

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139745.3 - phase: 0 /pseudo
(330 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
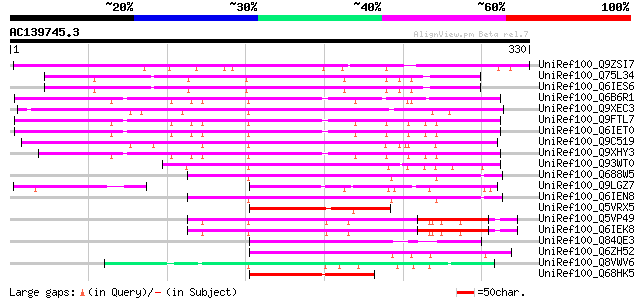
UniRef100_Q9ZSI7 Probable WRKY transcription factor 47 [Arabidop... 206 7e-52
UniRef100_Q75L34 Hypothetical protein OJ1127_B08.13 [Oryza sativa] 162 9e-39
UniRef100_Q6IES6 WRKY transcription factor 5 [Oryza sativa] 159 1e-37
UniRef100_Q6B6R1 Transcription factor WRKY10 [Oryza sativa] 149 8e-35
UniRef100_Q9XEC3 Probable WRKY transcription factor 42 [Arabidop... 149 1e-34
UniRef100_Q9FTL7 Putative WRKY DNA binding protein [Oryza sativa] 147 4e-34
UniRef100_Q6IET0 WRKY transcription factor 1 [Oryza sativa] 147 4e-34
UniRef100_Q9C519 WRKY transcription factor 6 [Arabidopsis thaliana] 145 1e-33
UniRef100_Q9XHY3 Putative WRKY DNA binding protein [Oryza sativa] 143 7e-33
UniRef100_Q93WT0 Probable WRKY transcription factor 31 [Arabidop... 133 6e-30
UniRef100_Q688W5 Putative WRKY transcription factor [Oryza sativa] 129 1e-28
UniRef100_Q9LGZ7 Similar to Arabidopsis thaliana chromosome 4 BA... 128 2e-28
UniRef100_Q6IEN8 WRKY transcription factor 43 [Oryza sativa] 126 9e-28
UniRef100_Q5VRX5 WRKY transcription factor 6-like [Oryza sativa] 92 3e-17
UniRef100_Q5VP49 Putative WRKY transcription factor [Oryza sativa] 90 7e-17
UniRef100_Q6IEK8 WRKY transcription factor 73 [Oryza sativa] 90 7e-17
UniRef100_Q84QE3 Avr9/Cf-9 rapidly elicited protein 126 [Nicotia... 89 2e-16
UniRef100_Q6ZH52 Putative WRKY transcription factor [Oryza sativa] 86 2e-15
UniRef100_Q8VWV6 Probable WRKY transcription factor 61 [Arabidop... 85 3e-15
UniRef100_Q68HK5 WRKY6-1 [Pelargonium hortorum] 84 4e-15
>UniRef100_Q9ZSI7 Probable WRKY transcription factor 47 [Arabidopsis thaliana]
Length = 489
Score = 206 bits (524), Expect = 7e-52
Identities = 163/415 (39%), Positives = 210/415 (50%), Gaps = 95/415 (22%)
Query: 3 SSAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQ 62
S S + + ++T++S L+ EL R+ EENHKL+ +L+++++SY+ LQ ++ + Q Q
Sbjct: 82 SCHGTSSNDGDDKTKTQISRLKLELERLHEENHKLKHLLDEVSESYNDLQRRVLLARQTQ 141
Query: 63 KPN-HGQNMEENHGMVSEQIFLN------NNNASVSDGKQACPHD------------HPA 103
H + E+ S Q N N+ + K+ P D P
Sbjct: 142 VEGLHHKQHEDVPQAGSSQALENRRPKDMNHETPATTLKRRSPDDVDGRDMHRGSPKTPR 201
Query: 104 EDSSHSSKLEEPTQ--DLIPFKKARVSIRARSEA--------------------PLPTI- 140
D + S+ EE D +P++KARVS+RARS+A P P
Sbjct: 202 IDQNKSTNHEEQQNPHDQLPYRKARVSVRARSDATTVNDGCQWRKYGQKMAKGNPCPRAY 261
Query: 141 ----VALWL*DVQFANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSST 196
+A+ + RCAED TIL TTYEGNHNHPLPP+ATA+A TTSAAAAMLLS S+
Sbjct: 262 YRCTMAVGCPVRKQVQRCAEDTTILTTTYEGNHNHPLPPSATAMAATTSAAAAMLLSGSS 321
Query: 197 SS----TLRKESATGYLS--NSFPYATMATSTLSASQPFPTITLDFTQ------------ 238
SS TL SAT S ++FPY T +TLSAS PFPTITLD T
Sbjct: 322 SSNLHQTLSSPSATSSSSFYHNFPY-TSTIATLSASAPFPTITLDLTNPPRPLQPPPQFL 380
Query: 239 ------------NHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSSMVESVSAAISSDPN 286
N SM++N L L P L Q PP MV+SV AAI+ DPN
Sbjct: 381 SQYGPAAFLPNANQIRSMNNNNQQL-------LIPNLFGPQAPPREMVDSVRAAIAMDPN 433
Query: 287 FTTALAAAISSIIGPQRSGDGNN-----NLAGVVPG------SPQLPQSCTTFST 330
FT ALAAAIS+IIG + + NN N G SPQLPQSCTTFST
Sbjct: 434 FTAALAAAISNIIGGGNNDNNNNTDINDNKVDAKSGGSSNGDSPQLPQSCTTFST 488
>UniRef100_Q75L34 Hypothetical protein OJ1127_B08.13 [Oryza sativa]
Length = 502
Score = 162 bits (411), Expect = 9e-39
Identities = 132/360 (36%), Positives = 171/360 (46%), Gaps = 88/360 (24%)
Query: 23 LESELRRVQEENHKLRIMLEQITKSYSQLQ------AQLFITLQKQKPNHGQNMEENHGM 76
+E ELR+ EEN +LR LE++T SY L QL Q+Q P G + +
Sbjct: 99 VEGELRQAGEENRRLRRRLEELTSSYGALYHQLVQAQQLHTKHQQQAPIAGVQLLDALAA 158
Query: 77 VSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKL------------------------ 112
S A+V DG + D D + S L
Sbjct: 159 ASPASHRRRAAAAV-DGDRTADSDGGEGDENVSPSLGSKRPAAAATLTRLTPESGSGGEN 217
Query: 113 -----EEPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQ------------------ 149
+ P ++ P +KARVS+RARSEAP+ + W Q
Sbjct: 218 NGGGEQAPAAEMAPCRKARVSVRARSEAPMISDGCQWRKYGQKMAKGNPCPRAYYRCTMA 277
Query: 150 -------FANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRK 202
RCAEDK+ILITTYEG HNHPLPPAA A+A TTSAAAAMLLS S
Sbjct: 278 SQCPVRKQVQRCAEDKSILITTYEGTHNHPLPPAAAAMAKTTSAAAAMLLSGPAVSRDAL 337
Query: 203 ESATGYLSN-----SFPYATMATSTLSASQPFPTITLDFTQ----------NHNLSMHH- 246
+A ++ PYA +TLSAS PFPTITLD TQ L +H
Sbjct: 338 FAAHHHVVAPPPFFHHPYAGSTMATLSASAPFPTITLDLTQPPPTTTTTAAAAMLQLHRP 397
Query: 247 ---NRVPLPLF----FSHKLPPLLQLGQPPPSSMVESVSAAISSDPNFTTALAAAISSII 299
+ +P ++ SH+ P +L PPPSS+VE+++AAI+ DPNFTTA+AAA+SSI+
Sbjct: 398 YAFSSLPFSMYGAGGGSHRPPVVL----PPPSSVVETMTAAITRDPNFTTAVAAALSSIM 453
>UniRef100_Q6IES6 WRKY transcription factor 5 [Oryza sativa]
Length = 502
Score = 159 bits (402), Expect = 1e-37
Identities = 130/360 (36%), Positives = 171/360 (47%), Gaps = 88/360 (24%)
Query: 23 LESELRRVQEENHKLRIMLEQITKSYSQLQ------AQLFITLQKQKPNHGQNMEENHGM 76
+E ELR+ EEN +LR LE++T SY L QL Q+Q P G + +
Sbjct: 99 VEGELRQAGEENRRLRRRLEELTSSYGALYHQLVQAQQLHTKHQQQAPIAGVQLLDALAA 158
Query: 77 VSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKL------------------------ 112
S A+V DG + D D + S L
Sbjct: 159 ASPASHRRRAAAAV-DGDRTADSDGGEGDENVSPSLGSKRPAAAATLTRLTPESGSGGEN 217
Query: 113 -----EEPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQ------------------ 149
+ P ++ P +KARVS+RARSEAP+ + W Q
Sbjct: 218 NGGGEQAPAAEMAPCRKARVSVRARSEAPMISDGCQWRKYGQKMAKGNPCPRAYYRCTMA 277
Query: 150 -------FANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRK 202
RCA+DK+ILITTYEG H+HPLPPAA A+A TTSAAAAMLLS S
Sbjct: 278 SQCPVRKQVQRCAKDKSILITTYEGTHSHPLPPAAAAMAKTTSAAAAMLLSGPAVSRDAL 337
Query: 203 ESATGYLSN-----SFPYATMATSTLSASQPFPTITLDFTQ----------NHNLSMHH- 246
+A ++ PYA +TLSAS PFPTITLD TQ L +H
Sbjct: 338 FAAHHHVVAPPPFFHHPYAGSTMATLSASAPFPTITLDLTQPPPTTTTTAAAAMLQLHRP 397
Query: 247 ---NRVPLPLF----FSHKLPPLLQLGQPPPSSMVESVSAAISSDPNFTTALAAAISSII 299
+ +P ++ SH+ P +L PPPSS+VE+++AAI+ DPNFTTA+AAA+SSI+
Sbjct: 398 YAFSSLPFSMYGAGGGSHRPPVVL----PPPSSVVETMTAAITRDPNFTTAVAAALSSIM 453
>UniRef100_Q6B6R1 Transcription factor WRKY10 [Oryza sativa]
Length = 580
Score = 149 bits (377), Expect = 8e-35
Identities = 142/446 (31%), Positives = 190/446 (41%), Gaps = 145/446 (32%)
Query: 4 SAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQK 63
+A+ E S E++ +++EL R+ EEN +LR ML Q+T SY LQ L +Q++
Sbjct: 114 AASRPDHEEKSRSSNELAAMQAELGRMNEENQRLRGMLTQVTTSYQALQMHLVALMQQRP 173
Query: 64 ------------PNHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDH-------PAE 104
P H E G V + FL+ +S + G+ A + P
Sbjct: 174 QMMQPPTQPEPPPPHQDGKAE--GAVVPRQFLDLGPSSGAGGEAAEEPSNSSTEAGSPRR 231
Query: 105 DSSHSSKLEE---------------PTQDLIP--------------------FKKARVSI 129
SS +K +E P + + P +KARVS+
Sbjct: 232 SSSTGNKDQERGDSPDAPSTAAAWLPGRAMAPQMGAAGAAGKSHDQQAQDANMRKARVSV 291
Query: 130 RARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKTILITT 164
RARSEAP+ W Q RCAED++ILITT
Sbjct: 292 RARSEAPIIADGCQWRKYGQKMAKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITT 351
Query: 165 YEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATM-----A 219
YEG HNHPLPPAA A+A TTSAAA+MLLS S S + A G +S++F T+ +
Sbjct: 352 YEGTHNHPLPPAAMAMASTTSAAASMLLSGSMPSA---DGAAGLMSSNFLARTVLPCSSS 408
Query: 220 TSTLSASQPFPTITLDFTQNHNLSMHHNRVPL----------------------PLF--- 254
+T+SAS PFPT+TLD T H N VPL P F
Sbjct: 409 MATISASAPFPTVTLDLT--HAPPGAPNAVPLNAARPGAPAPQFQVPLPGGGMAPAFAVP 466
Query: 255 ----------------------------FSHKLPPLLQLGQPPPSSMVESVSAAISSDPN 286
F+ PP+ QL P S V + +AAI++DPN
Sbjct: 467 PQVLYNQSKFSGLQMSSDSAEAAAAAAQFAQPRPPIGQL-PGPLSDTVSAAAAAITADPN 525
Query: 287 FTTALAAAISSIIGPQRSGDGNNNLA 312
FT ALAAAI+SIIG Q + N+ A
Sbjct: 526 FTVALAAAITSIIGGQHAAAAGNSNA 551
>UniRef100_Q9XEC3 Probable WRKY transcription factor 42 [Arabidopsis thaliana]
Length = 528
Score = 149 bits (376), Expect = 1e-34
Identities = 135/424 (31%), Positives = 188/424 (43%), Gaps = 120/424 (28%)
Query: 6 AVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPN 65
+V EE + ++ E + L EL++ E+N +L+ ML Q T +++ LQ QL +++Q+ +
Sbjct: 101 SVDMEE--KRTKCENAQLREELKKASEDNQRLKQMLSQTTNNFNSLQMQLVAVMRQQEDH 158
Query: 66 HGQNMEENHG----------MVSEQIF------------------------LNNNNASVS 91
H EN+ MV Q L ++S
Sbjct: 159 HHLATTENNDNVKNRHEVPEMVPRQFIDLGPHSDEVSSEERTTVRSGSPPSLLEKSSSRQ 218
Query: 92 DGKQACPHDHPAEDSSH--------------------------SSKLEEPTQDLIPFKKA 125
+GK+ + E S+ SSK+ E +KA
Sbjct: 219 NGKRVLVREESPETESNGWRNPNKVPKHHASSSICGGNGSENASSKVIEQAAAEATMRKA 278
Query: 126 RVSIRARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKTI 160
RVS+RARSEAP+ + W Q RCAED+TI
Sbjct: 279 RVSVRARSEAPMLSDGCQWRKYGQKMAKGNPCPRAYYRCTMAVGCPVRKQVQRCAEDRTI 338
Query: 161 LITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTL-RKESATGYLSNSFPYATMA 219
LITTYEGNHNHPLPPAA +A TT+AAA+MLLS ST S + T L+ + + +
Sbjct: 339 LITTYEGNHNHPLPPAAMNMASTTTAAASMLLSGSTMSNQDGLMNPTNLLARTILPCSSS 398
Query: 220 TSTLSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQ------------ 267
+T+SAS PFPTITLD T++ N +N PL + L++L Q
Sbjct: 399 MATISASAPFPTITLDLTESPN---GNNPTNNPLMQFSQRSGLVELNQSVLPHMMGQALY 455
Query: 268 -------------PPPSSMVESVS---AAISSDPNFTTALAAAISSII-GPQRSGDGNNN 310
P + ESVS AAI+S+PNF ALAAAI+SII G +GNNN
Sbjct: 456 YNQQSKFSGLHMPSQPLNAGESVSAATAAIASNPNFAAALAAAITSIINGSNNQQNGNNN 515
Query: 311 LAGV 314
+ V
Sbjct: 516 NSNV 519
>UniRef100_Q9FTL7 Putative WRKY DNA binding protein [Oryza sativa]
Length = 635
Score = 147 bits (371), Expect = 4e-34
Identities = 139/446 (31%), Positives = 190/446 (42%), Gaps = 142/446 (31%)
Query: 4 SAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQK 63
+A+ E S E++ +++EL R+ EEN +LR ML Q+T SY LQ L +Q++
Sbjct: 166 AASRPDHEEKSRSSNELAAMQAELGRMNEENQRLRGMLTQVTTSYQALQMHLVALMQQRP 225
Query: 64 ------------PNHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDH-------PAE 104
P H E G V + FL+ +S + G+ A + P
Sbjct: 226 QMMQPPTQPEPPPPHQDGKAE--GAVVPRQFLDLGPSSGAGGEAAEEPSNSSTEAGSPRR 283
Query: 105 DSSHSSKLEE---------------PTQDLIP--------------------FKKARVSI 129
SS +K +E P + + P +KARVS+
Sbjct: 284 SSSTGNKDQERGDSPDAPSTAAAWLPGRAMAPQMGAAGAAGKSHDQQAQDANMRKARVSV 343
Query: 130 RARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKTILITT 164
RARSEAP+ W Q RCAED++ILITT
Sbjct: 344 RARSEAPIIADGCQWRKYGQKMAKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITT 403
Query: 165 YEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATM-----A 219
YEG HNHPLPPAA A+A TTSAAA+MLLS S S + A G +S++F T+ +
Sbjct: 404 YEGTHNHPLPPAAMAMASTTSAAASMLLSGSMPSA---DGAAGLMSSNFLARTVLPCSSS 460
Query: 220 TSTLSASQPFPTITLDFTQ--------------NHNLSMHHNRVPLP---LFFSHKLPPL 262
+T+SAS PFPT+TLD T +VPLP + + +PP
Sbjct: 461 MATISASAPFPTVTLDLTHAPPGAPNAVPLNAARPGAPAPQFQVPLPGGGMAPAFAVPPQ 520
Query: 263 L---------------------------QLGQPPP---------SSMVESVSAAISSDPN 286
+ Q QP P S V + +AAI++DPN
Sbjct: 521 VLYNQSKFSGLQMSSDSAEAAAAAAAAAQFAQPRPPIGQLPGPLSDTVSAAAAAITADPN 580
Query: 287 FTTALAAAISSIIGPQRSGDGNNNLA 312
FT ALAAAI+SIIG Q + N+ A
Sbjct: 581 FTVALAAAITSIIGGQHAAAAGNSNA 606
>UniRef100_Q6IET0 WRKY transcription factor 1 [Oryza sativa]
Length = 593
Score = 147 bits (371), Expect = 4e-34
Identities = 139/446 (31%), Positives = 190/446 (42%), Gaps = 142/446 (31%)
Query: 4 SAAVSKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQK 63
+A+ E S E++ +++EL R+ EEN +LR ML Q+T SY LQ L +Q++
Sbjct: 124 AASRPDHEEKSRSSNELAAMQAELGRMNEENQRLRGMLTQVTTSYQALQMHLVALMQQRP 183
Query: 64 ------------PNHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDH-------PAE 104
P H E G V + FL+ +S + G+ A + P
Sbjct: 184 QMMQPPTQPEPPPPHQDGKAE--GAVVPRQFLDLGPSSGAGGEAAEEPSNSSTEAGSPRR 241
Query: 105 DSSHSSKLEE---------------PTQDLIP--------------------FKKARVSI 129
SS +K +E P + + P +KARVS+
Sbjct: 242 SSSTGNKDQERGDSPDAPSTAAAWLPGRAMAPQMGAAGAAGKSHDQQAQDANMRKARVSV 301
Query: 130 RARSEAPLPTIVALWL*DVQF-------------------------ANRCAEDKTILITT 164
RARSEAP+ W Q RCAED++ILITT
Sbjct: 302 RARSEAPIIADGCQWRKYGQKMAKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITT 361
Query: 165 YEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATM-----A 219
YEG HNHPLPPAA A+A TTSAAA+MLLS S S + A G +S++F T+ +
Sbjct: 362 YEGTHNHPLPPAAMAMASTTSAAASMLLSGSMPSA---DGAAGLMSSNFLARTVLPCSSS 418
Query: 220 TSTLSASQPFPTITLDFTQ--------------NHNLSMHHNRVPLP---LFFSHKLPPL 262
+T+SAS PFPT+TLD T +VPLP + + +PP
Sbjct: 419 MATISASAPFPTVTLDLTHAPPGAPNAVPLNAARPGAPAPQFQVPLPGGGMAPAFAVPPQ 478
Query: 263 L---------------------------QLGQPPP---------SSMVESVSAAISSDPN 286
+ Q QP P S V + +AAI++DPN
Sbjct: 479 VLYNQSKFSGLQMSSDSAEAAAAAAAAAQFAQPRPPIGQLPGPLSDTVSAAAAAITADPN 538
Query: 287 FTTALAAAISSIIGPQRSGDGNNNLA 312
FT ALAAAI+SIIG Q + N+ A
Sbjct: 539 FTVALAAAITSIIGGQHAAAAGNSNA 564
>UniRef100_Q9C519 WRKY transcription factor 6 [Arabidopsis thaliana]
Length = 553
Score = 145 bits (367), Expect = 1e-33
Identities = 127/405 (31%), Positives = 178/405 (43%), Gaps = 102/405 (25%)
Query: 8 SKEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHG 67
S E + ++ E+ L+ EL+++ +N KLR +L Q++ SY+ LQ L +Q+Q+ +
Sbjct: 145 SSEMEDKRAKNELVKLQDELKKMTMDNQKLRELLTQVSNSYTSLQMHLVSLMQQQQQQNN 204
Query: 68 QNMEENHG-----------------MVSEQIFLNNNNASV-------------SDGKQAC 97
+ +E V E ++N+++ S+GK+
Sbjct: 205 KVIEAAEKPEETIVPRQFIDLGPTRAVGEAEDVSNSSSEDRTRSGGSSAAERRSNGKRLG 264
Query: 98 PHDHPAEDSSHSSKLEEPTQDLIP------FKKARVSIRARSEAPLPTIVALWL*DVQF- 150
+ P +S+ K+ T +KARVS+RARSEAP+ + W Q
Sbjct: 265 REESPETESNKIQKVNSTTPTTFDQTAEATMRKARVSVRARSEAPMISDGCQWRKYGQKM 324
Query: 151 ------------------------ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSA 186
RCAED++ILITTYEGNHNHPLPPAA A+A TT+A
Sbjct: 325 AKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITTYEGNHNHPLPPAAVAMASTTTA 384
Query: 187 AAAMLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFT--------- 237
AA MLLS S SS + T L+ + + + +T+SAS PFPT+TLD T
Sbjct: 385 AANMLLSGSMSSHDGMMNPTNLLARAVLPCSTSMATISASAPFPTVTLDLTHSPPPPNGS 444
Query: 238 ---------QNHNLSMHH---------NRVP--LP------LFFSHKLPPLLQLGQPP-- 269
NHN M N P LP L+ K L G P
Sbjct: 445 NPSSSAATNNNHNSLMQRPQQQQQQMTNLPPGMLPHVIGQALYNQSKFSGLQFSGGSPST 504
Query: 270 ----PSSMVESVSAAISSDPNFTTALAAAISSIIGPQRSGDGNNN 310
S V A+++DPNFT ALAA ISS+I DG N
Sbjct: 505 AAFSQSHAVADTITALTADPNFTAALAAVISSMINGTNHHDGEGN 549
>UniRef100_Q9XHY3 Putative WRKY DNA binding protein [Oryza sativa]
Length = 470
Score = 143 bits (360), Expect = 7e-33
Identities = 135/431 (31%), Positives = 185/431 (42%), Gaps = 142/431 (32%)
Query: 19 EMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQK------------PNH 66
+++ +++EL R+ EEN +LR ML Q+T SY LQ L +Q++ P H
Sbjct: 16 QLAAMQAELGRMNEENQRLRGMLTQVTTSYQALQMHLVALMQQRPQMMQPPTQPEPPPPH 75
Query: 67 GQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDH-------PAEDSSHSSKLEE----- 114
E G V + FL+ +S + G+ A + P SS +K +E
Sbjct: 76 QDGKAE--GAVVPRQFLDLGPSSGAGGEAAEEPSNSSTEAGSPRRSSSTGNKDQERGDSP 133
Query: 115 ----------PTQDLIP--------------------FKKARVSIRARSEAPLPTIVALW 144
P + + P +KARVS+RARSEAP+ W
Sbjct: 134 DAPSTAAAWLPGRAMAPQMGAAGAAGKSHDQQAQDANMRKARVSVRARSEAPIIADGCQW 193
Query: 145 L*DVQF-------------------------ANRCAEDKTILITTYEGNHNHPLPPAATA 179
Q RCAED++ILITTYEG HNHPLPPAA A
Sbjct: 194 RKYGQKMAKGNPCPRAYYRCTMATGCPVRKQVQRCAEDRSILITTYEGTHNHPLPPAAMA 253
Query: 180 IAHTTSAAAAMLLSSSTSSTLRKESATGYLSNSFPYATM-----ATSTLSASQPFPTITL 234
+A TTSAAA+MLLS S S + A G +S++F T+ + +T+SAS PFPT+TL
Sbjct: 254 MASTTSAAASMLLSGSMPSA---DGAAGLMSSNFLARTVLPCSSSMATISASAPFPTVTL 310
Query: 235 DFTQ--------------NHNLSMHHNRVPLP---LFFSHKLPPLL-------------- 263
D T +VPLP + + +PP +
Sbjct: 311 DLTHAPPGAPNAVPLNAARPGAPAPQFQVPLPGGGMAPAFAVPPQVLYNQSKFSGLQMSS 370
Query: 264 -------------QLGQPPP---------SSMVESVSAAISSDPNFTTALAAAISSIIGP 301
Q QP P S V + +AAI++DPNFT ALAAAI+SIIG
Sbjct: 371 DSAEAAAAAAAAAQFAQPRPPIGQLPGPLSDTVSAAAAAITADPNFTVALAAAITSIIGG 430
Query: 302 QRSGDGNNNLA 312
Q + N+ A
Sbjct: 431 QHAAAAGNSNA 441
>UniRef100_Q93WT0 Probable WRKY transcription factor 31 [Arabidopsis thaliana]
Length = 538
Score = 133 bits (335), Expect = 6e-30
Identities = 109/279 (39%), Positives = 144/279 (51%), Gaps = 65/279 (23%)
Query: 98 PHDHPAEDSSHSSK---LEEPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQF---- 150
P +P+ +S+ ++ + + + +KARVS+RARSEA + + W Q
Sbjct: 253 PKHNPSSSNSNGNRNGNVIDQSAAEATMRKARVSVRARSEAAMISDGCQWRKYGQKMAKG 312
Query: 151 ---------------------ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAA 189
RCAED++ILITTYEGNHNHPLPPAATA+A TT+AAA+
Sbjct: 313 NPCPRAYYRCTMAGGCPVRKQVQRCAEDRSILITTYEGNHNHPLPPAATAMASTTTAAAS 372
Query: 190 MLLSSSTSSTLRKESATGYLSNSFPYATMATSTLSASQPFPTITLDFTQN---HNLSMHH 246
MLLS S SS + T L+ + + + +T+SAS PFPTITLD T + +N +M
Sbjct: 373 MLLSGSMSSQDGLMNPTNLLARAILPCSSSMATISASAPFPTITLDLTNSPNGNNPNMTT 432
Query: 247 NRVPL------PLFFSHKLPPL----------------LQLGQPP-----PSSMVESVSA 279
N PL P F LP + LQL P SS+ ESVSA
Sbjct: 433 NN-PLMQFAQRPGFNPAVLPQVVGQAMYNNQQQSKFSGLQLPAQPLQIAATSSVAESVSA 491
Query: 280 A---ISSDPNFTTALAAAISSII---GPQRSGDGNNNLA 312
A I+SDPNF ALAAAI+SI+ Q + NNN+A
Sbjct: 492 ASAAIASDPNFAAALAAAITSIMNGSSHQNNNTNNNNVA 530
Score = 40.0 bits (92), Expect = 0.087
Identities = 22/71 (30%), Positives = 42/71 (58%), Gaps = 8/71 (11%)
Query: 9 KEENLENSETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQ 68
K +EN++ L+ EL++++ EN +LR ML Q T +++ LQ QL +++Q+ +
Sbjct: 105 KRAKIENAQ-----LQEELKKMKIENQRLRDMLSQATTNFNALQMQLVAVMRQQEQ---R 156
Query: 69 NMEENHGMVSE 79
N ++H + E
Sbjct: 157 NSSQDHLLAQE 167
>UniRef100_Q688W5 Putative WRKY transcription factor [Oryza sativa]
Length = 625
Score = 129 bits (324), Expect = 1e-28
Identities = 96/261 (36%), Positives = 123/261 (46%), Gaps = 62/261 (23%)
Query: 114 EPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQF----------------------- 150
EP + +KARVS+RARS+AP+ + W Q
Sbjct: 339 EPVPEAATMRKARVSVRARSDAPMISDGCQWRKYGQKMAKGNPCPRAYYRCTMAAGCPVR 398
Query: 151 --ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESA-TG 207
RCAED+T+LITTYEGNHNHPLPPAA A+A TT+AAA+MLLS S S A +
Sbjct: 399 KQVQRCAEDRTVLITTYEGNHNHPLPPAAMAMASTTAAAASMLLSGSMPSADGSLMAGSN 458
Query: 208 YLSNSFPYATMATSTLSASQPFPTITLDFTQNHN-----------------------LSM 244
+L+ + + +T+SAS PFPT+TLD TQ L+
Sbjct: 459 FLARAVLPCSSTVATISASAPFPTVTLDLTQTAPPPPPASSTQPQPPRPEPAQLQAALAE 518
Query: 245 HHNRVPLPLFFSHKLPPLLQLGQPPP------------SSMVESVSAAISSDPNFTTALA 292
V LP F KL +L + V + +AAI+SDPNFT LA
Sbjct: 519 AARPVALPQLFGQKLYDQSKLSAVQAVAGTKGSDGGALADTVNAATAAIASDPNFTAVLA 578
Query: 293 AAISSIIGPQRSGDGNNNLAG 313
AA++S IG RSG G G
Sbjct: 579 AALTSYIG-SRSGSGGAGAGG 598
>UniRef100_Q9LGZ7 Similar to Arabidopsis thaliana chromosome 4 BAC T15B16 [Oryza
sativa]
Length = 594
Score = 128 bits (322), Expect = 2e-28
Identities = 94/215 (43%), Positives = 117/215 (53%), Gaps = 62/215 (28%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSN- 211
RCAEDKT+LITTYEGNHNH LPPAAT +A+TTSAAAAMLLS +S R +A L +
Sbjct: 346 RCAEDKTVLITTYEGNHNHQLPPAATTMANTTSAAAAMLLSGPAAS--RDGAAAALLGHH 403
Query: 212 -----------SFPYATMATSTLSASQPFPTITLDFTQN-----------HNLS----MH 245
SFPYA+ +TLSAS PFPTITLD TQ H L +H
Sbjct: 404 HHHHPAAMFHQSFPYAS-TMATLSASAPFPTITLDLTQTPAGGAGAASLLHALHRPPVIH 462
Query: 246 HNRVPLPLFFSHKLPPLLQL---------------GQPPPSSMVESVSAAISSDPNFTTA 290
+ F+ +PP L + G S++E+V+AA+++DPNFTTA
Sbjct: 463 PGAAAQAMPFA--VPPQLAMYLPQQRAAAAGLGGAGAARQPSVMETVTAALAADPNFTTA 520
Query: 291 LAAAISSIIG-----------PQRS----GDGNNN 310
LAAAISS++ P+ S GDGN N
Sbjct: 521 LAAAISSVVAGGAHHQALSTTPRGSAAGAGDGNGN 555
Score = 47.0 bits (110), Expect = 7e-04
Identities = 29/87 (33%), Positives = 45/87 (51%), Gaps = 12/87 (13%)
Query: 3 SSAAVSKEENLEN--SETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQ 60
++A +E+ +N E + ++ ELRRV EEN +LR ML+++ +SYS L Q Q
Sbjct: 119 AAAGAGEEDTGKNHRKEATTAAVDVELRRVVEENRRLRGMLDELNRSYSALYHQYLQVTQ 178
Query: 61 KQKPNHGQNMEENHGMVSEQIFLNNNN 87
+Q NH + +NNNN
Sbjct: 179 QQ----------NHRHPDHHLIMNNNN 195
>UniRef100_Q6IEN8 WRKY transcription factor 43 [Oryza sativa]
Length = 618
Score = 126 bits (316), Expect = 9e-28
Identities = 95/261 (36%), Positives = 122/261 (46%), Gaps = 62/261 (23%)
Query: 114 EPTQDLIPFKKARVSIRARSEAPLPTIVALWL*DVQF----------------------- 150
EP + +KARVS+RARS+AP+ + W Q
Sbjct: 332 EPIPEAATMRKARVSVRARSDAPMISDGCQWRKYGQKMAKGNPCPRAYYRCTMAAGCPVR 391
Query: 151 --ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESA-TG 207
RCAED+T+LITTYEGNHNHPLPPAA A+A TT+AAA+MLLS S S A +
Sbjct: 392 KQVQRCAEDRTVLITTYEGNHNHPLPPAAMAMASTTAAAASMLLSGSMPSADGSLMAGSN 451
Query: 208 YLSNSFPYATMATSTLSASQPFPTITLDFTQNHN-----------------------LSM 244
+L+ + + +T+SAS PFPT+TLD TQ L+
Sbjct: 452 FLARAVLPCSSTVATISASAPFPTVTLDLTQTAPPPPPASSTQPQPPRPEPAQLQAALAE 511
Query: 245 HHNRVPLPLFFSHKLPPLLQLGQPPP------------SSMVESVSAAISSDPNFTTALA 292
V LP F KL +L + V + +AAI+SDPNFT LA
Sbjct: 512 AARPVALPQLFGQKLYDQSKLSAVQAVAGTKGSDGGALADTVNAATAAIASDPNFTAVLA 571
Query: 293 AAISSIIGPQRSGDGNNNLAG 313
AA++S IG SG G G
Sbjct: 572 AALTSYIG-SSSGSGGGGGGG 591
>UniRef100_Q5VRX5 WRKY transcription factor 6-like [Oryza sativa]
Length = 507
Score = 91.7 bits (226), Expect = 3e-17
Identities = 49/100 (49%), Positives = 66/100 (66%), Gaps = 13/100 (13%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS 212
RCA+D +ILITTYEG H+HPLPPAA A+A TTSAAAAML S ST+ST+ +G + +
Sbjct: 277 RCADDMSILITTYEGTHSHPLPPAAAAMASTTSAAAAMLTSGSTNSTMH---GSGGVHHH 333
Query: 213 FPYAT----------MATSTLSASQPFPTITLDFTQNHNL 242
P+A+ + +T+S + PT+TLD T H+L
Sbjct: 334 LPFASAVGGGGGVGLLGPTTISTATSCPTVTLDLTAPHSL 373
>UniRef100_Q5VP49 Putative WRKY transcription factor [Oryza sativa]
Length = 624
Score = 90.1 bits (222), Expect = 7e-17
Identities = 86/266 (32%), Positives = 119/266 (44%), Gaps = 58/266 (21%)
Query: 114 EPTQDLIP---FKKARVSIRARSEAPLPTIVALWL*DVQF-------------------- 150
E D+ P KKARVS+RAR +AP W Q
Sbjct: 248 EAEDDIAPQPQVKKARVSVRARCDAPTMNDGCQWRKYGQKIAKGNPCPRAYYRCTVAAGC 307
Query: 151 -----ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTL----- 200
RCA+D +ILITTYEG HNHPL +ATA+A TTSAAA+ML+S S+S++L
Sbjct: 308 PVRKQVQRCADDMSILITTYEGTHNHPLSVSATAMASTTSAAASMLISGSSSTSLAAYPA 367
Query: 201 -RKESATGYLSNSFP---------YATMATSTLSASQPFPTITLDFTQ-NHNLSMHHNRV 249
A + ++S P T A + ++++ +PTITLD T + H
Sbjct: 368 AAASPALAFDASSKPPLIGGRPFFLPTAAAAAITSTPSYPTITLDLTSPAAAATSSHAAF 427
Query: 250 PLPLFFSHKLPPLLQL---GQPPPSSM---VESVSAAISSDP------NFTTALAAAISS 297
L FSH P G P S+ S A++S+ P +++ AA+SS
Sbjct: 428 SLSNRFSHTRYPSTGFTFSGSGPSSAPWPGYLSYGASLSAHPYNAGGGKSSSSFEAALSS 487
Query: 298 IIGPQRSGDGNNNLAGVVPGSPQLPQ 323
I G ++ G G G P Q+ Q
Sbjct: 488 INGSRQQGGGGG--GGSAPPLYQMQQ 511
Score = 47.4 bits (111), Expect = 5e-04
Identities = 24/52 (46%), Positives = 36/52 (69%), Gaps = 7/52 (13%)
Query: 260 PPLLQLGQ-------PPPSSMVESVSAAISSDPNFTTALAAAISSIIGPQRS 304
PPL Q+ Q PPPS + ++++ AI++DP+F TALAAAI+S +G + S
Sbjct: 504 PPLYQMQQKAAAAAPPPPSVITDTIAKAITADPSFHTALAAAITSYVGKKGS 555
>UniRef100_Q6IEK8 WRKY transcription factor 73 [Oryza sativa]
Length = 527
Score = 90.1 bits (222), Expect = 7e-17
Identities = 86/266 (32%), Positives = 119/266 (44%), Gaps = 58/266 (21%)
Query: 114 EPTQDLIP---FKKARVSIRARSEAPLPTIVALWL*DVQF-------------------- 150
E D+ P KKARVS+RAR +AP W Q
Sbjct: 151 EAEDDIAPQPQVKKARVSVRARCDAPTMNDGCQWRKYGQKIAKGNPCPRAYYRCTVAAGC 210
Query: 151 -----ANRCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTL----- 200
RCA+D +ILITTYEG HNHPL +ATA+A TTSAAA+ML+S S+S++L
Sbjct: 211 PVRKQVQRCADDMSILITTYEGTHNHPLSVSATAMASTTSAAASMLISGSSSTSLAAYPA 270
Query: 201 -RKESATGYLSNSFP---------YATMATSTLSASQPFPTITLDFTQ-NHNLSMHHNRV 249
A + ++S P T A + ++++ +PTITLD T + H
Sbjct: 271 AAASPALAFDASSKPPLIGGRPFFLPTAAAAAITSTPSYPTITLDLTSPAAAATSSHAAF 330
Query: 250 PLPLFFSHKLPPLLQL---GQPPPSSM---VESVSAAISSDP------NFTTALAAAISS 297
L FSH P G P S+ S A++S+ P +++ AA+SS
Sbjct: 331 SLSNRFSHTRYPSTGFTFSGSGPSSAPWPGYLSYGASLSAHPYNAGGGKSSSSFEAALSS 390
Query: 298 IIGPQRSGDGNNNLAGVVPGSPQLPQ 323
I G ++ G G G P Q+ Q
Sbjct: 391 INGSRQQGGGGG--GGSAPPLYQMQQ 414
Score = 47.4 bits (111), Expect = 5e-04
Identities = 24/52 (46%), Positives = 36/52 (69%), Gaps = 7/52 (13%)
Query: 260 PPLLQLGQ-------PPPSSMVESVSAAISSDPNFTTALAAAISSIIGPQRS 304
PPL Q+ Q PPPS + ++++ AI++DP+F TALAAAI+S +G + S
Sbjct: 407 PPLYQMQQKAAAAAPPPPSVITDTIAKAITADPSFHTALAAAITSYVGKKGS 458
>UniRef100_Q84QE3 Avr9/Cf-9 rapidly elicited protein 126 [Nicotiana tabacum]
Length = 303
Score = 89.0 bits (219), Expect = 2e-16
Identities = 59/148 (39%), Positives = 77/148 (51%), Gaps = 20/148 (13%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS 212
RC ED ++LITTYEG HNH LP ATA+A TTSAAA+MLLS S+SS + +NS
Sbjct: 84 RCVEDMSVLITTYEGTHNHSLPIEATAMASTTSAAASMLLSGSSSSQSANKDLRNLPNNS 143
Query: 213 FPYATMATSTLSASQPFPTITLDFTQNHNLSMHHNRVPLPLFFSHKLPPLLQLGQPPPSS 272
++ S S PFPTITLDFT S F S P S
Sbjct: 144 KTTPLYLSNPPSNSNPFPTITLDFTTFPTTSS---------FTSFNFP-----------S 183
Query: 273 MVESVSAAISSDPNFTTALAAAISSIIG 300
+S + +S+ NF++ + +S I+G
Sbjct: 184 NFQSNTGFLSNSLNFSSPESDTLSKILG 211
>UniRef100_Q6ZH52 Putative WRKY transcription factor [Oryza sativa]
Length = 637
Score = 85.5 bits (210), Expect = 2e-15
Identities = 66/193 (34%), Positives = 95/193 (49%), Gaps = 26/193 (13%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS 212
RC ED +IL+TTYEG HNHPLP ATA+A TTSAAA +L SST+S+ A+ S+S
Sbjct: 442 RCLEDMSILVTTYEGTHNHPLPVGATAMASTTSAAATFMLLSSTTSSSSVSDASAAPSSS 501
Query: 213 FPYATMATSTLSASQPFPTITLDFTQNHNL---SMHHNRVPLPLF-------FSHKLPPL 262
+ + S P T Q+ NL S + + P +S PPL
Sbjct: 502 YLSPYLLNSASPLLMPGATGGGGGMQHLNLFGNSPSSSSLLAPQAPGSSKYPWSPNHPPL 561
Query: 263 LQLG--------------QPPPSSMVESVSAAISSDPNFTTALAAAISSIIGP--QRSGD 306
G +P P+++ E+V A +S F+ A+AAAI++ +G + S
Sbjct: 562 AGAGGNKRPFWSAGGDGDKPAPAALAENVGAVMSDPNKFSAAIAAAINNFMGKDGESSSG 621
Query: 307 GNNNLAGVVPGSP 319
+++ GVV P
Sbjct: 622 KSSSKWGVVESLP 634
Score = 33.9 bits (76), Expect = 6.3
Identities = 17/63 (26%), Positives = 36/63 (56%), Gaps = 2/63 (3%)
Query: 16 SETEMSILESELRRVQEENHKLRIMLEQITKSYSQLQAQLFITLQKQKPNHGQNMEENHG 75
++ E+S ++ E+ R++EEN LR ++++ + Y +LQ +L +Q+P +E
Sbjct: 202 AQDELSEMQEEMERMKEENRMLRRVVDKTVRDYYELQMKL--AAYQQQPAAADEPKETEV 259
Query: 76 MVS 78
+S
Sbjct: 260 FLS 262
>UniRef100_Q8VWV6 Probable WRKY transcription factor 61 [Arabidopsis thaliana]
Length = 480
Score = 84.7 bits (208), Expect = 3e-15
Identities = 94/359 (26%), Positives = 139/359 (38%), Gaps = 118/359 (32%)
Query: 61 KQKPNHGQNMEENHGMVSEQIFLNNNNASVSDGKQACPHDHPAEDSSHSSKLEEPTQDLI 120
K N + +E +H + + ++NNN S +D H + E Q+L+
Sbjct: 119 KALSNPNEKLEIDHNQETMSLEISNNNKIRSQNSFGFKND----GDDHEDEDEILPQNLV 174
Query: 121 PFKKARVSIRARSEAPLPTIVALWL*DVQF-------------------------ANRCA 155
KK RVS+R+R E P W Q RC+
Sbjct: 175 --KKTRVSVRSRCETPTMNDGCQWRKYGQKIAKGNPCPRAYYRCTIAASCPVRKQVQRCS 232
Query: 156 EDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSST---------------- 199
ED +ILI+TYEG HNHPLP +ATA+A TSAAA+MLLS ++SS+
Sbjct: 233 EDMSILISTYEGTHNHPLPMSATAMASATSAAASMLLSGASSSSSAAADLHGLNFSLSGN 292
Query: 200 ----------LRKESATGY----------LSNSFPYATMAT------STLSASQPFPTIT 233
L+ S++G+ S+ P+ +M S +S S +P+
Sbjct: 293 NITPKPKTHFLQSPSSSGHPTVTLDLTTSSSSQQPFLSMLNRFSSPPSNVSRSNSYPSTN 352
Query: 234 LDFTQNHNLSMH--------------------HNRVPLPLFF---------------SHK 258
L+F+ N N M+ H + P S
Sbjct: 353 LNFSNNTNTLMNWGGGGNPSDQYRAAYGNINTHQQSPYHKIIQTRTAGSSFDPFGRSSSS 412
Query: 259 LPPLLQL---------GQPPPSSMVESVSAAISSDPNFTTALAAAISSIIGPQRSGDGN 308
P + L PS E++ A I++DP+F +ALA A+SSI+G D N
Sbjct: 413 HSPQINLDHIGIKNIISHQVPSLPAETIKA-ITTDPSFQSALATALSSIMGGDLKIDHN 470
>UniRef100_Q68HK5 WRKY6-1 [Pelargonium hortorum]
Length = 113
Score = 84.3 bits (207), Expect = 4e-15
Identities = 44/80 (55%), Positives = 57/80 (71%), Gaps = 1/80 (1%)
Query: 153 RCAEDKTILITTYEGNHNHPLPPAATAIAHTTSAAAAMLLSSSTSSTLRKESATGYLSNS 212
RCA+D++ILITTYEGNHNHPLPPAA A+A TT+AAA+MLLS S S L+ +
Sbjct: 35 RCADDRSILITTYEGNHNHPLPPAAMAMASTTTAAASMLLSGSMPSA-DGIMNPNLLARA 93
Query: 213 FPYATMATSTLSASQPFPTI 232
+ + +T+SAS PFPT+
Sbjct: 94 MLPCSSSMATISASAPFPTV 113
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.313 0.126 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 510,548,476
Number of Sequences: 2790947
Number of extensions: 19592408
Number of successful extensions: 81611
Number of sequences better than 10.0: 452
Number of HSP's better than 10.0 without gapping: 153
Number of HSP's successfully gapped in prelim test: 307
Number of HSP's that attempted gapping in prelim test: 80623
Number of HSP's gapped (non-prelim): 1185
length of query: 330
length of database: 848,049,833
effective HSP length: 128
effective length of query: 202
effective length of database: 490,808,617
effective search space: 99143340634
effective search space used: 99143340634
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 75 (33.5 bits)
Medicago: description of AC139745.3


