

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139744.1 + phase: 0
(48 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
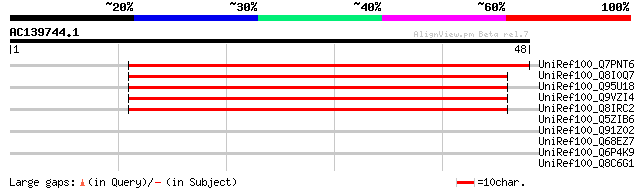
UniRef100_Q7PNT6 ENSANGP00000004511 [Anopheles gambiae str. PEST] 62 5e-09
UniRef100_Q8I0Q7 CG14995-PC, isoform C [Drosophila melanogaster] 56 2e-07
UniRef100_Q95U18 GH13848p [Drosophila melanogaster] 56 2e-07
UniRef100_Q9VZI4 CG14995-PA, isoform A [Drosophila melanogaster] 56 2e-07
UniRef100_Q8IRC2 CG14995-PD, isoform D [Drosophila melanogaster] 56 2e-07
UniRef100_Q5ZIB6 Hypothetical protein [Gallus gallus] 33 2.6
UniRef100_Q91Z02 1810043G02Rik protein [Mus musculus] 32 3.4
UniRef100_Q68EZ7 MGC83386 protein [Xenopus laevis] 32 5.9
UniRef100_Q6P4K9 Hypothetical protein MGC75902 [Xenopus tropicalis] 31 7.7
UniRef100_Q8C6G1 Mus musculus 10 day old male pancreas cDNA, RIK... 31 7.7
>UniRef100_Q7PNT6 ENSANGP00000004511 [Anopheles gambiae str. PEST]
Length = 438
Score = 61.6 bits (148), Expect = 5e-09
Identities = 29/37 (78%), Positives = 34/37 (91%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDELED 48
++SNLLSA LCLVKELDYPSLEVVE AV+CR+DEL +
Sbjct: 402 RNSNLLSATLCLVKELDYPSLEVVEHAVRCRIDELAE 438
>UniRef100_Q8I0Q7 CG14995-PC, isoform C [Drosophila melanogaster]
Length = 411
Score = 56.2 bits (134), Expect = 2e-07
Identities = 26/35 (74%), Positives = 32/35 (91%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
++SN+LSA LCLVKELDY SLEV+E AV+CR+DEL
Sbjct: 374 RNSNILSAALCLVKELDYASLEVLEHAVRCRIDEL 408
>UniRef100_Q95U18 GH13848p [Drosophila melanogaster]
Length = 389
Score = 56.2 bits (134), Expect = 2e-07
Identities = 26/35 (74%), Positives = 32/35 (91%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
++SN+LSA LCLVKELDY SLEV+E AV+CR+DEL
Sbjct: 352 RNSNILSAALCLVKELDYASLEVLEHAVRCRIDEL 386
>UniRef100_Q9VZI4 CG14995-PA, isoform A [Drosophila melanogaster]
Length = 454
Score = 56.2 bits (134), Expect = 2e-07
Identities = 26/35 (74%), Positives = 32/35 (91%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
++SN+LSA LCLVKELDY SLEV+E AV+CR+DEL
Sbjct: 417 RNSNILSAALCLVKELDYASLEVLEHAVRCRIDEL 451
>UniRef100_Q8IRC2 CG14995-PD, isoform D [Drosophila melanogaster]
Length = 354
Score = 56.2 bits (134), Expect = 2e-07
Identities = 26/35 (74%), Positives = 32/35 (91%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
++SN+LSA LCLVKELDY SLEV+E AV+CR+DEL
Sbjct: 317 RNSNILSAALCLVKELDYASLEVLEHAVRCRIDEL 351
>UniRef100_Q5ZIB6 Hypothetical protein [Gallus gallus]
Length = 254
Score = 32.7 bits (73), Expect = 2.6
Identities = 17/40 (42%), Positives = 25/40 (62%), Gaps = 5/40 (12%)
Query: 14 SNLLSAVLCLVKELDYPSLEVVEMAV-----KCRMDELED 48
+N+L+A+L L KELD LE+V+ V CR EL++
Sbjct: 214 NNVLNAILLLTKELDAEGLEIVQQTVGRRLQACRKRELQE 253
>UniRef100_Q91Z02 1810043G02Rik protein [Mus musculus]
Length = 249
Score = 32.3 bits (72), Expect = 3.4
Identities = 15/35 (42%), Positives = 23/35 (64%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
+S N+L+A+L L++ELD LE V+ V R+ L
Sbjct: 205 KSRNILTAILLLLRELDTEGLETVQQTVGSRLQAL 239
>UniRef100_Q68EZ7 MGC83386 protein [Xenopus laevis]
Length = 256
Score = 31.6 bits (70), Expect = 5.9
Identities = 14/34 (41%), Positives = 24/34 (70%)
Query: 14 SNLLSAVLCLVKELDYPSLEVVEMAVKCRMDELE 47
+N+L+AVL L+K+LD LE+V+ R+ +L+
Sbjct: 215 NNILNAVLLLLKDLDTEGLEIVQRTAGQRLHQLQ 248
>UniRef100_Q6P4K9 Hypothetical protein MGC75902 [Xenopus tropicalis]
Length = 212
Score = 31.2 bits (69), Expect = 7.7
Identities = 14/34 (41%), Positives = 24/34 (70%)
Query: 14 SNLLSAVLCLVKELDYPSLEVVEMAVKCRMDELE 47
+N+L+AVL L+K+LD LE+V+ R+ +L+
Sbjct: 171 NNILNAVLLLLKDLDTEGLEIVQRTAGQRLHQLK 204
>UniRef100_Q8C6G1 Mus musculus 10 day old male pancreas cDNA, RIKEN full-length
enriched library, clone:1810043G02 product:transient
receptor protein 7, full insert sequence [Mus musculus]
Length = 249
Score = 31.2 bits (69), Expect = 7.7
Identities = 14/35 (40%), Positives = 23/35 (65%)
Query: 12 QSSNLLSAVLCLVKELDYPSLEVVEMAVKCRMDEL 46
++ N+L+A+L L++ELD LE V+ V R+ L
Sbjct: 205 KNRNILTAILLLLRELDTEGLETVQQTVGSRLQAL 239
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.137 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,144,891
Number of Sequences: 2790947
Number of extensions: 1257927
Number of successful extensions: 4639
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 4630
Number of HSP's gapped (non-prelim): 10
length of query: 48
length of database: 848,049,833
effective HSP length: 24
effective length of query: 24
effective length of database: 781,067,105
effective search space: 18745610520
effective search space used: 18745610520
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC139744.1


