (PPR) repeat-contain... 117 8e-25
UniRef100_Q7XJT3 At2g17140 protein [Arabidopsis thaliana] 106 2e-21
UniRef100_Q76C99 Rf1 [Oryza sativa] 104 7e-21
UniRef100_Q8L8A4 Fertility restorer [Petunia hybrida] 103 1e-20
UniRef100_Q9FLL3 Salt-inducible protein-like [Arabidopsis thaliana] 102 4e-20
UniRef100_Q8W0G9 P0529E05.10 protein [Oryza sativa] 101 6e-20
UniRef100_Q6K7A0 Putative PPR protein [Oryza sativa] 101 8e-20
UniRef100_Q9SZ52 Hypothetical protein AT4g31850 [Arabidopsis tha... 100 2e-19
UniRef100_Q6ETX2 Putative pentatricopeptide (PPR) repeat-contain... 99 3e-19
UniRef100_Q76C26 Hypothetical protein PPR794 [Oryza sativa] 99 4e-19
UniRef100_Q8LHA7 Pentatricopeptide repeat protein-like [Oryza sa... 98 7e-19
UniRef100_Q8W3E4 Putative membrane-associated protein [Oryza sat... 97 1e-18
UniRef100_Q769C9 PPR protein [Oryza sativa] 97 1e-18
UniRef100_Q76C22 Hypothetical protein [Oryza sativa] 97 1e-18
UniRef100_Q6ATD8 Hypothetical protein OSJNBa0018H09.7 [Oryza sat... 97 1e-18
UniRef100_Q7XHS8 Putative pentatricopeptide (PPR) repeat-contain... 97 2e-18
>UniRef100_Q9FJW6 Gb|AAD25748.1 [Arabidopsis thaliana]
Length = 505
Score = 626 bits (1614), Expect = e-178
Identities = 301/512 (58%), Positives = 396/512 (76%), Gaps = 11/512 (2%)
Query: 108 LIGKPWEGVEKVDFLERIKV--NYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIE 165
++G PWEG+E+V E + E +LK+E+L ELK++ + +L+WV +DD++
Sbjct: 1 MVGNPWEGIERVKLKELVSGVRREEVSAGELKKENLKELKKILEK----DLRWVLDDDVD 56
Query: 166 INEVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQ 225
+ E D+ + ++ R+E + VR LVDRL +EI K WKF R+M SGL FTE Q
Sbjct: 57 VEEFDLDKEFDPAKRW---RNEGEAVRVLVDRLSGREINEKHWKFVRMMNQSGLQFTEDQ 113
Query: 226 LMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFN 285
++ I++ LG K+ WKQA +VV WVY+ K + +SRFVYTKLL+VLG ARRP+EALQIFN
Sbjct: 114 MLKIVDRLGRKQSWKQASAVVHWVYSDKKRKHLRSRFVYTKLLSVLGFARRPQEALQIFN 173
Query: 286 MMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL-KYMYRKNWDPTLEP 344
MLG+ ++YPDMAAYH IAVTLGQAGLLKELL ++E MRQKP L K + +KNWDP LEP
Sbjct: 174 QMLGDRQLYPDMAAYHCIAVTLGQAGLLKELLKVIERMRQKPTKLTKNLRQKNWDPVLEP 233
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D+V+YNA+LNACVP+ QWK VSWVF ++RK+ L+PNGATYGLAMEVML+SG +D VH+ F
Sbjct: 234 DLVVYNAILNACVPTLQWKAVSWVFVELRKNGLRPNGATYGLAMEVMLESGKFDRVHDFF 293
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
KM+ +GE P+A+TYKV+VR W+EGK++EAV+AVRDME++GV+GT SVYYELACCLCN
Sbjct: 294 RKMKSSGEAPKAITYKVLVRALWREGKIEEAVEAVRDMEQKGVIGTGSVYYELACCLCNN 353
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTV 524
GRW DA LEV ++KRL + +PLE+TFTG+I +S++GGH+DDC+ IF+YM+D C PN+GT
Sbjct: 354 GRWCDAMLEVGRMKRLENCRPLEITFTGLIAASLNGGHVDDCMAIFQYMKDKCDPNIGTA 413
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAK-SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
N MLKVY +NDMFS AK LFEE+ K + L P+ YTY+ MLEAS+R QWEYFEHVY+
Sbjct: 414 NMMLKVYGRNDMFSEAKELFEEIVSRKETHLVPNEYTYSFMLEASARSLQWEYFEHVYQT 473
Query: 584 MILSGYHLDQNKHLPLLVKASRAGKVDNILVI 615
M+LSGY +DQ KH +L++ASRAGKV N+ ++
Sbjct: 474 MVLSGYQMDQTKHASMLIEASRAGKVTNLCLV 505
>UniRef100_Q5QM38 Pentatricopeptide (PPR) repeat-containing protein-like [Oryza
sativa]
Length = 988
Score = 385 bits (990), Expect = e-105
Identities = 190/449 (42%), Positives = 295/449 (65%), Gaps = 19/449 (4%)
Query: 168 EVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLM 227
EV+ D N + + +Q L RL ++ +WKFS+++ + + F++ ++
Sbjct: 424 EVFTDVRNRPRVLQMELEDRIQK---LASRLNATDVNTPEWKFSKMIHDAKIKFSDHSIL 480
Query: 228 MIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMM 287
++++LG WK+ L VV+W+ + + + ++SR++YT +L VLGKA+RP EAL
Sbjct: 481 RVVQILGRYGNWKRVLQVVEWLQSRERFKSYKSRYIYTTVLDVLGKAKRPFEALN----- 535
Query: 288 LGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVV 347
+ YPDMAAYH IAVTLGQ+GL+KEL ++++ MR P+ K +NWDP LEPD++
Sbjct: 536 -DQLSSYPDMAAYHCIAVTLGQSGLVKELFDVIDRMRSPPKKFKLSPIQNWDPRLEPDLI 594
Query: 348 IYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKM 407
+YNAVLNACV KQW+G WV QQ+++ +++P TYGL MEVML G Y+LV+E F K+
Sbjct: 595 VYNAVLNACVQQKQWEGAFWVLQQLKEKNIRPTNTTYGLVMEVMLVCGKYNLVYEFFNKV 654
Query: 408 QRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRW 467
++ +P L YKV++ W+EGK+DEAV AV+ ME RG++G+AS+YY+LA CLC+ GR
Sbjct: 655 EKT-SIPGTLNYKVLINALWREGKIDEAVMAVKGMESRGIVGSASLYYDLARCLCSGGRC 713
Query: 468 QDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTM 527
++A L+VEKI ++ + KPL VT+TG+I++ +D G +++ IF+ M +C+PN T N M
Sbjct: 714 KEALLQVEKICKVAN-KPLVVTYTGLIQTCIDNGSMENAKYIFDEMCKYCSPNNITCNIM 772
Query: 528 LKVYSQNDMFSTAKVLFEEV-------KVAKSDLR-PDAYTYNLMLEASSRGHQWEYFEH 579
LK Y+++ MF AK L E + KV S D +T+N+ +EA + +W FE+
Sbjct: 773 LKSYTEHGMFEDAKDLLENILSGRIRSKVESSQKAIADKFTFNIFMEACAEAKRWNDFEY 832
Query: 580 VYKEMILSGYHLDQNKHLPLLVKASRAGK 608
+++M+ SGYH D+ +HL +++ A R GK
Sbjct: 833 AFRKMLSSGYHFDERRHLRMVLDAYRNGK 861
>UniRef100_Q9SA76 T5I8.6 [Arabidopsis thaliana]
Length = 1006
Score = 369 bits (947), Expect = e-100
Identities = 195/477 (40%), Positives = 289/477 (59%), Gaps = 42/477 (8%)
Query: 172 DENNNGKRKKTSKRSEVQV-VRFLVDRLCDKEIRAKDWKFSRLMKLSGLSFTEGQLMMII 230
DE+++ K + R E++ + L L +I +W+FS+ ++ + + +T+ +M +I
Sbjct: 417 DESSDIVDKPATSRVEMEDRIEKLAKVLNGADINMPEWQFSKAIRSAKIRYTDYTVMRLI 476
Query: 231 EMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGN 290
LG W++ L V++W+ ++ + R +YT L VLGK+RRP EAL +F+ ML
Sbjct: 477 HFLGKLGNWRRVLQVIEWLQRQDRYKSNKIRIIYTTALNVLGKSRRPVEALNVFHAMLLQ 536
Query: 291 IRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKP-ETLKYMYRKNWDPTLEPDVVIY 349
I YPDM AY SIAVTLGQAG +KEL +++ MR P + K + WDP LEPDVV+Y
Sbjct: 537 ISSYPDMVAYRSIAVTLGQAGHIKELFYVIDTMRSPPKKKFKPTTLEKWDPRLEPDVVVY 596
Query: 350 NAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR 409
NAVLNACV KQW+G WV QQ+++ KP+ TYGL MEVML Y+LVHE F KMQ+
Sbjct: 597 NAVLNACVQRKQWEGAFWVLQQLKQRGQKPSPVTYGLIMEVMLACEKYNLVHEFFRKMQK 656
Query: 410 NGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQD 469
+ +P AL Y+V+V T WKEGK DEAV V DME RG++G+A++YY+LA CLC+ GR +
Sbjct: 657 S-SIPNALAYRVLVNTLWKEGKSDEAVHTVEDMESRGIVGSAALYYDLARCLCSAGRCNE 715
Query: 470 A----------------------------TLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
+++KI R+ + KPL VT+TG+I++ +D G
Sbjct: 716 GLNMVNFVNPVVLKLIENLIYKADLVHTIQFQLKKICRVAN-KPLVVTYTGLIQACVDSG 774
Query: 502 HIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKV----------AK 551
+I + IF+ M+ C+PN+ T N MLK Y Q +F A+ LF+++ +
Sbjct: 775 NIKNAAYIFDQMKKVCSPNLVTCNIMLKAYLQGGLFEEARELFQKMSEDGNHIKNSSDFE 834
Query: 552 SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGK 608
S + PD YT+N ML+ + +W+ F + Y+EM+ GYH + +HL ++++ASRAGK
Sbjct: 835 SRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKRHLRMVLEASRAGK 891
Score = 69.7 bits (169), Expect = 3e-10
Identities = 65/286 (22%), Positives = 110/286 (37%), Gaps = 28/286 (9%)
Query: 244 SVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSI 303
++V + + Y L+ L K + EA+ M + A Y+ +
Sbjct: 645 NLVHEFFRKMQKSSIPNALAYRVLVNTLWKEGKSDEAVHTVEDMESR-GIVGSAALYYDL 703
Query: 304 AVTLGQAGLLKELLNIVECMR--------------QKPETLKYMYRKNWDPTLEPDVVIY 349
A L AG E LN+V + T+++ +K +P VV Y
Sbjct: 704 ARCLCSAGRCNEGLNMVNFVNPVVLKLIENLIYKADLVHTIQFQLKKICRVANKPLVVTY 763
Query: 350 NAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQR 409
++ ACV S K +++F QM+K PN T + ++ LQ G ++ ELF+KM
Sbjct: 764 TGLIQACVDSGNIKNAAYIFDQMKKVC-SPNLVTCNIMLKAYLQGGLFEEARELFQKMSE 822
Query: 410 NGE------------VPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
+G +P+ T+ M+ T ++ K D+ A R+M R G A + +
Sbjct: 823 DGNHIKNSSDFESRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKRHLRM 882
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
G+ + E ++R P + R G HI
Sbjct: 883 VLEASRAGKEEVMEATWEHMRRSNRIPPSPLIKERFFRKLEKGDHI 928
Score = 42.4 bits (98), Expect = 0.044
Identities = 32/179 (17%), Positives = 76/179 (41%), Gaps = 4/179 (2%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ + K A IF+ M P++ + + Q GL +E + + M
Sbjct: 763 YTGLIQACVDSGNIKNAAYIFDQMKKVCS--PNLVTCNIMLKAYLQGGLFEEARELFQKM 820
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ +K +++ + PD +N +L+ C ++W + +++M + N
Sbjct: 821 SEDGNHIKNS--SDFESRVLPDTYTFNTMLDTCAEQEKWDDFGYAYREMLRHGYHFNAKR 878
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
+ + ++G +++ +E M+R+ +P + K ++G A+ ++ D+
Sbjct: 879 HLRMVLEASRAGKEEVMEATWEHMRRSNRIPPSPLIKERFFRKLEKGDHISAISSLADL 937
>UniRef100_Q9LYZ9 Hypothetical protein F9G14_170 [Arabidopsis thaliana]
Length = 819
Score = 124 bits (312), Expect = 7e-27
Identities = 88/399 (22%), Positives = 173/399 (43%), Gaps = 20/399 (5%)
Query: 222 TEGQLMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEAL 281
T +L+ ++ LG + + AL W KD++ V ++++LGK R A
Sbjct: 134 TSSELLAFLKGLGFHKKFDLALRAFDWFMKQKDYQSMLDNSVVAIIISMLGKEGRVSSAA 193
Query: 282 QIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPT 341
+FN + + D+ +Y S+ +G +E +N+ ++K +
Sbjct: 194 NMFNGLQED-GFSLDVYSYTSLISAFANSGRYREAVNV--------------FKKMEEDG 238
Query: 342 LEPDVVIYNAVLNACVP-SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLV 400
+P ++ YN +LN W ++ + ++M+ + P+ TY + + +
Sbjct: 239 CKPTLITYNVILNVFGKMGTPWNKITSLVEKMKSDGIAPDAYTYNTLITCCKRGSLHQEA 298
Query: 401 HELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACC 460
++FE+M+ G + +TY ++ + K + EA+K + +M G + Y L
Sbjct: 299 AQVFEEMKAAGFSYDKVTYNALLDVYGKSHRPKEAMKVLNEMVLNGFSPSIVTYNSLISA 358
Query: 461 LCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAP 519
G +A +E++ KP T+T ++ G ++ + IFE M++ C P
Sbjct: 359 YARDGMLDEA-MELKNQMAEKGTKPDVFTYTTLLSGFERAGKVESAMSIFEEMRNAGCKP 417
Query: 520 NVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEH 579
N+ T N +K+Y F+ +F+E+ V L PD T+N +L +
Sbjct: 418 NICTFNAFIKMYGNRGKFTEMMKIFDEINVC--GLSPDIVTWNTLLAVFGQNGMDSEVSG 475
Query: 580 VYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIR 618
V+KEM +G+ ++ L+ SR G + + +Y R
Sbjct: 476 VFKEMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRR 514
Score = 92.0 bits (227), Expect = 5e-17
Identities = 93/415 (22%), Positives = 169/415 (40%), Gaps = 57/415 (13%)
Query: 261 RFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIV 320
+ Y LL V GK+ RPKEA+++ N M+ N P + Y+S+ + G+L E + +
Sbjct: 314 KVTYNALLDVYGKSHRPKEAMKVLNEMVLN-GFSPSIVTYNSLISAYARDGMLDEAMELK 372
Query: 321 ECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPN 380
M +K +PDV Y +L+ + + + +F++MR + KPN
Sbjct: 373 NQMAEKGT--------------KPDVFTYTTLLSGFERAGKVESAMSIFEEMRNAGCKPN 418
Query: 381 GATY--------------------------GLAMEVML---------QSGNYDLVHELFE 405
T+ GL+ +++ Q+G V +F+
Sbjct: 419 ICTFNAFIKMYGNRGKFTEMMKIFDEINVCGLSPDIVTWNTLLAVFGQNGMDSEVSGVFK 478
Query: 406 KMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCG 465
+M+R G VPE T+ ++ + + G ++A+ R M GV S Y + L G
Sbjct: 479 EMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAGVTPDLSTYNTVLAALARGG 538
Query: 466 RWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTV 524
W+ + + +++ KP E+T+ ++ + +G I + E + P +
Sbjct: 539 MWEQSEKVLAEMED-GRCKPNELTYCSLLHAYANGKEIGLMHSLAEEVYSGVIEPRAVLL 597
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ V S+ D+ A+ F E+K + PD T N M+ R V M
Sbjct: 598 KTLVLVCSKCDLLPEAERAFSELK--ERGFSPDITTLNSMVSIYGRRQMVAKANGVLDYM 655
Query: 585 ILSGYHLDQNKHLPLLVKASRA---GKVDNILVIYIRSQCLFYISAHSIISYKNC 636
G+ + L+ SR+ GK + IL + I +++ + Y C
Sbjct: 656 KERGFTPSMATYNSLMYMHSRSADFGKSEEILREILAKGIKPDIISYNTVIYAYC 710
Score = 73.6 bits (179), Expect = 2e-11
Identities = 76/359 (21%), Positives = 142/359 (39%), Gaps = 34/359 (9%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ + + G + E ++IF+ + + PD+ ++++ GQ G+ E+ + + M
Sbjct: 422 FNAFIKMYGNRGKFTEMMKIFDE-INVCGLSPDIVTWNTLLAVFGQNGMDSEVSGVFKEM 480
Query: 324 RQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
++ PE + +YR+ D + PD+ YN VL A W
Sbjct: 481 KRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAGVTPDLSTYNTVLAALARGGMW 540
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ V +M KPN TY + L+H L E++ P A+ K +
Sbjct: 541 EQSEKVLAEMEDGRCKPNELTYCSLLHAYANGKEIGLMHSLAEEVYSGVIEPRAVLLKTL 600
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQ-----DATLEVEKI 477
V K + EA +A +++ RG + + GR Q + L+ K
Sbjct: 601 VLVCSKCDLLPEAERAFSELKERGFSPDITTLNSMVSIY---GRRQMVAKANGVLDYMKE 657
Query: 478 KRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMF 537
+ + + M S D G ++ + E + P++ + NT++ Y +N
Sbjct: 658 RGFTPSMATYNSLMYMHSRSADFGKSEE--ILREILAKGIKPDIISYNTVIYAYCRNTRM 715
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKH 596
A +F E++ S + PD TYN + + + +E V + MI G +QN +
Sbjct: 716 RDASRIFSEMR--NSGIVPDVITYNTFIGSYAADSMFEEAIGVVRYMIKHGCRPNQNTY 772
Score = 57.4 bits (137), Expect = 1e-06
Identities = 37/190 (19%), Positives = 83/190 (43%), Gaps = 15/190 (7%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLN 354
PD+ +S+ G+ ++ + +++ M+++ T P + YN+++
Sbjct: 627 PDITTLNSMVSIYGRRQMVAKANGVLDYMKERGFT--------------PSMATYNSLMY 672
Query: 355 ACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
S + + +++ +KP+ +Y + ++ +F +M+ +G VP
Sbjct: 673 MHSRSADFGKSEEILREILAKGIKPDIISYNTVIYAYCRNTRMRDASRIFSEMRNSGIVP 732
Query: 415 EALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEV 474
+ +TY + ++ + +EA+ VR M + G + Y + C R +A L V
Sbjct: 733 DVITYNTFIGSYAADSMFEEAIGVVRYMIKHGCRPNQNTYNSIVDGYCKLNRKDEAKLFV 792
Query: 475 EKIKRL-PHA 483
E ++ L PHA
Sbjct: 793 EDLRNLDPHA 802
>UniRef100_Q6ZLH4 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 784
Score = 117 bits (294), Expect = 8e-25
Identities = 83/347 (23%), Positives = 171/347 (48%), Gaps = 20/347 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGL-LKELLNIVEC 322
YT L++ +A R ++A+ +F M+ + V P + Y+ + + + KE++ +V
Sbjct: 175 YTALVSAFSRAGRFRDAVAVFRRMVDS-GVQPAIVTYNVVLHVYSKMAVPWKEVVELVAS 233
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M++ + PD YN +++ C +K + VF +M+ S +P+
Sbjct: 234 MKEHG--------------VAPDRYTYNTLISCCRRRALYKEAAQVFDEMKASGFEPDKV 279
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
T+ ++V ++ +D E+ ++M+R G P +TY ++ ++ K+G +++AV ++M
Sbjct: 280 TFNSLLDVYGKARRHDEAIEVIQEMERVGCPPSVVTYNSLISSYVKDGLLEQAVALKQEM 339
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
E +G+ Y L L G+ A +E +++ R KP T+ +I+ G
Sbjct: 340 EVKGMKPDVVTYTTLISGLDRAGKIDAAIVEYDEMVR-NGCKPNLCTYNALIKMHGVRGK 398
Query: 503 IDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
+ + +F E+ P++ T NT+L V+ QN + S +F+E+K K+ P+ TY
Sbjct: 399 FPEMMAVFDEFRSAGFVPDIVTWNTLLAVFGQNGLDSEVSGVFKEMK--KAGYIPERDTY 456
Query: 562 NLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGK 608
++ + SR ++ +YK M+ +G + D + + +L +R G+
Sbjct: 457 VSLISSYSRCGLFDLAMQIYKRMMEAGIYPDVSTYNAVLSALARGGR 503
Score = 112 bits (280), Expect = 3e-23
Identities = 89/360 (24%), Positives = 165/360 (45%), Gaps = 28/360 (7%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
V + V+ +A R EA + + G PD AY ++ +AG ++ + +
Sbjct: 143 VLATAIRVMARAGRLAEASALLDAAPG-----PDAGAYTALVSAFSRAGRFRDAVAV--- 194
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQ---WKGVSWVFQQMRKSSLKP 379
+R+ D ++P +V YN VL+ V SK WK V + M++ + P
Sbjct: 195 -----------FRRMVDSGVQPAIVTYNVVLH--VYSKMAVPWKEVVELVASMKEHGVAP 241
Query: 380 NGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAV 439
+ TY + + Y ++F++M+ +G P+ +T+ ++ + K + DEA++ +
Sbjct: 242 DRYTYNTLISCCRRRALYKEAAQVFDEMKASGFEPDKVTFNSLLDVYGKARRHDEAIEVI 301
Query: 440 RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMD 499
++MER G + Y L G + A ++++ + KP VT+T +I
Sbjct: 302 QEMERVGCPPSVVTYNSLISSYVKDGLLEQAVALKQEME-VKGMKPDVVTYTTLISGLDR 360
Query: 500 GGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDA 558
G ID I ++ M ++ C PN+ T N ++K++ F +F+E + A PD
Sbjct: 361 AGKIDAAIVEYDEMVRNGCKPNLCTYNALIKMHGVRGKFPEMMAVFDEFRSA--GFVPDI 418
Query: 559 YTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIR 618
T+N +L + V+KEM +GY +++ ++ L+ SR G D + IY R
Sbjct: 419 VTWNTLLAVFGQNGLDSEVSGVFKEMKKAGYIPERDTYVSLISSYSRCGLFDLAMQIYKR 478
Score = 92.8 bits (229), Expect = 3e-17
Identities = 86/419 (20%), Positives = 172/419 (40%), Gaps = 63/419 (15%)
Query: 261 RFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNI- 319
+ + LL V GKARR EA+++ M + P + Y+S+ + + GLL++ + +
Sbjct: 278 KVTFNSLLDVYGKARRHDEAIEVIQEM-ERVGCPPSVVTYNSLISSYVKDGLLEQAVALK 336
Query: 320 --VECMRQKPETLKYM------------------YRKNWDPTLEPDVVIYNAVLNACVPS 359
+E KP+ + Y Y + +P++ YNA++
Sbjct: 337 QEMEVKGMKPDVVTYTTLISGLDRAGKIDAAIVEYDEMVRNGCKPNLCTYNALIKMHGVR 396
Query: 360 KQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTY 419
++ + VF + R + P+ T+ + V Q+G V +F++M++ G +PE TY
Sbjct: 397 GKFPEMMAVFDEFRSAGFVPDIVTWNTLLAVFGQNGLDSEVSGVFKEMKKAGYIPERDTY 456
Query: 420 KVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA-TLEVEKIK 478
++ ++ + G D A++ + M G+ S Y + L GRW+ A L E +
Sbjct: 457 VSLISSYSRCGLFDLAMQIYKRMMEAGIYPDVSTYNAVLSALARGGRWEQAEKLFAEMEE 516
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICI----------------------------- 509
R KP E +++ ++ + + +D +
Sbjct: 517 R--DCKPDEYSYSSLLHAYANAKRLDKMKALSDDIYSERIEPHNWLVKTLVLVNSKVNNL 574
Query: 510 -------FEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
E Q C+ ++ +N M+ +Y +N M + + +K +S + A TYN
Sbjct: 575 AEAEKAFLELRQKRCSLDINVLNAMVSIYGKNRMVRKVEKILSLMK--ESAINLSAATYN 632
Query: 563 LMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQC 621
++ SR E E++ E+ SG D+ + ++ R G++ ++ +C
Sbjct: 633 SLMHMYSRLGDCEKCENILTEIKSSGVRPDRYSYNTVIYAYGRKGQMKEASRLFSEMKC 691
Score = 90.1 bits (222), Expect = 2e-16
Identities = 78/352 (22%), Positives = 153/352 (43%), Gaps = 26/352 (7%)
Query: 239 WKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMA 298
WK+ VV+ V + K+H R+ Y L++ + KEA Q+F+ M + PD
Sbjct: 224 WKE---VVELVASMKEHGVAPDRYTYNTLISCCRRRALYKEAAQVFDEMKAS-GFEPDKV 279
Query: 299 AYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVP 358
++S+ G+A E + +++ M + P VV YN+++++ V
Sbjct: 280 TFNSLLDVYGKARRHDEAIEVIQEMER--------------VGCPPSVVTYNSLISSYVK 325
Query: 359 SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALT 418
+ + Q+M +KP+ TY + + ++G D +++M RNG P T
Sbjct: 326 DGLLEQAVALKQEMEVKGMKPDVVTYTTLISGLDRAGKIDAAIVEYDEMVRNGCKPNLCT 385
Query: 419 YKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
Y +++ GK E + + G + + L G + + +++K
Sbjct: 386 YNALIKMHGVRGKFPEMMAVFDEFRSAGFVPDIVTWNTLLAVFGQNGLDSEVSGVFKEMK 445
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMF 537
+ + P T+ +I S G D + I++ M + P+V T N +L ++ +
Sbjct: 446 KAGYI-PERDTYVSLISSYSRCGLFDLAMQIYKRMMEAGIYPDVSTYNAVLSALARGGRW 504
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYF----EHVYKEMI 585
A+ LF E++ + D +PD Y+Y+ +L A + + + + +Y E I
Sbjct: 505 EQAEKLFAEME--ERDCKPDEYSYSSLLHAYANAKRLDKMKALSDDIYSERI 554
Score = 72.4 bits (176), Expect = 4e-11
Identities = 75/371 (20%), Positives = 148/371 (39%), Gaps = 32/371 (8%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ + G + E + +F+ + PD+ ++++ GQ GL E+ + + M
Sbjct: 386 YNALIKMHGVRGKFPEMMAVFDEFR-SAGFVPDIVTWNTLLAVFGQNGLDSEVSGVFKEM 444
Query: 324 RQK---PETLKYM------------------YRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
++ PE Y+ Y++ + + PDV YNAVL+A +W
Sbjct: 445 KKAGYIPERDTYVSLISSYSRCGLFDLAMQIYKRMMEAGIYPDVSTYNAVLSALARGGRW 504
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ +F +M + KP+ +Y + + D + L + + P K +
Sbjct: 505 EQAEKLFAEMEERDCKPDEYSYSSLLHAYANAKRLDKMKALSDDIYSERIEPHNWLVKTL 564
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH 482
V K + EA KA ++ ++ S+ + + + +VEKI L
Sbjct: 565 VLVNSKVNNLAEAEKAFLELRQK----RCSLDINVLNAMVSIYGKNRMVRKVEKILSLMK 620
Query: 483 AKPLEV---TFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFS 538
+ + T+ ++ G + C I E P+ + NT++ Y +
Sbjct: 621 ESAINLSAATYNSLMHMYSRLGDCEKCENILTEIKSSGVRPDRYSYNTVIYAYGRKGQMK 680
Query: 539 TAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLP 598
A LF E+K S L+PD TYN+ +++ +E + + M+ G ++ +
Sbjct: 681 EASRLFSEMKC--SGLKPDVVTYNIFVKSYVSNSMFEEAIELVRYMVTQGCKPNERTYNS 738
Query: 599 LLVKASRAGKV 609
++ R GK+
Sbjct: 739 IVEGYCRNGKL 749
Score = 52.4 bits (124), Expect = 4e-05
Identities = 36/199 (18%), Positives = 86/199 (43%), Gaps = 37/199 (18%)
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D+ + NA+++ ++ + V + M++S++ + ATY M + + G+ + +
Sbjct: 592 DINVLNAMVSIYGKNRMVRKVEKILSLMKESAINLSAATYNSLMHMYSRLGDCEKCENIL 651
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
+++ +G P+ +Y ++ + ++G++ EA + +M+ G+
Sbjct: 652 TEIKSSGVRPDRYSYNTVIYAYGRKGQMKEASRLFSEMKCSGL----------------- 694
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGT 523
KP VT+ ++S + ++ I + YM C PN T
Sbjct: 695 -------------------KPDVVTYNIFVKSYVSNSMFEEAIELVRYMVTQGCKPNERT 735
Query: 524 VNTMLKVYSQNDMFSTAKV 542
N++++ Y +N + AK+
Sbjct: 736 YNSIVEGYCRNGKLTDAKI 754
Score = 50.8 bits (120), Expect = 1e-04
Identities = 40/173 (23%), Positives = 77/173 (44%), Gaps = 15/173 (8%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
V ++++ GK R ++ +I ++M + + A Y+S+ + G ++ NI+
Sbjct: 595 VLNAMVSIYGKNRMVRKVEKILSLMKESA-INLSAATYNSLMHMYSRLGDCEKCENILTE 653
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
++ + PD YN V+ A Q K S +F +M+ S LKP+
Sbjct: 654 IKSSG--------------VRPDRYSYNTVIYAYGRKGQMKEASRLFSEMKCSGLKPDVV 699
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEA 435
TY + ++ + + ++ EL M G P TY +V + + GK+ +A
Sbjct: 700 TYNIFVKSYVSNSMFEEAIELVRYMVTQGCKPNERTYNSIVEGYCRNGKLTDA 752
Score = 49.7 bits (117), Expect = 3e-04
Identities = 25/113 (22%), Positives = 53/113 (46%)
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+ +++ S ++P+ +Y + + G LF +M+ +G P+ +TY + V+++
Sbjct: 650 ILTEIKSSGVRPDRYSYNTVIYAYGRKGQMKEASRLFSEMKCSGLKPDVVTYNIFVKSYV 709
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
+EA++ VR M +G Y + C G+ DA + V + +L
Sbjct: 710 SNSMFEEAIELVRYMVTQGCKPNERTYNSIVEGYCRNGKLTDAKIFVSNLPQL 762
>UniRef100_Q7XJT3 At2g17140 protein [Arabidopsis thaliana]
Length = 903
Score = 106 bits (265), Expect = 2e-21
Identities = 81/312 (25%), Positives = 143/312 (44%), Gaps = 13/312 (4%)
Query: 287 MLGNIRVYPDMAAYHSIAVTLG-QAGLLKELLNIVECMRQKPETLK----YMYRKNWDPT 341
+L +++ ++ H++ ++ Q L LL++V + K + ++ P
Sbjct: 48 ILVRAKMHEEIQELHNLILSSSIQKTKLSSLLSVVSIFAKSNHIDKAFPQFQLVRSRFPE 107
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+P V +YN +L +C+ ++ + VSW+++ M + P T+ L + + S D
Sbjct: 108 NKPSVYLYNLLLESCIKERRVEFVSWLYKDMVLCGIAPQTYTFNLLIRALCDSSCVDAAR 167
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
ELF++M G P T+ ++VR + K G D+ ++ + ME GV+ +Y +
Sbjct: 168 ELFDEMPEKGCKPNEFTFGILVRGYCKAGLTDKGLELLNAMESFGVLPNKVIYNTIVSSF 227
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ-----DH 516
C GR D+ VEK+ R P VTF I + G + D IF M+
Sbjct: 228 CREGRNDDSEKMVEKM-REEGLVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLGL 286
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
PN T N MLK + + + AK LFE ++ ++D +YN+ L+ R ++
Sbjct: 287 PRPNSITYNLMLKGFCKVGLLEDAKTLFESIR--ENDDLASLQSYNIWLQGLVRHGKFIE 344
Query: 577 FEHVYKEMILSG 588
E V K+M G
Sbjct: 345 AETVLKQMTDKG 356
Score = 92.0 bits (227), Expect = 5e-17
Identities = 104/496 (20%), Positives = 197/496 (38%), Gaps = 72/496 (14%)
Query: 144 LKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEI 203
L+ +ER+++ + W+++D + N + S V R L D + +K
Sbjct: 119 LESCIKERRVEFVSWLYKDMVLCGIAPQTYTFNLLIRALCDSSCVDAARELFDEMPEKGC 178
Query: 204 RAKDWKFSRLMK---LSGLSFTEGQLMMIIEMLGV---KRCWKQALSVVQWVYNSKDHRK 257
+ ++ F L++ +GL+ +L+ +E GV K + +S + D K
Sbjct: 179 KPNEFTFGILVRGYCKAGLTDKGLELLNAMESFGVLPNKVIYNTIVSSFCREGRNDDSEK 238
Query: 258 FQSRF----------VYTKLLAVLGKARRPKEALQIFNMM-----LGNIRVYPDMAAYHS 302
+ + ++ L K + +A +IF+ M LG R P+ Y+
Sbjct: 239 MVEKMREEGLVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLGLPR--PNSITYNL 296
Query: 303 IAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNW---------------------DPT 341
+ + GLL++ + E +R+ + W D
Sbjct: 297 MLKGFCKVGLLEDAKTLFESIRENDDLASLQSYNIWLQGLVRHGKFIEAETVLKQMTDKG 356
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+ P + YN +++ + M+++ + P+ TYG + G D
Sbjct: 357 IGPSIYSYNILMDGLCKLGMLSDAKTIVGLMKRNGVCPDAVTYGCLLHGYCSVGKVDAAK 416
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
L ++M RN +P A T +++ + WK G++ EA + +R M +G Y L
Sbjct: 417 SLLQEMMRNNCLPNAYTCNILLHSLWKMGRISEAEELLRKMNEKG--------YGLDTVT 468
Query: 462 CN------CGRWQ-DATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ 514
CN CG + D +E+ K R+ + L I G +DD + ++
Sbjct: 469 CNIIVDGLCGSGELDKAIEIVKGMRVHGSAALGNLGNSYI------GLVDDSL-----IE 517
Query: 515 DHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQW 574
++C P++ T +T+L + F+ AK LF E+ K L+PD+ YN+ + + +
Sbjct: 518 NNCLPDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEK--LQPDSVAYNIFIHHFCKQGKI 575
Query: 575 EYFEHVYKEMILSGYH 590
V K+M G H
Sbjct: 576 SSAFRVLKDMEKKGCH 591
Score = 67.8 bits (164), Expect = 1e-09
Identities = 73/337 (21%), Positives = 134/337 (39%), Gaps = 56/337 (16%)
Query: 260 SRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNI 319
+ F + L+ KA + L++ N M + V P+ Y++I + + G + +
Sbjct: 181 NEFTFGILVRGYCKAGLTDKGLELLNAM-ESFGVLPNKVIYNTIVSSFCREGRNDDSEKM 239
Query: 320 VECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL-- 377
VE MR++ L PD+V +N+ ++A + S +F M
Sbjct: 240 VEKMREEG--------------LVPDIVTFNSRISALCKEGKVLDASRIFSDMELDEYLG 285
Query: 378 --KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEA 435
+PN TY L ++ + G + LFE ++ N ++ +Y + ++ + GK EA
Sbjct: 286 LPRPNSITYNLMLKGFCKVGLLEDAKTLFESIRENDDLASLQSYNIWLQGLVRHGKFIEA 345
Query: 436 VKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIR 495
++ M +G+ + Y L LC G DA V +KR
Sbjct: 346 ETVLKQMTDKGIGPSIYSYNILMDGLCKLGMLSDAKTIVGLMKR---------------- 389
Query: 496 SSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
+ +C P+ T +L Y AK L +E+ +++
Sbjct: 390 ---------NGVC----------PDAVTYGCLLHGYCSVGKVDAAKSLLQEMM--RNNCL 428
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
P+AYT N++L + + + E + ++M GY LD
Sbjct: 429 PNAYTCNILLHSLWKMGRISEAEELLRKMNEKGYGLD 465
Score = 57.8 bits (138), Expect = 1e-06
Identities = 39/182 (21%), Positives = 85/182 (46%), Gaps = 2/182 (1%)
Query: 365 VSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVR 424
+ V + +++ P+ TY + + ++G + LF +M P+++ Y + +
Sbjct: 508 IGLVDDSLIENNCLPDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEKLQPDSVAYNIFIH 567
Query: 425 TFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAK 484
F K+GK+ A + ++DME++G + Y L L + + ++++K
Sbjct: 568 HFCKQGKISSAFRVLKDMEKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKE-KGIS 626
Query: 485 PLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVL 543
P T+ I+ +G ++D + E MQ + APNV + +++ + + F A+ +
Sbjct: 627 PNICTYNTAIQYLCEGEKVEDATNLLDEMMQKNIAPNVFSFKYLIEAFCKVPDFDMAQEV 686
Query: 544 FE 545
FE
Sbjct: 687 FE 688
Score = 52.8 bits (125), Expect = 3e-05
Identities = 48/264 (18%), Positives = 102/264 (38%), Gaps = 3/264 (1%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
PD++ Y+ +LN + ++ +F +M L+P+ Y + + + G +
Sbjct: 522 PDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEKLQPDSVAYNIFIHHFCKQGKISSAFRV 581
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
+ M++ G TY ++ + ++ E + +M+ +G+ Y LC
Sbjct: 582 LKDMEKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKEKGISPNICTYNTAIQYLCE 641
Query: 464 CGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGT 523
+ +DAT ++++ + + P +F +I + D +FE C G
Sbjct: 642 GEKVEDATNLLDEMMQ-KNIAPNVFSFKYLIEAFCKVPDFDMAQEVFETAVSICGQKEGL 700
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
+ M A L E V +L + Y ++E+ + + E + +
Sbjct: 701 YSLMFNELLAAGQLLKATELLEAVLDRGFEL--GTFLYKDLVESLCKKDELEVASGILHK 758
Query: 584 MILSGYHLDQNKHLPLLVKASRAG 607
MI GY D +P++ + G
Sbjct: 759 MIDRGYGFDPAALMPVIDGLGKMG 782
Score = 42.7 bits (99), Expect = 0.033
Identities = 46/220 (20%), Positives = 90/220 (40%), Gaps = 23/220 (10%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ LL L KA R EA +F M+G ++ PD AY+ + G + +++ M
Sbjct: 527 YSTLLNGLCKAGRFAEAKNLFAEMMGE-KLQPDSVAYNIFIHHFCKQGKISSAFRVLKDM 585
Query: 324 RQKP-----ETLKYMYR----KNW------------DPTLEPDVVIYNAVLNACVPSKQW 362
+K ET + KN + + P++ YN + ++
Sbjct: 586 EKKGCHKSLETYNSLILGLGIKNQIFEIHGLMDEMKEKGISPNICTYNTAIQYLCEGEKV 645
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ + + +M + ++ PN ++ +E + ++D+ E+FE E L Y +M
Sbjct: 646 EDATNLLDEMMQKNIAPNVFSFKYLIEAFCKVPDFDMAQEVFETAVSICGQKEGL-YSLM 704
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
G++ +A + + + RG +Y +L LC
Sbjct: 705 FNELLAAGQLLKATELLEAVLDRGFELGTFLYKDLVESLC 744
Score = 40.8 bits (94), Expect = 0.13
Identities = 34/155 (21%), Positives = 67/155 (42%), Gaps = 18/155 (11%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNI-VECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVL 353
PD+ Y ++ L +AG E N+ E M +K L+PD V YN +
Sbjct: 522 PDLITYSTLLNGLCKAGRFAEAKNLFAEMMGEK---------------LQPDSVAYNIFI 566
Query: 354 NACVPSKQWKGVSWVFQQMRKSSLKPNGATY-GLAMEVMLQSGNYDLVHELFEKMQRNGE 412
+ + V + M K + TY L + + +++ ++ +H L ++M+ G
Sbjct: 567 HHFCKQGKISSAFRVLKDMEKKGCHKSLETYNSLILGLGIKNQIFE-IHGLMDEMKEKGI 625
Query: 413 VPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGV 447
P TY ++ + KV++A + +M ++ +
Sbjct: 626 SPNICTYNTAIQYLCEGEKVEDATNLLDEMMQKNI 660
>UniRef100_Q76C99 Rf1 [Oryza sativa]
Length = 791
Score = 104 bits (260), Expect = 7e-21
Identities = 103/412 (25%), Positives = 170/412 (41%), Gaps = 57/412 (13%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y ++A L KA+ +A+++ N M+ N V PD Y+SI +G KE + ++ M
Sbjct: 234 YNSIIAALCKAQAMDKAMEVLNTMVKN-GVMPDCMTYNSILHGYCSSGQPKEAIGFLKKM 292
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
R +EPDVV Y+ +++ + + +F M K LKP T
Sbjct: 293 RSDG--------------VEPDVVTYSLLMDYLCKNGRCMEARKIFDSMTKRGLKPEITT 338
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ G +H L + M RNG P+ + +++ + K+GKVD+A+ M
Sbjct: 339 YGTLLQGYATKGALVEMHGLLDLMVRNGIHPDHYVFSILICAYAKQGKVDQAMLVFSKMR 398
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKI----------------------KRLP 481
++G+ A Y + LC GR +DA L E++ +
Sbjct: 399 QQGLNPNAVTYGAVIGILCKSGRVEDAMLYFEQMIDEGLSPGNIVYNSLIHGLCTCNKWE 458
Query: 482 HAKPL------------EVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTML 528
A+ L + F +I S G + + +FE M + PNV T NT++
Sbjct: 459 RAEELILEMLDRGICLNTIFFNSIIDSHCKEGRVIESEKLFELMVRIGVKPNVITYNTLI 518
Query: 529 KVYS-QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILS 587
Y M K+L V V L+P+ TY+ ++ + + E ++KEM S
Sbjct: 519 NGYCLAGKMDEAMKLLSGMVSVG---LKPNTVTYSTLINGYCKISRMEDALVLFKEMESS 575
Query: 588 GYHLD---QNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYKNC 636
G D N L L + R + V S +S ++II + C
Sbjct: 576 GVSPDIITYNIILQGLFQTRRTAAAKELYVRITESGTQIELSTYNIILHGLC 627
Score = 75.9 bits (185), Expect = 4e-12
Identities = 79/337 (23%), Positives = 136/337 (39%), Gaps = 26/337 (7%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIR--VYPDMAAYHSIAVTLGQAGLLKELLNI 319
F Y LL L R +EAL++ +MM + PD+ +Y ++ G KE
Sbjct: 159 FSYNILLKGLCDENRSQEALELLHMMADDRGGGSPPDVVSYTTVI-----NGFFKE---- 209
Query: 320 VECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKP 379
+ Y + D + PDVV YN+++ A ++ V M K+ + P
Sbjct: 210 -----GDSDKAYSTYHEMLDRGILPDVVTYNSIIAALCKAQAMDKAMEVLNTMVKNGVMP 264
Query: 380 NGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAV 439
+ TY + SG +KM+ +G P+ +TY +++ K G+ EA K
Sbjct: 265 DCMTYNSILHGYCSSGQPKEAIGFLKKMRSDGVEPDVVTYSLLMDYLCKNGRCMEARKIF 324
Query: 440 RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL---PHAKPLEVTFTGMIRS 496
M +RG+ + Y L G A +E+ + L P F+ +I +
Sbjct: 325 DSMTKRGLKPEITTYGTLLQGYATKG----ALVEMHGLLDLMVRNGIHPDHYVFSILICA 380
Query: 497 SMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
G +D + +F M Q PN T ++ + ++ A + FE+ + L
Sbjct: 381 YAKQGKVDQAMLVFSKMRQQGLNPNAVTYGAVIGILCKSGRVEDAMLYFEQ--MIDEGLS 438
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
P YN ++ ++WE E + EM+ G L+
Sbjct: 439 PGNIVYNSLIHGLCTCNKWERAEELILEMLDRGICLN 475
Score = 56.6 bits (135), Expect = 2e-06
Identities = 63/319 (19%), Positives = 134/319 (41%), Gaps = 22/319 (6%)
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY 332
K R E+ ++F +M+ I V P++ Y+++ AG + E + ++ M
Sbjct: 488 KEGRVIESEKLFELMV-RIGVKPNVITYNTLINGYCLAGKMDEAMKLLSGMVSVG----- 541
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVML 392
L+P+ V Y+ ++N + + +F++M S + P+ TY + ++ +
Sbjct: 542 ---------LKPNTVTYSTLINGYCKISRMEDALVLFKEMESSGVSPDIITYNIILQGLF 592
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
Q+ EL+ ++ +G E TY +++ K D+A++ +++ + A
Sbjct: 593 QTRRTAAAKELYVRITESGTQIELSTYNIILHGLCKNKLTDDALQMFQNLCLMDLKLEAR 652
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY 512
+ + L GR D ++ P T+ M + + G +++ +F
Sbjct: 653 TFNIMIDALLKVGR-NDEAKDLFVAFSSNGLVPNYWTYRLMAENIIGQGLLEELDQLFLS 711
Query: 513 MQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRG 571
M+D+ C + G +N +++ Q + A + L +A T +L ++ S G
Sbjct: 712 MEDNGCTVDSGMLNFIVRELLQRGEITRAGTYLSMIDEKHFSL--EASTASLFIDLLSGG 769
Query: 572 HQWEYFEHV---YKEMILS 587
EY+ + YK I S
Sbjct: 770 KYQEYYRFLPEKYKSFIES 788
Score = 53.5 bits (127), Expect = 2e-05
Identities = 43/183 (23%), Positives = 77/183 (41%), Gaps = 15/183 (8%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ L+ K R ++AL +F M + V PD+ Y+ I L Q
Sbjct: 549 YSTLINGYCKISRMEDALVLFKEMESS-GVSPDIITYNIILQGLFQT------------- 594
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
++ K +Y + + + ++ YN +L+ +K +FQ + LK T
Sbjct: 595 -RRTAAAKELYVRITESGTQIELSTYNIILHGLCKNKLTDDALQMFQNLCLMDLKLEART 653
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
+ + ++ +L+ G D +LF NG VP TY++M +G ++E + ME
Sbjct: 654 FNIMIDALLKVGRNDEAKDLFVAFSSNGLVPNYWTYRLMAENIIGQGLLEELDQLFLSME 713
Query: 444 RRG 446
G
Sbjct: 714 DNG 716
>UniRef100_Q8L8A4 Fertility restorer [Petunia hybrida]
Length = 592
Score = 103 bits (258), Expect = 1e-20
Identities = 76/325 (23%), Positives = 151/325 (46%), Gaps = 26/325 (8%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ LGK + ++ +F M+ ++ +YPD+ ++S+ L + G +++ I+ M
Sbjct: 251 YTSLIDGLGKLSQWEKVRTLFLEMI-HLNIYPDVCTFNSVIDGLCKEGKVEDAEEIMTYM 309
Query: 324 RQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
+K P + Y ++ D +EPD++ Y A++N V K+
Sbjct: 310 IEKGVEPNEITYNVVMDGYCLRGQMGRARRIFDSMIDKGIEPDIISYTALINGYVEKKKM 369
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+F+++ ++ LKP+ T + + + + G + F++MQ G +P T+ +
Sbjct: 370 DKAMQLFREISQNGLKPSIVTCSVLLRGLFEVGRTECAKIFFDEMQAAGHIPNLYTHCTL 429
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH 482
+ ++K G V+EA+ +ERR +Y + LC G+ A EK+ L
Sbjct: 430 LGGYFKNGLVEEAMSHFHKLERRREDTNIQIYTAVINGLCKNGKLDKAHATFEKLP-LIG 488
Query: 483 AKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAK 541
P +T+T MI G +D+ + M+D+ C P+ T N +++ + ++ S K
Sbjct: 489 LHPDVITYTAMISGYCQEGLLDEAKDMLRKMEDNGCLPDNRTYNVIVRGFFRSSKVSEMK 548
Query: 542 VLFEEVKVAKSDLRPDAYTYNLMLE 566
+E +A +A T L+++
Sbjct: 549 AFLKE--IAGKSFSFEAATVELLMD 571
Score = 73.6 bits (179), Expect = 2e-11
Identities = 68/328 (20%), Positives = 143/328 (42%), Gaps = 26/328 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T L+ L + K+A+ +F ++ PD Y ++ L + G ++ +++ M
Sbjct: 145 FTTLIRGLFAENKVKDAVHLFKKLVRENICEPDEVMYGTVMDGLCKKGHTQKAFDLLRLM 204
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
Q +PD IYN V++A G + + +M++ ++ P+ T
Sbjct: 205 EQG--------------ITKPDTCIYNIVIDAFCKDGMLDGATSLLNEMKQKNIPPDIIT 250
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ + + ++ V LF +M P+ T+ ++ KEGKV++A + + M
Sbjct: 251 YTSLIDGLGKLSQWEKVRTLFLEMIHLNIYPDVCTFNSVIDGLCKEGKVEDAEEIMTYMI 310
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
+GV Y + C G+ A + + +P +++T +I ++ +
Sbjct: 311 EKGVEPNEITYNVVMDGYCLRGQMGRARRIFDSMID-KGIEPDIISYTALINGYVEKKKM 369
Query: 504 DDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
D + +F E Q+ P++ T + +L+ + AK+ F+E++ A P+ YT+
Sbjct: 370 DKAMQLFREISQNGLKPSIVTCSVLLRGLFEVGRTECAKIFFDEMQAAGH--IPNLYTHC 427
Query: 563 LMLEASSRGHQWEYFEHVYKEMILSGYH 590
+L YF++ E +S +H
Sbjct: 428 TLLGG--------YFKNGLVEEAMSHFH 447
Score = 42.7 bits (99), Expect = 0.033
Identities = 49/253 (19%), Positives = 98/253 (38%), Gaps = 44/253 (17%)
Query: 398 DLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
D LF +M +P A+++ +++ V R++ + + A +
Sbjct: 54 DDAFSLFRQMVTTKPLPSAVSFSKLLKALVHMKHYSSVVSIFREIHKLRIPVDAFALSTV 113
Query: 458 --ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-- 513
+CCL + + L + K +P+ EVTFT +IR + D + +F+ +
Sbjct: 114 VNSCCLMHRTDLGFSVLAIHFKKGIPYN---EVTFTTLIRGLFAENKVKDAVHLFKKLVR 170
Query: 514 QDHCAPN---VGTV--------------------------------NTMLKVYSQNDMFS 538
++ C P+ GTV N ++ + ++ M
Sbjct: 171 ENICEPDEVMYGTVMDGLCKKGHTQKAFDLLRLMEQGITKPDTCIYNIVIDAFCKDGMLD 230
Query: 539 TAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLP 598
A L E+K + ++ PD TY +++ + QWE ++ EMI + D
Sbjct: 231 GATSLLNEMK--QKNIPPDIITYTSLIDGLGKLSQWEKVRTLFLEMIHLNIYPDVCTFNS 288
Query: 599 LLVKASRAGKVDN 611
++ + GKV++
Sbjct: 289 VIDGLCKEGKVED 301
>UniRef100_Q9FLL3 Salt-inducible protein-like [Arabidopsis thaliana]
Length = 527
Score = 102 bits (254), Expect = 4e-20
Identities = 86/358 (24%), Positives = 160/358 (44%), Gaps = 24/358 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T L+ R +EA+ + N M+ + + PD+ Y +I +L + G + L++ + M
Sbjct: 145 FTSLINGFCLGNRMEEAMSMVNQMV-EMGIKPDVVMYTTIIDSLCKNGHVNYALSLFDQM 203
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+N+ + PDVV+Y +++N S +W+ + + M K +KP+ T
Sbjct: 204 ------------ENYG--IRPDVVMYTSLVNGLCNSGRWRDADSLLRGMTKRKIKPDVIT 249
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
+ ++ ++ G + EL+ +M R P TY ++ F EG VDEA + ME
Sbjct: 250 FNALIDAFVKEGKFLDAEELYNEMIRMSIAPNIFTYTSLINGFCMEGCVDEARQMFYLME 309
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPL---EVTFTGMIRSSMDG 500
+G Y L C C + DA KI K L +T+T +I+
Sbjct: 310 TKGCFPDVVAYTSLINGFCKCKKVDDAM----KIFYEMSQKGLTGNTITYTTLIQGFGQV 365
Query: 501 GHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSD-LRPDA 558
G + +F +M PN+ T N +L N A ++FE+++ + D + P+
Sbjct: 366 GKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQKREMDGVAPNI 425
Query: 559 YTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
+TYN++L + E V+++M + + ++ +AGKV N + ++
Sbjct: 426 WTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIITYTIIIQGMCKAGKVKNAVNLF 483
Score = 74.7 bits (182), Expect = 8e-12
Identities = 67/289 (23%), Positives = 122/289 (42%), Gaps = 20/289 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ L+ K + +A +++N M+ + + P++ Y S+ G + E + M
Sbjct: 250 FNALIDAFVKEGKFLDAEELYNEMI-RMSIAPNIFTYTSLINGFCMEGCVDEARQMFYLM 308
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
K PDVV Y +++N K+ +F +M + L N T
Sbjct: 309 ETKG--------------CFPDVVAYTSLINGFCKCKKVDDAMKIFYEMSQKGLTGNTIT 354
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ Q G ++ E+F M G P TY V++ GKV +A+ DM+
Sbjct: 355 YTTLIQGFGQVGKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQ 414
Query: 444 RRGVMGTAS---VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDG 500
+R + G A Y L LC G+ + A + E +++ + +T+T +I+
Sbjct: 415 KREMDGVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGI-ITYTIIIQGMCKA 473
Query: 501 GHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVK 548
G + + + +F + PNV T TM+ + + A VLF ++K
Sbjct: 474 GKVKNAVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
Score = 72.8 bits (177), Expect = 3e-11
Identities = 75/365 (20%), Positives = 152/365 (41%), Gaps = 21/365 (5%)
Query: 254 DHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLL 313
+ R S +TKLL V+ K ++ F++++ M H +
Sbjct: 65 ESRPLPSIIDFTKLLNVIAKMKK-------FDVVINLCDHLQIMGVSHDLYTC------- 110
Query: 314 KELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA-CVPSKQWKGVSWVFQQM 372
LL C +P K EPD+V + +++N C+ ++ + +S V QM
Sbjct: 111 -NLLMNCFCQSSQPYLASSFLGKMMKLGFEPDIVTFTSLINGFCLGNRMEEAMSMV-NQM 168
Query: 373 RKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKV 432
+ +KP+ Y ++ + ++G+ + LF++M+ G P+ + Y +V G+
Sbjct: 169 VEMGIKPDVVMYTTIIDSLCKNGHVNYALSLFDQMENYGIRPDVVMYTSLVNGLCNSGRW 228
Query: 433 DEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTG 492
+A +R M +R + + L G++ DA ++ R+ A P T+T
Sbjct: 229 RDADSLLRGMTKRKIKPDVITFNALIDAFVKEGKFLDAEELYNEMIRMSIA-PNIFTYTS 287
Query: 493 MIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAK 551
+I G +D+ +F M+ C P+V +++ + + A +F E +++
Sbjct: 288 LINGFCMEGCVDEARQMFYLMETKGCFPDVVAYTSLINGFCKCKKVDDAMKIFYE--MSQ 345
Query: 552 SDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDN 611
L + TY +++ + + + V+ M+ G + + LL GKV
Sbjct: 346 KGLTGNTITYTTLIQGFGQVGKPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKK 405
Query: 612 ILVIY 616
L+I+
Sbjct: 406 ALMIF 410
Score = 61.2 bits (147), Expect = 9e-08
Identities = 46/184 (25%), Positives = 83/184 (45%), Gaps = 12/184 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ G+ +P A ++F+ M+ V P++ Y+ + L G +K+ L I E M
Sbjct: 355 YTTLIQGFGQVGKPNVAQEVFSHMVSR-GVPPNIRTYNVLLHCLCYNGKVKKALMIFEDM 413
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+++ + P++ YN +L+ + + + VF+ MRK + T
Sbjct: 414 QKREMD-----------GVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIIT 462
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + ++ M ++G LF + G P +TY M+ ++EG EA R M+
Sbjct: 463 YTIIIQGMCKAGKVKNAVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
Query: 444 RRGV 447
GV
Sbjct: 523 EDGV 526
Score = 51.2 bits (121), Expect = 9e-05
Identities = 47/224 (20%), Positives = 90/224 (39%), Gaps = 18/224 (8%)
Query: 258 FQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELL 317
F YT L+ K ++ +A++IF M + + Y ++ GQ G
Sbjct: 314 FPDVVAYTSLINGFCKCKKVDDAMKIFYEM-SQKGLTGNTITYTTLIQGFGQVG------ 366
Query: 318 NIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
KP + ++ + P++ YN +L+ + + K +F+ M+K +
Sbjct: 367 --------KPNVAQEVFSHMVSRGVPPNIRTYNVLLHCLCYNGKVKKALMIFEDMQKREM 418
Query: 378 K---PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDE 434
PN TY + + + +G + +FE M++ +TY ++++ K GKV
Sbjct: 419 DGVAPNIWTYNVLLHGLCYNGKLEKALMVFEDMRKREMDIGIITYTIIIQGMCKAGKVKN 478
Query: 435 AVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
AV + +GV Y + L G +A + K+K
Sbjct: 479 AVNLFCSLPSKGVKPNVVTYTTMISGLFREGLKHEAHVLFRKMK 522
>UniRef100_Q8W0G9 P0529E05.10 protein [Oryza sativa]
Length = 703
Score = 101 bits (252), Expect = 6e-20
Identities = 70/282 (24%), Positives = 137/282 (47%), Gaps = 31/282 (10%)
Query: 292 RVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNA 351
+V PD Y ++ L + L L++++ M + ++PDVV YNA
Sbjct: 188 QVAPDRITYSTLMCGLAKQDRLDHALDLLDEMPRS--------------RVQPDVVCYNA 233
Query: 352 VLNACVPSKQWKGVSWVFQQMRKS-SLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRN 410
+L C + +++ V V+ ++ K +PN ATY + ++ + + G + V E++E+M N
Sbjct: 234 LLGGCFKAGEFEKVMRVWDKLVKDPGARPNLATYNVMLDGLCKFGRFKEVGEVWERMVAN 293
Query: 411 GEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA 470
P+ +TY +++ + G VD A + ++ + G++ A++Y L C GR Q+A
Sbjct: 294 NLQPDVITYGILIHGLCRSGDVDGAARVYSEIIKTGLVIDAAMYNSLVKGFCQAGRVQEA 353
Query: 471 -----TLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH--CAPNVGT 523
+ ++ L T+ MI+ D G +D+ I +++ ++ C P+ T
Sbjct: 354 WKFWDSAGFAGLRNLR-------TYNIMIKGLFDSGMVDEAIELWDLLEKDVACIPDTVT 406
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLML 565
T++ QN + A +FEE +V+ L D ++Y+ M+
Sbjct: 407 FGTLIHGLCQNGFANKAFTIFEEARVSGKQL--DVFSYSSMI 446
Score = 95.1 bits (235), Expect = 6e-18
Identities = 94/430 (21%), Positives = 191/430 (43%), Gaps = 62/430 (14%)
Query: 261 RFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIV 320
R Y+ L+ L K R AL + + M + RV PD+ Y+++ +AG ++++ +
Sbjct: 193 RITYSTLMCGLAKQDRLDHALDLLDEMPRS-RVQPDVVCYNALLGGCFKAGEFEKVMRVW 251
Query: 321 ECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPN 380
+ + + DP P++ YN +L+ ++K V V+++M ++L+P+
Sbjct: 252 DKLVK-------------DPGARPNLATYNVMLDGLCKFGRFKEVGEVWERMVANNLQPD 298
Query: 381 GATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK--- 437
TYG+ + + +SG+ D ++ ++ + G V +A Y +V+ F + G+V EA K
Sbjct: 299 VITYGILIHGLCRSGDVDGAARVYSEIIKTGLVIDAAMYNSLVKGFCQAGRVQEAWKFWD 358
Query: 438 ---------------AVRDMERRGVMGTASVYYEL-----ACC------------LCNCG 465
++ + G++ A ++L AC LC G
Sbjct: 359 SAGFAGLRNLRTYNIMIKGLFDSGMVDEAIELWDLLEKDVACIPDTVTFGTLIHGLCQNG 418
Query: 466 RWQDATLEVEKIKRLPHAKPLEV-TFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGT 523
A E+ + K L+V +++ MI + G + D + ++E M +D C PN
Sbjct: 419 FANKAFTIFEEAR--VSGKQLDVFSYSSMINGLCNVGRLVDAVKVYEKMDKDGCKPNSHI 476
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
N ++ + Q ++ T+ + K+A + P TYN +++ + +++ V +E
Sbjct: 477 YNALISGFCQ--VYRTSDAVRIYSKMADNGCSPTVITYNTLIDGLCKAEKYQEASSVARE 534
Query: 584 MILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQCLFY-----ISAHSIISYKNCYI 638
M+ +G+ D + L+ K+D+ L I+ Q L+ + H+I+ + C
Sbjct: 535 MVENGFTPDITTYGSLIRGLFSDKKIDDALSIW--KQILYKGLKVDVMMHNILIHGLCSA 592
Query: 639 QTMLTRVHTF 648
+ +H F
Sbjct: 593 GKVDEALHVF 602
Score = 73.2 bits (178), Expect = 2e-11
Identities = 59/311 (18%), Positives = 128/311 (40%), Gaps = 24/311 (7%)
Query: 279 EALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY------ 332
EA+++++++ ++ PD + ++ L Q G + I E R + L
Sbjct: 386 EAIELWDLLEKDVACIPDTVTFGTLIHGLCQNGFANKAFTIFEEARVSGKQLDVFSYSSM 445
Query: 333 ---------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
+Y K +P+ IYNA+++ + ++ +M +
Sbjct: 446 INGLCNVGRLVDAVKVYEKMDKDGCKPNSHIYNALISGFCQVYRTSDAVRIYSKMADNGC 505
Query: 378 KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK 437
P TY ++ + ++ Y + +M NG P+ TY ++R + + K+D+A+
Sbjct: 506 SPTVITYNTLIDGLCKAEKYQEASSVAREMVENGFTPDITTYGSLIRGLFSDKKIDDALS 565
Query: 438 AVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSS 497
+ + +G+ ++ L LC+ G+ +A +K + P VT+ ++
Sbjct: 566 IWKQILYKGLKVDVMMHNILIHGLCSAGKVDEALHVFSDMKEKKNCPPNLVTYNTLMDGL 625
Query: 498 MDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRP 556
+ G+ID ++ +D P++ + NT +K D L +E V + P
Sbjct: 626 YETGYIDKAATLWTSITEDGLEPDIISYNTRIKGLCSCDRIHEGIQLLDE--VLSRGIIP 683
Query: 557 DAYTYNLMLEA 567
T+N+++ A
Sbjct: 684 TVITWNILVRA 694
Score = 65.9 bits (159), Expect = 4e-09
Identities = 55/270 (20%), Positives = 109/270 (40%), Gaps = 40/270 (14%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSS----LKPNGATYGLAMEVMLQSGNYDL 399
P + +NA+L+A V ++++ F + + + PN TY + + + G+ D
Sbjct: 117 PGIRSHNALLDAFVRARRFSDADAFFASLSHGAFGRRIAPNLQTYNIVLRSLCARGDLDR 176
Query: 400 VHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELAC 459
LF+ ++R P+ +TY ++ K+ ++D A+ + +M R V Y L
Sbjct: 177 AVTLFDSLRRRQVAPDRITYSTLMCGLAKQDRLDHALDLLDEMPRSRVQPDVVCYNALLG 236
Query: 460 CLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAP 519
G ++ +K+ + P A+ P
Sbjct: 237 GCFKAGEFEKVMRVWDKLVKDPGAR----------------------------------P 262
Query: 520 NVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEH 579
N+ T N ML + F ++E ++ ++L+PD TY +++ R +
Sbjct: 263 NLATYNVMLDGLCKFGRFKEVGEVWE--RMVANNLQPDVITYGILIHGLCRSGDVDGAAR 320
Query: 580 VYKEMILSGYHLDQNKHLPLLVKASRAGKV 609
VY E+I +G +D + L+ +AG+V
Sbjct: 321 VYSEIIKTGLVIDAAMYNSLVKGFCQAGRV 350
Score = 48.5 bits (114), Expect = 6e-04
Identities = 48/230 (20%), Positives = 101/230 (43%), Gaps = 12/230 (5%)
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+ P++ YN VL + +F +R+ + P+ TY M + + D
Sbjct: 154 IAPNLQTYNIVLRSLCARGDLDRAVTLFDSLRRRQVAPDRITYSTLMCGLAKQDRLDHAL 213
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAV----KAVRDMERRGVMGTASVYYEL 457
+L ++M R+ P+ + Y ++ +K G+ ++ + K V+D R + T +V +
Sbjct: 214 DLLDEMPRSRVQPDVVCYNALLGGCFKAGEFEKVMRVWDKLVKDPGARPNLATYNVMLD- 272
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDH 516
LC GR+++ EV + + +P +T+ +I G +D ++ E ++
Sbjct: 273 --GLCKFGRFKEVG-EVWERMVANNLQPDVITYGILIHGLCRSGDVDGAARVYSEIIKTG 329
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLE 566
+ N+++K + Q A ++ A LR + TYN+M++
Sbjct: 330 LVIDAAMYNSLVKGFCQAGRVQEAWKFWDSAGFA--GLR-NLRTYNIMIK 376
Score = 37.7 bits (86), Expect = 1.1
Identities = 37/173 (21%), Positives = 80/173 (45%), Gaps = 22/173 (12%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMML-GNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
Y L+ L ++ +AL I+ +L ++V D+ ++ + L AG + E L++
Sbjct: 547 YGSLIRGLFSDKKIDDALSIWKQILYKGLKV--DVMMHNILIHGLCSAGKVDEALHVFSD 604
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M++K KN P L V YN +++ + + ++ + + L+P+
Sbjct: 605 MKEK---------KNCPPNL----VTYNTLMDGLYETGYIDKAATLWTSITEDGLEPDII 651
Query: 383 TYGLAMEVMLQSGNYDLVHE---LFEKMQRNGEVPEALTYKVMVRTFWKEGKV 432
+Y ++ + + D +HE L +++ G +P +T+ ++VR K G +
Sbjct: 652 SYNTRIKGLC---SCDRIHEGIQLLDEVLSRGIIPTVITWNILVRAVIKYGPI 701
>UniRef100_Q6K7A0 Putative PPR protein [Oryza sativa]
Length = 649
Score = 101 bits (251), Expect = 8e-20
Identities = 67/293 (22%), Positives = 136/293 (45%), Gaps = 29/293 (9%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLN 354
PD+ Y ++ + G + + L+++ M KP T V YNA L
Sbjct: 258 PDVVIYSTLINGFSEQGHVDQALDLLNTMLCKPNT-----------------VCYNAALK 300
Query: 355 ACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
+++W+ + + +M + PN AT+ + + + Q+ D E+ E+M++ G P
Sbjct: 301 GLCIAERWEDIGELMAEMVRKGCSPNEATFSMLISSLCQNNLVDSAVEVLEQMEKYGCEP 360
Query: 415 EALTYKVMVRTFWKEGKVDEAVKAVRDME-RRGVMGTASVYYELACCLCNCGRWQDATLE 473
+ + Y +++ + + G+VD+A++ + M + +G +V C RW DA+
Sbjct: 361 DTVNYNIIINSLSERGRVDDALRLLNSMVCKPDALGFNAVLKG----FCRAERWHDASEL 416
Query: 474 VEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYS 532
+ ++ R +E+TF +I G ++ +FE M + C P++ T +++L +S
Sbjct: 417 IAQMFR-DDCPLIEMTFNILIDMLCQNGLVNYATQVFEQMPRYRCTPDIVTYSSLLNGFS 475
Query: 533 QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMI 585
+ + A LF + +PD ++YN +L+ R +WE + EM+
Sbjct: 476 EQGLVEVAIQLFRSM-----PCKPDIFSYNAVLKGLCRAARWEDAGELIAEMV 523
Score = 96.7 bits (239), Expect = 2e-18
Identities = 73/293 (24%), Positives = 133/293 (44%), Gaps = 29/293 (9%)
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLN 354
PD Y+ I +L + G + + L ++ M KP+ L + NAVL
Sbjct: 360 PDTVNYNIIINSLSERGRVDDALRLLNSMVCKPDALGF-----------------NAVLK 402
Query: 355 ACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
+++W S + QM + T+ + ++++ Q+G + ++FE+M R P
Sbjct: 403 GFCRAERWHDASELIAQMFRDDCPLIEMTFNILIDMLCQNGLVNYATQVFEQMPRYRCTP 462
Query: 415 EALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA-TLE 473
+ +TY ++ F ++G V+ A++ R M + + + Y + LC RW+DA L
Sbjct: 463 DIVTYSSLLNGFSEQGLVEVAIQLFRSMPCKPDIFS---YNAVLKGLCRAARWEDAGELI 519
Query: 474 VEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYS 532
E + + P EVTF +I S G +D I + E M ++ P++ T N ++ +S
Sbjct: 520 AEMVGK--DCPPNEVTFNILINSLCQKGLVDRAIEVLEQMPNYGSTPDIFTYNALINGFS 577
Query: 533 QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMI 585
+ A L + +PDA +YN L+ R +W+ E + EM+
Sbjct: 578 EQGRLDDALKLLSTMSC-----KPDAISYNSTLKGLCRAERWQDAEELVAEML 625
Score = 86.3 bits (212), Expect = 3e-15
Identities = 60/244 (24%), Positives = 107/244 (43%), Gaps = 14/244 (5%)
Query: 344 PDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
PD YN VL +K+W+ + +M ++ PN T+ + Q+G D +L
Sbjct: 188 PDTYTYNTVLKGLCIAKKWEEAEELMAEMIRNRCPPNEVTFATQIRSFCQNGLLDRAVQL 247
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELAC-CLC 462
++M R G P+ + Y ++ F ++G VD+A+ + M + +V Y A LC
Sbjct: 248 LDQMPRYGCTPDVVIYSTLINGFSEQGHVDQALDLLNTM----LCKPNTVCYNAALKGLC 303
Query: 463 NCGRWQD-ATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPN 520
RW+D L E +++ P E TF+ +I S +D + + E M+ + C P+
Sbjct: 304 IAERWEDIGELMAEMVRK--GCSPNEATFSMLISSLCQNNLVDSAVEVLEQMEKYGCEPD 361
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHV 580
N ++ S+ A L + +PDA +N +L+ R +W +
Sbjct: 362 TVNYNIIINSLSERGRVDDALRLLNSMV-----CKPDALGFNAVLKGFCRAERWHDASEL 416
Query: 581 YKEM 584
+M
Sbjct: 417 IAQM 420
Score = 80.9 bits (198), Expect = 1e-13
Identities = 57/211 (27%), Positives = 90/211 (42%), Gaps = 22/211 (10%)
Query: 280 ALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWD 339
A Q+F M R PD+ Y S+ + GL++ + + M KP
Sbjct: 448 ATQVFEQM-PRYRCTPDIVTYSSLLNGFSEQGLVEVAIQLFRSMPCKP------------ 494
Query: 340 PTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDL 399
D+ YNAVL + +W+ + +M PN T+ + + + Q G D
Sbjct: 495 -----DIFSYNAVLKGLCRAARWEDAGELIAEMVGKDCPPNEVTFNILINSLCQKGLVDR 549
Query: 400 VHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELAC 459
E+ E+M G P+ TY ++ F ++G++D+A+K + M + A Y
Sbjct: 550 AIEVLEQMPNYGSTPDIFTYNALINGFSEQGRLDDALKLLSTMSCK---PDAISYNSTLK 606
Query: 460 CLCNCGRWQDATLEVEKIKRLPHAKPLEVTF 490
LC RWQDA V ++ R P EVTF
Sbjct: 607 GLCRAERWQDAEELVAEMLR-NKCTPNEVTF 636
Score = 56.2 bits (134), Expect = 3e-06
Identities = 42/174 (24%), Positives = 76/174 (43%), Gaps = 12/174 (6%)
Query: 417 LTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDAT-LEVE 475
+TY ++ + + G++D+A++ + M V Y + LC +W++A L E
Sbjct: 159 VTYTALIDGYCRSGRLDDALRLIASMP---VAPDTYTYNTVLKGLCIAKKWEEAEELMAE 215
Query: 476 KIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQN 534
I+ P EVTF IRS G +D + + + M + C P+V +T++ +S+
Sbjct: 216 MIRN--RCPPNEVTFATQIRSFCQNGLLDRAVQLLDQMPRYGCTPDVVIYSTLINGFSEQ 273
Query: 535 DMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
A L + +P+ YN L+ +WE + EM+ G
Sbjct: 274 GHVDQALDLLNTMLC-----KPNTVCYNAALKGLCIAERWEDIGELMAEMVRKG 322
Score = 37.7 bits (86), Expect = 1.1
Identities = 32/125 (25%), Positives = 50/125 (39%), Gaps = 7/125 (5%)
Query: 461 LCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPN 520
LC R DA ++ +K A V+ ++ G + D + + A N
Sbjct: 100 LCATRRLADAERVLDALKAAAAADA--VSHNTLVAGYCRDGRLADAERVLGAARATGAAN 157
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHV 580
V T ++ Y ++ A L + VA PD YTYN +L+ +WE E +
Sbjct: 158 VVTYTALIDGYCRSGRLDDALRLIASMPVA-----PDTYTYNTVLKGLCIAKKWEEAEEL 212
Query: 581 YKEMI 585
EMI
Sbjct: 213 MAEMI 217
Score = 35.8 bits (81), Expect = 4.1
Identities = 32/149 (21%), Positives = 64/149 (42%), Gaps = 19/149 (12%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE 321
F Y +L L +A R ++A ++ M+G P+ ++ + +L Q GL+ + ++E
Sbjct: 497 FSYNAVLKGLCRAARWEDAGELIAEMVGK-DCPPNEVTFNILINSLCQKGLVDRAIEVLE 555
Query: 322 CMRQ---KPETLKYMYRKNW-------DPTLE--------PDVVIYNAVLNACVPSKQWK 363
M P+ Y N D L+ PD + YN+ L +++W+
Sbjct: 556 QMPNYGSTPDIFTYNALINGFSEQGRLDDALKLLSTMSCKPDAISYNSTLKGLCRAERWQ 615
Query: 364 GVSWVFQQMRKSSLKPNGATYGLAMEVML 392
+ +M ++ PN T+ A +++
Sbjct: 616 DAEELVAEMLRNKCTPNEVTFKYANHLLM 644
>UniRef100_Q9SZ52 Hypothetical protein AT4g31850 [Arabidopsis thaliana]
Length = 1112
Score = 100 bits (248), Expect = 2e-19
Identities = 105/422 (24%), Positives = 182/422 (42%), Gaps = 45/422 (10%)
Query: 227 MMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQ---SRFVYTKLLAVLGKARRPKEALQI 283
+ I + L VK KQA Y + R+F + + Y L+ +L K+R EA+++
Sbjct: 157 LTIFKSLSVKGGLKQA------PYALRKMREFGFVLNAYSYNGLIHLLLKSRFCTEAMEV 210
Query: 284 FN-MMLGNIRVYPDMAAYHSIAVTLGQA-------GLLKELLNIVECMRQKPETLKY--- 332
+ M+L R P + Y S+ V LG+ GLLKE+ E + KP +
Sbjct: 211 YRRMILEGFR--PSLQTYSSLMVGLGKRRDIDSVMGLLKEM----ETLGLKPNVYTFTIC 264
Query: 333 ---------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSL 377
+ ++ D PDVV Y +++A +++ VF++M+
Sbjct: 265 IRVLGRAGKINEAYEILKRMDDEGCGPDVVTYTVLIDALCTARKLDCAKEVFEKMKTGRH 324
Query: 378 KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVK 437
KP+ TY ++ + + D V + + +M+++G VP+ +T+ ++V K G EA
Sbjct: 325 KPDRVTYITLLDRFSDNRDLDSVKQFWSEMEKDGHVPDVVTFTILVDALCKAGNFGEAFD 384
Query: 438 AVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSS 497
+ M +G++ Y L C L R DA LE+ KP T+ I
Sbjct: 385 TLDVMRDQGILPNLHTYNTLICGLLRVHRLDDA-LELFGNMESLGVKPTAYTYIVFIDYY 443
Query: 498 MDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRP 556
G + FE M+ APN+ N L ++ AK +F +K L P
Sbjct: 444 GKSGDSVSALETFEKMKTKGIAPNIVACNASLYSLAKAGRDREAKQIFYGLK--DIGLVP 501
Query: 557 DAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
D+ TYN+M++ S+ + + + EM+ +G D L+ +A +VD ++
Sbjct: 502 DSVTYNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDEAWKMF 561
Query: 617 IR 618
+R
Sbjct: 562 MR 563
Score = 83.6 bits (205), Expect = 2e-14
Identities = 70/329 (21%), Positives = 138/329 (41%), Gaps = 21/329 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L +A + A +F + + + PD+A Y+ + G++G + EL +
Sbjct: 788 YNLLIGGLLEADMIEIAQDVF-LQVKSTGCIPDVATYNFLLDAYGKSGKIDELFEL---- 842
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWK-GVSWVFQQMRKSSLKPNGA 382
Y++ E + + +N V++ V + + + M P
Sbjct: 843 ----------YKEMSTHECEANTITHNIVISGLVKAGNVDDALDLYYDLMSDRDFSPTAC 892
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TYG ++ + +SG +LFE M G P Y +++ F K G+ D A + M
Sbjct: 893 TYGPLIDGLSKSGRLYEAKQLFEGMLDYGCRPNCAIYNILINGFGKAGEADAACALFKRM 952
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGH 502
+ GV Y L CLC GR + +++K P V + +I
Sbjct: 953 VKEGVRPDLKTYSVLVDCLCMVGRVDEGLHYFKELKE-SGLNPDVVCYNLIINGLGKSHR 1011
Query: 503 IDDCICIFEYMQDH--CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYT 560
+++ + +F M+ P++ T N+++ M A ++ E++ ++ L P+ +T
Sbjct: 1012 LEEALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIYNEIQ--RAGLEPNVFT 1069
Query: 561 YNLMLEASSRGHQWEYFEHVYKEMILSGY 589
+N ++ S + E+ VY+ M+ G+
Sbjct: 1070 FNALIRGYSLSGKPEHAYAVYQTMVTGGF 1098
Score = 83.6 bits (205), Expect = 2e-14
Identities = 83/349 (23%), Positives = 146/349 (41%), Gaps = 26/349 (7%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLL---KELLN 318
+ +T + VLG+A + EA +I M + PD+ Y + L A L KE+
Sbjct: 259 YTFTICIRVLGRAGKINEAYEILKRM-DDEGCGPDVVTYTVLIDALCTARKLDCAKEVFE 317
Query: 319 IVECMRQKPETLKYM--------------YRKNWDPTLE----PDVVIYNAVLNACVPSK 360
++ R KP+ + Y+ ++ W + PDVV + +++A +
Sbjct: 318 KMKTGRHKPDRVTYITLLDRFSDNRDLDSVKQFWSEMEKDGHVPDVVTFTILVDALCKAG 377
Query: 361 QWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
+ MR + PN TY + +L+ D ELF M+ G P A TY
Sbjct: 378 NFGEAFDTLDVMRDQGILPNLHTYNTLICGLLRVHRLDDALELFGNMESLGVKPTAYTYI 437
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRL 480
V + + K G A++ M+ +G+ L GR ++A +K +
Sbjct: 438 VFIDYYGKSGDSVSALETFEKMKTKGIAPNIVACNASLYSLAKAGRDREAKQIFYGLKDI 497
Query: 481 PHAKPLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFST 539
P VT+ M++ G ID+ I + E M++ C P+V VN+++ + D
Sbjct: 498 GLV-PDSVTYNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDE 556
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
A +F +K K L+P TYN +L + + + +++ M+ G
Sbjct: 557 AWKMFMRMKEMK--LKPTVVTYNTLLAGLGKNGKIQEAIELFEGMVQKG 603
Score = 75.1 bits (183), Expect = 6e-12
Identities = 78/382 (20%), Positives = 153/382 (39%), Gaps = 52/382 (13%)
Query: 271 LGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETL 330
L KA R +EA QIF L +I + PD Y+ + + G + E + ++ M +
Sbjct: 478 LAKAGRDREAKQIF-YGLKDIGLVPDSVTYNMMMKCYSKVGEIDEAIKLLSEMMENG--- 533
Query: 331 KYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEV 390
EPDV++ N+++N + + +F +M++ LKP TY +
Sbjct: 534 -----------CEPDVIVVNSLINTLYKADRVDEAWKMFMRMKEMKLKPTVVTYNTLLAG 582
Query: 391 MLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGT 450
+ ++G ELFE M + G P +T+ + K +V A+K + M G +
Sbjct: 583 LGKNGKIQEAIELFEGMVQKGCPPNTITFNTLFDCLCKNDEVTLALKMLFKMMDMGCVPD 642
Query: 451 ASVYYELACCLCNCGRWQDATLEVEKIKRLPHA--------------------------- 483
Y + L G+ ++A ++K+L +
Sbjct: 643 VFTYNTIIFGLVKNGQVKEAMCFFHQMKKLVYPDFVTLCTLLPGVVKASLIEDAYKIITN 702
Query: 484 -------KPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH--CAPNVGTVNTMLKVYSQN 534
+P + + +I S + ID+ + E + + C + +++ ++
Sbjct: 703 FLYNCADQPANLFWEDLIGSILAEAGIDNAVSFSERLVANGICRDGDSILVPIIRYSCKH 762
Query: 535 DMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQN 594
+ S A+ LFE+ ++P TYNL++ E + V+ ++ +G D
Sbjct: 763 NNVSGARTLFEKF-TKDLGVQPKLPTYNLLIGGLLEADMIEIAQDVFLQVKSTGCIPDVA 821
Query: 595 KHLPLLVKASRAGKVDNILVIY 616
+ LL ++GK+D + +Y
Sbjct: 822 TYNFLLDAYGKSGKIDELFELY 843
Score = 74.7 bits (182), Expect = 8e-12
Identities = 69/296 (23%), Positives = 119/296 (39%), Gaps = 37/296 (12%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L + R +AL++F M ++ V P Y G++G L E M
Sbjct: 401 YNTLICGLLRVHRLDDALELFGNM-ESLGVKPTAYTYIVFIDYYGKSGDSVSALETFEKM 459
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ K + P++V NA L + + + + +F ++ L P+ T
Sbjct: 460 KTKG--------------IAPNIVACNASLYSLAKAGRDREAKQIFYGLKDIGLVPDSVT 505
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + M+ + G D +L +M NG P+ + ++ T +K +VDEA K M+
Sbjct: 506 YNMMMKCYSKVGEIDEAIKLLSEMMENGCEPDVIVVNSLINTLYKADRVDEAWKMFMRMK 565
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
+ T Y L L G+ Q+A +E+ + P +TF +
Sbjct: 566 EMKLKPTVVTYNTLLAGLGKNGKIQEA-IELFEGMVQKGCPPNTITFNTLF--------- 615
Query: 504 DDCIC-----------IFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVK 548
DC+C +F+ M C P+V T NT++ +N A F ++K
Sbjct: 616 -DCLCKNDEVTLALKMLFKMMDMGCVPDVFTYNTIIFGLVKNGQVKEAMCFFHQMK 670
Score = 70.9 bits (172), Expect = 1e-10
Identities = 66/281 (23%), Positives = 120/281 (42%), Gaps = 6/281 (2%)
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
D ++P + YN ++ + + + VF Q++ + P+ ATY ++ +SG D
Sbjct: 778 DLGVQPKLPTYNLLIGGLLEADMIEIAQDVFLQVKSTGCIPDVATYNFLLDAYGKSGKID 837
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD-MERRGVMGTASVYYEL 457
+ EL+++M + +T+ +++ K G VD+A+ D M R TA Y L
Sbjct: 838 ELFELYKEMSTHECEANTITHNIVISGLVKAGNVDDALDLYYDLMSDRDFSPTACTYGPL 897
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDH 516
L GR +A E + +P + +I G D +F+ M ++
Sbjct: 898 IDGLSKSGRLYEAKQLFEGMLDY-GCRPNCAIYNILINGFGKAGEADAACALFKRMVKEG 956
Query: 517 CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEY 576
P++ T + ++ F+E+K +S L PD YNL++ + H+ E
Sbjct: 957 VRPDLKTYSVLVDCLCMVGRVDEGLHYFKELK--ESGLNPDVVCYNLIINGLGKSHRLEE 1014
Query: 577 FEHVYKEMILS-GYHLDQNKHLPLLVKASRAGKVDNILVIY 616
++ EM S G D + L++ AG V+ IY
Sbjct: 1015 ALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIY 1055
Score = 56.2 bits (134), Expect = 3e-06
Identities = 53/217 (24%), Positives = 96/217 (43%), Gaps = 25/217 (11%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAG-------LLKEL 316
Y L+ L K+ R EA Q+F ML + P+ A Y+ + G+AG L K +
Sbjct: 894 YGPLIDGLSKSGRLYEAKQLFEGML-DYGCRPNCAIYNILINGFGKAGEADAACALFKRM 952
Query: 317 LN------------IVECM---RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQ 361
+ +V+C+ + E L Y +++ + L PDVV YN ++N S +
Sbjct: 953 VKEGVRPDLKTYSVLVDCLCMVGRVDEGLHY-FKELKESGLNPDVVCYNLIINGLGKSHR 1011
Query: 362 WKGVSWVFQQMRKS-SLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYK 420
+ +F +M+ S + P+ TY + + +G + +++ ++QR G P T+
Sbjct: 1012 LEEALVLFNEMKTSRGITPDLYTYNSLILNLGIAGMVEEAGKIYNEIQRAGLEPNVFTFN 1071
Query: 421 VMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYEL 457
++R + GK + A + M G Y +L
Sbjct: 1072 ALIRGYSLSGKPEHAYAVYQTMVTGGFSPNTGTYEQL 1108
Score = 52.4 bits (124), Expect = 4e-05
Identities = 60/306 (19%), Positives = 114/306 (36%), Gaps = 45/306 (14%)
Query: 291 IRVYPD----MAAYHSIAVTLGQAGLLKELLNIVECMRQ--KPETLKYMYRKNWDPTLEP 344
++ +PD + + S+A L + ++E +R K E + Y++ ++
Sbjct: 92 LKSFPDTDSSFSYFKSVAGNLNLVHTTETCNYMLEALRVDGKLEEMAYVFDLMQKRIIKR 151
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D Y + + K + ++MR+ N +Y + ++L+S E++
Sbjct: 152 DTNTYLTIFKSLSVKGGLKQAPYALRKMREFGFVLNAYSYNGLIHLLLKSRFCTEAMEVY 211
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
+M G P TY ++ K +D + +++ME G+
Sbjct: 212 RRMILEGFRPSLQTYSSLMVGLGKRRDIDSVMGLLKEMETLGL----------------- 254
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGT 523
KP TFT IR G I++ I + M D C P+V T
Sbjct: 255 -------------------KPNVYTFTICIRVLGRAGKINEAYEILKRMDDEGCGPDVVT 295
Query: 524 VNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKE 583
++ AK +FE++K + +PD TY +L+ S + + + E
Sbjct: 296 YTVLIDALCTARKLDCAKEVFEKMKTGRH--KPDRVTYITLLDRFSDNRDLDSVKQFWSE 353
Query: 584 MILSGY 589
M G+
Sbjct: 354 MEKDGH 359
>UniRef100_Q6ETX2 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 632
Score = 99.4 bits (246), Expect = 3e-19
Identities = 91/355 (25%), Positives = 153/355 (42%), Gaps = 31/355 (8%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE 321
F YT+L+ LGKA R EA ++ M PD +++ LG+AG L + L + E
Sbjct: 301 FTYTELIRGLGKAGRIDEAYHFYHEMQRE-DCKPDTVVMNNMINFLGKAGRLDDGLKLFE 359
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKG--VSWVFQQMRKSSLKP 379
M P+VV YN ++ A SK SW F++M+ S + P
Sbjct: 360 EMGVSH--------------CIPNVVTYNTIIKALFESKSRVSEVFSW-FERMKGSGISP 404
Query: 380 NGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAV 439
+ TY + ++ ++ + L E+M G P Y ++ K + D A +
Sbjct: 405 SPFTYSILIDGFCKTNRIEKAMMLLEEMDEKGFPPCPAAYCSLIDALGKAKRYDLACELF 464
Query: 440 RDMERRGVMGTASVYYELACCLCNCGRWQDATL---EVEKIKRLPHAKPLEVTFTGMIRS 496
++++ +A VY + L GR DA E+ K+ P+ +G+ R+
Sbjct: 465 QELKENCGSSSARVYAVMIKHLGKAGRLDDAINLFDEMSKLGCTPNVYAYNALMSGLARA 524
Query: 497 SMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
M +D+ + MQ+H C P++ + N +L ++ A + +K S ++
Sbjct: 525 CM----LDEALTTMRKMQEHGCLPDINSYNIILNGLAKTGGPHRAMEMLTNMK--NSTIK 578
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVD 610
PDA +YN +L A S +E + KEM G+ D + +L GKVD
Sbjct: 579 PDAVSYNTVLSALSHAGMFEEAAELMKEMNALGFEYDLITYSSIL---EAIGKVD 630
Score = 79.3 bits (194), Expect = 3e-13
Identities = 70/327 (21%), Positives = 134/327 (40%), Gaps = 54/327 (16%)
Query: 265 TKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMR 324
++++ +LG A+ +A+ IF + + P AY+S+ + L G +++ + M
Sbjct: 163 SQVIRMLGNAKMIGKAITIFYQIKAR-KCQPTAQAYNSMIIMLIHEGQYEKVHELYNEMS 221
Query: 325 QKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATY 384
+ PD V Y+A+++A + + +M+++ ++P Y
Sbjct: 222 NEGHC-------------HPDTVTYSALISAFCKLGRQDSAIRLLNEMKENGMQPTAKIY 268
Query: 385 GLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMER 444
+ + + + N LFE+M+ P+ TY ++R K G++DEA +M+R
Sbjct: 269 TMIISLFFKLDNVHGALSLFEEMRYMYCRPDVFTYTELIRGLGKAGRIDEAYHFYHEMQR 328
Query: 445 RGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHID 504
KP V MI G +D
Sbjct: 329 E------------------------------------DCKPDTVVMNNMINFLGKAGRLD 352
Query: 505 DCICIFEYM-QDHCAPNVGTVNTMLK-VYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
D + +FE M HC PNV T NT++K ++ S FE +K S + P +TY+
Sbjct: 353 DGLKLFEEMGVSHCIPNVVTYNTIIKALFESKSRVSEVFSWFERMK--GSGISPSPFTYS 410
Query: 563 LMLEASSRGHQWEYFEHVYKEMILSGY 589
++++ + ++ E + +EM G+
Sbjct: 411 ILIDGFCKTNRIEKAMMLLEEMDEKGF 437
Score = 64.3 bits (155), Expect = 1e-08
Identities = 63/281 (22%), Positives = 116/281 (40%), Gaps = 28/281 (9%)
Query: 156 LKWVFEDDIEINEV--WFDENNNGKRKKTSKRSEVQVVRFLVDRLCDKEIRAKDWKFSRL 213
+K +FE ++EV WF+ + K + L+D C K
Sbjct: 377 IKALFESKSRVSEVFSWFE-----RMKGSGISPSPFTYSILIDGFCKTNRIEKAMMLLEE 431
Query: 214 MKLSGLSFTEGQLMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGK 273
M G +I+ LG + + A + Q + K++ S VY ++ LGK
Sbjct: 432 MDEKGFPPCPAAYCSLIDALGKAKRYDLACELFQEL---KENCGSSSARVYAVMIKHLGK 488
Query: 274 ARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYM 333
A R +A+ +F+ M + P++ AY+++ L +A +L E L +
Sbjct: 489 AGRLDDAINLFDEM-SKLGCTPNVYAYNALMSGLARACMLDEALTTM------------- 534
Query: 334 YRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQ 393
RK + PD+ YN +LN + + M+ S++KP+ +Y + +
Sbjct: 535 -RKMQEHGCLPDINSYNIILNGLAKTGGPHRAMEMLTNMKNSTIKPDAVSYNTVLSALSH 593
Query: 394 SGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDE 434
+G ++ EL ++M G + +TY ++ GKVD+
Sbjct: 594 AGMFEEAAELMKEMNALGFEYDLITYSSILEAI---GKVDQ 631
>UniRef100_Q76C26 Hypothetical protein PPR794 [Oryza sativa]
Length = 794
Score = 99.0 bits (245), Expect = 4e-19
Identities = 99/415 (23%), Positives = 166/415 (39%), Gaps = 63/415 (15%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y ++A L KA+ +A+++ M+ N V P+ Y+SI +G KE + ++ M
Sbjct: 237 YNSIIAALCKAQAMDKAMEVLTSMVKN-GVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKM 295
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+EPDVV YN++++ + + +F M K LKP T
Sbjct: 296 HSDG--------------VEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTKRGLKPEITT 341
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ G +H L + M RNG P + +++ + K+GKVD+A+ M
Sbjct: 342 YGTLLQGYATKGALVEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFSKMR 401
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKI--KRLPHAKPLEVTFTGMIRS----- 496
++G+ Y + LC GR +DA E++ +RL P + + +I S
Sbjct: 402 QQGLNPDTVTYGTVIGILCKSGRVEDAMRYFEQMIDERL---SPGNIVYNSLIHSLCIFD 458
Query: 497 -----------SMDGGHIDDCICIFEYMQDHC--------------------APNVGTVN 525
+D G D I + HC PN+ T +
Sbjct: 459 KWDKAKELILEMLDRGICLDTIFFNSIIDSHCKEGRVIESEKLFDLMVRIGVKPNIITYS 518
Query: 526 TMLKVYS-QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ Y M K+L V V ++PD TYN ++ + + E +++EM
Sbjct: 519 TLIDGYCLAGKMDEATKLLASMVSVG---MKPDCVTYNTLINGYCKISRMEDALVLFREM 575
Query: 585 ILSGYHLD---QNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYKNC 636
SG D N L L + R + V S +S ++II + C
Sbjct: 576 ESSGVSPDIITYNIILQGLFQTRRTAAAKELYVGITESGTQLELSTYNIILHGLC 630
Score = 74.7 bits (182), Expect = 8e-12
Identities = 81/374 (21%), Positives = 149/374 (39%), Gaps = 22/374 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T LL L +R +A+ I + + P++ +Y+ + L +E L +++ M
Sbjct: 129 FTPLLKGLCADKRTSDAMDIVLRRMTQLGCIPNVFSYNILLKGLCDENRSQEALELLQMM 188
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
PDVV Y V+N + +M + PN T
Sbjct: 189 PDD------------GGDCPPDVVSYTTVINGFFKEGDLDKAYGTYHEMLDRGILPNVVT 236
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + ++ D E+ M +NG +P TY +V + G+ EA+ ++ M
Sbjct: 237 YNSIIAALCKAQAMDKAMEVLTSMVKNGVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKMH 296
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV Y L LC GR +A + + + KP T+ +++ G +
Sbjct: 297 SDGVEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTK-RGLKPEITTYGTLLQGYATKGAL 355
Query: 504 DDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
+ + + M ++ PN + ++ Y++ A ++F K+ + L PD TY
Sbjct: 356 VEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFS--KMRQQGLNPDTVTYG 413
Query: 563 LMLEASSRGHQWE----YFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV-DNILVIYI 617
++ + + E YFE + E + G ++ N + L + K + IL +
Sbjct: 414 TVIGILCKSGRVEDAMRYFEQMIDERLSPG-NIVYNSLIHSLCIFDKWDKAKELILEMLD 472
Query: 618 RSQCLFYISAHSII 631
R CL I +SII
Sbjct: 473 RGICLDTIFFNSII 486
Score = 59.7 bits (143), Expect = 3e-07
Identities = 65/320 (20%), Positives = 135/320 (41%), Gaps = 24/320 (7%)
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY 332
K R E+ ++F++M+ I V P++ Y ++ AG + E ++ M
Sbjct: 491 KEGRVIESEKLFDLMV-RIGVKPNIITYSTLIDGYCLAGKMDEATKLLASMVSVG----- 544
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVML 392
++PD V YN ++N + + +F++M S + P+ TY + ++ +
Sbjct: 545 ---------MKPDCVTYNTLINGYCKISRMEDALVLFREMESSGVSPDIITYNIILQGLF 595
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
Q+ EL+ + +G E TY +++ K DEA++ +++ +
Sbjct: 596 QTRRTAAAKELYVGITESGTQLELSTYNIILHGLCKNNLTDEALRMFQNLCLTDLQLETR 655
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY 512
+ + L GR D ++ P T++ M + ++ G +++ +F
Sbjct: 656 TFNIMIGALLKVGR-NDEAKDLFAALSANGLVPDVRTYSLMAENLIEQGLLEELDDLFLS 714
Query: 513 MQDH-CAPNVGTVNTML-KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSR 570
M+++ C N +N+++ K+ + D+ LF + + +A T +L L+ S
Sbjct: 715 MEENGCTANSRMLNSIVRKLLQRGDITRAGTYLF---MIDEKHFSLEASTASLFLDLLSG 771
Query: 571 GHQWEYFEHV---YKEMILS 587
G EY + YK I S
Sbjct: 772 GKYQEYHRFLPEKYKSFIES 791
Score = 43.5 bits (101), Expect = 0.020
Identities = 43/190 (22%), Positives = 80/190 (41%), Gaps = 11/190 (5%)
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
PN TYG+ + +G DL + + G +A+ + +++ + + +A+
Sbjct: 89 PNLCTYGILIGSCCCAGRLDLGFAALGNVIKKGFRVDAIAFTPLLKGLCADKRTSDAMDI 148
Query: 439 V-RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLP----HAKPLEVTFTGM 493
V R M + G + Y L LC+ R Q+A +E ++ +P P V++T +
Sbjct: 149 VLRRMTQLGCIPNVFSYNILLKGLCDENRSQEA---LELLQMMPDDGGDCPPDVVSYTTV 205
Query: 494 IRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
I G +D + M D PNV T N+++ + A + + K+
Sbjct: 206 INGFFKEGDLDKAYGTYHEMLDRGILPNVVTYNSIIAALCKAQAMDKAMEVL--TSMVKN 263
Query: 553 DLRPDAYTYN 562
+ P+ TYN
Sbjct: 264 GVMPNCRTYN 273
>UniRef100_Q8LHA7 Pentatricopeptide repeat protein-like [Oryza sativa]
Length = 624
Score = 98.2 bits (243), Expect = 7e-19
Identities = 85/349 (24%), Positives = 155/349 (44%), Gaps = 19/349 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ L R +A+ + M + V D+ Y ++ L A E+ VE M
Sbjct: 118 YTVLMRALCADRLADQAVGLLRSMR-SAGVRADVVTYGTLIRGLCDAA---EVDKAVELM 173
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+ E+ +EP+VV+Y+++L S +W+ V VF +M + ++P+
Sbjct: 174 GEMCES-----------GIEPNVVVYSSLLQGYCKSGRWEDVGKVFVEMSEKGIEPDVVM 222
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ + + G H + + M R G P +TY V++ KEG V EA+ ++ M
Sbjct: 223 YTGLIDSLCKVGKAKKAHGVMDMMVRRGLEPNVVTYNVLINCMCKEGSVKEAIGVLKKMS 282
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH-AKPLEVTFTGMIRSSMDGGH 502
+GV Y L L + +A +E++ R + KP VTF +I+ D G
Sbjct: 283 EKGVAPDVVTYNTLIKGLSDVLEMDEAMWLLEEMVRGKNIVKPNVVTFNSVIQGLCDIGR 342
Query: 503 IDDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTY 561
+ + M++ C N+ T N ++ + A L +E + L PD++TY
Sbjct: 343 MRQAFQVRAMMEETGCMVNLVTYNLLIGGLLRVHKVRKAMELMDE--MTSLGLEPDSFTY 400
Query: 562 NLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVD 610
+++++ + Q + E + M G + ++PLLV G ++
Sbjct: 401 SILIKGFCKMWQVDRAEDLLSTMRDRGIEPELFHYIPLLVAMCEQGMME 449
Score = 79.7 bits (195), Expect = 2e-13
Identities = 74/354 (20%), Positives = 155/354 (42%), Gaps = 22/354 (6%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
+YT L+ L K + K+A + +MM+ + P++ Y+ + + + G +KE + +++
Sbjct: 222 MYTGLIDSLCKVGKAKKAHGVMDMMVRR-GLEPNVVTYNVLINCMCKEGSVKEAIGVLKK 280
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQM--RKSSLKPN 380
M +K + PDVV YN ++ + W+ ++M K+ +KPN
Sbjct: 281 MSEKG--------------VAPDVVTYNTLIKGLSDVLEMDEAMWLLEEMVRGKNIVKPN 326
Query: 381 GATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVR 440
T+ ++ + G ++ M+ G + +TY +++ + KV +A++ +
Sbjct: 327 VVTFNSVIQGLCDIGRMRQAFQVRAMMEETGCMVNLVTYNLLIGGLLRVHKVRKAMELMD 386
Query: 441 DMERRGVMGTASVYYELACCLCNCGRWQ-DATLEVEKIKRLPHAKPLEVTFTGMIRSSMD 499
+M G+ + Y L C WQ D ++ R +P + ++ + +
Sbjct: 387 EMTSLGLEPDSFTYSILIKGFCKM--WQVDRAEDLLSTMRDRGIEPELFHYIPLLVAMCE 444
Query: 500 GGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAY 559
G ++ +F M ++ +V +TM+ + TAK L + + L PDA
Sbjct: 445 QGMMERARNLFNEMDNNFPLDVVAYSTMIHGACKAGDLKTAKELLKSI--VDEGLTPDAV 502
Query: 560 TYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNIL 613
TY++++ ++ E V K+M SG+ D L+ S G+++ +L
Sbjct: 503 TYSIVINMFAKSGDMEAANGVLKQMTASGFLPDVAVFDSLIQGYSTKGEINKVL 556
Score = 76.6 bits (187), Expect = 2e-12
Identities = 58/272 (21%), Positives = 118/272 (43%), Gaps = 13/272 (4%)
Query: 296 DMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA 355
D +Y+++ L + G ++ M +P P P+ V Y ++ A
Sbjct: 76 DAVSYNTVLTALCRRGHHDRAGALLRAMSLEPH-----------PACRPNAVSYTVLMRA 124
Query: 356 CVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPE 415
+ + + MR + ++ + TYG + + + D EL +M +G P
Sbjct: 125 LCADRLADQAVGLLRSMRSAGVRADVVTYGTLIRGLCDAAEVDKAVELMGEMCESGIEPN 184
Query: 416 ALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVE 475
+ Y +++ + K G+ ++ K +M +G+ +Y L LC G+ + A ++
Sbjct: 185 VVVYSSLLQGYCKSGRWEDVGKVFVEMSEKGIEPDVVMYTGLIDSLCKVGKAKKAHGVMD 244
Query: 476 KIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQN 534
+ R +P VT+ +I G + + I + + M + AP+V T NT++K S
Sbjct: 245 MMVR-RGLEPNVVTYNVLINCMCKEGSVKEAIGVLKKMSEKGVAPDVVTYNTLIKGLSDV 303
Query: 535 DMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLE 566
A L EE+ K+ ++P+ T+N +++
Sbjct: 304 LEMDEAMWLLEEMVRGKNIVKPNVVTFNSVIQ 335
>UniRef100_Q8W3E4 Putative membrane-associated protein [Oryza sativa]
Length = 1219
Score = 97.4 bits (241), Expect = 1e-18
Identities = 98/415 (23%), Positives = 167/415 (39%), Gaps = 63/415 (15%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ ++A L KA+ +A+++ M+ N V P+ Y+SI +G KE + ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKN-GVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKM 295
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+EPDVV YN++++ + + +F M K LKP T
Sbjct: 296 HSDG--------------VEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTKRGLKPEITT 341
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ G +H L + M RNG P + +++ + K+GKVD+A+ M
Sbjct: 342 YGTLLQGYATKGALVEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFSKMR 401
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKI--KRLPHAKPLEVTFTGMIRS----- 496
++G+ Y + LC GR +DA E++ +RL P + + +I S
Sbjct: 402 QQGLNPDTVTYGTVIGILCKSGRVEDAMRYFEQMIDERL---SPGNIVYNSLIHSLCIFD 458
Query: 497 -----------SMDGGHIDDCICIFEYMQDHC--------------------APNVGTVN 525
+D G D I + HC P++ T +
Sbjct: 459 KWDKAKELILEMLDRGICLDTIFFNSIIDSHCKEGRVIESEKLFDLMVRIGVKPDIITYS 518
Query: 526 TMLKVYS-QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ Y M K+L V V ++PD TYN ++ + + E +++EM
Sbjct: 519 TLIDGYCLAGKMDEATKLLASMVSVG---MKPDCVTYNTLINGYCKISRMEDALVLFREM 575
Query: 585 ILSGYHLD---QNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYKNC 636
SG D N L L + R + V S +S ++II + C
Sbjct: 576 ESSGVSPDIITYNIILQGLFQTRRTAAAKELYVGITESGTQLELSTYNIILHGLC 630
Score = 74.7 bits (182), Expect = 8e-12
Identities = 81/374 (21%), Positives = 149/374 (39%), Gaps = 22/374 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T LL L +R +A+ I + + P++ +Y+ + L +E L +++ M
Sbjct: 129 FTPLLKGLCADKRTSDAMDIVLRRMTQLGCIPNVFSYNILLKGLCDENRSQEALELLQMM 188
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
PDVV Y V+N + +M + PN T
Sbjct: 189 PDD------------GGDCPPDVVSYTTVINGFFKEGDLDKAYGTYHEMLDRGILPNVVT 236
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + ++ D E+ M +NG +P TY +V + G+ EA+ ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKNGVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKMH 296
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV Y L LC GR +A + + + KP T+ +++ G +
Sbjct: 297 SDGVEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTK-RGLKPEITTYGTLLQGYATKGAL 355
Query: 504 DDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
+ + + M ++ PN + ++ Y++ A ++F K+ + L PD TY
Sbjct: 356 VEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFS--KMRQQGLNPDTVTYG 413
Query: 563 LMLEASSRGHQWE----YFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV-DNILVIYI 617
++ + + E YFE + E + G ++ N + L + K + IL +
Sbjct: 414 TVIGILCKSGRVEDAMRYFEQMIDERLSPG-NIVYNSLIHSLCIFDKWDKAKELILEMLD 472
Query: 618 RSQCLFYISAHSII 631
R CL I +SII
Sbjct: 473 RGICLDTIFFNSII 486
Score = 60.8 bits (146), Expect = 1e-07
Identities = 62/306 (20%), Positives = 130/306 (42%), Gaps = 21/306 (6%)
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY 332
K R E+ ++F++M+ I V PD+ Y ++ AG + E ++ M
Sbjct: 491 KEGRVIESEKLFDLMV-RIGVKPDIITYSTLIDGYCLAGKMDEATKLLASMVSVG----- 544
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVML 392
++PD V YN ++N + + +F++M S + P+ TY + ++ +
Sbjct: 545 ---------MKPDCVTYNTLINGYCKISRMEDALVLFREMESSGVSPDIITYNIILQGLF 595
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
Q+ EL+ + +G E TY +++ K DEA++ +++ +
Sbjct: 596 QTRRTAAAKELYVGITESGTQLELSTYNIILHGLCKNNLTDEALRMFQNLCLTDLQLETR 655
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY 512
+ + L GR D ++ P T++ M + ++ G +++ +F
Sbjct: 656 TFNIMIGALLKVGR-NDEAKDLFAALSANGLVPDVRTYSLMAENLIEQGLLEELDDLFLS 714
Query: 513 MQDH-CAPNVGTVNTML-KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSR 570
M+++ C N +N+++ K+ + D+ LF + + +A T +L L+ S
Sbjct: 715 MEENGCTANSRMLNSIVRKLLQRGDITRAGTYLF---MIDEKHFSLEASTASLFLDLLSG 771
Query: 571 GHQWEY 576
G EY
Sbjct: 772 GKYQEY 777
Score = 41.6 bits (96), Expect = 0.074
Identities = 42/190 (22%), Positives = 80/190 (42%), Gaps = 11/190 (5%)
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
PN TYG+ + +G DL + + G +A+ + +++ + + +A+
Sbjct: 89 PNLCTYGILIGSCCCAGRLDLGFAALGNVIKKGFRVDAIAFTPLLKGLCADKRTSDAMDI 148
Query: 439 V-RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLP----HAKPLEVTFTGM 493
V R M + G + Y L LC+ R Q+A +E ++ +P P V++T +
Sbjct: 149 VLRRMTQLGCIPNVFSYNILLKGLCDENRSQEA---LELLQMMPDDGGDCPPDVVSYTTV 205
Query: 494 IRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
I G +D + M D PNV T ++++ + A + + K+
Sbjct: 206 INGFFKEGDLDKAYGTYHEMLDRGILPNVVTYSSIIAALCKAQAMDKAMEVL--TSMVKN 263
Query: 553 DLRPDAYTYN 562
+ P+ TYN
Sbjct: 264 GVMPNCRTYN 273
>UniRef100_Q769C9 PPR protein [Oryza sativa]
Length = 794
Score = 97.4 bits (241), Expect = 1e-18
Identities = 98/415 (23%), Positives = 167/415 (39%), Gaps = 63/415 (15%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ ++A L KA+ +A+++ M+ N V P+ Y+SI +G KE + ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKN-GVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKM 295
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+EPDVV YN++++ + + +F M K LKP T
Sbjct: 296 HSDG--------------VEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTKRGLKPEITT 341
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ G +H L + M RNG P + +++ + K+GKVD+A+ M
Sbjct: 342 YGTLLQGYATKGALVEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFSKMR 401
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKI--KRLPHAKPLEVTFTGMIRS----- 496
++G+ Y + LC GR +DA E++ +RL P + + +I S
Sbjct: 402 QQGLNPDTVTYGTVIGILCKSGRVEDAMRYFEQMIDERL---SPGNIVYNSLIHSLCIFD 458
Query: 497 -----------SMDGGHIDDCICIFEYMQDHC--------------------APNVGTVN 525
+D G D I + HC P++ T +
Sbjct: 459 KWDKAKELILEMLDRGICLDTIFFNSIIDSHCKEGRVIESEKLFDLMVRIGVKPDIITYS 518
Query: 526 TMLKVYS-QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ Y M K+L V V ++PD TYN ++ + + E +++EM
Sbjct: 519 TLIDGYCLAGKMDEATKLLASMVSVG---MKPDCVTYNTLINGYCKISRMEDALVLFREM 575
Query: 585 ILSGYHLD---QNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYKNC 636
SG D N L L + R + V S +S ++II + C
Sbjct: 576 ESSGVSPDIITYNIILQGLFQTRRTAAAKELYVGITESGTQLELSTYNIILHGLC 630
Score = 73.6 bits (179), Expect = 2e-11
Identities = 80/374 (21%), Positives = 149/374 (39%), Gaps = 22/374 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T +L L +R +A+ I + + P++ +Y+ + L +E L +++ M
Sbjct: 129 FTPMLKGLCADKRTSDAMDIVLRRMTQLGCIPNVFSYNILLKGLCDDNRSQEALELLQMM 188
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
PDVV Y V+N + +M + PN T
Sbjct: 189 PDD------------GGDCPPDVVSYTTVINGFFKEGDLDKAYGTYHEMLDRGILPNVVT 236
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + ++ D E+ M +NG +P TY +V + G+ EA+ ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKNGVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKMH 296
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV Y L LC GR +A + + + KP T+ +++ G +
Sbjct: 297 SDGVEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTK-RGLKPEITTYGTLLQGYATKGAL 355
Query: 504 DDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
+ + + M ++ PN + ++ Y++ A ++F K+ + L PD TY
Sbjct: 356 VEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFS--KMRQQGLNPDTVTYG 413
Query: 563 LMLEASSRGHQWE----YFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV-DNILVIYI 617
++ + + E YFE + E + G ++ N + L + K + IL +
Sbjct: 414 TVIGILCKSGRVEDAMRYFEQMIDERLSPG-NIVYNSLIHSLCIFDKWDKAKELILEMLD 472
Query: 618 RSQCLFYISAHSII 631
R CL I +SII
Sbjct: 473 RGICLDTIFFNSII 486
Score = 61.6 bits (148), Expect = 7e-08
Identities = 66/320 (20%), Positives = 135/320 (41%), Gaps = 24/320 (7%)
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY 332
K R E+ ++F++M+ I V PD+ Y ++ AG + E ++ M
Sbjct: 491 KEGRVIESEKLFDLMV-RIGVKPDIITYSTLIDGYCLAGKMDEATKLLASMVSVG----- 544
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVML 392
++PD V YN ++N + + +F++M S + P+ TY + ++ +
Sbjct: 545 ---------MKPDCVTYNTLINGYCKISRMEDALVLFREMESSGVSPDIITYNIILQGLF 595
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
Q+ EL+ + +G E TY +++ K DEA++ +++ +
Sbjct: 596 QTRRTAAAKELYVGITESGTQLELSTYNIILHGLCKNNLTDEALRMFQNLCLTDLQLETR 655
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY 512
+ + L GR D ++ P T++ M + ++ G +++ +F
Sbjct: 656 TFNIMIGALLKVGR-NDEAKDLFAALSANGLVPDVRTYSLMAENLIEQGLLEELDDLFLS 714
Query: 513 MQDH-CAPNVGTVNTML-KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSR 570
M+++ C N +N+++ K+ + D+ LF + + +A T +L L+ S
Sbjct: 715 MEENGCTANSRMLNSIVRKLLQRGDITRAGTYLF---MIDEKHFSLEASTASLFLDLLSG 771
Query: 571 GHQWEYFEHV---YKEMILS 587
G EY + YK I S
Sbjct: 772 GKYQEYHRFLPEKYKSFIES 791
Score = 47.0 bits (110), Expect = 0.002
Identities = 44/190 (23%), Positives = 81/190 (42%), Gaps = 11/190 (5%)
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
PN TYG+ M +G DL + + G + +A+ + M++ + + +A+
Sbjct: 89 PNLCTYGILMGSCCCAGRLDLGFAALGNVIKKGFIVDAIAFTPMLKGLCADKRTSDAMDI 148
Query: 439 V-RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLP----HAKPLEVTFTGM 493
V R M + G + Y L LC+ R Q+A +E ++ +P P V++T +
Sbjct: 149 VLRRMTQLGCIPNVFSYNILLKGLCDDNRSQEA---LELLQMMPDDGGDCPPDVVSYTTV 205
Query: 494 IRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
I G +D + M D PNV T ++++ + A + + K+
Sbjct: 206 INGFFKEGDLDKAYGTYHEMLDRGILPNVVTYSSIIAALCKAQAMDKAMEVL--TSMVKN 263
Query: 553 DLRPDAYTYN 562
+ P+ TYN
Sbjct: 264 GVMPNCRTYN 273
>UniRef100_Q76C22 Hypothetical protein [Oryza sativa]
Length = 794
Score = 97.4 bits (241), Expect = 1e-18
Identities = 98/415 (23%), Positives = 167/415 (39%), Gaps = 63/415 (15%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ ++A L KA+ +A+++ M+ N V P+ Y+SI +G KE + ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKN-GVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKM 295
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+EPDVV YN++++ + + +F M K LKP T
Sbjct: 296 HSDG--------------VEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTKRGLKPEITT 341
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ G +H L + M RNG P + +++ + K+GKVD+A+ M
Sbjct: 342 YGTLLQGYATKGALVEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFSKMR 401
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKI--KRLPHAKPLEVTFTGMIRS----- 496
++G+ Y + LC GR +DA E++ +RL P + + +I S
Sbjct: 402 QQGLNPDTVTYGTVIGILCKSGRVEDAMRYFEQMIDERL---SPGNIVYNSLIHSLCIFD 458
Query: 497 -----------SMDGGHIDDCICIFEYMQDHC--------------------APNVGTVN 525
+D G D I + HC P++ T +
Sbjct: 459 KWDKAKELILEMLDRGICLDTIFFNSIIDSHCKEGRVIESEKLFDLMVRIGVKPDIITYS 518
Query: 526 TMLKVYS-QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
T++ Y M K+L V V ++PD TYN ++ + + E +++EM
Sbjct: 519 TLIDGYCLAGKMDEATKLLASMVSVG---MKPDCVTYNTLINGYCKISRMEDALVLFREM 575
Query: 585 ILSGYHLD---QNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYKNC 636
SG D N L L + R + V S +S ++II + C
Sbjct: 576 ESSGVSPDIITYNIILQGLFQTRRTAAAKELYVGITESGTQLELSTYNIILHGLC 630
Score = 74.7 bits (182), Expect = 8e-12
Identities = 81/374 (21%), Positives = 149/374 (39%), Gaps = 22/374 (5%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+T LL L +R +A+ I + + P++ +Y+ + L +E L +++ M
Sbjct: 129 FTPLLKGLCADKRTSDAMDIVLRRMTQLGCIPNVFSYNILLKGLCDENRSQEALELLQMM 188
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
PDVV Y V+N + +M + PN T
Sbjct: 189 PDD------------GGDCPPDVVSYTTVINGFFKEGDLDKAYGTYHEMLDRGILPNVVT 236
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + ++ D E+ M +NG +P TY +V + G+ EA+ ++ M
Sbjct: 237 YSSIIAALCKAQAMDKAMEVLTSMVKNGVMPNCRTYNSIVHGYCSSGQPKEAIGFLKKMH 296
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV Y L LC GR +A + + + KP T+ +++ G +
Sbjct: 297 SDGVEPDVVTYNSLMDYLCKNGRCTEARKMFDSMTK-RGLKPEITTYGTLLQGYATKGAL 355
Query: 504 DDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYN 562
+ + + M ++ PN + ++ Y++ A ++F K+ + L PD TY
Sbjct: 356 VEMHGLLDLMVRNGIHPNHYVFSILICAYAKQGKVDQAMLVFS--KMRQQGLNPDTVTYG 413
Query: 563 LMLEASSRGHQWE----YFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKV-DNILVIYI 617
++ + + E YFE + E + G ++ N + L + K + IL +
Sbjct: 414 TVIGILCKSGRVEDAMRYFEQMIDERLSPG-NIVYNSLIHSLCIFDKWDKAKELILEMLD 472
Query: 618 RSQCLFYISAHSII 631
R CL I +SII
Sbjct: 473 RGICLDTIFFNSII 486
Score = 61.6 bits (148), Expect = 7e-08
Identities = 66/320 (20%), Positives = 135/320 (41%), Gaps = 24/320 (7%)
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKY 332
K R E+ ++F++M+ I V PD+ Y ++ AG + E ++ M
Sbjct: 491 KEGRVIESEKLFDLMV-RIGVKPDIITYSTLIDGYCLAGKMDEATKLLASMVSVG----- 544
Query: 333 MYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVML 392
++PD V YN ++N + + +F++M S + P+ TY + ++ +
Sbjct: 545 ---------MKPDCVTYNTLINGYCKISRMEDALVLFREMESSGVSPDIITYNIILQGLF 595
Query: 393 QSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS 452
Q+ EL+ + +G E TY +++ K DEA++ +++ +
Sbjct: 596 QTRRTAAAKELYVGITESGTQLELSTYNIILHGLCKNNLTDEALRMFQNLCLTDLQLETR 655
Query: 453 VYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEY 512
+ + L GR D ++ P T++ M + ++ G +++ +F
Sbjct: 656 TFNIMIGALLKVGR-NDEAKDLFAALSANGLVPDVRTYSLMAENLIEQGLLEELDDLFLS 714
Query: 513 MQDH-CAPNVGTVNTML-KVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSR 570
M+++ C N +N+++ K+ + D+ LF + + +A T +L L+ S
Sbjct: 715 MEENGCTANSRMLNSIVRKLLQRGDITRAGTYLF---MIDEKHFSLEASTASLFLDLLSG 771
Query: 571 GHQWEYFEHV---YKEMILS 587
G EY + YK I S
Sbjct: 772 GKYQEYHRFLPEKYKSFIES 791
Score = 41.6 bits (96), Expect = 0.074
Identities = 42/190 (22%), Positives = 80/190 (42%), Gaps = 11/190 (5%)
Query: 379 PNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKA 438
PN TYG+ + +G DL + + G +A+ + +++ + + +A+
Sbjct: 89 PNLCTYGILIGSCCCAGRLDLGFAALGNVIKKGFRVDAIAFTPLLKGLCADKRTSDAMDI 148
Query: 439 V-RDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLP----HAKPLEVTFTGM 493
V R M + G + Y L LC+ R Q+A +E ++ +P P V++T +
Sbjct: 149 VLRRMTQLGCIPNVFSYNILLKGLCDENRSQEA---LELLQMMPDDGGDCPPDVVSYTTV 205
Query: 494 IRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
I G +D + M D PNV T ++++ + A + + K+
Sbjct: 206 INGFFKEGDLDKAYGTYHEMLDRGILPNVVTYSSIIAALCKAQAMDKAMEVL--TSMVKN 263
Query: 553 DLRPDAYTYN 562
+ P+ TYN
Sbjct: 264 GVMPNCRTYN 273
>UniRef100_Q6ATD8 Hypothetical protein OSJNBa0018H09.7 [Oryza sativa]
Length = 793
Score = 97.4 bits (241), Expect = 1e-18
Identities = 74/316 (23%), Positives = 140/316 (43%), Gaps = 23/316 (7%)
Query: 296 DMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNA 355
D+ AY ++ L +AG + L + +R++ + P +V YN VL+
Sbjct: 179 DVRAYTTVLHALSRAGRYERALQLFAELRRQG--------------VVPTIVTYNVVLDV 224
Query: 356 CVP-SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVP 414
+ W + + ++MR + ++P+ T + + G D FE ++ G VP
Sbjct: 225 YGRMGRSWPRIVALLEEMRAAGVEPDDFTASTVIAACGRDGLLDQAVAFFEDLKARGHVP 284
Query: 415 EALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDAT--L 472
+TY +++ F K G EA++ +++ME G A Y ELA G +++A L
Sbjct: 285 CVVTYNALLQVFGKAGNYTEALRVLKEMEDSGCQPDAVTYNELAGTYARAGFFEEAAKCL 344
Query: 473 EVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVY 531
+ K L P T+ ++ + + G +D+ + +F+ M+ + PNV T N + +
Sbjct: 345 DTMTSKGL---LPNTFTYNTVMTAYANVGRVDEALALFDRMKKNGYVPNVNTYNLIFGML 401
Query: 532 SQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHL 591
+ F+ + EE +++S P+ T+N ML + +Y V M G L
Sbjct: 402 GKKSRFTAMLEMLEE--MSRSGCTPNRVTWNTMLAVCGKRGMEDYVTRVLNGMKSCGVEL 459
Query: 592 DQNKHLPLLVKASRAG 607
++ + L+ R G
Sbjct: 460 SRDTYNTLISAYGRCG 475
Score = 90.9 bits (224), Expect = 1e-16
Identities = 86/348 (24%), Positives = 147/348 (41%), Gaps = 24/348 (6%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAG-LLKELLNIVEC 322
YT +L L +A R + ALQ+F L V P + Y+ + G+ G ++ ++E
Sbjct: 183 YTTVLHALSRAGRYERALQLF-AELRRQGVVPTIVTYNVVLDVYGRMGRSWPRIVALLEE 241
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
MR +EPD + V+ AC F+ ++ P
Sbjct: 242 MRA--------------AGVEPDDFTASTVIAACGRDGLLDQAVAFFEDLKARGHVPCVV 287
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY ++V ++GNY + ++M+ +G P+A+TY + T+ + G +EA K + M
Sbjct: 288 TYNALLQVFGKAGNYTEALRVLKEMEDSGCQPDAVTYNELAGTYARAGFFEEAAKCLDTM 347
Query: 443 ERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKR---LPHAKPLEVTFTGMIRSSMD 499
+G++ Y + N GR +A +++K+ +P+ + F + + S
Sbjct: 348 TSKGLLPNTFTYNTVMTAYANVGRVDEALALFDRMKKNGYVPNVNTYNLIFGMLGKKSRF 407
Query: 500 GGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAY 559
++ + E + C PN T NTML V + M + +K +L D
Sbjct: 408 TAMLE---MLEEMSRSGCTPNRVTWNTMLAVCGKRGMEDYVTRVLNGMKSCGVELSRD-- 462
Query: 560 TYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAG 607
TYN ++ A R Y +Y EMI SG+ + LL SR G
Sbjct: 463 TYNTLISAYGRCGSRTYAFKMYDEMISSGFTPCLTTYNALLNVLSRQG 510
Score = 83.2 bits (204), Expect = 2e-14
Identities = 74/321 (23%), Positives = 125/321 (38%), Gaps = 52/321 (16%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y LL V GKA EAL++ M + PD Y+ +A T +AG +E ++ M
Sbjct: 289 YNALLQVFGKAGNYTEALRVLKEMEDS-GCQPDAVTYNELAGTYARAGFFEEAAKCLDTM 347
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
K L P+ YN V+ A + +F +M+K+ PN T
Sbjct: 348 TSKG--------------LLPNTFTYNTVMTAYANVGRVDEALALFDRMKKNGYVPNVNT 393
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y L ++ + + + E+ E+M R+G P +T+ M+ K G D + + M+
Sbjct: 394 YNLIFGMLGKKSRFTAMLEMLEEMSRSGCTPNRVTWNTMLAVCGKRGMEDYVTRVLNGMK 453
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHI 503
GV + Y L CG T+ + M
Sbjct: 454 SCGVELSRDTYNTLISAYGRCG---------------------SRTYAFKMYDEMISSGF 492
Query: 504 DDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNL 563
C+ T N +L V S+ +STA+ + K+ K+ +P+ +Y+L
Sbjct: 493 TPCLT--------------TYNALLNVLSRQGDWSTAQSIVS--KMLKNGFKPNDQSYSL 536
Query: 564 MLEASSRGHQWEYFEHVYKEM 584
+L+ ++G E + KE+
Sbjct: 537 LLQCYAKGGNAAGIESIEKEV 557
Score = 65.1 bits (157), Expect = 6e-09
Identities = 72/356 (20%), Positives = 143/356 (39%), Gaps = 40/356 (11%)
Query: 262 FVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE 321
F Y ++ R EAL +F+ M N V P++ Y+ I LG+ +L ++E
Sbjct: 357 FTYNTVMTAYANVGRVDEALALFDRMKKNGYV-PNVNTYNLIFGMLGKKSRFTAMLEMLE 415
Query: 322 CMRQKPETLKYMYRKNWDPTLE-------PDVVI-----------------YNAVLNACV 357
M + T R W+ L D V YN +++A
Sbjct: 416 EMSRSGCTPN---RVTWNTMLAVCGKRGMEDYVTRVLNGMKSCGVELSRDTYNTLISAYG 472
Query: 358 PSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEAL 417
++ +M S P TY + V+ + G++ + KM +NG P
Sbjct: 473 RCGSRTYAFKMYDEMISSGFTPCLTTYNALLNVLSRQGDWSTAQSIVSKMLKNGFKPNDQ 532
Query: 418 TYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTAS----VYYELACCLCNCGRWQDATLE 473
+Y ++++ + K G + +E+ +GT + L C R +
Sbjct: 533 SYSLLLQCYAKGGNA----AGIESIEKEVYVGTIFPSWVILRTLVIANFKCRRLEGVEKA 588
Query: 474 VEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYS 532
+++K + KP V F M+ G +F+ + Q +P++ T N+++ +Y+
Sbjct: 589 FQEVKAQGY-KPDLVIFNSMLAMYAKNGLYSKATEMFDSIKQSGLSPDLITYNSLMDMYA 647
Query: 533 QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
+++ A+ + +++K S ++PD +YN ++ + + + + EMI G
Sbjct: 648 KSNESWEAEKILKQLK--SSQVKPDVVSYNTVINGFCKQGLIKEAQRILSEMIADG 701
Score = 61.2 bits (147), Expect = 9e-08
Identities = 60/281 (21%), Positives = 125/281 (44%), Gaps = 13/281 (4%)
Query: 351 AVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQS----GNYDLVHELFEK 406
++L A S W+ W +R +S GA A+E+++++ G +D+V +L ++
Sbjct: 115 SLLKALELSGHWE---WALALLRWAS--DEGAADAAALEMVVRALGREGQHDVVCDLLDE 169
Query: 407 MQRN-GEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCG 465
M G + Y ++ + G+ + A++ ++ R+GV+ T Y + G
Sbjct: 170 MPLPPGSRLDVRAYTTVLHALSRAGRYERALQLFAELRRQGVVPTIVTYNVVLDVYGRMG 229
Query: 466 RWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHC-APNVGTV 524
R + + + R +P + T + +I + G +D + FE ++ P V T
Sbjct: 230 RSWPRIVALLEEMRAAGVEPDDFTASTVIAACGRDGLLDQAVAFFEDLKARGHVPCVVTY 289
Query: 525 NTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
N +L+V+ + ++ A + +E++ S +PDA TYN + +R +E M
Sbjct: 290 NALLQVFGKAGNYTEALRVLKEME--DSGCQPDAVTYNELAGTYARAGFFEEAAKCLDTM 347
Query: 585 ILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQCLFYI 625
G + + ++ + G+VD L ++ R + Y+
Sbjct: 348 TSKGLLPNTFTYNTVMTAYANVGRVDEALALFDRMKKNGYV 388
Score = 59.3 bits (142), Expect = 3e-07
Identities = 79/416 (18%), Positives = 156/416 (36%), Gaps = 57/416 (13%)
Query: 214 MKLSGLSFTEGQLMMIIEMLGVKRCWKQALSVVQWVYNSKDHRKFQSRFVYTKLLAVLGK 273
MK +G +I MLG K + L +++ + S +R + +LAV GK
Sbjct: 382 MKKNGYVPNVNTYNLIFGMLGKKSRFTAMLEMLEEMSRSGCT---PNRVTWNTMLAVCGK 438
Query: 274 ARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYM 333
++ N M + V Y+++ G+ G + K M
Sbjct: 439 RGMEDYVTRVLNGMK-SCGVELSRDTYNTLISAYGRCG-------------SRTYAFK-M 483
Query: 334 YRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQ 393
Y + P + YNA+LN W + +M K+ KPN +Y L ++ +
Sbjct: 484 YDEMISSGFTPCLTTYNALLNVLSRQGDWSTAQSIVSKMLKNGFKPNDQSYSLLLQCYAK 543
Query: 394 SGN-----------------------------------YDLVHELFEKMQRNGEVPEALT 418
GN + V + F++++ G P+ +
Sbjct: 544 GGNAAGIESIEKEVYVGTIFPSWVILRTLVIANFKCRRLEGVEKAFQEVKAQGYKPDLVI 603
Query: 419 YKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
+ M+ + K G +A + +++ G+ Y L +A ++++K
Sbjct: 604 FNSMLAMYAKNGLYSKATEMFDSIKQSGLSPDLITYNSLMDMYAKSNESWEAEKILKQLK 663
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMF 537
KP V++ +I G I + I E + D AP V T +T++ Y+ +MF
Sbjct: 664 S-SQVKPDVVSYNTVINGFCKQGLIKEAQRILSEMIADGMAPCVVTYHTLVGGYASLEMF 722
Query: 538 STAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQ 593
+ A+ + + +L+P TY ++++ + +++ E+ + + DQ
Sbjct: 723 NEAREVVN--YMIHHNLKPMELTYRRVVDSYCKAKRYDEAREFLSEISDTDQNFDQ 776
Score = 56.2 bits (134), Expect = 3e-06
Identities = 34/163 (20%), Positives = 79/163 (47%), Gaps = 1/163 (0%)
Query: 343 EPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
+PD+VI+N++L + + + +F +++S L P+ TY M++ +S +
Sbjct: 598 KPDLVIFNSMLAMYAKNGLYSKATEMFDSIKQSGLSPDLITYNSLMDMYAKSNESWEAEK 657
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
+ ++++ + P+ ++Y ++ F K+G + EA + + +M G+ Y+ L
Sbjct: 658 ILKQLKSSQVKPDVVSYNTVINGFCKQGLIKEAQRILSEMIADGMAPCVVTYHTLVGGYA 717
Query: 463 NCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDD 505
+ + +A EV + KP+E+T+ ++ S D+
Sbjct: 718 SLEMFNEAR-EVVNYMIHHNLKPMELTYRRVVDSYCKAKRYDE 759
>UniRef100_Q7XHS8 Putative pentatricopeptide (PPR) repeat-containing protein [Oryza
sativa]
Length = 882
Score = 96.7 bits (239), Expect = 2e-18
Identities = 78/338 (23%), Positives = 156/338 (46%), Gaps = 25/338 (7%)
Query: 257 KFQSRF-VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKE 315
KF+ F YT L+ L +ARRP+ AL++ M + + + ++ L + G + +
Sbjct: 174 KFRPAFSAYTVLIGALAEARRPERALELLRQM-QEVGYEVGVHLFTTLVRALAREGQVAD 232
Query: 316 LLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSW-VFQQMRK 374
L +V+ ++ LEPD+V+YN ++ C ++W F +++
Sbjct: 233 ALALVDEVK--------------GSCLEPDIVLYNVCID-CFGKAGNVDMAWKFFHELKA 277
Query: 375 SSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDE 434
LKP+ +Y + V+ ++G ELF +M+ VP A Y M+ + G+ ++
Sbjct: 278 QGLKPDDVSYTSMIWVLCKAGRLGEAEELFAQMEAERSVPCAYAYNTMIMGYGSAGRFED 337
Query: 435 AVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMI 494
A K + + RG + + + + CL + +A E +K+ A+P T+ +I
Sbjct: 338 AYKLLERLRERGCIPSVVSFNSILTCLGKKRKVDEALSLFEVMKK--DAEPNSSTYNIII 395
Query: 495 RSSMDGGHIDDCICIFEYMQDHCA--PNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
GG +++ I + M +H + PN+ TVN M+ + A +FE ++
Sbjct: 396 DMLCLGGRVEEAYRILDEM-EHASLFPNLLTVNIMVDRLCKARKLEEAYKIFE--SASQR 452
Query: 553 DLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYH 590
PD TY +++ + Q + ++++M+ +G++
Sbjct: 453 GCNPDCVTYCSLIDGLGKKGQVDEAYRLFEKMLDAGHN 490
Score = 83.6 bits (205), Expect = 2e-14
Identities = 72/344 (20%), Positives = 150/344 (42%), Gaps = 18/344 (5%)
Query: 267 LLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQK 326
L A L +ARR +A+ +M ++ P +AY + L +A ++
Sbjct: 150 LAAALVRARRLDDAVLAVAVMR-RLKFRPAFSAYTVLIGALAEA--------------RR 194
Query: 327 PETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGL 386
PE + R+ + E V ++ ++ A Q + +++ S L+P+ Y +
Sbjct: 195 PERALELLRQMQEVGYEVGVHLFTTLVRALAREGQVADALALVDEVKGSCLEPDIVLYNV 254
Query: 387 AMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRG 446
++ ++GN D+ + F +++ G P+ ++Y M+ K G++ EA + ME
Sbjct: 255 CIDCFGKAGNVDMAWKFFHELKAQGLKPDDVSYTSMIWVLCKAGRLGEAEELFAQMEAER 314
Query: 447 VMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDC 506
+ A Y + + GR++DA +E++ R P V+F ++ +D+
Sbjct: 315 SVPCAYAYNTMIMGYGSAGRFEDAYKLLERL-RERGCIPSVVSFNSILTCLGKKRKVDEA 373
Query: 507 ICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLE 566
+ +FE M+ PN T N ++ + A + +E++ A L P+ T N+M++
Sbjct: 374 LSLFEVMKKDAEPNSSTYNIIIDMLCLGGRVEEAYRILDEMEHA--SLFPNLLTVNIMVD 431
Query: 567 ASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVD 610
+ + E +++ G + D + L+ + G+VD
Sbjct: 432 RLCKARKLEEAYKIFESASQRGCNPDCVTYCSLIDGLGKKGQVD 475
Score = 81.6 bits (200), Expect = 6e-14
Identities = 78/373 (20%), Positives = 158/373 (41%), Gaps = 32/373 (8%)
Query: 271 LGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQ----- 325
L KAR+ +EA +IF PD Y S+ LG+ G + E + E M
Sbjct: 433 LCKARKLEEAYKIFESA-SQRGCNPDCVTYCSLIDGLGKKGQVDEAYRLFEKMLDAGHNA 491
Query: 326 ----------------KPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVF 369
+ E ++++ +PD+ + N ++ + + + +F
Sbjct: 492 NPVVYTSLIRNFFIHGRKEDGHKIFKELIRRGCKPDLTLLNTYMDCVFKAGEVEKGRMIF 551
Query: 370 QQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKE 429
+ +R P+ +Y + + + ++G +F M++ G +A Y +V F K
Sbjct: 552 EDIRSYGFLPDVRSYSILIHGLTKAGQARETSNIFHAMKQQGFALDARAYNAVVDGFCKS 611
Query: 430 GKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLE-- 487
GKV +A + + +M+ + V T + Y + L R +A + E+ K +K +E
Sbjct: 612 GKVHKAYEILEEMKEKCVQPTVATYGAIVDGLAKIDRLDEAYMLFEEAK----SKGIELN 667
Query: 488 -VTFTGMIRSSMDGGHIDDCICIF-EYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFE 545
V ++ +I G ID+ I E M+ PNV T N++L + + + A V F+
Sbjct: 668 VVLYSSLIDGFGKVGRIDEAYLILEEMMKKGLTPNVYTWNSLLDALVKAEEINEALVCFQ 727
Query: 546 EVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASR 605
+K K P+ YTY++++ R ++ +++M G + + ++ ++
Sbjct: 728 SMKEMKCP--PNTYTYSILINGLCRVQKYNKAFVFWQDMQKQGLVPNVVTYTTMISGLAK 785
Query: 606 AGKVDNILVIYIR 618
G + + ++ R
Sbjct: 786 VGNITDAYSLFER 798
Score = 75.5 bits (184), Expect = 5e-12
Identities = 69/356 (19%), Positives = 152/356 (42%), Gaps = 26/356 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
+ +L LGK R+ EAL +F +M + P+ + Y+ I L G ++E I++ M
Sbjct: 357 FNSILTCLGKKRKVDEALSLFEVMKKDAE--PNSSTYNIIIDMLCLGGRVEEAYRILDEM 414
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+L P+++ N +++ +++ + +F+ + P+ T
Sbjct: 415 EHA--------------SLFPNLLTVNIMVDRLCKARKLEEAYKIFESASQRGCNPDCVT 460
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y ++ + + G D + LFEKM G + Y ++R F+ G+ ++ K +++
Sbjct: 461 YCSLIDGLGKKGQVDEAYRLFEKMLDAGHNANPVVYTSLIRNFFIHGRKEDGHKIFKELI 520
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKR---LPHAKPLEVTFTGMIRSSMDG 500
RRG ++ C+ G + + E I+ LP + + G+ ++
Sbjct: 521 RRGCKPDLTLLNTYMDCVFKAGEVEKGRMIFEDIRSYGFLPDVRSYSILIHGLTKA---- 576
Query: 501 GHIDDCICIFEYM-QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAY 559
G + IF M Q A + N ++ + ++ A + EE+K + ++P
Sbjct: 577 GQARETSNIFHAMKQQGFALDARAYNAVVDGFCKSGKVHKAYEILEEMK--EKCVQPTVA 634
Query: 560 TYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVI 615
TY +++ ++ + + +++E G L+ + L+ + G++D +I
Sbjct: 635 TYGAIVDGLAKIDRLDEAYMLFEEAKSKGIELNVVLYSSLIDGFGKVGRIDEAYLI 690
Score = 60.5 bits (145), Expect = 2e-07
Identities = 48/207 (23%), Positives = 87/207 (41%), Gaps = 15/207 (7%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y ++ L K R EA +F + ++ Y S+ G+ G + E I+E M
Sbjct: 636 YGAIVDGLAKIDRLDEAYMLFEEAKSK-GIELNVVLYSSLIDGFGKVGRIDEAYLILEEM 694
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
+K L P+V +N++L+A V +++ FQ M++ PN T
Sbjct: 695 MKKG--------------LTPNVYTWNSLLDALVKAEEINEALVCFQSMKEMKCPPNTYT 740
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + + + + Y+ ++ MQ+ G VP +TY M+ K G + +A +
Sbjct: 741 YSILINGLCRVQKYNKAFVFWQDMQKQGLVPNVVTYTTMISGLAKVGNITDAYSLFERFK 800
Query: 444 RRGVMGTASVYYELACCLCNCGRWQDA 470
G + A+ + L + N R +A
Sbjct: 801 ANGGIPDAASFNALIEGMSNANRAMEA 827
Score = 56.2 bits (134), Expect = 3e-06
Identities = 40/184 (21%), Positives = 79/184 (42%), Gaps = 15/184 (8%)
Query: 263 VYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVEC 322
+Y+ L+ GK R EA I M+ + P++ ++S+ L +A + E L +
Sbjct: 670 LYSSLIDGFGKVGRIDEAYLILEEMMKK-GLTPNVYTWNSLLDALVKAEEINEALVCFQS 728
Query: 323 MRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGA 382
M++ P+ Y+ ++N +++ +Q M+K L PN
Sbjct: 729 MKEMK--------------CPPNTYTYSILINGLCRVQKYNKAFVFWQDMQKQGLVPNVV 774
Query: 383 TYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
TY + + + GN + LFE+ + NG +P+A ++ ++ + EA + +
Sbjct: 775 TYTTMISGLAKVGNITDAYSLFERFKANGGIPDAASFNALIEGMSNANRAMEAYQVFEET 834
Query: 443 ERRG 446
RG
Sbjct: 835 RLRG 838
Score = 53.9 bits (128), Expect = 1e-05
Identities = 58/312 (18%), Positives = 131/312 (41%), Gaps = 25/312 (8%)
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ L+ L KA + +E IF+ M D AY+++ ++G + + I+E M
Sbjct: 566 YSILIHGLTKAGQARETSNIFHAMKQQGFAL-DARAYNAVVDGFCKSGKVHKAYEILEEM 624
Query: 324 RQK---PETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
++K P Y ++ + +E +VV+Y+++++ +
Sbjct: 625 KEKCVQPTVATYGAIVDGLAKIDRLDEAYMLFEEAKSKGIELNVVLYSSLIDGFGKVGRI 684
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
+ ++M K L PN T+ ++ ++++ + F+ M+ P TY ++
Sbjct: 685 DEAYLILEEMMKKGLTPNVYTWNSLLDALVKAEEINEALVCFQSMKEMKCPPNTYTYSIL 744
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPH 482
+ + K ++A +DM+++G++ Y + L G DA E+ K
Sbjct: 745 INGLCRVQKYNKAFVFWQDMQKQGLVPNVVTYTTMISGLAKVGNITDAYSLFERFK-ANG 803
Query: 483 AKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ-DHCAPNVGTVNTMLKVYSQNDMFSTAK 541
P +F +I + + +FE + C N+ + ++L ++++ A
Sbjct: 804 GIPDAASFNALIEGMSNANRAMEAYQVFEETRLRGCRINIKSCISLLDALNKSECLEQAA 863
Query: 542 VLFEEVK-VAKS 552
++ ++ +AKS
Sbjct: 864 IVGAVLREIAKS 875
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,081,848,692
Number of Sequences: 2790947
Number of extensions: 45586275
Number of successful extensions: 149621
Number of sequences better than 10.0: 1113
Number of HSP's better than 10.0 without gapping: 547
Number of HSP's successfully gapped in prelim test: 591
Number of HSP's that attempted gapping in prelim test: 139900
Number of HSP's gapped (non-prelim): 5188
length of query: 653
length of database: 848,049,833
effective HSP length: 134
effective length of query: 519
effective length of database: 474,062,935
effective search space: 246038663265
effective search space used: 246038663265
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC140720.4


