

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140544.4 - phase: 0 /pseudo
(305 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
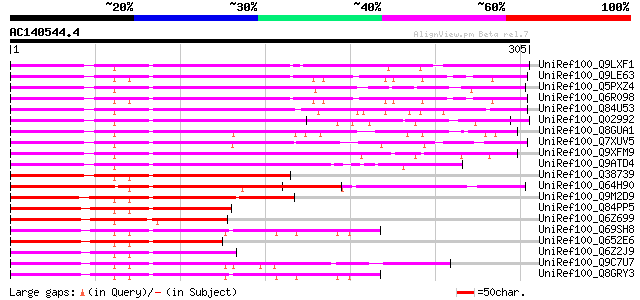
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LXF1 Myb-related protein-like [Arabidopsis thaliana] 243 5e-63
UniRef100_Q9LE63 Putative transcription factor [Arabidopsis thal... 237 3e-61
UniRef100_Q5PXZ4 MYB transcription factor MIXTA-like 2 [Antirrhi... 234 2e-60
UniRef100_Q6R098 MYB transcription factor [Arabidopsis thaliana] 233 5e-60
UniRef100_Q84U53 MYB1 [Dendrobium sp. XMW-2002-1] 223 4e-57
UniRef100_Q02992 Protein 1 [Petunia hybrida] 213 7e-54
UniRef100_Q8GUA1 MYB27 protein [Oryza sativa] 209 7e-53
UniRef100_Q7XUV5 OSJNBa0072F16.11 protein [Oryza sativa] 208 1e-52
UniRef100_Q9XFM9 Myb-related transcription factor mixta-like 1 [... 207 2e-52
UniRef100_Q9ATD4 GHMYB25 [Gossypium hirsutum] 201 2e-50
UniRef100_Q38739 Mixta protein [Antirrhinum majus] 188 1e-46
UniRef100_Q64H90 MYB transcription factor MYBML3 [Antirrhinum ma... 183 6e-45
UniRef100_Q9M2D9 Putative transcription factor [Arabidopsis thal... 146 8e-34
UniRef100_Q84PP5 Transcription factor MYB101 [Lotus japonicus] 124 2e-27
UniRef100_Q6Z699 Putative P-type R2R3 Myb protein [Oryza sativa] 122 9e-27
UniRef100_Q69SH8 Putative myb-related transcription factor [Oryz... 122 2e-26
UniRef100_Q652E6 Putative myb factor [Oryza sativa] 120 5e-26
UniRef100_Q6Z2J9 Putative Myb51 protein [Oryza sativa] 119 1e-25
UniRef100_Q9C7U7 Myb-related transcription factor, putative; 176... 118 2e-25
UniRef100_Q8GRY3 Myb13 protein [Oryza sativa] 118 2e-25
>UniRef100_Q9LXF1 Myb-related protein-like [Arabidopsis thaliana]
Length = 326
Score = 243 bits (620), Expect = 5e-63
Identities = 162/339 (47%), Positives = 193/339 (56%), Gaps = 48/339 (14%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+K+GLKKGPWT EEDQKLL+YIEEHGHGSWRSLP KA G+ R L
Sbjct: 1 MGRSPCCDKLGLKKGPWTPEEDQKLLAYIEEHGHGSWRSLPEKA-----GLHRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFNLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPE-NHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDPVTHKPKNET L SKN A LSH AQWESARLEAEARL RESKL L
Sbjct: 114 KRLVKMGIDPVTHKPKNETPLSSLGLSKNAAILSHTAQWESARLEAEARLARESKL-LHL 172
Query: 170 QHQFGSNTSVFPSQSSSSSSNQVL-NIKEEGEKEWKGYEDSTHLLEFKDLMEN------- 221
QH + + TS P + +L N + ++ + E T + F ++ E+
Sbjct: 173 QH-YQTKTSSQPHHHHGFTHKSLLPNWTTKPHEDQQQLESPTSTVSFSEMKESIPAKIEF 231
Query: 222 --SSMAFSSTLQHEMTMINVA-------------EEGFTNLLLDNSNSGDLSLSPESGGE 266
SS + + E IN EEGFT LLL G S+ G+
Sbjct: 232 VGSSTGVTLMKEPEHDWINSTMHEFETTQMGEGIEEGFTGLLL-----GGDSIDRSFSGD 286
Query: 267 CNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPSDSSM 305
N G SGG ++ EDNKNY ++I N V+ SPSDS M
Sbjct: 287 KNETAGESSGGDCNYYEDNKNYLDSIFNFVDPSPSDSPM 325
>UniRef100_Q9LE63 Putative transcription factor [Arabidopsis thaliana]
Length = 345
Score = 237 bits (604), Expect = 3e-61
Identities = 166/360 (46%), Positives = 202/360 (56%), Gaps = 74/360 (20%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+K GLKKGPWT EEDQKLL+YIEEHGHGSWRSLP KA G+ R L
Sbjct: 1 MGRSPCCDKAGLKKGPWTPEEDQKLLAYIEEHGHGSWRSLPEKA-----GLQRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFTVQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPE-NHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDPVTHK KNETL SKN A LSH AQWESARLEAEARL RESKL L
Sbjct: 114 KRLIKMGIDPVTHKHKNETLSSSTGQSKNAATLSHMAQWESARLEAEARLARESKL-LHL 172
Query: 170 QHQFGSNT---SVFPSQ---SSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLM---- 219
QH +N S P Q + +S+N + G+++ E T + F + +
Sbjct: 173 QHYQNNNNLNKSAAPQQHCFTQKTSTNWTKPNQGNGDQQ---LESPTSTVTFSENLLMPL 229
Query: 220 ---------------ENSSM---AFSSTLQHEMTMINVAE---------------EGFTN 246
E+S+M A SS+ +++++ E EGFT+
Sbjct: 230 GIPTDSSRNRNNNNNESSAMIELAVSSSTSSDVSLVKEHEHDWIRQINCGSGGIGEGFTS 289
Query: 247 LLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFN--EDNKNYWNNILNLVNSSPSDSS 304
LL+ +S L P E +G G S++N EDNKNYWN+ILNLV+SSPSDS+
Sbjct: 290 LLIGDSVGRGL---PTGKNEAT----AGVGNESEYNYYEDNKNYWNSILNLVDSSPSDSA 342
>UniRef100_Q5PXZ4 MYB transcription factor MIXTA-like 2 [Antirrhinum majus]
Length = 352
Score = 234 bits (597), Expect = 2e-60
Identities = 160/371 (43%), Positives = 201/371 (54%), Gaps = 90/371 (24%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+KVGLKKGPWT EEDQKLL+YIEEHGHGSWR+LP +A G+ R L
Sbjct: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPARA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+I L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKISLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLP-ENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDPVTHKPK++TL+ + SKN ANLSH AQWESARLEAEARLVR+SKL+
Sbjct: 114 KRLAKMGIDPVTHKPKSDTLMSNDGQSKNAANLSHMAQWESARLEAEARLVRQSKLQPPS 173
Query: 170 QHQFGSNTS---------------------------------------------VFPSQS 184
+ F ++TS P+ +
Sbjct: 174 ANSFQASTSNPVQKSMGQARCLDVLKAWNGVVWGKNDEAAGVSVTVTTGIVGELGSPTST 233
Query: 185 SSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAEEGF 244
SS++ +KEE E+EWK + E +D + N++ F+S + E +N + E
Sbjct: 234 LSSAAGTNGIMKEESEEEWK------QMPENRDEIGNAT-PFTSNWESEQVALN-SSEAC 285
Query: 245 TNLLLDNSNSGDLSLSPESGGECNTC------------DGSGSGGGSDFNEDNKNYWNNI 292
++ G ++ EC C G G SD+ EDNKNY N+I
Sbjct: 286 SSREFRGKLYGLVT-------ECFLCRWRGMRIVGYTNSGGNDVGVSDYYEDNKNYGNSI 338
Query: 293 LNLVNSSPSDS 303
LNLVNSSPS S
Sbjct: 339 LNLVNSSPSHS 349
>UniRef100_Q6R098 MYB transcription factor [Arabidopsis thaliana]
Length = 345
Score = 233 bits (594), Expect = 5e-60
Identities = 165/360 (45%), Positives = 201/360 (55%), Gaps = 74/360 (20%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSP C+K GLKKGPWT EEDQKLL+YIEEHGHGSWRSLP KA G+ R L
Sbjct: 1 MGRSPSCDKAGLKKGPWTPEEDQKLLAYIEEHGHGSWRSLPEKA-----GLQRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFTVQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPE-NHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDPVTHK KNETL SKN A LSH AQWESARLEAEARL RESKL L
Sbjct: 114 KRLIKMGIDPVTHKHKNETLSSSTGQSKNAATLSHMAQWESARLEAEARLARESKL-LHL 172
Query: 170 QHQFGSNT---SVFPSQ---SSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLM---- 219
QH +N S P Q + +S+N + G+++ E T + F + +
Sbjct: 173 QHYQNNNNLNKSAAPQQHCFTQKTSTNWTKPNQGNGDQQ---LESPTSTVTFSENLLMPL 229
Query: 220 ---------------ENSSM---AFSSTLQHEMTMINVAE---------------EGFTN 246
E+S+M A SS+ +++++ E EGFT+
Sbjct: 230 GIPTDSSRNRNNNNNESSAMIELAVSSSTSSDVSLVKEHEHDWIRQINCGSGGIGEGFTS 289
Query: 247 LLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFN--EDNKNYWNNILNLVNSSPSDSS 304
LL+ +S L P E +G G S++N EDNKNYWN+ILNLV+SSPSDS+
Sbjct: 290 LLIGDSVGRGL---PTGKNEAT----AGVGNESEYNYYEDNKNYWNSILNLVDSSPSDSA 342
>UniRef100_Q84U53 MYB1 [Dendrobium sp. XMW-2002-1]
Length = 367
Score = 223 bits (569), Expect = 4e-57
Identities = 153/374 (40%), Positives = 196/374 (51%), Gaps = 82/374 (21%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+KVGLKKGPWT EEDQKLL+YIE+HGHGSWR+LP KA G+ R L
Sbjct: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIEKHGHGSWRALPNKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ ++ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFSMQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETL-LPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDP+THKPK++ L + HSK ANL+H AQWESARLEAEARLVRESKLRS
Sbjct: 114 KRLAKMGIDPITHKPKSDNLSSADGHSKCTANLNHMAQWESARLEAEARLVRESKLRSS- 172
Query: 170 QHQFGSNTSVF----------PSQSSSSSSNQVLNIKEEGEKEW----------KGY--- 206
NT+ P+ + +++ L++ + W +GY
Sbjct: 173 --SISXNTTTIFTSKNQPLLPPAVPMTPAASPCLDVLSAWQGAWTKPAANNSQGRGYTID 230
Query: 207 -EDSTHLLEFKDLM-----ENSSMAFSSTLQHE---------MTMINVA----------- 240
E T L F D M +A ST + + +M A
Sbjct: 231 LESPTSTLSFSDNMMAPTISGMGLAEDSTREDQEWKCLRKTGFSMHTAAPFVSTEASSWL 290
Query: 241 --------EEGFTNLLLDNSN--SGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWN 290
GFT +L D +N + + + EC +C G + ++NKNYW+
Sbjct: 291 SESSGGDFAAGFTGMLSDKANEQNSEDGCNDSDNAECGSCVDVEESEGEE--DENKNYWS 348
Query: 291 NILNLVNSSPSDSS 304
+ILNLVNSS +S
Sbjct: 349 SILNLVNSSSPPNS 362
>UniRef100_Q02992 Protein 1 [Petunia hybrida]
Length = 421
Score = 213 bits (541), Expect = 7e-54
Identities = 142/317 (44%), Positives = 177/317 (55%), Gaps = 34/317 (10%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+KVGLKKGPWT EEDQKLL+YIEEHGHGSWR+LP KA G+ R L
Sbjct: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPAKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFTLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLP-ENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRL KMGIDPVTHKPKN+ LL + SKN ANLSH AQWESARLEAEARLVR+SKLRS
Sbjct: 114 KRLVKMGIDPVTHKPKNDALLSHDGQSKNAANLSHMAQWESARLEAEARLVRQSKLRSNS 173
Query: 170 QHQFGSNTSVFPSQSSSSSSNQ--VLNIKEEGE-------KEWKGYEDSTHLLEFKDLME 220
++ +F S + SS ++ V K G K W G D++
Sbjct: 174 FQNPLASHELFTSPTPSSPLHKPIVTPTKAPGSPRCLDVLKAWNG----VWTKPMNDVLH 229
Query: 221 -NSSMAFSSTLQHEMTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDG--SGSGG 277
+ S + S+T+ +++ T +N+ + E+ G SGS
Sbjct: 230 ADGSTSASATVSVNALGLDLESPTSTLSYFENAQHISTGMIQENSTSLFEFVGNSSGSSE 289
Query: 278 GSDFNEDNKNYWNNILN 294
G NE+++ W N
Sbjct: 290 GGIMNEESEEDWKGFGN 306
Score = 90.1 bits (222), Expect = 7e-17
Identities = 62/154 (40%), Positives = 88/154 (56%), Gaps = 29/154 (18%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDST--HLLEFKDLMENSSMAFSSTLQH 232
++TS+F +SS S++ + EE E++WKG+ +S+ HL E+KD + +SM+ +STLQ
Sbjct: 273 NSTSLFEFVGNSSGSSEGGIMNEESEEDWKGFGNSSTGHLPEYKDGINENSMSLTSTLQ- 331
Query: 233 EMTMI--------------------NVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDG 272
++TM N E FT+LLL S G LS G ++ +G
Sbjct: 332 DLTMPMDTTWTAESLRSNAEDISHGNNFVETFTDLLLSTSGDGGLS-----GNGTDSDNG 386
Query: 273 SGSGGG-SDFNEDNKNYWNNILNLVNSSPSDSSM 305
GSG S+ DNKNYWN+I NLVNSSPSDS+M
Sbjct: 387 GGSGNDPSETCGDNKNYWNSIFNLVNSSPSDSAM 420
>UniRef100_Q8GUA1 MYB27 protein [Oryza sativa]
Length = 357
Score = 209 bits (532), Expect = 7e-53
Identities = 160/373 (42%), Positives = 188/373 (49%), Gaps = 101/373 (27%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK+GLKKGPWT EEDQKLL+YIEEHGHGSWR+LP+KA G+ R L
Sbjct: 1 MGRSPCCEKIGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPSKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFSLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETL---LPENHSKNVANLSHKAQWESARLEAEARLVRESKLR- 166
KRL KMGIDPVTHK N TL + +K A+LSH AQWESARLEAEARL RESK+R
Sbjct: 114 KRLAKMGIDPVTHKAINGTLNNTAGDKSAKVTASLSHMAQWESARLEAEARLARESKMRI 173
Query: 167 -----SQLQHQ-----------------------------------FGSNTSVFP----- 181
S+L Q GSN S+ P
Sbjct: 174 AASTPSKLHAQSTNPPASTPSPCFDVLNAWQSAKIDLESPTSTLTFAGSNASMLPFSTTT 233
Query: 182 ----SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQH----- 232
S+S+S+ Q + E E EWK F + M T +H
Sbjct: 234 ALELSESNSNVWQQRSDELEGEESEWK----------FVSKQQLQGMHGKETEEHFIGCE 283
Query: 233 EMTMINVAE--EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGG---SDFNED--N 285
E A GFT +LLD SN D S EC D S +G S +ED N
Sbjct: 284 ESWFPGTANIGAGFTGMLLDGSNMHDTS-------EC--WDESSNGQDEQRSQVSEDAEN 334
Query: 286 KNYWNNILNLVNS 298
KNYWN I ++VNS
Sbjct: 335 KNYWNGIFSMVNS 347
>UniRef100_Q7XUV5 OSJNBa0072F16.11 protein [Oryza sativa]
Length = 345
Score = 208 bits (530), Expect = 1e-52
Identities = 145/355 (40%), Positives = 182/355 (50%), Gaps = 78/355 (21%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK GLKKGPWT EEDQKLL+YIE+HGHG WRSLP+KA G+ R L
Sbjct: 1 MGRSPCCEKEGLKKGPWTPEEDQKLLAYIEQHGHGCWRSLPSKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFSLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNET-------------LLPENHSKNVANLSHKAQWESARLEAEA 157
KRL KMGIDPVTHKP+++ H+K A+LSH AQWESARLEAEA
Sbjct: 114 KRLAKMGIDPVTHKPRSDVAGAGGGGGGAAGGAAGAQHAKAAAHLSHTAQWESARLEAEA 173
Query: 158 RLVRESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKD 217
RL RE+KLR+ L P+ +S+++ + G + T L F +
Sbjct: 174 RLAREAKLRA-LAASATPGAPHLPAPPASAAAAAAAH----------GLDSPTSTLSFSE 222
Query: 218 ------LMENSSMAFSSTLQHEMTMINVAEE----------------------GFTNLLL 249
++E A ++ + M + +E GFT LLL
Sbjct: 223 SAVLATVLEAHGAAAAAAARAAMQPMQAYDEACKDQHWGDVDAADVGFPGAGAGFTGLLL 282
Query: 250 DNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPSDSS 304
+ G L+ P G DG E+ KNYWN+ILNLVNSS + S
Sbjct: 283 E----GSLNQIPRPAGRDAEADGE-----FQETEEEKNYWNSILNLVNSSSAPMS 328
>UniRef100_Q9XFM9 Myb-related transcription factor mixta-like 1 [Antirrhinum majus]
Length = 359
Score = 207 bits (528), Expect = 2e-52
Identities = 149/362 (41%), Positives = 184/362 (50%), Gaps = 74/362 (20%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+KV LK+GPWT EEDQKLLSYI+EHGHGSWR+LP+KA G+ R L
Sbjct: 1 MGRSPCCDKVSLKRGPWTPEEDQKLLSYIQEHGHGSWRALPSKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWSNYLRPDIKRGKFSLQEEQAIIQLHAFLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQ 170
KRLTKMGIDP+THKPK+ +L K VANLSH AQWESARL+AEARLVRES+L S
Sbjct: 114 KRLTKMGIDPMTHKPKSHDVLGCGQPKVVANLSHMAQWESARLQAEARLVRESRLVSHHY 173
Query: 171 HQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKG-------------------YEDSTH 211
H N + + + VL + G E T
Sbjct: 174 HSQLLNRATAITHPTLPPCLDVLKVWHGAWTTRPGKDIITSAMFNGFFASNNGNLESPTS 233
Query: 212 LLEFKDLMENSSMAFSSTLQHEMTMI----------NVAEEGFTNLLLDNSNSGDLSLSP 261
+L+ D M N+S S L HE I NV E+ + ++DNSN + + P
Sbjct: 234 ILKSSDNMLNAST--SVGLLHENPFITDMSYVGKPSNVYED-WVKGIMDNSNELNKIIEP 290
Query: 262 ESG------------------GECNTCDGSGSGGGSD-------FNEDNKNYWNNILNLV 296
G N + G D F E++ NYW N+LN+V
Sbjct: 291 IDSTHYVAHDDDSIGFPGFMEGSTNLVTSTVRTNGPDDDNVVGVFEENDINYWRNVLNVV 350
Query: 297 NS 298
NS
Sbjct: 351 NS 352
>UniRef100_Q9ATD4 GHMYB25 [Gossypium hirsutum]
Length = 309
Score = 201 bits (511), Expect = 2e-50
Identities = 132/279 (47%), Positives = 167/279 (59%), Gaps = 28/279 (10%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEKVGLKKGPWT EEDQKLL+YIE+HGHGSWR+LP KA G+ R L
Sbjct: 1 MGRSPCCEKVGLKKGPWTPEEDQKLLAYIEQHGHGSWRALPLKA-----GLQRCGKSCRL 55
Query: 61 ---------II*GRI-LRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
I G+ L+ E + + + L N + A AT LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWINYLRPDIKRGKFSLQEEQTIIQLHALLGN--RWSAIATHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETL-LPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQL 169
KRLTKMGIDPVTHKPK + L + + ANLSH AQWESARLEAEARLVRESKL
Sbjct: 114 KRLTKMGIDPVTHKPKTDALGSTTGNPIDAANLSHMAQWESARLEAEARLVRESKLVPSN 173
Query: 170 QHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSST 229
Q T+V PS + ++ Q L++ K W+G L F ++ N+ + +ST
Sbjct: 174 PPQSNHFTAVAPSPTPATRP-QCLDVL----KAWQGV--VCGLFTF-NMDNNNLQSPTST 225
Query: 230 L--QHEMTMINVAEEGFTNLLLDNSNSGDLSLSPESGGE 266
L T + ++ N + + + + S++P G+
Sbjct: 226 LNFMENTTTLPMSSSSSVNGMFNENFGWNSSINPCESGD 264
>UniRef100_Q38739 Mixta protein [Antirrhinum majus]
Length = 321
Score = 188 bits (478), Expect = 1e-46
Identities = 104/175 (59%), Positives = 121/175 (68%), Gaps = 17/175 (9%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M RSPCC+KVG+KKGPWT +EDQKLL+YIEEHGHGSWRSLP KA G+ R L
Sbjct: 1 MVRSPCCDKVGVKKGPWTVDEDQKLLAYIEEHGHGSWRSLPLKA-----GLQRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + + + L N + A A+ LPKRTDNEIKNYWNTHLK
Sbjct: 56 RWANYLRPDIKRGPFSLQEEQTIIQLHALLGN--RWSAIASHLPKRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKL 165
KRLT+MGIDPVTHKP +L K+VANL+H AQWESARL+AE RLVRES+L
Sbjct: 114 KRLTRMGIDPVTHKPHTHNILGHGQPKDVANLNHIAQWESARLQAERRLVRESRL 168
>UniRef100_Q64H90 MYB transcription factor MYBML3 [Antirrhinum majus]
Length = 219
Score = 183 bits (464), Expect = 6e-45
Identities = 105/205 (51%), Positives = 132/205 (64%), Gaps = 13/205 (6%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSP C+K GLK+GPWT EEDQKLL+ I+EHGHG+WR+LP KA + G + +
Sbjct: 1 MGRSPFCDKTGLKRGPWTPEEDQKLLACIQEHGHGNWRALPAKAGLERCGKSCRLRWNNY 60
Query: 61 II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLT 114
+ I RG E + + + L N + A AT L +RTDNEIKNYWNTH+KK+L
Sbjct: 61 LR-PDIKRGKFSSQEEQTIIQLHALLGN--RWSAIATHLSRRTDNEIKNYWNTHIKKKLA 117
Query: 115 KMGIDPVTHKPKNETLLPENH----SKNVANLSHKAQWESARLEAEARLVRESKLRSQLQ 170
KMGIDPVTHKP+ + L N+ SKN AN+SH AQWESARLEAEARL R+SKL++ +
Sbjct: 118 KMGIDPVTHKPQRDHALSSNNAHVQSKNAANMSHLAQWESARLEAEARLARQSKLQANSE 177
Query: 171 HQFGSNTSVFPSQSSSSSSNQVLNI 195
GS+T S + N NI
Sbjct: 178 CGGGSDTGGGGSDCCGDNKNYWDNI 202
Score = 54.3 bits (129), Expect = 4e-06
Identities = 44/148 (29%), Positives = 66/148 (43%), Gaps = 12/148 (8%)
Query: 161 RESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVLN-----IKEEGEKEWKGYEDSTHLLEF 215
+E + QL G+ S + S + N++ N IK++ K G + TH +
Sbjct: 73 QEEQTIIQLHALLGNRWSAIATHLSRRTDNEIKNYWNTHIKKKLAK--MGIDPVTHKPQR 130
Query: 216 KDLMENSSMAFSSTLQHEMTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGS 275
+ +++ S M+ + E + L + E GG +T
Sbjct: 131 DHALSSNNAHVQSKNAANMSHLAQWESARLEAEARLARQSKLQANSECGGGSDT-----G 185
Query: 276 GGGSDFNEDNKNYWNNILNLVNSSPSDS 303
GGGSD DNKNYW+NIL+LVN SPSDS
Sbjct: 186 GGGSDCCGDNKNYWDNILDLVNFSPSDS 213
>UniRef100_Q9M2D9 Putative transcription factor [Arabidopsis thaliana]
Length = 299
Score = 146 bits (368), Expect = 8e-34
Identities = 88/177 (49%), Positives = 111/177 (61%), Gaps = 18/177 (10%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCC+K+GLKKGPWT EED+ L+++I+++GHGSWR+LP L G+ R L
Sbjct: 1 MGRTPCCDKIGLKKGPWTPEEDEVLVAHIKKNGHGSWRTLP-----KLAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E LV + + L N + A A +LP RTDNEIKN WNTHLK
Sbjct: 56 RWTNYLRPDIKRGPFTADEEKLVIQLHAILGN--RWAAIAAQLPGRTDNEIKNLWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRS 167
KRL MG+DP TH+P L + + + H AQWESAR+EAEARL RES L S
Sbjct: 114 KRLLSMGLDPRTHEPLPSYGLAK-QAPSSPTTRHMAQWESARVEAEARLSRESMLFS 169
>UniRef100_Q84PP5 Transcription factor MYB101 [Lotus japonicus]
Length = 348
Score = 124 bits (312), Expect = 2e-27
Identities = 69/140 (49%), Positives = 89/140 (63%), Gaps = 17/140 (12%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCC+++GLKKGPWT EED L+ YI++HGHGSWR+LP L G+ R L
Sbjct: 1 MGRTPCCDEIGLKKGPWTPEEDAILVDYIQKHGHGSWRALPK-----LAGLNRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E L+ + L N + A A LP RTDNEIKN+WNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFTEEEEQLIINLHAVLGN--KWSAIAGHLPGRTDNEIKNFWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNETL 130
K+L +MG+DPVTH+P+ + L
Sbjct: 114 KKLLQMGLDPVTHRPRLDHL 133
>UniRef100_Q6Z699 Putative P-type R2R3 Myb protein [Oryza sativa]
Length = 377
Score = 122 bits (307), Expect = 9e-27
Identities = 68/138 (49%), Positives = 88/138 (63%), Gaps = 17/138 (12%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC++ GLKKGPWT+EED+KL+ YI+++GHGSWR+LP L G+ R L
Sbjct: 1 MGRSPCCDENGLKKGPWTTEEDEKLMEYIQKNGHGSWRALPK-----LAGLNRCGKSCRL 55
Query: 61 ----II*GRILRGESLVCKKNKPLFNFMH------F*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG+ +K+ L +H + A A LP RTDNEIKNYWNTHLK
Sbjct: 56 RWTNYLRPDIKRGKFTSAEKDTILQ--LHAVLGNKWSAIAKHLPGRTDNEIKNYWNTHLK 113
Query: 111 KRLTKMGIDPVTHKPKNE 128
K L + GIDP TH+P+ +
Sbjct: 114 KDLIQKGIDPTTHRPRTD 131
>UniRef100_Q69SH8 Putative myb-related transcription factor [Oryza sativa]
Length = 285
Score = 122 bits (305), Expect = 2e-26
Identities = 88/246 (35%), Positives = 130/246 (52%), Gaps = 37/246 (15%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+K+G+K+GPWT+EED+KL+S+I +GH WR++P L G+ R L
Sbjct: 1 MGRQPCCDKLGVKRGPWTAEEDKKLMSFILTNGHCCWRAVPK-----LAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E LV + L N + A +LP RTDNEIKN+WNTH+K
Sbjct: 56 RWTNYLRPDLKRGLLTDAEEQLVIDLHAKLGN--RWSKIAAKLPGRTDNEIKNHWNTHIK 113
Query: 111 KRLTKMGIDPVTHKP---KNETLLPENHSKNVANLSHKAQWESARLEA---EARLVRESK 164
K+L KMGIDPVTH+P K E+ P S++ E+ R ++ + +VR+
Sbjct: 114 KKLIKMGIDPVTHEPLDRKQES--PATTSQSTVTAESSKSGEATRQQSRQLDDAVVRDMS 171
Query: 165 LRS--QLQHQFGSNTSVFPSQSSSSSSNQ-----VLNIKEE-----GEKEWKGYEDSTHL 212
+ + + +NT+ SSSSSS+ V + EE G++ W + S +
Sbjct: 172 VSAGGDSPPESSTNTASTAGGSSSSSSSHHQDPLVKWLLEEDLLPTGDEPWLNFTASNDV 231
Query: 213 LEFKDL 218
EF +
Sbjct: 232 DEFSSI 237
>UniRef100_Q652E6 Putative myb factor [Oryza sativa]
Length = 267
Score = 120 bits (301), Expect = 5e-26
Identities = 68/135 (50%), Positives = 84/135 (61%), Gaps = 17/135 (12%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+KVGLKKGPWT+EEDQKL++++ HGH WR +P L G+ R L
Sbjct: 1 MGRQPCCDKVGLKKGPWTAEEDQKLVAFLLTHGHCCWRVVPK-----LAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E LV + L N + A RLP RTDNEIKN+WNTH+K
Sbjct: 56 RWTNYLRPDLKRGLLSDDEERLVIDLHAQLGN--RWSKIAARLPGRTDNEIKNHWNTHIK 113
Query: 111 KRLTKMGIDPVTHKP 125
K+L KMGIDPVTH+P
Sbjct: 114 KKLRKMGIDPVTHQP 128
>UniRef100_Q6Z2J9 Putative Myb51 protein [Oryza sativa]
Length = 255
Score = 119 bits (298), Expect = 1e-25
Identities = 67/143 (46%), Positives = 87/143 (59%), Gaps = 17/143 (11%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCCEKVGLKKGPWT++EDQKL++++ +GH WR +P L G+ R L
Sbjct: 1 MGRQPCCEKVGLKKGPWTADEDQKLVTFLLSNGHCCWRLVPK-----LAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E LV ++ L N + A RLP RTDNEIKN+WNTH+K
Sbjct: 56 RWTNYLRPDLKRGLLSEEEEKLVIDLHEQLGN--RWSKIAARLPGRTDNEIKNHWNTHIK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPE 133
K+L KMG+DPVTH+P P+
Sbjct: 114 KKLKKMGLDPVTHRPVMSLAQPD 136
>UniRef100_Q9C7U7 Myb-related transcription factor, putative; 17635-18559
[Arabidopsis thaliana]
Length = 282
Score = 118 bits (296), Expect = 2e-25
Identities = 95/298 (31%), Positives = 145/298 (47%), Gaps = 56/298 (18%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+KVGLKKGPWT+EED+KL+++I +G WR++P L G+ R L
Sbjct: 1 MGRQPCCDKVGLKKGPWTAEEDRKLINFILTNGQCCWRAVPK-----LSGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E +V + L N + A+ LP RTDNEIKN+WNTH+K
Sbjct: 56 RWTNYLRPDLKRGLLSDYEEKMVIDLHSQLGN--RWSKIASHLPGRTDNEIKNHWNTHIK 113
Query: 111 KRLTKMGIDPVTHKP---------------KNETL--------LPENHSKNVANLSHKA- 146
K+L KMGIDP+THKP +N T+ EN+ KN++ L
Sbjct: 114 KKLRKMGIDPLTHKPLSIVEKEDEEPLKKLQNNTVPFQETMERPLENNIKNISRLEESLG 173
Query: 147 --QWESARLE---AEARLVRESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEK 201
Q+ LE + L+ L + S+++ S SS+ SS L + E
Sbjct: 174 DDQFMEINLEYGVEDVPLIETESLDLICSNSTMSSSTSTSSHSSNDSS--FLKDLQFPEF 231
Query: 202 EWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAEEGFTNLLLDNSNSGDLSL 259
EW Y +S + +N++ + + M++ +++ +LLL++ +S L
Sbjct: 232 EWSDYGNSNN--------DNNNGVDNIIENNMMSLWEISDFSSLDLLLNDESSSTFGL 281
>UniRef100_Q8GRY3 Myb13 protein [Oryza sativa]
Length = 285
Score = 118 bits (295), Expect = 2e-25
Identities = 87/246 (35%), Positives = 129/246 (52%), Gaps = 37/246 (15%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PC +K+G+K+GPWT+EED+KL+S+I +GH WR++P L G+ R L
Sbjct: 1 MGRQPCSDKLGVKRGPWTAEEDKKLMSFILTNGHCCWRAVPK-----LAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E LV + L N + A +LP RTDNEIKN+WNTH+K
Sbjct: 56 RWTNYLRPDLKRGLLTDAEEQLVIDLHAKLGN--RWSKIAAKLPGRTDNEIKNHWNTHIK 113
Query: 111 KRLTKMGIDPVTHKP---KNETLLPENHSKNVANLSHKAQWESARLEA---EARLVRESK 164
K+L KMGIDPVTH+P K E+ P S++ E+ R ++ + +VR+
Sbjct: 114 KKLIKMGIDPVTHEPLDRKQES--PATTSQSTVTAESSKSGEATRQQSRQLDDAVVRDMS 171
Query: 165 LRS--QLQHQFGSNTSVFPSQSSSSSSNQ-----VLNIKEE-----GEKEWKGYEDSTHL 212
+ + + +NT+ SSSSSS+ V + EE G++ W + S +
Sbjct: 172 VSAGGDSPPESSTNTASTAGGSSSSSSSHHQDPLVKWLLEEDLLPTGDEPWLNFTASNDV 231
Query: 213 LEFKDL 218
EF +
Sbjct: 232 DEFSSI 237
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.131 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 520,602,974
Number of Sequences: 2790947
Number of extensions: 21643554
Number of successful extensions: 95183
Number of sequences better than 10.0: 993
Number of HSP's better than 10.0 without gapping: 822
Number of HSP's successfully gapped in prelim test: 178
Number of HSP's that attempted gapping in prelim test: 92166
Number of HSP's gapped (non-prelim): 2451
length of query: 305
length of database: 848,049,833
effective HSP length: 127
effective length of query: 178
effective length of database: 493,599,564
effective search space: 87860722392
effective search space used: 87860722392
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC140544.4


