

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139853.10 + phase: 0
(342 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
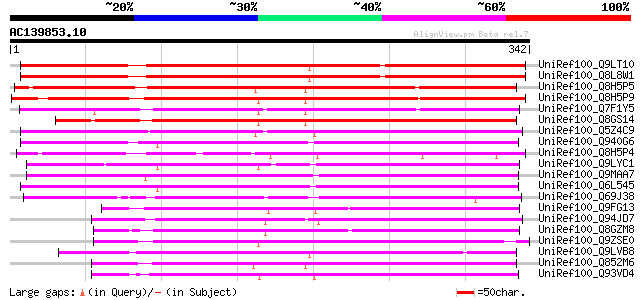
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LT10 Similarity to unknown protein [Arabidopsis thal... 381 e-104
UniRef100_Q8L8W1 Hypothetical protein [Arabidopsis thaliana] 378 e-103
UniRef100_Q8H5P5 Putative esterase [Oryza sativa] 303 4e-81
UniRef100_Q8H5P9 Putative esterase [Oryza sativa] 278 1e-73
UniRef100_Q7F1Y5 Putative esterase [Oryza sativa] 261 2e-68
UniRef100_Q8GS14 Carboxylesterase-like protein [Oryza sativa] 256 5e-67
UniRef100_Q5Z4C9 Putative esterase [Oryza sativa] 255 1e-66
UniRef100_Q940G6 Hypothetical protein At5g27320 [Arabidopsis tha... 245 1e-63
UniRef100_Q8H5P4 Putative esterase [Oryza sativa] 237 4e-61
UniRef100_Q9LYC1 Hypothetical protein T20O10_110 [Arabidopsis th... 229 1e-58
UniRef100_Q9MAA7 T12H1.8 protein [Arabidopsis thaliana] 228 2e-58
UniRef100_Q6L545 Hypothetical protein OJ1657_H11.10 [Oryza sativa] 227 3e-58
UniRef100_Q69J38 Putative PrMC3 [Oryza sativa] 220 4e-56
UniRef100_Q9FG13 Gb|AAD04946.2 [Arabidopsis thaliana] 176 6e-43
UniRef100_Q94JD7 Putative PrMC3 [Oryza sativa] 173 5e-42
UniRef100_Q8GZM8 Putative serine hydrolase [Vitis vinifera] 169 1e-40
UniRef100_Q9ZSE0 PrMC3 [Pinus radiata] 164 2e-39
UniRef100_Q9LVB8 HSR203J protein-like protein [Arabidopsis thali... 164 2e-39
UniRef100_Q852M6 Putative esterase [Oryza sativa] 162 9e-39
UniRef100_Q93VD4 Putative PrMC3 [Oryza sativa] 162 1e-38
>UniRef100_Q9LT10 Similarity to unknown protein [Arabidopsis thaliana]
Length = 335
Score = 381 bits (979), Expect = e-104
Identities = 182/337 (54%), Positives = 237/337 (70%), Gaps = 18/337 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
KL PLKTR+++++I T+TD A R +G++NRR L D + P+N VST D VD
Sbjct: 10 KLTLPLKTRIALTVISTMTDNAQRPDGTINRRFLRLFDFRAPPNPKPVNIVSTSDFVVDQ 69
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRE 127
+WFRL+TP + +PVV+FFHGGGF F+SP + YD +CRRF+R+
Sbjct: 70 SRDLWFRLYTP-----------HVSGDKIPVVVFFHGGGFAFLSPNAYPYDNVCRRFARK 118
Query: 128 LNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENK-SVLPENVDVSKCFLAGDSAGANL 186
L V+SVNYR PE+RYP QY+DG ALK+++EN S+LP N D+S+CF AGDSAG N+
Sbjct: 119 LPAYVISVNYRLAPEHRYPAQYDDGFDALKYIEENHGSILPANADLSRCFFAGDSAGGNI 178
Query: 187 AHHVAVRACK---AGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKV 243
AH+VA+R C+ + +++ GLIS+QPFFGGEERTEAE +L G+ ++S RTDW WK
Sbjct: 179 AHNVAIRICREPRSSFTAVKLIGLISIQPFFGGEERTEAEKQLVGAPLVSPDRTDWCWKA 238
Query: 244 FLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQ 303
G NRDH A NV GPNA D+S LDYP+T+V V G D L DWQ+ YYEWLK+ GKKA
Sbjct: 239 M---GLNRDHEAVNVGGPNAVDISGLDYPETMVVVAGFDPLKDWQRSYYEWLKLCGKKAT 295
Query: 304 LIEYPNMMHGFYAFPNVPEASQLILQIKDFINNRVSN 340
LIEYPNM H FY FP +PEA QLI++IKDF++ RV++
Sbjct: 296 LIEYPNMFHAFYIFPELPEAGQLIMRIKDFVDERVAS 332
>UniRef100_Q8L8W1 Hypothetical protein [Arabidopsis thaliana]
Length = 335
Score = 378 bits (971), Expect = e-103
Identities = 181/337 (53%), Positives = 236/337 (69%), Gaps = 18/337 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
KL PLKTR+++++I T+TD A R +G++NRR L D + P+N VST D VD
Sbjct: 10 KLTLPLKTRIALTVISTMTDNAQRPDGTINRRFLRLFDFRAPPNPKPVNIVSTSDFVVDQ 69
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRE 127
+WFRL+TP + +PVV+FFHGGGF F+SP + YD +CRRF+R+
Sbjct: 70 SRDLWFRLYTP-----------HVSGDKIPVVVFFHGGGFAFLSPNAYPYDNVCRRFARK 118
Query: 128 LNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENK-SVLPENVDVSKCFLAGDSAGANL 186
L V+SVNYR PE+RYP QY+DG ALK+++EN S+LP N D+S+CF AGDSAG N+
Sbjct: 119 LPAYVISVNYRLAPEHRYPAQYDDGFDALKYIEENHGSILPANADLSRCFFAGDSAGGNI 178
Query: 187 AHHVAVRACK---AGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKV 243
AH+VA+R C+ + +++ GLIS+QPFFGGEERTEAE +L G+ ++S RTDW WK
Sbjct: 179 AHNVAIRICREPRSSFTAVKLIGLISIQPFFGGEERTEAEKQLVGAPLVSPDRTDWCWKA 238
Query: 244 FLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQ 303
G NRDH A NV GPNA D+S LDYP+T+V V G D L DWQ+ YYEWLK+ GKKA
Sbjct: 239 M---GLNRDHEAVNVGGPNAVDISGLDYPETMVVVAGFDPLKDWQRSYYEWLKLCGKKAT 295
Query: 304 LIEYPNMMHGFYAFPNVPEASQLILQIKDFINNRVSN 340
LIEY NM H FY FP +PEA QLI++IKDF++ RV++
Sbjct: 296 LIEYSNMFHAFYIFPELPEAGQLIMRIKDFVDERVAS 332
>UniRef100_Q8H5P5 Putative esterase [Oryza sativa]
Length = 355
Score = 303 bits (776), Expect = 4e-81
Identities = 157/340 (46%), Positives = 222/340 (65%), Gaps = 20/340 (5%)
Query: 4 PKKPKLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP-INGVSTKD 62
P+ P LP+ + RL + ++T D R +G+VNR L + D +++A A P +GV + D
Sbjct: 17 PEPPALPWTV--RLQLFALVTAVDIVQRGDGTVNRFLFSLADRQSAAAARPDAHGVRSGD 74
Query: 63 ITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICR 122
+TVDA +W R+F+P +A E+ LPVV++FHGGGF ++ AS YD +CR
Sbjct: 75 VTVDASRGLWARVFSPASSSA-------VESPPLPVVVYFHGGGFALLTAASSQYDALCR 127
Query: 123 RFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLD----ENKSVLPENVDVSKCFLA 178
R REL VVVSVNYR PE+RYP Y+DG L+ L V VD+++CFL
Sbjct: 128 RLCRELRAVVVSVNYRLAPEHRYPAAYDDGVDVLRHLATVGLPADVVAAVPVDLTRCFLV 187
Query: 179 GDSAGANLAHHVAVR---ACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEG-SLMISM 234
GDSAG N+AHHVA R A + +R+R+AG++ +QPFFGGEERTEAE+RL+G ++SM
Sbjct: 188 GDSAGGNIAHHVAHRWAAATTSSSRRVRLAGVVLLQPFFGGEERTEAELRLDGVGPVVSM 247
Query: 235 ARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEW 294
AR DW W+ FLPEG++RDH AA+V+G NAE ++P +V VGG D L DWQ+RY
Sbjct: 248 ARADWCWRAFLPEGADRDHPAAHVTGENAELAE--EFPPAMVVVGGYDTLQDWQRRYAGM 305
Query: 295 LKISGKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFI 334
L+ +GK Q++EYP +H FY FP + ++ +L+ ++K F+
Sbjct: 306 LRRNGKAVQVVEYPAAIHSFYVFPELADSGELVKEMKAFM 345
>UniRef100_Q8H5P9 Putative esterase [Oryza sativa]
Length = 345
Score = 278 bits (712), Expect = 1e-73
Identities = 151/349 (43%), Positives = 211/349 (60%), Gaps = 26/349 (7%)
Query: 2 SAPKKPKLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP-INGVST 60
S+ P LP+ ++ +L+ AA RS+GSV R L D +A P GV +
Sbjct: 10 SSSSPPPLPWTVRVQLAA------LSAAHRSDGSVRRLLFYLGDLHAAASPRPDAAGVRS 63
Query: 61 KDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTI 120
D+T+DA +W R+F P +NT LPVV++FHGGGF S AS YD +
Sbjct: 64 VDVTIDASRGLWARVFCPP---------TNTAAVKLPVVVYFHGGGFVLFSAASRPYDAL 114
Query: 121 CRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN-----KSVLPENVDVSKC 175
CRR SR + VVVSVNYR PE+R+P Y+DG AL++LD N + L VD+S+C
Sbjct: 115 CRRISRGVGAVVVSVNYRLAPEHRFPAAYDDGLAALRYLDANGLAEAAAELGAAVDLSRC 174
Query: 176 FLAGDSAGANLAHHVAVR---ACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLE-GSLM 231
FLAGDSAG N+ HHVA R + + +R+AG + + PFFGGEERTE E+ L+ SL
Sbjct: 175 FLAGDSAGGNIVHHVAQRWAASTTSPSSSLRLAGAVLISPFFGGEERTEEEVGLDKASLS 234
Query: 232 ISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRY 291
+S+ARTD+ W+ FLPEG+ RDH AA V G +L+ +P +V +GG D L WQ RY
Sbjct: 235 LSLARTDYFWREFLPEGATRDHAAARVCGGERVELAEA-FPPAMVVIGGFDLLKGWQARY 293
Query: 292 YEWLKISGKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFINNRVSN 340
L+ GK +++EYP+ +HGF+AFP + ++ +L+ ++K F+ SN
Sbjct: 294 VAALREKGKAVRVVEYPDAIHGFHAFPELADSGKLVEEMKQFVQEHSSN 342
>UniRef100_Q7F1Y5 Putative esterase [Oryza sativa]
Length = 362
Score = 261 bits (667), Expect = 2e-68
Identities = 141/346 (40%), Positives = 209/346 (59%), Gaps = 28/346 (8%)
Query: 7 PKLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFL-DNKTSAKATP-----INGVST 60
P P PL RL + + DA R +G+VNR L + L ++ S +A P V +
Sbjct: 21 PPPPMPLGLRLQLIGLSAAIDAVERRDGTVNRALYSVLVEHLMSVRADPSPDAATGAVRS 80
Query: 61 KDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTI 120
D T+DA +W R+F P +A +PV++++HGGGF SPA +D +
Sbjct: 81 FDFTIDAARGLWARVFAPAAAAQAAA--------PMPVMVYYHGGGFALFSPAVAPFDGV 132
Query: 121 CRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN------KSVLPENVDVSK 174
CRR ++ VVVVSVNYR PE+RYP Y+DG AL+FLD N V+P VD++
Sbjct: 133 CRRLCGDVGVVVVSVNYRLAPEHRYPAAYDDGVDALRFLDGNGIPGLDGDVVP--VDLAS 190
Query: 175 CFLAGDSAGANLAHHVAVR---ACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEG-SL 230
CFLAG+SAG N+ H VA R + + +R+AG+I +QP+FGGEERT +E+ L+G +
Sbjct: 191 CFLAGESAGGNIVHQVANRWAATWQPTAKNLRLAGMIPVQPYFGGEERTPSELALDGVAP 250
Query: 231 MISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKR 290
++++ R+D+ WK FLP G++RDH AA+V+ NAE +P +V +GG D L DWQ+R
Sbjct: 251 VVNLRRSDFSWKAFLPVGADRDHPAAHVTDENAELAEA--FPPAMVVIGGFDPLQDWQRR 308
Query: 291 YYEWLKISGKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFINN 336
Y + L+ GK ++ E+P+ HGFY FP + +A +++ IK F+ +
Sbjct: 309 YVDVLRRKGKAVEVAEFPDAFHGFYGFPELADAGKVLQDIKVFVQS 354
>UniRef100_Q8GS14 Carboxylesterase-like protein [Oryza sativa]
Length = 347
Score = 256 bits (655), Expect = 5e-67
Identities = 140/316 (44%), Positives = 192/316 (60%), Gaps = 21/316 (6%)
Query: 31 RSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSN 90
R +GSV R + + LD AK GV + D+T+DA +W R+F+P A
Sbjct: 32 RRDGSVRRLVFSLLDIHVRAKRRA--GVRSVDVTIDASRGLWARVFSPPPTKGEAA---- 85
Query: 91 TETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRT-PEYRYPTQY 149
+LPVV+FFHGGGF S AS YD +CRR REL VVVSVNYR P R+P Y
Sbjct: 86 ---QALPVVVFFHGGGFVLFSAASCYYDRLCRRICRELRAVVVSVNYRLAGPARRFPAAY 142
Query: 150 EDGETALKFLDEN---KSVLPENVDVSKCFLAGDSAGANLAHHVAVR------ACKAGLQ 200
+DG AL++LD N ++ VD+S CFLAGDSAG N+ HHVA R A +
Sbjct: 143 DDGLAALRYLDANGLAEAAGVAAVDLSSCFLAGDSAGGNMVHHVAQRWAAASAASPSSST 202
Query: 201 RIRVAGLISMQPFFGGEERTEAEIRLE-GSLMISMARTDWMWKVFLPEGSNRDHNAANVS 259
+R+AG + +QPFFGGEERTE E+ L+ +L +S+ARTD+ W+ FLPEG+ RDH AA+V
Sbjct: 203 TLRLAGAVLIQPFFGGEERTEEELELDKAALTLSLARTDYYWREFLPEGATRDHPAAHVC 262
Query: 260 GPNAEDLSRLD-YPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAFP 318
G D+ + +P +V +GG D L WQ RY E L+ GK +++EYP +HGF FP
Sbjct: 263 GGGEHDVEVAEAFPAAMVAIGGFDLLKGWQARYVEALRGKGKAVRVVEYPGAIHGFCLFP 322
Query: 319 NVPEASQLILQIKDFI 334
+ ++ + + ++K F+
Sbjct: 323 ELADSGEFVEEMKLFV 338
>UniRef100_Q5Z4C9 Putative esterase [Oryza sativa]
Length = 360
Score = 255 bits (651), Expect = 1e-66
Identities = 140/347 (40%), Positives = 207/347 (59%), Gaps = 19/347 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP-INGVSTKDITVD 66
++ P RL + L+ DA R +G++NR L + D + A P GVS+ D+TVD
Sbjct: 10 RVALPCAVRLRLCLLEAAIDATQRRDGAINRPLFSLYDRRAPADPRPDAAGVSSTDVTVD 69
Query: 67 AESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSR 126
A +W R+FTPT S+T TT PV+++FHGGGF S AS +DT CR
Sbjct: 70 ASRGLWARVFTPTAPEHEHSSSSST-TTPRPVIVYFHGGGFAMFSAASRPFDTHCRTLCA 128
Query: 127 ELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLD----ENKSVLPENVDVSKCFLAGDSA 182
+ VVVSV+YR PE+R+P Y+DGE L++L ++ +P VD+S CFLAGDSA
Sbjct: 129 GVGAVVVSVDYRLAPEHRFPAAYDDGEAVLRYLATTGLRDEHGVP--VDLSACFLAGDSA 186
Query: 183 GANLAHHVAVRACKAGL--------QRIRVAGLISMQPFFGGEERTEAEIRLEG-SLMIS 233
G N+AHHVA R + +AG+I ++P+FGGEERT+AE LEG + +++
Sbjct: 187 GGNIAHHVAQRWTTTSAATPPPPSDNPVHLAGVILLEPYFGGEERTKAERALEGVAPVVN 246
Query: 234 MARTDWMWKVFLPEGSNRDHNAANVSG-PNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYY 292
+ R+D W+ FLPEG++R+H AA+V+G E + +P +V VGGLD L DW +RY
Sbjct: 247 IRRSDRWWRAFLPEGADRNHPAAHVTGDAGPEPELQEAFPPAMVVVGGLDPLQDWDRRYA 306
Query: 293 EWLKISGKKAQLIEYPNMMHGFYAFPN-VPEASQLILQIKDFINNRV 338
L+ GK +++E+P +H FY FP + +L+ +I+ F+ + +
Sbjct: 307 GMLRRKGKAVRVVEFPEAIHAFYFFPEFAGDIRKLVGEIRAFVEDSI 353
>UniRef100_Q940G6 Hypothetical protein At5g27320 [Arabidopsis thaliana]
Length = 344
Score = 245 bits (625), Expect = 1e-63
Identities = 134/337 (39%), Positives = 190/337 (55%), Gaps = 18/337 (5%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
K PL T + IS + R +G+ NR L FLD K A A P+NGV + D+ +D
Sbjct: 13 KTVVPLNTWVLISNFKLAYNLLRRPDGTFNRHLAEFLDRKVPANANPVNGVFSFDVIIDR 72
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSL---------PVVIFFHGGGFTFMSPASLSYD 118
++ + R++ P A G++ T L PV++FFHGG F S S YD
Sbjct: 73 QTNLLSRVYRP------ADAGTSPSITDLQNPVDGEIVPVIVFFHGGSFAHSSANSAIYD 126
Query: 119 TICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENVDVSKCFLA 178
T+CRR VVVSVNYRR PE RYP Y+DG LK+++ + + + + FLA
Sbjct: 127 TLCRRLVGLCGAVVVSVNYRRAPENRYPCAYDDGWAVLKWVNSSSWLRSKKDSKVRIFLA 186
Query: 179 GDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTD 238
GDS+G N+ H+VAVRA ++ RI V G I + P FGG ERTE+E RL+G +++ D
Sbjct: 187 GDSSGGNIVHNVAVRAVES---RIDVLGNILLNPMFGGTERTESEKRLDGKYFVTVRDRD 243
Query: 239 WMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKIS 298
W W+ FLPEG +R+H A + GP ++ L L +P +LV V GLD + DWQ +Y E LK +
Sbjct: 244 WYWRAFLPEGEDREHPACSPFGPRSKSLEGLSFPKSLVVVAGLDLIQDWQLKYAEGLKKA 303
Query: 299 GKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFIN 335
G++ +L+ GFY PN ++ +I F+N
Sbjct: 304 GQEVKLLYLEQATIGFYLLPNNNHFHTVMDEIAAFVN 340
>UniRef100_Q8H5P4 Putative esterase [Oryza sativa]
Length = 346
Score = 237 bits (604), Expect = 4e-61
Identities = 141/353 (39%), Positives = 205/353 (57%), Gaps = 37/353 (10%)
Query: 5 KKPKLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP-INGVSTKDI 63
++ LP+P++ RL + DA R +GSVNR L + D + A P GVS+ DI
Sbjct: 9 RRVALPWPVRLRLCV--FEAAIDATQRRDGSVNRFLFSLFDRRAPADPRPDAAGVSSTDI 66
Query: 64 TVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRR 123
TVDA +W R+F + + PVV++FHGGGFT S AS +YD +CR
Sbjct: 67 TVDASRGLWARVFY------------SPSPSPRPVVVYFHGGGFTLFSAASRAYDALCRT 114
Query: 124 FSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENV---DVSKCFLAGD 180
L VVVSV+YR PE+R P Y+DGE L++L + LP++V DVS CF+ GD
Sbjct: 115 ----LCAVVVSVDYRLAPEHRAPAAYDDGEAVLRYL--GATGLPDHVGPVDVSTCFVVGD 168
Query: 181 SAGANLAHHVAVRACKAGLQR--------IRVAGLISMQPFFGGEERTEAEIRLEG-SLM 231
SAG N+AHHVA R + +AG+I +QP F GEERTE+E L+G + +
Sbjct: 169 SAGGNIAHHVAQRWTATATTTTTTTDNPVVHLAGVILIQPCFSGEERTESERALDGVAPV 228
Query: 232 ISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLD--YPDTLVFVGGLDGLYDWQK 289
++ R+D WK FLPEG++R+H AA+V + +D + L +P +V VGGLD L DW +
Sbjct: 229 LNTRRSDLSWKAFLPEGADRNHPAAHVVTGDDDDDAELHEAFPPAMVVVGGLDPLQDWDR 288
Query: 290 RYYEWLKISGKKAQLIEYPNMMHGFYAFPN--VPEASQLILQIKDFINNRVSN 340
RY L+ GK A+++E+P +H FY FP + +L+ +I+ F+ +++
Sbjct: 289 RYAAMLRRKGKAARVVEFPEAIHSFYFFPEFLADDHRKLVGEIRAFVEECITS 341
>UniRef100_Q9LYC1 Hypothetical protein T20O10_110 [Arabidopsis thaliana]
Length = 358
Score = 229 bits (583), Expect = 1e-58
Identities = 133/334 (39%), Positives = 187/334 (55%), Gaps = 16/334 (4%)
Query: 12 PLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKI 71
PL T + IS R +GS NR L FLD K A + P++GV + D VD+ + +
Sbjct: 17 PLNTWVLISNFKLAYKVLRRPDGSFNRDLAEFLDRKVPANSFPLDGVFSFD-HVDSTTNL 75
Query: 72 WFRLFTPTGINASAGGGSNTETTSL------PVVIFFHGGGFTFMSPASLSYDTICRRFS 125
R++ P + G+ T L PV+IFFHGG FT S S YDT CRR
Sbjct: 76 LTRIYQPASLLHQTRHGTLELTKPLSTTEIVPVLIFFHGGSFTHSSANSAIYDTFCRRLV 135
Query: 126 RELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN---KSVLPENVDVSKCFLAGDSA 182
VVVVSV+YRR+PE+RYP Y+DG AL ++ +S NV V +LAGDS+
Sbjct: 136 TICGVVVVSVDYRRSPEHRYPCAYDDGWNALNWVKSRVWLQSGKDSNVYV---YLAGDSS 192
Query: 183 GANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWK 242
G N+AH+VAVRA G ++V G I + P FGG+ERT++E L+G +++ DW W+
Sbjct: 193 GGNIAHNVAVRATNEG---VKVLGNILLHPMFGGQERTQSEKTLDGKYFVTIQDRDWYWR 249
Query: 243 VFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKA 302
+LPEG +RDH A N GP + L +++P +LV V GLD + DWQ Y + LK +G +
Sbjct: 250 AYLPEGEDRDHPACNPFGPRGQSLKGVNFPKSLVVVAGLDLVQDWQLAYVDGLKKTGLEV 309
Query: 303 QLIEYPNMMHGFYAFPNVPEASQLILQIKDFINN 336
L+ GFY PN L+ ++ F+++
Sbjct: 310 NLLYLKQATIGFYFLPNNDHFHCLMEELNKFVHS 343
>UniRef100_Q9MAA7 T12H1.8 protein [Arabidopsis thaliana]
Length = 345
Score = 228 bits (581), Expect = 2e-58
Identities = 124/329 (37%), Positives = 179/329 (53%), Gaps = 8/329 (2%)
Query: 12 PLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKI 71
PL T + IS + R +G+ NR L +LD K +A A P++GV + D+ +D +
Sbjct: 17 PLNTWVLISNFKVAYNILRRPDGTFNRHLAEYLDRKVTANANPVDGVFSFDVLIDRRINL 76
Query: 72 WFRLFTPTGINASAGGG-----SNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSR 126
R++ P + + +PV++FFHGG F S S YDT+CRR
Sbjct: 77 LSRVYRPAYADQEQPPSILDLEKPVDGDIVPVILFFHGGSFAHSSANSAIYDTLCRRLVG 136
Query: 127 ELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENVDVSKCFLAGDSAGANL 186
VVVSVNYRR PE YP Y+DG AL +++ + + FLAGDS+G N+
Sbjct: 137 LCKCVVVSVNYRRAPENPYPCAYDDGWIALNWVNSRSWLKSKKDSKVHIFLAGDSSGGNI 196
Query: 187 AHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLP 246
AH+VA+RA ++G+ V G I + P FGG ERTE+E L+G +++ DW WK FLP
Sbjct: 197 AHNVALRAGESGID---VLGNILLNPMFGGNERTESEKSLDGKYFVTVRDRDWYWKAFLP 253
Query: 247 EGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQLIE 306
EG +R+H A N P + L + +P +LV V GLD + DWQ Y E LK +G++ +L+
Sbjct: 254 EGEDREHPACNPFSPRGKSLEGVSFPKSLVVVAGLDLIRDWQLAYAEGLKKAGQEVKLMH 313
Query: 307 YPNMMHGFYAFPNVPEASQLILQIKDFIN 335
GFY PN ++ +I F+N
Sbjct: 314 LEKATVGFYLLPNNNHFHNVMDEISAFVN 342
>UniRef100_Q6L545 Hypothetical protein OJ1657_H11.10 [Oryza sativa]
Length = 354
Score = 227 bits (579), Expect = 3e-58
Identities = 128/340 (37%), Positives = 183/340 (53%), Gaps = 15/340 (4%)
Query: 8 KLPFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDA 67
K PL T + IS + R++G+ R L +LD + A A P+ GVS+ D +D
Sbjct: 13 KTVVPLHTWVLISNFKLSYNILRRADGTFERDLGEYLDRRVPANARPLEGVSSFDHIIDQ 72
Query: 68 ESKIWFRLFTPTGINASAGGGSNTETTSL------------PVVIFFHGGGFTFMSPASL 115
+ R++ + G + L PV+IFFHGG F S +S
Sbjct: 73 SVGLEVRIYRAAAEGDAEEGAAAVTRPILEFLTDAPAAEPFPVIIFFHGGSFVHSSASST 132
Query: 116 SYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPENVDVSKC 175
YD++CRRF + VVVSVNYRR PE+RYP Y+DG TALK++ + ++
Sbjct: 133 IYDSLCRRFVKLSKGVVVSVNYRRAPEHRYPCAYDDGWTALKWVMSQPFMRSGGDAQARV 192
Query: 176 FLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMA 235
FL+GDS+G N+AHHVAVRA G ++V G I + FGG ERTE+E RL+G +++
Sbjct: 193 FLSGDSSGGNIAHHVAVRAADEG---VKVCGNILLNAMFGGTERTESERRLDGKYFVTLQ 249
Query: 236 RTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYYEWL 295
DW WK +LPE ++RDH A N GPN L L + +L+ V GLD D Q Y + L
Sbjct: 250 DRDWYWKAYLPEDADRDHPACNPFGPNGRRLGGLPFAKSLIIVSGLDLTCDRQLAYADAL 309
Query: 296 KISGKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFIN 335
+ G ++++ N GFY PN +++ +I DF+N
Sbjct: 310 REDGHHVKVVQCENATVGFYLLPNTVHYHEVMEEISDFLN 349
>UniRef100_Q69J38 Putative PrMC3 [Oryza sativa]
Length = 367
Score = 220 bits (561), Expect = 4e-56
Identities = 128/333 (38%), Positives = 189/333 (56%), Gaps = 21/333 (6%)
Query: 10 PFPLKTRLSISLIMTLTDAACRSNGSVNRRLLNFLDNKTSAKATP-INGVSTKDITVDAE 68
P P + RL + + L A+ R +G+VNR LL+ D P GV++ D V +
Sbjct: 15 PLPWRARLLVGAVSVLHSASLRRDGTVNRFLLSLFDRVVPPNPAPDAAGVASSDHAVSDD 74
Query: 69 SKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSREL 128
++ R+F P G A GGG + LPVV++FHGGGF F S AS +D +CRRF+ +
Sbjct: 75 LRV--RMFFP-GAAARDGGGDH-----LPVVVYFHGGGFVFHSVASAQFDALCRRFASAI 126
Query: 129 NVVVVSVNYRRTPEYRYPTQYEDGETALKF-LDENKSVLPENVDVSKCFLAGDSAGANLA 187
VV SV++R PE+ +P Y+DG+ AL++ L LP + F+AGDSAG N+A
Sbjct: 127 PAVVASVDFRLAPEHGFPAPYDDGKAALRWVLAGAGGALPS--PPATVFVAGDSAGGNVA 184
Query: 188 HHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLPE 247
HHV R + V+GLI++QPFF GE T +E RL + S R W+W+ FLP
Sbjct: 185 HHVVARTPSS------VSGLIALQPFFAGETPTASEQRLRDAPFGSPERISWLWRAFLPP 238
Query: 248 GSNRDHNAANVSGPNAEDLS-RLDYPDTLVFVGGLDGLYDWQKRYYEWLKISGKKAQLI- 305
G+ RDH AANV D R +P T+V VGG D D Q+ Y + L+ +G +++
Sbjct: 239 GATRDHEAANVPAALRRDAERRRAFPPTMVCVGGWDAHQDRQRDYADALRAAGGAEEVVV 298
Query: 306 -EYPNMMHGFYAFPNVPEASQLILQIKDFINNR 337
E+P+ +H FY F ++ ++ +L+ ++ F+N R
Sbjct: 299 AEFPDAIHAFYIFDDLADSKRLLTEVTAFVNRR 331
>UniRef100_Q9FG13 Gb|AAD04946.2 [Arabidopsis thaliana]
Length = 329
Score = 176 bits (447), Expect = 6e-43
Identities = 107/288 (37%), Positives = 152/288 (52%), Gaps = 22/288 (7%)
Query: 61 KDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASLSYDTI 120
KD + + RL+ P S + T+LPVV+FFHGGGF F S + +
Sbjct: 50 KDSIYHKPNNLHLRLYKPI---------SASNRTALPVVVFFHGGGFCFGSRSWPHFHNF 100
Query: 121 CRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLPEN--------VDV 172
C + LN +VVS +YR PE+R P +ED E L +L + N VD
Sbjct: 101 CLTLASSLNALVVSPDYRLAPEHRLPAAFEDAEAVLTWLWDQAVSDGVNHWFEDGTDVDF 160
Query: 173 SKCFLAGDSAGANLAHHVAVRACKAGLQ--RIRVAGLISMQPFFGGEERTEAEIRLEGSL 230
+ F+ GDS+G N+AH +AVR ++ +RV G + M PFFGGEERT +E
Sbjct: 161 DRVFVVGDSSGGNIAHQLAVRFGSGSIELTPVRVRGYVLMGPFFGGEERTNSE-NGPSEA 219
Query: 231 MISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKR 290
++S+ D W++ LP G+ RDH+ AN GP + L + LV VGG + L D K
Sbjct: 220 LLSLDLLDKFWRLSLPNGATRDHHMANPFGPTSPTLESISLEPMLVIVGGSELLRDRAKE 279
Query: 291 Y-YEWLKISGKKAQLIEYPNMMHGFYA-FPNVPEASQLILQIKDFINN 336
Y Y+ K+ GK+ IE+ N HGFY+ +P+ A Q++ I DF+NN
Sbjct: 280 YAYKLKKMGGKRVDYIEFENKEHGFYSNYPSSEAAEQVLRIIGDFMNN 327
>UniRef100_Q94JD7 Putative PrMC3 [Oryza sativa]
Length = 361
Score = 173 bits (439), Expect = 5e-42
Identities = 109/297 (36%), Positives = 153/297 (50%), Gaps = 20/297 (6%)
Query: 55 INGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPAS 114
+ GV KD DA + R+F A GG LPV+++FHGGG+ +
Sbjct: 67 VPGVQWKDAVYDATHGLRVRVFKLAAAAAGDDGGK------LPVLVYFHGGGYCIGALDQ 120
Query: 115 LSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLP---ENVD 171
+ T C R + EL VV+SV YR PE+R PT +DG +L S P E+ +
Sbjct: 121 SPFHTFCLRAADELPAVVLSVQYRLAPEHRLPTAIDDGAAFFSWLRGAGSADPWLAESAE 180
Query: 172 VSKCFLAGDSAGANLAHHVAVRACKAGLQRI--------RVAGLISMQPFFGGEERTEAE 223
+++ F++G SAGANLAHHVAVR +G Q + RVAG + + FFGG ERT AE
Sbjct: 181 LARTFISGVSAGANLAHHVAVRVA-SGRQPVVDDVDPVVRVAGYVLLDAFFGGVERTAAE 239
Query: 224 IRLEGSL-MISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLD 282
+ ++++ D W++ LP G+ RDH AN GP + L + P LV G D
Sbjct: 240 ANPPADVSLLTVEMADQFWRLALPAGATRDHPVANPFGPESPSLEAVALPPALVVASGGD 299
Query: 283 GLYDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAF-PNVPEASQLILQIKDFINNRV 338
LYD Y LK GK +L+E+ HGF P PE S++I +K F++ +
Sbjct: 300 VLYDRVVGYAARLKEMGKAVELVEFEGAQHGFSVIQPWSPETSEVIQVLKRFVHKAI 356
>UniRef100_Q8GZM8 Putative serine hydrolase [Vitis vinifera]
Length = 310
Score = 169 bits (428), Expect = 1e-40
Identities = 102/285 (35%), Positives = 156/285 (53%), Gaps = 15/285 (5%)
Query: 56 NGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASL 115
NG +KD+ +++ I R+F P + S+G LPV+++FHGGGF S
Sbjct: 36 NGYKSKDVIINSTKPISARIFLPD-VPGSSG--------RLPVLVYFHGGGFCLGSTTWF 86
Query: 116 SYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENKSVLP--ENVDVS 173
Y T F+ +V+SV+YR PE R P Y+D ++L++L S P E D+S
Sbjct: 87 GYHTFLGDFAVASQSIVLSVDYRHAPENRLPIAYDDCYSSLEWLSCQVSSEPWLERADLS 146
Query: 174 KCFLAGDSAGANLAHHVAVRACKA-GLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLMI 232
+ FL+GDSAG N+ H+VA+R + ++++ GL+ + PFFG EER E E G
Sbjct: 147 RVFLSGDSAGGNIVHNVALRTIQEQSCDQVKIKGLLLIHPFFGSEERIEKE--RAGGEAE 204
Query: 233 SMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRYY 292
++A TDWMWK+ LPEGSNRDH N +P +V+V GLD L + Y
Sbjct: 205 NLALTDWMWKLSLPEGSNRDHYWCNYEMAELSRAEWCRFPPAVVYVAGLDFLKERGVMYA 264
Query: 293 EWLKISGKKAQLIEYPNMMHGFYAFPNVPEASQLI-LQIKDFINN 336
+L+ +G + +L+E H ++ EA++L+ Q+ +FI+N
Sbjct: 265 AFLEKNGVEVKLVEAEGEKHVYHMLHPESEATRLLQKQMSEFIHN 309
>UniRef100_Q9ZSE0 PrMC3 [Pinus radiata]
Length = 319
Score = 164 bits (416), Expect = 2e-39
Identities = 98/291 (33%), Positives = 148/291 (50%), Gaps = 20/291 (6%)
Query: 56 NGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPASL 115
+G +KD+ +D I RLF P + + LP++ +FHGGGF + A
Sbjct: 40 DGARSKDVVIDPVKGISARLFLPAELPLAQ---------KLPLLFYFHGGGFCIGTTAWE 90
Query: 116 SYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDEN----KSVLPENVD 171
Y + +V+SV+YR PE+R P Y+D A++++ + L + D
Sbjct: 91 GYHLFLSLLAATTRALVISVDYRLAPEHRLPAAYDDCFDAVEWVASGGGKAEPWLDAHAD 150
Query: 172 VSKCFLAGDSAGANLAHHVAVRACKAGLQRIRVAGLISMQPFFGGEERTEAEIRLEGSLM 231
+CFLAG+SAG N+AH V R L +++ GLI + P+FG EER E E G
Sbjct: 151 YGRCFLAGESAGGNIAHVVGSRTADQDLGPLKIRGLIVIHPYFGSEERIECEKVAAGDDA 210
Query: 232 ISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLYDWQKRY 291
++ D W++ LP GS+RD+ N GP + DL ++ P LV V GLD L Y
Sbjct: 211 AALELNDLFWRLALPPGSDRDYPTCNPRGPRSADLRKVPLPPVLVTVAGLDLLKTRGLLY 270
Query: 292 YEWLKISGKKAQLIEYPNMMHGFYAFPNVPEASQLILQIKDFINNRVSNFI 342
YE L+ GK+A+L+E +H ++ F EA++L + R+S FI
Sbjct: 271 YELLQSCGKEAELMEAEGEIHAYHVFHPRSEATRL-------LQERMSQFI 314
>UniRef100_Q9LVB8 HSR203J protein-like protein [Arabidopsis thaliana]
Length = 327
Score = 164 bits (416), Expect = 2e-39
Identities = 107/309 (34%), Positives = 164/309 (52%), Gaps = 13/309 (4%)
Query: 33 NGSVNRRLLNFLDNKTSAKATPINGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTE 92
+GS+ R L NF + +P+N +KD+ V+ W RL+ P+ SA N
Sbjct: 21 DGSITRDLSNFPCTAATPDPSPLNPAVSKDLPVNQLKSTWLRLYLPS----SAVNEGNVS 76
Query: 93 TTSLPVVIFFHGGGFTFMSPASLSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDG 152
+ LP+V+++HGGGF S + C +R+LN +VVS +YR PE+R P Y+DG
Sbjct: 77 SQKLPIVVYYHGGGFILCSVDMQLFHDFCSEVARDLNAIVVSPSYRLAPEHRLPAAYDDG 136
Query: 153 ETALKFL-DENKSVLPENVDVSKCFLAGDSAGANLAHHVAVRACK--AGLQRIRVAGLIS 209
AL ++ + + + D S FL G SAG NLA++V +R+ + L +++ GLI
Sbjct: 137 VEALDWIKTSDDEWIKSHADFSNVFLMGTSAGGNLAYNVGLRSVDSVSDLSPLQIRGLIL 196
Query: 210 MQPFFGGEERTEAEIRLEGSLMISMARTDWMWKVFLPEGSNRDHNAANVS-GPNAEDLSR 268
PFFGGEER+E+EIRL + TD MW + LP G +RDH +N + G +E L +
Sbjct: 197 HHPFFGGEERSESEIRLMNDQVCPPIVTDVMWDLSLPVGVDRDHEYSNPTVGDGSEKLEK 256
Query: 269 LD-YPDTLVFVGGLDG-LYDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAFPNVP-EASQ 325
+ ++ +GG D + D QK + +K G +++E+ H A P +
Sbjct: 257 IGRLRWKVMMIGGEDDPMIDLQKDVAKLMKKKG--VEVVEHYTGGHVHGAEIRDPSKRKT 314
Query: 326 LILQIKDFI 334
L L IK+FI
Sbjct: 315 LFLSIKNFI 323
>UniRef100_Q852M6 Putative esterase [Oryza sativa]
Length = 342
Score = 162 bits (411), Expect = 9e-39
Identities = 94/291 (32%), Positives = 152/291 (51%), Gaps = 19/291 (6%)
Query: 55 INGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPAS 114
++ V KD+ DA + R++ P + LPV+++FHGGG+ S
Sbjct: 44 LSTVQWKDVVYDAGRGLKLRVYRPPAATVAG--------EKLPVLVYFHGGGYFIGSFEM 95
Query: 115 LSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFL--------DENKSVL 166
++ C R + EL VV+S +YR PE+R P ++D TA+ ++ D L
Sbjct: 96 DNFHACCLRLAHELPAVVLSADYRLAPEHRLPAAHDDAATAMSWVRDQAVASGDAADPWL 155
Query: 167 PENVDVSKCFLAGDSAGANLAHHVAVR--ACKAGLQRIRVAGLISMQPFFGGEERTEAEI 224
E+ D + F++GDSAGA + HHVA+R + + + RVAG + P+FGGEERT +E
Sbjct: 156 AESADFGRVFVSGDSAGAGIVHHVALRLGSGQIAVDPARVAGCALLFPYFGGEERTRSEA 215
Query: 225 RLEGSLMISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGL 284
+++ +D W++ LP G+ RDH AN GP + + + P LV V LD L
Sbjct: 216 EYPPGPFLTLPFSDQGWRLALPRGATRDHPLANPFGPESPAMDAVALPPLLVVVAQLDLL 275
Query: 285 YDWQKRYYEWLKISGKKAQLIEYPNMMHGFYAFPNVPEA-SQLILQIKDFI 334
D Y L+ GK+ +++E+ HGF+A + +A S+L+ ++ F+
Sbjct: 276 RDRDVDYAARLRAMGKQVEMVEFEGQHHGFFAVEPLGDAGSELVRVVRRFV 326
>UniRef100_Q93VD4 Putative PrMC3 [Oryza sativa]
Length = 337
Score = 162 bits (410), Expect = 1e-38
Identities = 101/291 (34%), Positives = 152/291 (51%), Gaps = 19/291 (6%)
Query: 55 INGVSTKDITVDAESKIWFRLFTPTGINASAGGGSNTETTSLPVVIFFHGGGFTFMSPAS 114
+ GV KD+ DA + R++ P +AG + LPV++ FHGGG+ +
Sbjct: 48 VPGVQWKDLVYDATHGLKLRVYRPP----TAG-----DAERLPVLVCFHGGGYCLGTFEK 98
Query: 115 LSYDTICRRFSRELNVVVVSVNYRRTPEYRYPTQYEDGETALKFLDENK-------SVLP 167
S+ C+R + EL VV+S +YR PE+R P +DG L +L + S L
Sbjct: 99 PSFHCCCQRLASELRAVVLSADYRLGPEHRLPAAIDDGAAVLSWLRDQAMSGPGADSWLA 158
Query: 168 ENVDVSKCFLAGDSAGANLAHHVAVRACKAGL--QRIRVAGLISMQPFFGGEERTEAEIR 225
E+ D ++ F+AG+SAG N++HHVAV L +RVAG + + PFFGG ER +E
Sbjct: 159 ESADFARVFVAGESAGGNMSHHVAVLIGSGQLTVDPLRVAGYMLLTPFFGGVERAPSEAE 218
Query: 226 LEGSLMISMARTDWMWKVFLPEGSNRDHNAANVSGPNAEDLSRLDYPDTLVFVGGLDGLY 285
+ +D +W++ LPEG+ RDH AN GP++ L+ + +P LV V G D L+
Sbjct: 219 PPAGAFFTPDMSDKLWRLSLPEGATRDHPVANPFGPDSPSLAAVAFPPVLVVVAGRDILH 278
Query: 286 DWQKRYYEWLKISGKKAQLIEYPNMMHGFYAF-PNVPEASQLILQIKDFIN 335
D Y LK K +L+ + H F + P A++LI +K FI+
Sbjct: 279 DRTVHYAARLKEMEKPVELVTFEEEKHLFLSLQPWSEPANELIRVMKRFIH 329
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 580,654,121
Number of Sequences: 2790947
Number of extensions: 24335978
Number of successful extensions: 58125
Number of sequences better than 10.0: 1274
Number of HSP's better than 10.0 without gapping: 407
Number of HSP's successfully gapped in prelim test: 867
Number of HSP's that attempted gapping in prelim test: 56173
Number of HSP's gapped (non-prelim): 1374
length of query: 342
length of database: 848,049,833
effective HSP length: 128
effective length of query: 214
effective length of database: 490,808,617
effective search space: 105033044038
effective search space used: 105033044038
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC139853.10


