

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140025.10 - phase: 0
(344 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
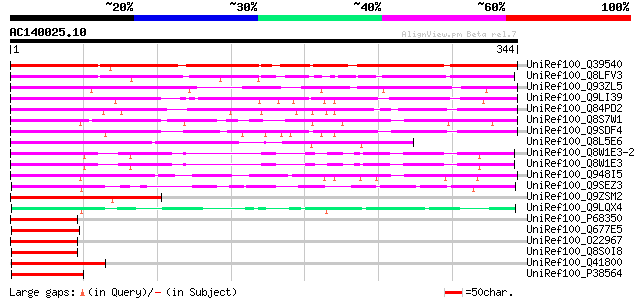
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q39540 AOBP [Cucurbita maxima] 309 8e-83
UniRef100_Q8LFV3 Dof zinc finger protein DOF3.3 [Arabidopsis tha... 261 3e-68
UniRef100_Q93ZL5 Dof zinc finger protein DOF5.2 [Arabidopsis tha... 253 4e-66
UniRef100_Q9LI39 ESTs C23582 [Oryza sativa] 235 2e-60
UniRef100_Q84PD2 Putative ascorbate oxidase promoter-binding pro... 221 2e-56
UniRef100_Q8S7W1 Putative zinc finger DNA-binding protein [Oryza... 215 1e-54
UniRef100_Q9SDF4 Similar to H-protein promoter binding factor-2a... 181 3e-44
UniRef100_Q8L5E6 Dof zinc finger protein [Hordeum vulgare] 177 4e-43
UniRef100_Q8W1E3-2 Splice isoform 2 of Q8W1E3 [Arabidopsis thali... 176 6e-43
UniRef100_Q8W1E3 Dof zinc finger protein DOF5.5 [Arabidopsis tha... 176 6e-43
UniRef100_Q948I5 Putative H-protein promoter binding factor-2a [... 173 7e-42
UniRef100_Q9SEZ3 Dof zinc finger protein DOF1.10 [Arabidopsis th... 157 4e-37
UniRef100_Q9ZSM2 Putative DNA-binding protein [Dendrobium grex M... 140 5e-32
UniRef100_Q9LQX4 Hypothetical Dof zinc finger protein DOF1.3 [Ar... 120 5e-26
UniRef100_P68350 Dof zinc finger protein DOF1.5 [Arabidopsis tha... 99 2e-19
UniRef100_Q677E5 H-protein promoter binding factor [Hyacinthus o... 97 6e-19
UniRef100_O22967 Dof zinc finger protein DOF2.3 [Arabidopsis tha... 97 6e-19
UniRef100_Q8S0I8 PutativeDof zinc finger protein [Oryza sativa] 97 8e-19
UniRef100_Q41800 Dof2 [Zea mays] 95 3e-18
UniRef100_P38564 Dof zinc finger protein MNB1A [Zea mays] 95 3e-18
>UniRef100_Q39540 AOBP [Cucurbita maxima]
Length = 380
Score = 309 bits (791), Expect = 8e-83
Identities = 180/354 (50%), Positives = 220/354 (61%), Gaps = 31/354 (8%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
M+TKFCYYNNYNVNQPRHFCK CQRYWT GGT+RNVPVGAGRRKNK+S+SHYR IT+SEA
Sbjct: 48 METKFCYYNNYNVNQPRHFCKACQRYWTEGGTIRNVPVGAGRRKNKNSASHYRHITISEA 107
Query: 61 TLQNSRI-------HPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHE 113
L+ ++I H + K NG +L F + PVCESM +V A++ + N TRN + +
Sbjct: 108 -LRAAQIDVPIEVNHLASKGNGRVLNFSVSPPVCESMVNVSHPAERKVLNGTRNEFEGAK 166
Query: 114 ELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTP 173
P E G+D S+ SSVT + S + QE M N + FP Q P G P
Sbjct: 167 G-----PCEGGETGDDCSSASSVTMSSSMKNGARRFPQEPHMQNINGFPSQIPCLP-GVP 220
Query: 174 WHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIP-RQAQPSSPNGANH 232
W W ++PIPPPA CPPG + FYPA TYW C+ +WNIP QP P
Sbjct: 221 WPCSW-----TAPIPPPALCPPGVPLSFYPA-TYWSCSASGSWNIPWVTPQPCPPIPG-- 272
Query: 233 DSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEK-SLWFPKTLRIDDSEEAEKSSFWTTL 291
PNSPTLGKHSR+ + L++ + K ++ S+ PKTLRIDD EA KSS W TL
Sbjct: 273 ---PNSPTLGKHSRDGDELQADNSEMKDPPKQKNGSVLVPKTLRIDDPNEAAKSSIWETL 329
Query: 292 GIKNNNADSVPPRRLFQAFPSKCDEKNHLVQV-SSVLQANPAALSRSLHFHETS 344
GIKN DS+ L F SK D K+++ +V S VLQANPAALSRSL FHE S
Sbjct: 330 GIKN---DSIKAVDLSNVFQSKGDLKSNVSEVLSPVLQANPAALSRSLTFHERS 380
>UniRef100_Q8LFV3 Dof zinc finger protein DOF3.3 [Arabidopsis thaliana]
Length = 448
Score = 261 bits (666), Expect = 3e-68
Identities = 167/354 (47%), Positives = 208/354 (58%), Gaps = 37/354 (10%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
M+TKFCYYNNYN+NQPRHFCK CQRYWTAGGTMRNVPVGAGRRKNKSSSSHYR IT+SEA
Sbjct: 118 METKFCYYNNYNINQPRHFCKACQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRHITISEA 177
Query: 61 TLQNSRIHPSVKCNGTILTFG---SNSPVCESMASVLK-HADKTMQNYTRNGYHKHEELR 116
L+ +R+ P ++ N +L+FG V M V+K D+ + N RN +H + R
Sbjct: 178 -LEAARLDPGLQANTRVLSFGLEAQQQHVAAPMTPVMKLQEDQKVSNGARNRFHGLADQR 236
Query: 117 ICVPHTSEEQGEDQSNKSSVTSTKS---TEGATTNVSQEQAMWNDHSFPPQGGY--FPHG 171
+ E G+D S+ SSVT++ + E + S +A N+++ GY P G
Sbjct: 237 LV---ARVENGDDCSSGSSVTTSNNHSVDESRAQSGSVVEAQMNNNNNNNMNGYACIP-G 292
Query: 172 TPWHLPWNPVQMSSPIPPPAFC-PPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGA 230
PW WNP +PPP F PPG+ MPFYP YW M +P Q SSP
Sbjct: 293 VPWPYTWNPA-----MPPPGFYPPPGYPMPFYP---YWTIPM-----LPPH-QSSSPISQ 338
Query: 231 NHDSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEKS-LWFPKTLRIDDSEEAEKSSFWT 289
+T NSPTLGKH R++ SS+ D + E ++ + PKTLRIDD EA KSS WT
Sbjct: 339 KCSNT-NSPTLGKHPRDEG---SSKKDNETERKQKAGCVLVPKTLRIDDPNEAAKSSIWT 394
Query: 290 TLGIKNNNADSVPPRRLFQAFPSKCD-EKNHLVQVSSVLQANPAALSRSLHFHE 342
TLGIKN +F+ F K N + S VL ANPAALSRS +FHE
Sbjct: 395 TLGIKNE--AMCKAGGMFKGFDHKTKMYNNDKAENSPVLSANPAALSRSHNFHE 446
>UniRef100_Q93ZL5 Dof zinc finger protein DOF5.2 [Arabidopsis thaliana]
Length = 457
Score = 253 bits (647), Expect = 4e-66
Identities = 164/365 (44%), Positives = 199/365 (53%), Gaps = 74/365 (20%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQ---ITV 57
M+TKFCYYNNYNVNQPRHFCK CQRYWTAGGTMRNVPVGAGRRKNKS +SHY + IT
Sbjct: 146 METKFCYYNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKSPASHYNRHVSITS 205
Query: 58 SEATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRI 117
+EA + +R +LTFGS+S +CESMAS L +K++ +E L+I
Sbjct: 206 AEAMQKVARTDLQHPNGANLLTFGSDSVLCESMASGLNLVEKSLLKTQTVLQEPNEGLKI 265
Query: 118 CVP--HTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTP-W 174
VP T+EE G S P+ FP P W
Sbjct: 266 TVPLNQTNEEAG------------------------------TVSPLPKVPCFPGPPPTW 295
Query: 175 HLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGC--TMPSAWN-IPRQAQPSSPNGAN 231
WN V + +PFYP YW C P AWN QP+SP+G+N
Sbjct: 296 PYAWNGVSWT-------------ILPFYPPPAYWSCPGVSPGAWNSFTWMPQPNSPSGSN 342
Query: 232 HDSTPNSPTLGKHSREDNMLK------SSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKS 285
PNSPTLGKHSR++N + +E G+++ E+ LW PKTLRIDD EEA KS
Sbjct: 343 ----PNSPTLGKHSRDENAAEPGTAFDETESLGREKSKPERCLWVPKTLRIDDPEEAAKS 398
Query: 286 SFWTTLGI-KNNNADSVPPRRLFQAFPSKCDEKNHLVQ-----VSSVLQANPAALSRSLH 339
S W TLGI K+ NAD+ F AF S EK+ L + LQANPAALSRS +
Sbjct: 399 SIWETLGIKKDENADT------FGAFRSSTKEKSSLSEGRLPGRRPELQANPAALSRSAN 452
Query: 340 FHETS 344
FHE+S
Sbjct: 453 FHESS 457
>UniRef100_Q9LI39 ESTs C23582 [Oryza sativa]
Length = 420
Score = 235 bits (599), Expect = 2e-60
Identities = 162/404 (40%), Positives = 197/404 (48%), Gaps = 88/404 (21%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRK+KSSS HYR + ++
Sbjct: 45 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKSKSSSLHYRHLLMAPD 104
Query: 61 TLQNSRIHPS------------------VKCNGTILTFGSNSPVCESMASVLKHADKTMQ 102
+ SR+ S + N T+L FG P+CESMASVL ++
Sbjct: 105 CMMGSRVEISKSMNPEAFASAHSTPIQPIGRNETVLKFGPEVPLCESMASVLNIQEQNGT 164
Query: 103 NYTRNGYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFP 162
N VP T E Q ED S SS+TS V + +
Sbjct: 165 N------------AAAVP-TGENQ-EDNSCISSITSHNVLPENAAQVDKNSTPVYCNGVG 210
Query: 163 PQGGYF---PHGTPWHLPWNPV---------------------------QMSSPIPPPA- 191
P Y+ P+ PW++ WN V M+SP+ P A
Sbjct: 211 PVPQYYLGAPYMYPWNIGWNNVPMMVPGTSMPESASQSESCSTSSAPWMNMNSPMMPVAS 270
Query: 192 -FCPPGFSMPFYPATTYWGCTM---PSAWNIP----RQAQPSSPNGANHDSTPNSPTLGK 243
P F P P WGC +AWNIP S + +N + N LGK
Sbjct: 271 RLSAPPFPYPLVP-PALWGCLSSWPATAWNIPWIRTNGGCMSPSSSSNSSCSGNGSPLGK 329
Query: 244 HSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSVPP 303
HSR+ ++ KE EEKSLW PKTLRIDD +EA KSS W TLGIK +
Sbjct: 330 HSRDSSL-------PLKEDKEEKSLWVPKTLRIDDPDEAAKSSIWATLGIKPGDPG---- 378
Query: 304 RRLFQAFPSKCDEKNHL---VQVSSVLQANPAALSRSLHFHETS 344
+F+ F SK + K + + L+ANPAALSRS F ETS
Sbjct: 379 --IFKPFQSKGESKGQAASETRPARALKANPAALSRSQSFQETS 420
>UniRef100_Q84PD2 Putative ascorbate oxidase promoter-binding protein AOBP [Oryza
sativa]
Length = 476
Score = 221 bits (564), Expect = 2e-56
Identities = 150/388 (38%), Positives = 197/388 (50%), Gaps = 71/388 (18%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
MDTKFCYYNNYN+NQPRHFCK+CQRYWTAGG+MRN+PVGAGRRK+KSS+++YR I ++ +
Sbjct: 116 MDTKFCYYNNYNINQPRHFCKSCQRYWTAGGSMRNLPVGAGRRKSKSSTANYRSILITGS 175
Query: 61 TL-----QNSRIHPSVKCN--GTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHE 113
L S+K + T + F +SP+C SMASVLK +++ +
Sbjct: 176 NLAAPAGDAPLYQLSIKGDQTATAVKFAPDSPLCNSMASVLKIGEQSKNAKPTS------ 229
Query: 114 ELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTN--VSQEQAMWNDHSFPPQGGYFP-- 169
GE Q+ +S T++ S N VS Q HS P P
Sbjct: 230 -------TAQPRNGETQTCPASGTTSDSPRNEPVNGAVSGHQNGIVGHSGVPPMHPIPCF 282
Query: 170 HGTPWHLPWNPVQMSSPIPPPAFC----------------------PPGFSMP-FYPA-- 204
G P+ PW+P P P C PP +P ++P
Sbjct: 283 PGPPFVYPWSPAWNGIPAMAPPVCTAPAEPANSSDNGSTASVQWSMPPVMPVPGYFPVIP 342
Query: 205 TTYWGCTMP---SAWNIP-----RQAQPSSPNGANHDSTPNSPTLGKHSREDNMLKSSEG 256
++ W P AW+ P SSP + S SP LGKHSR+ +G
Sbjct: 343 SSVWPFISPWPNGAWSSPWIQPNCSVSASSPTSTSTCSDNGSPVLGKHSRD----SKPQG 398
Query: 257 DGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSVPPRRLFQAFPSKCDE 316
D K EK+LW PKTLRIDD +EA KSS WTTLGI+ + R +F++F SK +
Sbjct: 399 DDK----AEKNLWIPKTLRIDDPDEAAKSSIWTTLGIEPGD------RSMFRSFQSKPES 448
Query: 317 KNHLVQVSSVLQANPAALSRSLHFHETS 344
+ + + VLQANPAALSRS F ET+
Sbjct: 449 REQISGAARVLQANPAALSRSQSFQETT 476
>UniRef100_Q8S7W1 Putative zinc finger DNA-binding protein [Oryza sativa]
Length = 440
Score = 215 bits (548), Expect = 1e-54
Identities = 145/365 (39%), Positives = 197/365 (53%), Gaps = 59/365 (16%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK--SSSSHYRQITVS 58
MDTKFCYYNNYN+NQPRHFCKNCQRYWTAGG MRNVPVGAGRRK+K S++SH+ Q V
Sbjct: 114 MDTKFCYYNNYNINQPRHFCKNCQRYWTAGGAMRNVPVGAGRRKSKSVSAASHFLQ-RVR 172
Query: 59 EATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRI- 117
A + ++ VK NGT+L+FGS+ + + DK + G +E+ +
Sbjct: 173 AALPGDPPLYAPVKTNGTVLSFGSDLSTLDLTEQMKHLKDKFIPT---TGIKNTDEMPVG 229
Query: 118 -CVPHTSEEQGEDQSN-KSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWH 175
C S+ + +Q+N K V++ +S NV+Q M G +P G
Sbjct: 230 LCAEGLSKTEESNQTNLKEKVSADRS-----PNVAQHPCM-------NGGAMWPFGV--- 274
Query: 176 LPWNPVQMSSPIPPPAFCPPGFSMPFYPA-----TTYWGCTMPSAWNIPRQAQP-----S 225
PPPA+ ++PFYPA YWGC +P AWN P Q S
Sbjct: 275 -----------APPPAYYTSSIAIPFYPAAAAAVAAYWGCMVPGAWNAPWPPQSQSQSVS 323
Query: 226 SPNGANHDST-PNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEK 284
S + A+ ST N LGKH R+ + S+G+GK +W PKT+RIDD +E +
Sbjct: 324 SSSAASPVSTMTNCFRLGKHPRDGDEELDSKGNGK--------VWVPKTVRIDDVDEVAR 375
Query: 285 SSFWTTLGIKNN--NADSVPPRRLFQAFPSKCDEK-NHLVQVSSV--LQANPAALSRSLH 339
SS W+ +GIK + AD +L + F SK + K + +SS+ +Q NPAAL+RS+
Sbjct: 376 SSIWSLIGIKGDKVGADHGRGCKLAKVFESKDEAKASTHTAISSLPFMQGNPAALTRSVT 435
Query: 340 FHETS 344
F E S
Sbjct: 436 FQEGS 440
>UniRef100_Q9SDF4 Similar to H-protein promoter binding factor-2a [Oryza sativa]
Length = 454
Score = 181 bits (458), Expect = 3e-44
Identities = 137/403 (33%), Positives = 182/403 (44%), Gaps = 93/403 (23%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
M+TKFCY+NNYNV+QPRHFC+NCQRYWTAGG MRNVPVGAGRR+NK S + + +
Sbjct: 86 METKFCYFNNYNVHQPRHFCRNCQRYWTAGGAMRNVPVGAGRRRNKHVSKYCQAMMTCNN 145
Query: 61 TLQ----------------NSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNY 104
T+ +S + ++K N T F S P C+S AS+L + +
Sbjct: 146 TVAPGDVSDVVHHQVITHGSSLLPATLKENETPTEFISEVPPCKSSASILDIGEPNDTD- 204
Query: 105 TRNGYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMW---NDHSF 161
VP S + E++S SSV + +E N+ + A+ N+ S
Sbjct: 205 -------------LVPLASGDNKEEKSCASSVVVSSCSE----NLMPDNAIMKEPNNRSG 247
Query: 162 PPQGGYFPHGT------PWHLPWNPVQM----------------SSPIPP---------- 189
G P T PW L WN V + P PP
Sbjct: 248 CCNGVALPFPTGPALVLPWSLGWNSVALMPATQCSMQPVLGLKDGIPCPPSWPPQLMVPA 307
Query: 190 PAFCPPGFSMPFYPATTYWGC--TMPSA-WNIP-----RQAQPSSPNGANHDSTPNSPTL 241
P C P +P P W C P+ WN PS+ S +S L
Sbjct: 308 PGICTPVVPIPLVP--PLWSCFPGWPNGMWNAQCPGGNTTVLPSTAPNKISCSGSSSLVL 365
Query: 242 GKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSV 301
GKHSRE+++ ++E LW PKTLRIDD EA KSS W TLGIK ++
Sbjct: 366 GKHSREESL--------QEEEKTRNYLWVPKTLRIDDPAEAAKSSIWATLGIKPDD---- 413
Query: 302 PPRRLFQAFPSKCDEKNHLVQVSSVLQANPAALSRSLHFHETS 344
+ +F++F + + LQANPAA SRS F ET+
Sbjct: 414 --KGIFKSFQPNVAKNGTAPESPQALQANPAAFSRSQSFQETT 454
>UniRef100_Q8L5E6 Dof zinc finger protein [Hordeum vulgare]
Length = 270
Score = 177 bits (449), Expect = 4e-43
Identities = 102/279 (36%), Positives = 140/279 (49%), Gaps = 32/279 (11%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
MDTKFCYYNNYN+NQPRHFCKNCQRYWTAGG MRNVPVGAGRRK+KS S+ + A
Sbjct: 16 MDTKFCYYNNYNINQPRHFCKNCQRYWTAGGAMRNVPVGAGRRKSKSISAASHFLQRIRA 75
Query: 61 TLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRICVP 120
L + V NGT+L+FGS++ + ++ +KH K + + TR + C
Sbjct: 76 ALPGDPLCTPVNTNGTVLSFGSDASTLDVVSEQMKHM-KELSSVTRTENTDAPSVGSCAE 134
Query: 121 HTSE-EQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWN 179
++ E+ +++ V + +S A AMW P+
Sbjct: 135 GWAKGEESSQMNSRERVAADRSPNFAQHPCMNGAAMW----------------PF----- 173
Query: 180 PVQMSSPIPPPAFCPPGFSMPFYP--ATTYWGCTMPSAWNIPRQAQPSSPNGANHDSTPN 237
S P PA+ P ++PFYP A YWGC +P AWN P Q QP + S P+
Sbjct: 174 -----SCAPSPAYFTPNVAIPFYPAAAAAYWGCMVPGAWNTPWQPQPQPQSQCQSSSPPS 228
Query: 238 --SPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTL 274
SP S + +GD +++ +W PKT+
Sbjct: 229 AASPVSTMSSCFQSRKHPRDGDEERDTKGNGKVWVPKTI 267
>UniRef100_Q8W1E3-2 Splice isoform 2 of Q8W1E3 [Arabidopsis thaliana]
Length = 237
Score = 176 bits (447), Expect = 6e-43
Identities = 130/356 (36%), Positives = 161/356 (44%), Gaps = 135/356 (37%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSS--SHYRQITVS 58
M+TKFCYYNNYNVNQPRHFCK CQRYWT+GGTMR+VP+GAGRRKNK++S SHY +T+S
Sbjct: 1 METKFCYYNNYNVNQPRHFCKACQRYWTSGGTMRSVPIGAGRRKNKNNSPTSHYHHVTIS 60
Query: 59 EATLQNSRIHPSVKCNGTILTF--------GSNSPVCESMASVLKHADKTMQNYTRNGYH 110
E NG +L+F SN + + + +++ D+ N T NG +
Sbjct: 61 ET-------------NGPVLSFSLGDDQKVSSNRFGNQKLVARIENNDERSNNNTSNGLN 107
Query: 111 KHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPH 170
C P
Sbjct: 108 -------CFP-------------------------------------------------- 110
Query: 171 GTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGA 230
G W WNP FYP Y W++P + P S
Sbjct: 111 GVSWPYTWNPA-------------------FYPVYPY--------WSMPVLSSPVS---- 139
Query: 231 NHDSTPNSPTLGKHSR-EDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWT 289
S+P S TLGKHSR ED +K + +G S+ PKTLRIDD EA KSS WT
Sbjct: 140 ---SSPTS-TLGKHSRDEDETVKQKQRNG--------SVLVPKTLRIDDPNEAAKSSIWT 187
Query: 290 TLGIKNNNADSVPPRRLFQAFPSKCDEK---NHLVQVSSVLQANPAALSRSLHFHE 342
TLGIKN +F F SK + K + S VL ANPAALSRS++FHE
Sbjct: 188 TLGIKN--------EVMFNGFGSKKEVKLSNKEETETSLVLCANPAALSRSINFHE 235
>UniRef100_Q8W1E3 Dof zinc finger protein DOF5.5 [Arabidopsis thaliana]
Length = 298
Score = 176 bits (447), Expect = 6e-43
Identities = 130/356 (36%), Positives = 161/356 (44%), Gaps = 135/356 (37%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSS--SHYRQITVS 58
M+TKFCYYNNYNVNQPRHFCK CQRYWT+GGTMR+VP+GAGRRKNK++S SHY +T+S
Sbjct: 62 METKFCYYNNYNVNQPRHFCKACQRYWTSGGTMRSVPIGAGRRKNKNNSPTSHYHHVTIS 121
Query: 59 EATLQNSRIHPSVKCNGTILTF--------GSNSPVCESMASVLKHADKTMQNYTRNGYH 110
E NG +L+F SN + + + +++ D+ N T NG +
Sbjct: 122 ET-------------NGPVLSFSLGDDQKVSSNRFGNQKLVARIENNDERSNNNTSNGLN 168
Query: 111 KHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPH 170
C P
Sbjct: 169 -------CFP-------------------------------------------------- 171
Query: 171 GTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGA 230
G W WNP FYP Y W++P + P S
Sbjct: 172 GVSWPYTWNPA-------------------FYPVYPY--------WSMPVLSSPVS---- 200
Query: 231 NHDSTPNSPTLGKHSR-EDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWT 289
S+P S TLGKHSR ED +K + +G S+ PKTLRIDD EA KSS WT
Sbjct: 201 ---SSPTS-TLGKHSRDEDETVKQKQRNG--------SVLVPKTLRIDDPNEAAKSSIWT 248
Query: 290 TLGIKNNNADSVPPRRLFQAFPSKCDEK---NHLVQVSSVLQANPAALSRSLHFHE 342
TLGIKN +F F SK + K + S VL ANPAALSRS++FHE
Sbjct: 249 TLGIKN--------EVMFNGFGSKKEVKLSNKEETETSLVLCANPAALSRSINFHE 296
>UniRef100_Q948I5 Putative H-protein promoter binding factor-2a [Oryza sativa]
Length = 486
Score = 173 bits (438), Expect = 7e-42
Identities = 128/368 (34%), Positives = 175/368 (46%), Gaps = 57/368 (15%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK--SSSSHYRQITVS 58
MDTKFCY+NNYNVNQPRHFCK+CQRYWTAGG MRNVPVGAGRRKNK ++++H+ +
Sbjct: 152 MDTKFCYFNNYNVNQPRHFCKHCQRYWTAGGAMRNVPVGAGRRKNKNATAAAHFLHRVRA 211
Query: 59 EATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRIC 118
A P N T+L+FG ++ L ADK M + G H
Sbjct: 212 CAAAAAMPAAPHDATNATVLSFGGGGGGHDAPPVTLDLADK-MTRLGKEGLVAH------ 264
Query: 119 VPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPW 178
+ + S V+S + E V++ P G H P H
Sbjct: 265 -----ARNADAAAACSEVSSNRDDEQIGNTVAK-----------PANGLQQHPPPPHHHH 308
Query: 179 NPVQMSSPIPPPAFCPPGFSMPFYPAT-TYWGCTM--PSAWNIP----RQAQPSSPNGAN 231
+ I P + G ++P YPA YWGC + P AW++P Q+Q S +
Sbjct: 309 HSAMNGGGIWP--YYTSGIAIPIYPAAPAYWGCMIPPPGAWSLPWPATVQSQAISSSSPP 366
Query: 232 HDSTP--NSPTLGKHSRE--DNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSF 287
+TP +S TLGKH RE D+ + G+GK +W PKT+RID+++E +SS
Sbjct: 367 TSATPSVSSFTLGKHPREGGDHEARDHHGNGK--------VWVPKTIRIDNADEVARSSI 418
Query: 288 WTTLGIK-------NNNADSVPPRRL-FQAFPSKCDE---KNHLVQVSSVLQANPAALSR 336
+ + NN+ D +L F K D K+ + +L NP AL+R
Sbjct: 419 RSLFAFRGGDKADDNNDDDGTGVHKLATTVFEPKRDSKTAKHPAITSLPLLHTNPVALTR 478
Query: 337 SLHFHETS 344
S F E S
Sbjct: 479 SATFQEGS 486
>UniRef100_Q9SEZ3 Dof zinc finger protein DOF1.10 [Arabidopsis thaliana]
Length = 399
Score = 157 bits (397), Expect = 4e-37
Identities = 118/346 (34%), Positives = 161/346 (46%), Gaps = 94/346 (27%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKS--SSSHYRQITVSE 59
+TKFCYYNNYNVNQPR+FC+NCQRYWTAGG+MRNVPVG+GRRKNK SS+HY Q+T +
Sbjct: 141 NTKFCYYNNYNVNQPRYFCRNCQRYWTAGGSMRNVPVGSGRRKNKGWPSSNHYLQVTSED 200
Query: 60 ATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRICV 119
NS GTIL+FGS S +SV
Sbjct: 201 CDNNNS---------GTILSFGS------SESSV-------------------------- 219
Query: 120 PHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWN 179
E G+ QS ++ S S VSQE + PPQ + +PW W+
Sbjct: 220 ----TETGKHQSGDTAKISADS-------VSQENKSYQGF-LPPQVMLPNNSSPWPYQWS 267
Query: 180 PVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGANHDSTPNSP 239
P G + FYP YWGCT+P + ++ S
Sbjct: 268 PT--------------GPNASFYPVPFYWGCTVPI-----------------YPTSETSS 296
Query: 240 TLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNAD 299
LGK SR+ + D I+ ++ ++LR+ + EA KS+ W+ L K
Sbjct: 297 CLGKRSRDQT--EGRINDTNTTITTTRARLVSESLRM--NIEASKSAVWSKLPTKPEK-- 350
Query: 300 SVPPRRLFQAFPSK--CDEKNHLVQVSSVLQANPAALSRSLHFHET 343
LF F +K + + + + S LQANPAA+SR+++F E+
Sbjct: 351 KTQGFSLFNGFDTKGNSNRSSLVSETSHSLQANPAAMSRAMNFRES 396
>UniRef100_Q9ZSM2 Putative DNA-binding protein [Dendrobium grex Madame Thong-In]
Length = 179
Score = 140 bits (353), Expect = 5e-32
Identities = 65/113 (57%), Positives = 81/113 (71%), Gaps = 10/113 (8%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGG+MRNVPVGAGRRKNK + S YR ++
Sbjct: 35 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGSMRNVPVGAGRRKNKHTGSVYRHTVITPD 94
Query: 61 TLQNSRIH----------PSVKCNGTILTFGSNSPVCESMASVLKHADKTMQN 103
+L + ++ K NGTIL FG ++P+CESMAS+L ++ + +
Sbjct: 95 SLASLQVDGPDLVDHKPLSPFKVNGTILKFGPDAPLCESMASILNLGEQNLSS 147
>UniRef100_Q9LQX4 Hypothetical Dof zinc finger protein DOF1.3 [Arabidopsis thaliana]
Length = 366
Score = 120 bits (301), Expect = 5e-26
Identities = 102/346 (29%), Positives = 139/346 (39%), Gaps = 98/346 (28%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKS--SSSHYRQITVSE 59
DTKFCYYNNYNVNQPRHFC+ CQRYWTAGG+MR VPVG+GRRKNK SS Y IT +
Sbjct: 114 DTKFCYYNNYNVNQPRHFCRKCQRYWTAGGSMRIVPVGSGRRKNKGWVSSDQYLHITSED 173
Query: 60 ATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRICV 119
NS + IL+F S+ + T H+ E++I
Sbjct: 174 TDNYNS-------SSTKILSFESSDSL-----------------VTERPKHQSNEVKINA 209
Query: 120 PHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTPWHLPWN 179
S+E Q PPQ P PW
Sbjct: 210 EPVSQEPNNFQG----------------------------LLPPQAS--PVSPPW----- 234
Query: 180 PVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMP--SAWNIPRQAQPSSPNGANHDSTPN 237
P PP S FY YWGC +P S + + + +H++
Sbjct: 235 ----------PYQYPPNPS--FYHMPVYWGCAIPVWSTLDTSTCLGKRTRDETSHETVKE 282
Query: 238 SPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNN 297
S R +L+S + ++ +W+P + + ++E S F K++N
Sbjct: 283 SK--NAFERTSLLLESQSIKNETSMATNNHVWYPVPMTREKTQEF--SFFSNGAETKSSN 338
Query: 298 ADSVPPRRLFQAFPSKCDEKNHLVQVSSVLQANPAALSRSLHFHET 343
VP L LQANPAA++RS++F E+
Sbjct: 339 NRFVPETYL-------------------NLQANPAAMARSMNFRES 365
>UniRef100_P68350 Dof zinc finger protein DOF1.5 [Arabidopsis thaliana]
Length = 175
Score = 99.0 bits (245), Expect = 2e-19
Identities = 40/46 (86%), Positives = 44/46 (94%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCY+NNYNVNQPRHFCK CQRYWTAGG +RNVPVGAGRRK+K
Sbjct: 70 METKFCYFNNYNVNQPRHFCKGCQRYWTAGGALRNVPVGAGRRKSK 115
>UniRef100_Q677E5 H-protein promoter binding factor [Hyacinthus orientalis]
Length = 84
Score = 97.1 bits (240), Expect = 6e-19
Identities = 38/46 (82%), Positives = 44/46 (95%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKS 47
+TKFCY+NNYNVNQPRHFC+NC RYWTAGGT+R VPVGAGRRKN++
Sbjct: 18 ETKFCYFNNYNVNQPRHFCRNCHRYWTAGGTLRRVPVGAGRRKNRT 63
>UniRef100_O22967 Dof zinc finger protein DOF2.3 [Arabidopsis thaliana]
Length = 170
Score = 97.1 bits (240), Expect = 6e-19
Identities = 39/46 (84%), Positives = 43/46 (92%)
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCY+NNYNVNQPRHFCK C RYWTAGG +RNVPVGAGRRK+K
Sbjct: 66 METKFCYFNNYNVNQPRHFCKGCHRYWTAGGALRNVPVGAGRRKSK 111
>UniRef100_Q8S0I8 PutativeDof zinc finger protein [Oryza sativa]
Length = 205
Score = 96.7 bits (239), Expect = 8e-19
Identities = 39/45 (86%), Positives = 42/45 (92%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK 46
DTKFCY+NNYNVNQPRHFCK C RYWTAGG +RNVPVGAGRRKN+
Sbjct: 112 DTKFCYFNNYNVNQPRHFCKACHRYWTAGGALRNVPVGAGRRKNR 156
>UniRef100_Q41800 Dof2 [Zea mays]
Length = 225
Score = 94.7 bits (234), Expect = 3e-18
Identities = 40/64 (62%), Positives = 47/64 (72%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEAT 61
DTKFCYYNNYN +QPRH CK+C+RYWT GG++RNVPVG G RK+ SSSS T + T
Sbjct: 27 DTKFCYYNNYNTSQPRHLCKSCRRYWTKGGSLRNVPVGGGTRKSSSSSSSSSAATTTTTT 86
Query: 62 LQNS 65
S
Sbjct: 87 TSTS 90
>UniRef100_P38564 Dof zinc finger protein MNB1A [Zea mays]
Length = 238
Score = 94.7 bits (234), Expect = 3e-18
Identities = 39/49 (79%), Positives = 42/49 (85%)
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSS 50
DTKFCYYNNYN +QPRHFCK C+RYWT GGT+RNVPVG G RK SSSS
Sbjct: 56 DTKFCYYNNYNTSQPRHFCKGCRRYWTKGGTLRNVPVGGGTRKKPSSSS 104
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.127 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 653,417,761
Number of Sequences: 2790947
Number of extensions: 29971526
Number of successful extensions: 74348
Number of sequences better than 10.0: 362
Number of HSP's better than 10.0 without gapping: 112
Number of HSP's successfully gapped in prelim test: 261
Number of HSP's that attempted gapping in prelim test: 73786
Number of HSP's gapped (non-prelim): 663
length of query: 344
length of database: 848,049,833
effective HSP length: 128
effective length of query: 216
effective length of database: 490,808,617
effective search space: 106014661272
effective search space used: 106014661272
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC140025.10


