

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
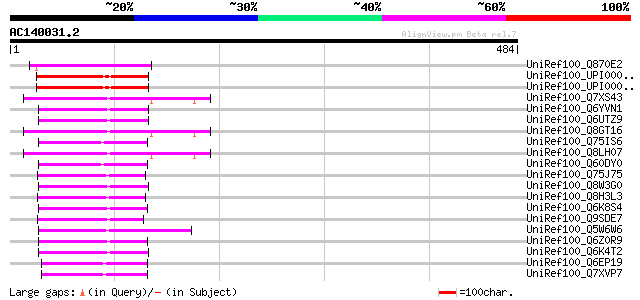
Sequences producing significant alignments: (bits) Value
UniRef100_Q870E2 Transposase [Fusarium oxysporum f. sp. melonis] 94 9e-18
UniRef100_UPI000042F827 UPI000042F827 UniRef100 entry 90 2e-16
UniRef100_UPI000042CB10 UPI000042CB10 UniRef100 entry 90 2e-16
UniRef100_Q7XS43 OSJNBa0074L08.5 protein [Oryza sativa] 69 4e-10
UniRef100_Q6YVN1 Putative far-red impaired response protein [Ory... 68 5e-10
UniRef100_Q6UTZ9 Putative transposase [Oryza sativa] 68 5e-10
UniRef100_Q8GT16 Putative far-red impaired response protein [Ory... 67 1e-09
UniRef100_Q75IS6 Putative Mutator-like transposase [Oryza sativa] 67 1e-09
UniRef100_Q8LH07 Far-red impaired response-like protein [Oryza s... 67 1e-09
UniRef100_Q60DY0 Hypothetical protein OSJNBa0090H02.8 [Oryza sat... 67 1e-09
UniRef100_Q75J75 Putative far-red impaired response protein [Ory... 67 1e-09
UniRef100_Q8W3G0 Putative far-red impaired response protein [Ory... 66 3e-09
UniRef100_Q8H3L3 Putative far-red impaired response protein [Ory... 65 4e-09
UniRef100_Q6K8S4 Putative far-red impaired response protein [Ory... 65 4e-09
UniRef100_Q9SDE7 Similar to Arabidopsis thaliana DNA chromosome ... 65 6e-09
UniRef100_Q5W6W6 Hypothetical protein OSJNBa0065C11.11 [Oryza sa... 64 7e-09
UniRef100_Q6Z0R9 Putative far-red impaired response protein [Ory... 64 1e-08
UniRef100_Q6K4T2 Putative far-red impaired response protein [Ory... 63 2e-08
UniRef100_Q6EP19 Putative far-red impaired response protein [Ory... 63 2e-08
UniRef100_Q7XVP7 OSJNBa0050F15.2 protein [Oryza sativa] 63 2e-08
>UniRef100_Q870E2 Transposase [Fusarium oxysporum f. sp. melonis]
Length = 836
Score = 94.0 bits (232), Expect = 9e-18
Identities = 45/119 (37%), Positives = 71/119 (58%), Gaps = 3/119 (2%)
Query: 20 LIQHP--LSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEEN 77
L HP L + +P VL++DSTY+TN +KMP+ +IVGV + T+ + F F++ E+E +
Sbjct: 213 LFAHPKSLEYLKTYPEVLILDSTYKTNRFKMPLLDIVGVDACQRTFCIAFAFLSGEEEGD 272
Query: 78 FVWVLQMMCKLLTS-RMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRC 135
F W LQ + + + +P V++TD+ MNAVS+ P S C +H+ K V+ C
Sbjct: 273 FTWALQALRSVYEDHNIGLPSVILTDRCLACMNAVSSCFPGSALFLCLWHINKAVQSYC 331
Score = 43.9 bits (102), Expect = 0.010
Identities = 22/70 (31%), Positives = 39/70 (55%), Gaps = 4/70 (5%)
Query: 284 YSWLGGYMSRYALGNIVLEEERCRGTLCMDKEICGCV*RKSYGLPCACFVATKIRDDKPI 343
YS + G++S AL + + ER L + +C V ++ GLPCA + ++ ++P+
Sbjct: 480 YSNIRGWISHEALRKVETQRER----LLKEVPVCTGVFTRTLGLPCAHSLQPLLKQNQPL 535
Query: 344 LLDEIHLHWN 353
LL+ H HW+
Sbjct: 536 LLNHFHSHWH 545
>UniRef100_UPI000042F827 UPI000042F827 UniRef100 entry
Length = 580
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>UniRef100_UPI000042CB10 UPI000042CB10 UniRef100 entry
Length = 932
Score = 89.7 bits (221), Expect = 2e-16
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 2/107 (1%)
Query: 26 SFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMM 85
+ + FP VL D TY+TN+Y+MPM IVG TST +TY+ G + E + L
Sbjct: 315 ALIHRFPEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSF 374
Query: 86 CKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+L+ + +V KVV+TD+D L+NA+ +VLP+++ +C++H+Q+NVK
Sbjct: 375 FELV-GKPDV-KVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>UniRef100_Q7XS43 OSJNBa0074L08.5 protein [Oryza sativa]
Length = 1061
Score = 68.6 bits (166), Expect = 4e-10
Identities = 46/184 (25%), Positives = 81/184 (44%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 507 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 566
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 567 TTETFKWVFETFLTAMGGKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRE 624
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + LKN Y+++ + K F
Sbjct: 625 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLKNIKYLEIMWRTRKNF 684
Query: 188 VMIF 191
+ ++
Sbjct: 685 IPVY 688
>UniRef100_Q6YVN1 Putative far-red impaired response protein [Oryza sativa]
Length = 1132
Score = 68.2 bits (165), Expect = 5e-10
Identities = 32/105 (30%), Positives = 52/105 (49%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ + + D+TY+TN Y MP VGV T G F+ E E F WV +
Sbjct: 624 YEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGTTCLFGCAFLGDETAETFKWVFETFAT 683
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD+D + +A++ V + NC FH++KN +
Sbjct: 684 AMGGKH--PKTIITDQDNAMRSAIAQVFQNTKHRNCLFHIKKNCR 726
>UniRef100_Q6UTZ9 Putative transposase [Oryza sativa]
Length = 1037
Score = 68.2 bits (165), Expect = 5e-10
Identities = 32/105 (30%), Positives = 52/105 (49%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ + + D+TY+TN Y MP VGV T G F+ E E F WV +
Sbjct: 529 YEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGTTCLFGCAFLGDETAETFKWVFETFAT 588
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD+D + +A++ V + NC FH++KN +
Sbjct: 589 AMGGKH--PKTIITDQDNAMRSAIAQVFQNTKHRNCLFHIKKNCR 631
>UniRef100_Q8GT16 Putative far-red impaired response protein [Oryza sativa]
Length = 1114
Score = 67.0 bits (162), Expect = 1e-09
Identities = 45/184 (24%), Positives = 81/184 (43%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 560 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 619
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 620 TTETFKWVFETFLTAMGGKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRE 677
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + L+N Y+++ + K F
Sbjct: 678 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLENIKYLEIMWRTRKNF 737
Query: 188 VMIF 191
+ ++
Sbjct: 738 IPVY 741
>UniRef100_Q75IS6 Putative Mutator-like transposase [Oryza sativa]
Length = 1510
Score = 66.6 bits (161), Expect = 1e-09
Identities = 33/104 (31%), Positives = 56/104 (53%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F VL +DSTY TN Y M GV + +G F+ EK E+FVW+ Q K
Sbjct: 930 YSVFGDVLSVDSTYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLK 989
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ P++++TD+D ++ A++ +LP + C +H+ + V
Sbjct: 990 --ATGGVAPRLIITDEDASMKAAIAQILPNTVHRLCMWHIMEKV 1031
>UniRef100_Q8LH07 Far-red impaired response-like protein [Oryza sativa]
Length = 772
Score = 66.6 bits (161), Expect = 1e-09
Identities = 45/184 (24%), Positives = 81/184 (43%), Gaps = 8/184 (4%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHE 73
+K+ F + + + D+TY+TN Y MP IVGVT F+ E
Sbjct: 245 VKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDE 304
Query: 74 KEENFVWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKV 133
E F WV + + + P+ ++TD+D + A+ V P S NC FH+ K +
Sbjct: 305 TTETFKWVFETFLTAMGRKH--PETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKYRE 362
Query: 134 R---CILDCRYPMAKKDGKEVKHGDVVK-KIMRVW*VIIESHPLKN--YIQMH*WSSKMF 187
R D R + ++ H + + + +W +I + L+N Y+++ + K F
Sbjct: 363 RSGNTFSDKRRKDLYAEFNDIVHNSLTRAEFESLWLQMIAQYNLENIKYLEIMWRTRKNF 422
Query: 188 VMIF 191
+ ++
Sbjct: 423 IPVY 426
>UniRef100_Q60DY0 Hypothetical protein OSJNBa0090H02.8 [Oryza sativa]
Length = 896
Score = 66.6 bits (161), Expect = 1e-09
Identities = 33/104 (31%), Positives = 56/104 (53%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F VL +DSTY TN Y M GV + +G F+ EK E+FVW+ Q K
Sbjct: 321 YSVFGDVLSVDSTYSTNQYNMKFVPFTGVNHHLQSVFLGASFLADEKIESFVWLFQTFLK 380
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ P++++TD+D ++ A++ +LP + C +H+ + V
Sbjct: 381 --ATGGVAPRLIITDEDASMKAAIAQILPNTVHRLCMWHIMEKV 422
>UniRef100_Q75J75 Putative far-red impaired response protein [Oryza sativa]
Length = 888
Score = 66.6 bits (161), Expect = 1e-09
Identities = 30/103 (29%), Positives = 55/103 (53%), Gaps = 2/103 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
++ + V+ D+TY TN Y +P VG++ T G F+ E E F W+ +
Sbjct: 528 WYEKYGDVVSFDTTYFTNRYNLPFAPFVGISGHGNTIVFGCAFLHDETSETFKWMFRTFL 587
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQK 129
K ++ + PK+++TD+D + +A++ V + NC+FH+ K
Sbjct: 588 KAMSQK--EPKIIITDQDGAMRSAIAQVFQNAKHGNCFFHIVK 628
>UniRef100_Q8W3G0 Putative far-red impaired response protein [Oryza sativa]
Length = 721
Score = 65.9 bits (159), Expect = 3e-09
Identities = 30/105 (28%), Positives = 57/105 (53%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ F V++ DSTY+ N Y +P VG T G +++E ++VW+LQ + +
Sbjct: 237 YEAFSDVVIFDSTYRVNRYNLPFVPFVGANHHRSTVIFGCGILSNESVSSYVWLLQTLLE 296
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
+ + PK ++TD D ++ A+ V+P + C +H+++N+K
Sbjct: 297 AMHQKH--PKSLITDGDASMAKAIRKVMPNTDHRLCSWHIEENMK 339
>UniRef100_Q8H3L3 Putative far-red impaired response protein [Oryza sativa]
Length = 1148
Score = 65.1 bits (157), Expect = 4e-09
Identities = 30/103 (29%), Positives = 53/103 (51%), Gaps = 2/103 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
++ + V+ D+TY TN Y +P VG+ T G F+ E E F W+ +
Sbjct: 587 WYEKYGDVVSFDTTYFTNKYNLPFAPFVGIYGHGNTIVFGCAFLHDETSETFKWLFRTFL 646
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQK 129
K ++ + PK ++TD+D + +A++ V + NC+FH+ K
Sbjct: 647 KAMSQK--EPKTIITDQDGAMRSAIAQVFQNAKHRNCFFHIVK 687
>UniRef100_Q6K8S4 Putative far-red impaired response protein [Oryza sativa]
Length = 880
Score = 65.1 bits (157), Expect = 4e-09
Identities = 31/104 (29%), Positives = 54/104 (51%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F V+V DSTY+ N Y +P VGV T G V EK + W+L+
Sbjct: 278 YDAFGDVVVFDSTYRVNRYNLPFIPFVGVNHHGSTIIFGCAIVADEKVATYEWILKQFLD 337
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ + P+ ++TD D + A++ V+P+S C +H+++N+
Sbjct: 338 CMYQKH--PRALITDGDNAMRRAIAAVMPDSEHWLCTWHIEQNM 379
>UniRef100_Q9SDE7 Similar to Arabidopsis thaliana DNA chromosome 4 [Oryza sativa]
Length = 861
Score = 64.7 bits (156), Expect = 6e-09
Identities = 30/101 (29%), Positives = 51/101 (49%), Gaps = 2/101 (1%)
Query: 27 FFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMC 86
+++ F + D+T+ TN Y +P VGV+ T G F+ E E F W+
Sbjct: 294 WYHKFGDCISFDTTFLTNRYNLPFAPFVGVSGHGNTILFGCAFLHDETTETFEWLFITFL 353
Query: 87 KLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHV 127
K + + PK ++TD+D + A+ N+ E+ NC+FH+
Sbjct: 354 KCMNGKQ--PKTIITDQDGAMRAAIKNIFREAKHRNCFFHI 392
>UniRef100_Q5W6W6 Hypothetical protein OSJNBa0065C11.11 [Oryza sativa]
Length = 655
Score = 64.3 bits (155), Expect = 7e-09
Identities = 39/148 (26%), Positives = 72/148 (48%), Gaps = 4/148 (2%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ F + ++ Y T Y +P I+G+ + T G + E EE F WV Q K
Sbjct: 171 YREFGDCVFFNTKYITTKYCLPFAPIIGMNNHGQTVLFGCVLLKAEIEETFEWVFQTFLK 230
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRC-ILDCRYPMAKK 146
+ + VPK ++TD+D + NA++NVLP + C +++ +N K + +L R +
Sbjct: 231 AMDGK--VPKSIMTDQDEAMENAIANVLPNTSHRRCSWYIWRNAKFKLGVLPSRLEGFED 288
Query: 147 DGKE-VKHGDVVKKIMRVW*VIIESHPL 173
D + + V++ R W +++ + L
Sbjct: 289 DLRHCIDESFNVEEFERRWAAVLDRYNL 316
>UniRef100_Q6Z0R9 Putative far-red impaired response protein [Oryza sativa]
Length = 690
Score = 63.9 bits (154), Expect = 1e-08
Identities = 32/104 (30%), Positives = 53/104 (50%), Gaps = 2/104 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F V+V DSTY+ N Y +P VGV T V EK + WVL+
Sbjct: 212 YDAFGDVVVFDSTYRVNKYNLPFIPFVGVNHHGSTIIFACAIVADEKIATYEWVLKRFLD 271
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ + PK ++TD D ++ A++ V+P S C +H+++N+
Sbjct: 272 CMCQKH--PKGLITDSDNSMRRAIATVMPNSEHRLCTWHIEQNM 313
>UniRef100_Q6K4T2 Putative far-red impaired response protein [Oryza sativa]
Length = 934
Score = 63.2 bits (152), Expect = 2e-08
Identities = 29/105 (27%), Positives = 57/105 (53%), Gaps = 2/105 (1%)
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
++ F +++ D+ Y T+MY MP I+G+ S + +G + EK E F W+L+ +
Sbjct: 489 YDIFGDLIMFDAAYSTDMYNMPFVPIIGINSHATPFLLGCALLKDEKVETFEWMLRTFLQ 548
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNVK 132
++ +M P+ V+T++DT++ A + ++P C HV +
Sbjct: 549 VMGGKM--PRAVITNQDTSMEKAFAELMPHVRLRFCKRHVMSKAQ 591
>UniRef100_Q6EP19 Putative far-red impaired response protein [Oryza sativa]
Length = 688
Score = 63.2 bits (152), Expect = 2e-08
Identities = 29/101 (28%), Positives = 55/101 (53%), Gaps = 2/101 (1%)
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLT 90
F V++ DSTY+ N Y +P +GV T G +++E ++ W+L+ + +
Sbjct: 238 FGGVVIFDSTYRVNKYNLPFIPFIGVNHHRSTTIFGCGILSNESVNSYCWLLETFLEAM- 296
Query: 91 SRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
R PK ++TD D + A+S V+P ++ C +H+++N+
Sbjct: 297 -RQVHPKSLITDGDLAMAKAISKVMPGAYHRLCTWHIEENM 336
>UniRef100_Q7XVP7 OSJNBa0050F15.2 protein [Oryza sativa]
Length = 688
Score = 63.2 bits (152), Expect = 2e-08
Identities = 29/101 (28%), Positives = 55/101 (53%), Gaps = 2/101 (1%)
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLLT 90
F V++ DSTY+ N Y +P +GV T G +++E ++ W+L+ + +
Sbjct: 238 FGGVVIFDSTYRVNKYNLPFIPFIGVNHHRSTTIFGCGILSNESVNSYCWLLETFLEAM- 296
Query: 91 SRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
R PK ++TD D + A+S V+P ++ C +H+++N+
Sbjct: 297 -RQVHPKSLITDGDLAMAKAISKVMPGAYHRLCTWHIEENM 336
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 765,743,844
Number of Sequences: 2790947
Number of extensions: 30282943
Number of successful extensions: 104480
Number of sequences better than 10.0: 277
Number of HSP's better than 10.0 without gapping: 93
Number of HSP's successfully gapped in prelim test: 184
Number of HSP's that attempted gapping in prelim test: 104258
Number of HSP's gapped (non-prelim): 323
length of query: 484
length of database: 848,049,833
effective HSP length: 131
effective length of query: 353
effective length of database: 482,435,776
effective search space: 170299828928
effective search space used: 170299828928
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 77 (34.3 bits)
Medicago: description of AC140031.2


