

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.4 - phase: 0 /pseudo
(105 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
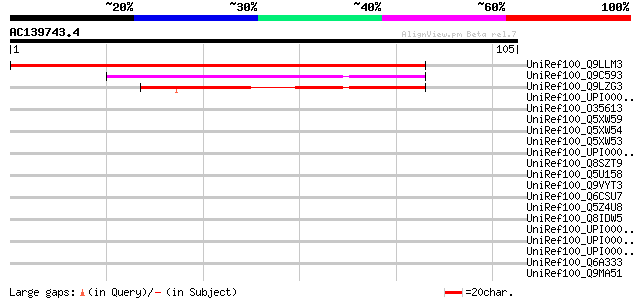
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LLM3 MTD1 [Medicago truncatula] 174 6e-43
UniRef100_Q9C593 Hypothetical protein At5g21940 [Arabidopsis tha... 54 1e-06
UniRef100_Q9LZG3 Hypothetical protein T28A8_140 [Arabidopsis tha... 52 3e-06
UniRef100_UPI00002DA9DC UPI00002DA9DC UniRef100 entry 40 0.013
UniRef100_O35613 Death domain-associated protein 6 [Mus musculus] 40 0.017
UniRef100_Q5XW59 Death-domain associated protein [Mus musculus] 39 0.038
UniRef100_Q5XW54 Death domain-associated protein [Mus musculus] 39 0.038
UniRef100_Q5XW53 Death domain-associated protein [Mus musculus] 39 0.038
UniRef100_UPI00002F1525 UPI00002F1525 UniRef100 entry 37 0.085
UniRef100_Q8SZT9 GH13968p [Drosophila melanogaster] 37 0.11
UniRef100_Q5U158 RE02581p [Drosophila melanogaster] 37 0.11
UniRef100_Q9VYT3 CG2025-PA [Drosophila melanogaster] 37 0.11
UniRef100_Q6CSU7 Kluyveromyces lactis strain NRRL Y-1140 chromos... 37 0.11
UniRef100_Q5Z4U8 Hypothetical protein P0545E05.31-1 [Oryza sativa] 37 0.15
UniRef100_Q8IDW5 Hypothetical protein PF13_0203 [Plasmodium falc... 37 0.15
UniRef100_UPI00003655FB UPI00003655FB UniRef100 entry 36 0.19
UniRef100_UPI00003655FA UPI00003655FA UniRef100 entry 36 0.19
UniRef100_UPI00003655F9 UPI00003655F9 UniRef100 entry 36 0.19
UniRef100_Q6A333 Always early 2 protein [Arabidopsis thaliana] 36 0.19
UniRef100_Q9MA51 F22F7.18 protein [Arabidopsis thaliana] 36 0.19
>UniRef100_Q9LLM3 MTD1 [Medicago truncatula]
Length = 231
Score = 174 bits (440), Expect = 6e-43
Identities = 86/86 (100%), Positives = 86/86 (100%)
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD
Sbjct: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
Query: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
ENEAESKYNGGALDCMEALEEVLPIR
Sbjct: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
>UniRef100_Q9C593 Hypothetical protein At5g21940 [Arabidopsis thaliana]
Length = 264
Score = 53.5 bits (127), Expect = 1e-06
Identities = 29/66 (43%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Query: 21 APRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALE 80
+P D S+++SSSIGRNSDD + ENE ES Y G L+ ME+LE
Sbjct: 24 SPSPSPSDSSSSPSSSASSSIGRNSDDGEKSSEDGGDDAGENEVESPYK-GPLEMMESLE 82
Query: 81 EVLPIR 86
+VLP+R
Sbjct: 83 QVLPVR 88
>UniRef100_Q9LZG3 Hypothetical protein T28A8_140 [Arabidopsis thaliana]
Length = 270
Score = 52.0 bits (123), Expect = 3e-06
Identities = 33/61 (54%), Positives = 37/61 (60%), Gaps = 12/61 (19%)
Query: 28 DQGKVC--STTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPI 85
DQ C S+TSS SIG NSDDD+ ENE ES YN G LD ME+LEE LPI
Sbjct: 15 DQEFACLSSSTSSDSIGENSDDDEG---------GENEIESSYN-GPLDMMESLEEALPI 64
Query: 86 R 86
+
Sbjct: 65 K 65
>UniRef100_UPI00002DA9DC UPI00002DA9DC UniRef100 entry
Length = 1679
Score = 40.0 bits (92), Expect = 0.013
Identities = 32/107 (29%), Positives = 52/107 (47%), Gaps = 16/107 (14%)
Query: 2 ELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSS----IGRNSDDDDDDEVSSER 57
+L T+ KR G R +A +++Q VC+ +S++ +G +SDDDDDD+ S+
Sbjct: 692 KLRTIYAKRTGKR-----EANMGVEENQS-VCTPSSAAKRKRKMGGDSDDDDDDDDDSDD 745
Query: 58 SMDENEAESKYNGGALDCMEALEEVLPIRLFQSFMVMQEKYLKFLQW 104
D+ E E + + +E +EVL + F YLK W
Sbjct: 746 QPDDEEEEEEEETKKVKKVETNDEVLVAPSPRIF------YLKLCSW 786
>UniRef100_O35613 Death domain-associated protein 6 [Mus musculus]
Length = 739
Score = 39.7 bits (91), Expect = 0.017
Identities = 28/95 (29%), Positives = 42/95 (43%), Gaps = 13/95 (13%)
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQG-------------KVCSTTSSSSIGRNSDD 47
M+ +T +R R +L AP+ D Q + C TTS + + DD
Sbjct: 388 MQDKTEEGERQKRRARLLGTAPQPSDPPQASSESGEGPSGMASQECPTTSKAETDDDDDD 447
Query: 48 DDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
DDDD+ +E S +E E E + D E LE++
Sbjct: 448 DDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 482
>UniRef100_Q5XW59 Death-domain associated protein [Mus musculus]
Length = 651
Score = 38.5 bits (88), Expect = 0.038
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 345 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 394
>UniRef100_Q5XW54 Death domain-associated protein [Mus musculus]
Length = 621
Score = 38.5 bits (88), Expect = 0.038
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 315 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 364
>UniRef100_Q5XW53 Death domain-associated protein [Mus musculus]
Length = 715
Score = 38.5 bits (88), Expect = 0.038
Identities = 19/50 (38%), Positives = 27/50 (54%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEV 82
C TTS + + DDDDDD+ +E S +E E E + D E LE++
Sbjct: 409 CPTTSKAETDDDDDDDDDDDEDNEESEEEEEEEEEEKEATEDEDEDLEQL 458
>UniRef100_UPI00002F1525 UPI00002F1525 UniRef100 entry
Length = 444
Score = 37.4 bits (85), Expect = 0.085
Identities = 24/84 (28%), Positives = 42/84 (49%), Gaps = 2/84 (2%)
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEAL 79
+ P + DD+ ST + + D+D+DD+ SE +++E ++K + E L
Sbjct: 233 EVPAEDGDDETADASTDDDDNEDSDEDEDEDDD--SEDDSEDDEPKTKTITKTVWDWERL 290
Query: 80 EEVLPIRLFQSFMVMQEKYLKFLQ 103
+V I L + V +E+Y KF Q
Sbjct: 291 NDVKAIWLRSASEVEEEEYTKFYQ 314
>UniRef100_Q8SZT9 GH13968p [Drosophila melanogaster]
Length = 343
Score = 37.0 bits (84), Expect = 0.11
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 230 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 273
Score = 32.7 bits (73), Expect = 2.1
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 235 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 273
>UniRef100_Q5U158 RE02581p [Drosophila melanogaster]
Length = 1147
Score = 37.0 bits (84), Expect = 0.11
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 1034 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
Score = 32.7 bits (73), Expect = 2.1
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 1039 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
>UniRef100_Q9VYT3 CG2025-PA [Drosophila melanogaster]
Length = 1147
Score = 37.0 bits (84), Expect = 0.11
Identities = 17/44 (38%), Positives = 22/44 (49%)
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNG 70
+D K S + S N DDDD+DE E S +E E E + G
Sbjct: 1034 EDTAKCASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
Score = 32.7 bits (73), Expect = 2.1
Identities = 14/39 (35%), Positives = 20/39 (50%)
Query: 33 CSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
C++ S +DDDDD++ E S +E E E K G
Sbjct: 1039 CASKSDDESESENDDDDDEDEEEEESSEEEEEEEKEKEG 1077
>UniRef100_Q6CSU7 Kluyveromyces lactis strain NRRL Y-1140 chromosome C of strain NRRL
Y- 1140 of Kluyveromyces lactis [Kluyveromyces lactis]
Length = 1194
Score = 37.0 bits (84), Expect = 0.11
Identities = 21/65 (32%), Positives = 34/65 (52%), Gaps = 1/65 (1%)
Query: 3 LETVAFKRVGSRTSV-LFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDE 61
+ETV R R +V L ++ D+DDD G + + S + + DDD +DE + DE
Sbjct: 97 VETVTSLRGRKRKNVSLRESDDDNDDDDGDISMSKRSRNRSKIEDDDSEDEYKDDGDDDE 156
Query: 62 NEAES 66
E ++
Sbjct: 157 EEEDN 161
>UniRef100_Q5Z4U8 Hypothetical protein P0545E05.31-1 [Oryza sativa]
Length = 228
Score = 36.6 bits (83), Expect = 0.15
Identities = 24/61 (39%), Positives = 34/61 (55%), Gaps = 13/61 (21%)
Query: 39 SSIGRNSDDDDDD---EVSSERSMDENEAESKYN----------GGALDCMEALEEVLPI 85
SSIG +S+D+ DD EV S+ S AE+ GGAL C++AL++ LPI
Sbjct: 85 SSIGADSEDEQDDGEEEVESKASSAAVAAETCRRKTKTKCGGGGGGALACLDALDDALPI 144
Query: 86 R 86
+
Sbjct: 145 K 145
>UniRef100_Q8IDW5 Hypothetical protein PF13_0203 [Plasmodium falciparum]
Length = 158
Score = 36.6 bits (83), Expect = 0.15
Identities = 16/50 (32%), Positives = 27/50 (54%)
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYN 69
D D DDD + TT++ + + DDDDDD+ + D+++ E + N
Sbjct: 90 DDDDDDDDDDDESIDTTNNDNDDNDDDDDDDDDDDDDDDDDDDDDEDEIN 139
>UniRef100_UPI00003655FB UPI00003655FB UniRef100 entry
Length = 704
Score = 36.2 bits (82), Expect = 0.19
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 2/48 (4%)
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDD--EVSSERSMDENEAESKYN 69
+ +D+Q K S SSS + DDDDDD E + +E+EAE K N
Sbjct: 637 EEEDEQEKASSEADSSSDDDDDDDDDDDNEEDDDDDEDEEDEAEDKEN 684
>UniRef100_UPI00003655FA UPI00003655FA UniRef100 entry
Length = 729
Score = 36.2 bits (82), Expect = 0.19
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 2/48 (4%)
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDD--EVSSERSMDENEAESKYN 69
+ +D+Q K S SSS + DDDDDD E + +E+EAE K N
Sbjct: 662 EEEDEQEKASSEADSSSDDDDDDDDDDDNEEDDDDDEDEEDEAEDKEN 709
>UniRef100_UPI00003655F9 UPI00003655F9 UniRef100 entry
Length = 746
Score = 36.2 bits (82), Expect = 0.19
Identities = 20/48 (41%), Positives = 27/48 (55%), Gaps = 2/48 (4%)
Query: 24 DHDDDQGKVCSTTSSSSIGRNSDDDDDD--EVSSERSMDENEAESKYN 69
+ +D+Q K S SSS + DDDDDD E + +E+EAE K N
Sbjct: 679 EEEDEQEKASSEADSSSDDDDDDDDDDDNEEDDDDDEDEEDEAEDKEN 726
>UniRef100_Q6A333 Always early 2 protein [Arabidopsis thaliana]
Length = 1051
Score = 36.2 bits (82), Expect = 0.19
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 311 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 368
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 369 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 404
>UniRef100_Q9MA51 F22F7.18 protein [Arabidopsis thaliana]
Length = 1055
Score = 36.2 bits (82), Expect = 0.19
Identities = 30/96 (31%), Positives = 45/96 (46%), Gaps = 9/96 (9%)
Query: 9 KRVGSRTSVLFDAPRDHDDDQGKVCSTTS---SSSIGRNSDDDDDDEVSSERSMDENEAE 65
KRV + + +A + DD G+ CS T S S R + + E S RS + +
Sbjct: 315 KRVYKKRVKVEEAECNDSDDNGEACSATQGLRSKSQRRKAAIEASREKYSPRS--PKKRD 372
Query: 66 SKYNGGALDCMEALEE----VLPIRLFQSFMVMQEK 97
K+ GA D ++AL E +LP L +S + Q K
Sbjct: 373 DKHTSGAFDALQALAELSASMLPANLMESELSAQLK 408
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.129 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 171,413,214
Number of Sequences: 2790947
Number of extensions: 6473256
Number of successful extensions: 69786
Number of sequences better than 10.0: 383
Number of HSP's better than 10.0 without gapping: 199
Number of HSP's successfully gapped in prelim test: 199
Number of HSP's that attempted gapping in prelim test: 66652
Number of HSP's gapped (non-prelim): 1859
length of query: 105
length of database: 848,049,833
effective HSP length: 81
effective length of query: 24
effective length of database: 621,983,126
effective search space: 14927595024
effective search space used: 14927595024
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC139743.4


