

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
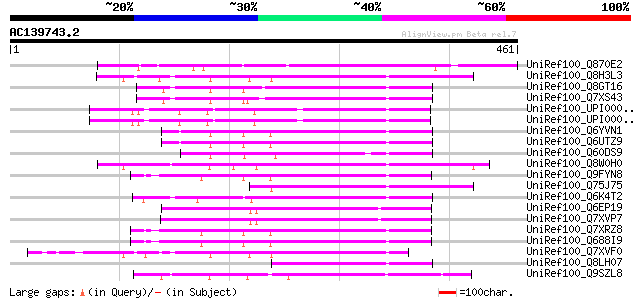
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q870E2 Transposase [Fusarium oxysporum f. sp. melonis] 142 2e-32
UniRef100_Q8H3L3 Putative far-red impaired response protein [Ory... 108 3e-22
UniRef100_Q8GT16 Putative far-red impaired response protein [Ory... 107 7e-22
UniRef100_Q7XS43 OSJNBa0074L08.5 protein [Oryza sativa] 106 1e-21
UniRef100_UPI000042F827 UPI000042F827 UniRef100 entry 105 4e-21
UniRef100_UPI000042CB10 UPI000042CB10 UniRef100 entry 105 4e-21
UniRef100_Q6YVN1 Putative far-red impaired response protein [Ory... 100 7e-20
UniRef100_Q6UTZ9 Putative transposase [Oryza sativa] 100 7e-20
UniRef100_Q60DS9 Hypothetical protein B1140B01.10 [Oryza sativa] 97 7e-19
UniRef100_Q8W0H0 P0529E05.9 protein [Oryza sativa] 95 5e-18
UniRef100_Q9FYN8 P0702D12.15 protein [Oryza sativa] 94 8e-18
UniRef100_Q75J75 Putative far-red impaired response protein [Ory... 93 1e-17
UniRef100_Q6K4T2 Putative far-red impaired response protein [Ory... 93 2e-17
UniRef100_Q6EP19 Putative far-red impaired response protein [Ory... 93 2e-17
UniRef100_Q7XVP7 OSJNBa0050F15.2 protein [Oryza sativa] 93 2e-17
UniRef100_Q7XRZ8 OSJNBb0002N06.8 protein [Oryza sativa] 92 2e-17
UniRef100_Q688I9 Hypothetical protein OSJNBb0012G21.8 [Oryza sat... 92 4e-17
UniRef100_Q7XVF0 OSJNBa0083D01.8 protein [Oryza sativa] 91 5e-17
UniRef100_Q8LH07 Far-red impaired response-like protein [Oryza s... 89 2e-16
UniRef100_Q9SZL8 Hypothetical protein F20D10.300 [Arabidopsis th... 87 1e-15
>UniRef100_Q870E2 Transposase [Fusarium oxysporum f. sp. melonis]
Length = 836
Score = 142 bits (358), Expect = 2e-32
Identities = 110/396 (27%), Positives = 187/396 (46%), Gaps = 26/396 (6%)
Query: 81 NRDELLEWVRRQANKAGFTIVTQRSS-LINPMFRLV--CERSGSHIVPEKKPKHANTGSR 137
+R+EL + A G+ V +RSS N ++ C+R G+ +P + T +R
Sbjct: 23 SREELRSAINAWAAPRGYAFVIKRSSKTANGRTHVIFNCDR-GAGRIPSLSDRRQTT-TR 80
Query: 138 KCGCLFMISGYQSKQTKEWGLNILNGVH--NHPMEPALE--GHILAGRLKEDDKKIVRDL 193
+ GCLF + +S W L G H H EP+ H +L ++ V L
Sbjct: 81 RTGCLFSVLAKESLCKTIWSLRHRPGPHFSQHNHEPSFSEVAHPTLRQLSRQEEITVNQL 140
Query: 194 TKSKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTY 253
T + + P+ I L+ + + + +YN + + + ++ L +L E +
Sbjct: 141 TNAGIAPKEIGSFLRITS-NTLATQQDIYNCIAKGRRDLSKGQSNIHALADQLNEEGF-- 197
Query: 254 YSRTQL-ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLT 312
++R L ES+ + + +AHP S++ +P VL++DSTYKTN ++MP+ ++VGV + T
Sbjct: 198 WNRICLDESSRVTAVLFAHPKSLEYLKTYPEVLILDSTYKTNRFKMPLLDIVGVDACQRT 257
Query: 313 YSVGFGFMTHEKEENFVWVLTMLRKLLSS-KMNMPKVIVTDRDMSLMKAVAHVFPESYAL 371
+ + F F++ E+E +F W L LR + + +P VI+TDR ++ M AV+ FP S
Sbjct: 258 FCIAFAFLSGEEEGDFTWALQALRSVYEDHNIGLPSVILTDRCLACMNAVSSCFPGSALF 317
Query: 372 NCFFHVQANVKQRC----VLNCKYPLGFKKDGKEVSNRDVVKKIMNAWKAMVESPNQQLY 427
C +H+ V+ C P G + +E K+ N W +V S + +Y
Sbjct: 318 LCLWHINKAVQSYCRPAFTRGKDNPQGLGGESEE------WKEFFNFWHEIVASTTEDIY 371
Query: 428 ANALVEFKDS-CSDFSIFVDYAMTT-LDEVKDKIVR 461
L +FK D+ V Y + T LD K V+
Sbjct: 372 NERLEKFKKRYIPDYINEVGYILETWLDLYKKSFVK 407
>UniRef100_Q8H3L3 Putative far-red impaired response protein [Oryza sativa]
Length = 1148
Score = 108 bits (270), Expect = 3e-22
Identities = 92/370 (24%), Positives = 154/370 (40%), Gaps = 36/370 (9%)
Query: 80 KNRDELLEWVRRQANKAGFTIVT----------QRSSLINPMFRLVCERSGSHIVPEKKP 129
KN E E+ A AGF+IV + + FR C R G K
Sbjct: 369 KNITEAHEFFNFYALLAGFSIVRAHNYHTTSKKRNGEVTRVTFR--CNRQGKPTSQSKSV 426
Query: 130 KHANT----------GSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILA 179
T + C C +IS ++ + W ++ + HNH + P E L
Sbjct: 427 SAEETVVSERNTNENDATDCKCALVIS----ERNEIWNISRVQLDHNHQLSPRDEVRFLK 482
Query: 180 GR--LKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMT---NVKQVYNERQQIWKANRG 234
+ ++K ++R L + + R++++ L R + K + N R I K
Sbjct: 483 SHKHMTTEEKMLIRTLKECNIPTRHMIVILSVLRGGLTSLPYTKKDISNVRTTINKETSS 542
Query: 235 DK--KSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYK 292
+ K+L+F K E+ + +Y ES ++++FW S + + + V+ D+TY
Sbjct: 543 NDIMKTLEFFRRKKEKDPHFFYEFDLDESKKVKNLFWTDGRSREWYEKYGDVVSFDTTYF 602
Query: 293 TNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTD 352
TN Y +P VG+ T G F+ E E F W+ K +S K PK I+TD
Sbjct: 603 TNKYNLPFAPFVGIYGHGNTIVFGCAFLHDETSETFKWLFRTFLKAMSQK--EPKTIITD 660
Query: 353 RDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEVSNRDVVKKIM 412
+D ++ A+A VF + NCFFH+ K G + +++ N V ++
Sbjct: 661 QDGAMRSAIAQVFQNAKHRNCFFHIVKKAFNLSGNLLKAKEGLYDEYEDIINNSVTEEEF 720
Query: 413 N-AWKAMVES 421
W+ M++S
Sbjct: 721 EYLWQEMIDS 730
>UniRef100_Q8GT16 Putative far-red impaired response protein [Oryza sativa]
Length = 1114
Score = 107 bits (267), Expect = 7e-22
Identities = 75/285 (26%), Positives = 123/285 (42%), Gaps = 23/285 (8%)
Query: 116 CERSGSHIVPEKKPKHANTGSRK--------CGCLFMISGYQSKQTKEWGLNILNGVHNH 167
C R G + +KK K R C C+ + + W + +N HNH
Sbjct: 401 CYRYGKNDDGQKKKKKKTQQQRNTQVIDRTDCKCMLTVR----LEGDVWRILSMNLNHNH 456
Query: 168 PMEPALEGHILAGR--LKEDDKKIVRDLTKSKMLPRNILIHLKNQR------PHCMTNVK 219
+ P E L + +++KK++R L + + RN++ L R P+ +V
Sbjct: 457 TLTPNREAKFLRSHKHMTDEEKKLIRTLKQCNIPTRNMISILSFLRGGIAALPYTRKDVS 516
Query: 220 QVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFN 279
V + N + +++L K + YY T E+N ++ +FW S +L+
Sbjct: 517 NVGTAINSETR-NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYE 575
Query: 280 NFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLL 339
+ + D+TYKTN Y MP +VGVT F+ E E F WV +
Sbjct: 576 QYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAM 635
Query: 340 SSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
K P+ I+TD+D+++ A+ VFP S NC FH+ ++R
Sbjct: 636 GGK--HPETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRER 678
>UniRef100_Q7XS43 OSJNBa0074L08.5 protein [Oryza sativa]
Length = 1061
Score = 106 bits (265), Expect = 1e-21
Identities = 76/288 (26%), Positives = 124/288 (42%), Gaps = 29/288 (10%)
Query: 116 CERSGSHIVPEKKPKHANTGSRK--------CGCLFMISGYQSKQTKEWGLNILNGVHNH 167
C R G + +KK K R C C+ + W + +N HNH
Sbjct: 348 CYRYGKNDDGQKKKKKKTQQQRNTQVIDRTDCKCMLTVR----LDGDVWRILSMNLNHNH 403
Query: 168 PMEPALEGHILAGR--LKEDDKKIVRDLTKSKMLPRNILIHLKNQR------PHC---MT 216
+ P E L + +++KK++R L + + RN++ L R P+ ++
Sbjct: 404 TLTPNREAKFLRSHKHMTDEEKKLIRTLKQCNIPTRNMISILSFLRGGIAALPYTGKDVS 463
Query: 217 NVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIK 276
NV N + N + +++L K + YY T E+N ++ +FW S +
Sbjct: 464 NVGTAINSETR----NTDMNQVMEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQ 519
Query: 277 LFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLR 336
L+ + + D+TYKTN Y MP +VGVT F+ E E F WV
Sbjct: 520 LYEQYGEFVSFDTTYKTNRYNMPFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFL 579
Query: 337 KLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
+ K P+ I+TD+D+++ A+ VFP S NC FH+ ++R
Sbjct: 580 TAMGGK--HPETIITDQDLAMRAAIRQVFPNSKHRNCLFHILKKCRER 625
>UniRef100_UPI000042F827 UPI000042F827 UniRef100 entry
Length = 580
Score = 105 bits (261), Expect = 4e-21
Identities = 90/341 (26%), Positives = 146/341 (42%), Gaps = 43/341 (12%)
Query: 73 FSDDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINP-----------MFRLV------ 115
F + +++DE V+ A GF V +RS P + +
Sbjct: 91 FDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSKRKKPTKTHKEGRELIQYSCIFGDEYR 150
Query: 116 CERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQT--------KEWGLNILNGVHNH 167
C R S K + + K GC F ++ + K W ++N HNH
Sbjct: 151 CTRGDSF----KDVRIRTIQTVKNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNH 206
Query: 168 PMEPALEGHIL--AGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQV---- 221
P + +L A R E K+++ L S+ +I+ + + P M + +
Sbjct: 207 PPVANIGIALLRRASRTPEA-KELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLA 265
Query: 222 YNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 281
+ R + + ++ L+S E H +R +L + + T+ L + F
Sbjct: 266 FRRRADLRAGSSASAACIRQLVSNDELHRPFVNTRDELIG-----LVYTKRTARALIHRF 320
Query: 282 PTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSS 341
P VL D TYKTN+YRMPM +VG TST +TY+ G M E + L +L+
Sbjct: 321 PEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGK 380
Query: 342 KMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVK 382
KV++TDRD +L+ A+ V P++Y +CF+H+Q NVK
Sbjct: 381 P--DVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>UniRef100_UPI000042CB10 UPI000042CB10 UniRef100 entry
Length = 932
Score = 105 bits (261), Expect = 4e-21
Identities = 90/341 (26%), Positives = 146/341 (42%), Gaps = 43/341 (12%)
Query: 73 FSDDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINP-----------MFRLV------ 115
F + +++DE V+ A GF V +RS P + +
Sbjct: 91 FDEQAYYQSKDECKGAVQAYAASQGFAAVIRRSKRKKPTKTHKEGRELIQYSCIFGDEYR 150
Query: 116 CERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQT--------KEWGLNILNGVHNH 167
C R S K + + K GC F ++ + K W ++N HNH
Sbjct: 151 CTRGDSF----KDVRIRTIQTVKNGCPFAAVAWEMAEAEVKEHGGDKCWTWKVVNASHNH 206
Query: 168 PMEPALEGHIL--AGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQV---- 221
P + +L A R E K+++ L S+ +I+ + + P M + +
Sbjct: 207 PPVANIGIALLRRASRTPEA-KELINTLIDSRTPFPSIISTMSQRFPTVMLTSEDLHRLA 265
Query: 222 YNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 281
+ R + + ++ L+S E H +R +L + + T+ L + F
Sbjct: 266 FRRRADLRAGSSASAACIRQLVSNDELHRPFVNTRDELIG-----LVYTKRTARALIHRF 320
Query: 282 PTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSS 341
P VL D TYKTN+YRMPM +VG TST +TY+ G M E + L +L+
Sbjct: 321 PEVLFFDCTYKTNLYRMPMLHIVGSTSTGMTYTAGVILMLRETTNWYTQALNSFFELVGK 380
Query: 342 KMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVK 382
KV++TDRD +L+ A+ V P++Y +CF+H+Q NVK
Sbjct: 381 P--DVKVVITDRDPALINALMSVLPKAYRFSCFWHLQENVK 419
>UniRef100_Q6YVN1 Putative far-red impaired response protein [Oryza sativa]
Length = 1132
Score = 100 bits (250), Expect = 7e-20
Identities = 70/253 (27%), Positives = 112/253 (43%), Gaps = 10/253 (3%)
Query: 139 CGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKS 196
C C+ M+ + W + L+ HNH + P E L + +K+++R L +
Sbjct: 479 CKCMMMVEKMFALGDV-WQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKEC 537
Query: 197 KMLPRNILIHLKNQR---PHCMTNVKQVYNERQQIWKANRGD--KKSLQFLISKLEEHNY 251
+ R++++ L R P K V N I R + K+ L +L K E
Sbjct: 538 NIPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNNDMKQVLAYLRKKEIEDPG 597
Query: 252 TYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDL 311
Y E+N + +FW S +L+ + + D+TY+TN Y MP VGV
Sbjct: 598 ISYKFKLDENNKVTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGT 657
Query: 312 TYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAL 371
T G F+ E E F WV + K PK I+TD+D ++ A+A VF +
Sbjct: 658 TCLFGCAFLGDETAETFKWVFETFATAMGGK--HPKTIITDQDNAMRSAIAQVFQNTKHR 715
Query: 372 NCFFHVQANVKQR 384
NC FH++ N +++
Sbjct: 716 NCLFHIKKNCREK 728
>UniRef100_Q6UTZ9 Putative transposase [Oryza sativa]
Length = 1037
Score = 100 bits (250), Expect = 7e-20
Identities = 70/253 (27%), Positives = 112/253 (43%), Gaps = 10/253 (3%)
Query: 139 CGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKS 196
C C+ M+ + W + L+ HNH + P E L + +K+++R L +
Sbjct: 384 CKCMMMVEKMFALGDV-WQIATLDLKHNHALCPRREAKFLKSHKNMTIKEKRLIRTLKEC 442
Query: 197 KMLPRNILIHLKNQR---PHCMTNVKQVYNERQQIWKANRGD--KKSLQFLISKLEEHNY 251
+ R++++ L R P K V N I R + K+ L +L K E
Sbjct: 443 NIPTRSMIVILSFLRGGLPALPYTKKDVSNVGTAINSETRNNDMKQVLAYLRKKEIEDPG 502
Query: 252 TYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDL 311
Y E+N + +FW S +L+ + + D+TY+TN Y MP VGV
Sbjct: 503 ISYKFKLDENNKVTSMFWTDGRSTQLYEEYGDCISFDTTYRTNRYNMPFAPFVGVIGHGT 562
Query: 312 TYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAL 371
T G F+ E E F WV + K PK I+TD+D ++ A+A VF +
Sbjct: 563 TCLFGCAFLGDETAETFKWVFETFATAMGGK--HPKTIITDQDNAMRSAIAQVFQNTKHR 620
Query: 372 NCFFHVQANVKQR 384
NC FH++ N +++
Sbjct: 621 NCLFHIKKNCREK 633
>UniRef100_Q60DS9 Hypothetical protein B1140B01.10 [Oryza sativa]
Length = 904
Score = 97.4 bits (241), Expect = 7e-19
Identities = 63/236 (26%), Positives = 109/236 (45%), Gaps = 14/236 (5%)
Query: 156 WGLNILNGVHNHPMEPALEGHILAGR--LKEDDKKIVRDLTKSKMLPRNILIHLKNQRP- 212
W + LN HNHP+ P E L + + +K ++R L K + R I+ L R
Sbjct: 431 WYITRLNLEHNHPLSPPEERKFLWSHKHMIDQEKLLIRTLNKINVPTRMIMSVLSYVRGG 490
Query: 213 -HCMTNVKQVYNERQQIWKANRGDKKSLQ---FLISKLEEHNYTYYSRTQLESNTIEDIF 268
+ K+ + + + G +Q F K+ E Y+ E+N ++ +F
Sbjct: 491 LFAVPYTKKAMSNYRDFVRRESGKNDMMQCLDFFEKKISEDPLFYFRFRTDENNVVKSLF 550
Query: 269 WAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENF 328
W+ S K + F ++ D+TYKTN Y +P VG+TS G+ F+ E
Sbjct: 551 WSDRNSRKFYEMFGDIVSFDTTYKTNRYDLPFAPFVGITSHGDNCLFGYAFLQDE----- 605
Query: 329 VWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
V+ + + K +P I+TD+D+++ A+ VFP++ NC FH+ +N + +
Sbjct: 606 TMVVQYVLDCMGGK--VPATIITDQDLAMKAAIVIVFPDTVHRNCMFHMLSNARDK 659
>UniRef100_Q8W0H0 P0529E05.9 protein [Oryza sativa]
Length = 773
Score = 94.7 bits (234), Expect = 5e-18
Identities = 89/371 (23%), Positives = 155/371 (40%), Gaps = 18/371 (4%)
Query: 81 NRDELLEWVRRQANKAGFTIVT----QRSSLINPMFRLVCERSGSHIVPEKKPKHANTGS 136
+ D E+ R A GF+I + S+ + + VC R G +
Sbjct: 159 DEDIAYEFYNRYAGDVGFSIRKFWHDKSSTNVIRTKKFVCSREGFNKRNTSDACQRKRAD 218
Query: 137 RKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAG--RLKEDDKKIVRDLT 194
+ GC+ ++ + T ++ + + HNH + + H+L R+ E K + L
Sbjct: 219 TRVGCMAEMT-IKITPTGKYAIASFSNTHNHELITPSKAHLLRSQRRMTEAQKAQIDILN 277
Query: 195 KSKMLPRN---ILIHLKNQRPHCMTNVKQVYN--ERQQIWKANRGDKKS-LQFLISKLEE 248
S + + ++ R K N +++ GD + LQ+L K E
Sbjct: 278 DSGVRSKEGHEVMSRQAGGRQSLTFTRKDYKNYLRSKRMKSIQEGDTGAILQYLQDKQME 337
Query: 249 HNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTS 308
+ +Y+ E + +IFWA S+ F+ F V+ D+TY+TN Y P VGV
Sbjct: 338 NPSFFYAIQVDEDEMMTNIFWADARSVLDFDYFGDVICFDTTYRTNNYGRPFALFVGVNH 397
Query: 309 TDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPES 368
T G + E F W+ ++ +S K P+ I+TD+ +++ A+ VFP S
Sbjct: 398 HKQTVVFGAALLYDETTSTFEWLFETFKRAMSGK--EPRTILTDQCAAIINAIGTVFPNS 455
Query: 369 YALNCFFHVQANVKQRCVLNCKYPLGFKKD-GKEVSNRDVVKKIMNAWKAMVE--SPNQQ 425
C +H+ N + FK D GK V + + V + ++AW M+E + N
Sbjct: 456 THRLCVWHMYQNAAVHLSHIFQGSKTFKNDFGKCVFDFEEVDEFISAWNKMIEEYNLNDN 515
Query: 426 LYANALVEFKD 436
+ + L E K+
Sbjct: 516 QWLHHLFEIKE 526
>UniRef100_Q9FYN8 P0702D12.15 protein [Oryza sativa]
Length = 832
Score = 94.0 bits (232), Expect = 8e-18
Identities = 70/281 (24%), Positives = 125/281 (43%), Gaps = 17/281 (6%)
Query: 111 MFRLVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPME 170
M ++ C G + KKP S + GC M+ +S+ W + + HNHPM+
Sbjct: 113 MRQICCSHQGRN-PKTKKP------SVRIGCPAMMKINRSRAGSGWSVTKVVSAHNHPMK 165
Query: 171 PAL---EGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQR--PHCMTNVKQVYNER 225
++ + + ++ E + I+ ++ S M N+ L P + ++ +
Sbjct: 166 KSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRL 225
Query: 226 QQIWKANRGD---KKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFP 282
+ + +K+L L + +YS E+ +++IFW+H S F +F
Sbjct: 226 AYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFG 285
Query: 283 TVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSK 342
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++ K
Sbjct: 286 DVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK 345
Query: 343 MNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+P I+TD S+ A+ VFP + C +HV K+
Sbjct: 346 --VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 384
>UniRef100_Q75J75 Putative far-red impaired response protein [Oryza sativa]
Length = 888
Score = 93.2 bits (230), Expect = 1e-17
Identities = 58/206 (28%), Positives = 97/206 (46%), Gaps = 5/206 (2%)
Query: 219 KQVYNERQQIWKANRGDK--KSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIK 276
K + N R I K + K+L+F K E+ + +Y ES ++++FW S +
Sbjct: 468 KDISNVRTTINKETSSNDMMKTLEFFRRKKEKDPHFFYEFDLDESKKVKNLFWTDGRSRE 527
Query: 277 LFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLR 336
+ + V+ D+TY TN Y +P VG++ T G F+ E E F W+
Sbjct: 528 WYEKYGDVVSFDTTYFTNRYNLPFAPFVGISGHGNTIVFGCAFLHDETSETFKWMFRTFL 587
Query: 337 KLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFK 396
K +S K PK+I+TD+D ++ A+A VF + NCFFH+ K G
Sbjct: 588 KAMSQK--EPKIIITDQDGAMRSAIAQVFQNAKHGNCFFHIVKKAFNLSGNLFKAKEGLY 645
Query: 397 KDGKEVSNRDVVKKIMN-AWKAMVES 421
+ +++ N V ++ W+ M++S
Sbjct: 646 DEYEDIINNSVTEEEFEYLWQEMIDS 671
>UniRef100_Q6K4T2 Putative far-red impaired response protein [Oryza sativa]
Length = 934
Score = 92.8 bits (229), Expect = 2e-17
Identities = 70/287 (24%), Positives = 128/287 (44%), Gaps = 21/287 (7%)
Query: 112 FRLVCERSG------SHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVH 165
F L C +SG S P KK + +C ++ EW + H
Sbjct: 314 FHLECNKSGKMKLTKSSENPMKKRRSNLVEKTQCKARVIVK----LDKGEWEFTAVRHEH 369
Query: 166 NHPM--EPALEGHILAGR-LKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNV---- 218
NHP+ P L I+ + + +K +R L ++++ P+ I+ + R C ++
Sbjct: 370 NHPLCPSPLLARFIVDHKQMSTGEKSFLRVLQQNRVPPKKIMKIFRKLRV-CFGDIPFEN 428
Query: 219 KQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTY-YSRTQLESNTIEDIFWAHPTSIKL 277
K +N Q + D +S ++L+ N + Y + E NT+ IFW
Sbjct: 429 KDEHNIAQTEHRKANSDVESALKHFTELQIQNPEFLYVMQKDEDNTVTSIFWTDARLRIE 488
Query: 278 FNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRK 337
++ F +++ D+ Y T+MY MP ++G+ S + +G + EK E F W+L +
Sbjct: 489 YDIFGDLIMFDAAYSTDMYNMPFVPIIGINSHATPFLLGCALLKDEKVETFEWMLRTFLQ 548
Query: 338 LLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
++ K MP+ ++T++D S+ KA A + P C HV + +++
Sbjct: 549 VMGGK--MPRAVITNQDTSMEKAFAELMPHVRLRFCKRHVMSKAQEK 593
>UniRef100_Q6EP19 Putative far-red impaired response protein [Oryza sativa]
Length = 688
Score = 92.8 bits (229), Expect = 2e-17
Identities = 63/253 (24%), Positives = 118/253 (45%), Gaps = 10/253 (3%)
Query: 139 CGCLFMISGYQSKQTKEWGLNILNGVHNHPM-EPALEGHILAGRLKEDDKKIVR-DLTKS 196
CGCL + ++++ W + + H H + P + R+ D KK +L S
Sbjct: 88 CGCLAKLEIERNEEKGVWFVKEFDNQHTHELANPDHVAFLGVHRVMSDSKKAQAVELRMS 147
Query: 197 KMLPRNILIHLKNQRPHCMTN---VKQVYN--ERQQIWKANRGDKKS-LQFLISKLEEHN 250
+ P ++ ++N +K +YN R ++ D + L++L K EE
Sbjct: 148 GLRPFQVMEVMENNHDELDEVGFVMKDLYNFFTRYEMKNIKGHDAEDVLKYLTRKQEEDA 207
Query: 251 YTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTD 310
++ T E + ++FWA S + F V++ DSTY+ N Y +P +GV
Sbjct: 208 EFFFKYTTDEEGRLRNVFWADAESRLDYAAFGGVVIFDSTYRVNKYNLPFIPFIGVNHHR 267
Query: 311 LTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYA 370
T G G +++E ++ W+L L + + PK ++TD D+++ KA++ V P +Y
Sbjct: 268 STTIFGCGILSNESVNSYCWLLETF--LEAMRQVHPKSLITDGDLAMAKAISKVMPGAYH 325
Query: 371 LNCFFHVQANVKQ 383
C +H++ N+ +
Sbjct: 326 RLCTWHIEENMSR 338
>UniRef100_Q7XVP7 OSJNBa0050F15.2 protein [Oryza sativa]
Length = 688
Score = 92.8 bits (229), Expect = 2e-17
Identities = 63/254 (24%), Positives = 118/254 (45%), Gaps = 10/254 (3%)
Query: 138 KCGCLFMISGYQSKQTKEWGLNILNGVHNHPM-EPALEGHILAGRLKEDDKKIVR-DLTK 195
+CGCL + ++++ W + + H H + P + R+ D KK +L
Sbjct: 87 RCGCLAKLEIERNEEKGVWFVKEFDNQHTHELANPDHVAFLGVHRVMSDSKKAQAVELRM 146
Query: 196 SKMLPRNILIHLKNQRPHCMTN---VKQVYN--ERQQIWKANRGDKKS-LQFLISKLEEH 249
S + P ++ ++N +K +YN R + D + L++L K EE
Sbjct: 147 SGLRPFQVMEVMENNHDELDEVGFVMKDLYNFFTRYDMKNIKGHDAEDVLKYLTRKQEED 206
Query: 250 NYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTST 309
++ T E + ++FWA S + F V++ DSTY+ N Y +P +GV
Sbjct: 207 AEFFFKYTTDEEGRLRNVFWADAESRLDYAAFGGVVIFDSTYRVNKYNLPFIPFIGVNHH 266
Query: 310 DLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESY 369
T G G +++E ++ W+L L + + PK ++TD D+++ KA++ V P +Y
Sbjct: 267 RSTTIFGCGILSNESVNSYCWLLETF--LEAMRQVHPKSLITDGDLAMAKAISKVMPGAY 324
Query: 370 ALNCFFHVQANVKQ 383
C +H++ N+ +
Sbjct: 325 HRLCTWHIEENMSR 338
>UniRef100_Q7XRZ8 OSJNBb0002N06.8 protein [Oryza sativa]
Length = 885
Score = 92.4 bits (228), Expect = 2e-17
Identities = 70/281 (24%), Positives = 124/281 (43%), Gaps = 17/281 (6%)
Query: 111 MFRLVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPME 170
M ++ C G + KKP S + GC M+ +S W + + HNHPM+
Sbjct: 166 MRQICCSHQGRN-PKTKKP------SVRIGCPAMMKINRSGAGSGWSVTKVVSTHNHPMK 218
Query: 171 PAL---EGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQR--PHCMTNVKQVYNER 225
++ + + ++ E + I+ ++ S M N+ L P + ++ +
Sbjct: 219 KSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRL 278
Query: 226 QQIWKANRGD---KKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFP 282
+ + +K+L L + +YS E+ +++IFW+H S F +F
Sbjct: 279 AYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFG 338
Query: 283 TVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSK 342
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++ K
Sbjct: 339 DVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK 398
Query: 343 MNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+P I+TD S+ A+ VFP + C +HV K+
Sbjct: 399 --VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 437
>UniRef100_Q688I9 Hypothetical protein OSJNBb0012G21.8 [Oryza sativa]
Length = 822
Score = 91.7 bits (226), Expect = 4e-17
Identities = 70/281 (24%), Positives = 123/281 (42%), Gaps = 17/281 (6%)
Query: 111 MFRLVCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPME 170
M ++ C G + KKP S + GC M+ S W + + HNHPM+
Sbjct: 166 MRQICCSHQGRN-PKTKKP------SVRIGCPAMMKINMSGAGSGWSVTKVVSAHNHPMK 218
Query: 171 PAL---EGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQR--PHCMTNVKQVYNER 225
++ + + ++ E + I+ ++ S M N+ L P + ++ +
Sbjct: 219 KSVGVTKNYQSHNQIDEGTRGIIEEMVDSSMSLTNMYGMLSGMHGGPSMVPFTRKAMDRV 278
Query: 226 QQIWKANRGD---KKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFP 282
+ + +K+L L + +YS E+ +++IFW+H S F +F
Sbjct: 279 AYAIRRDESSDDMQKTLDVLKDLQKRSKNFFYSIQVDEACRVKNIFWSHAVSRLNFEHFG 338
Query: 283 TVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSK 342
V+ D+TYKTN Y MP VGV + + G + E EE+F W+ ++ ++ K
Sbjct: 339 DVITFDTTYKTNKYNMPFAPFVGVNNHFQSTFFGCALLREETEESFTWLFNTFKECMNGK 398
Query: 343 MNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+P I+TD S+ A+ VFP + C +HV K+
Sbjct: 399 --VPIGILTDNCPSMAAAIRTVFPNTIHRVCKWHVLKKAKE 437
>UniRef100_Q7XVF0 OSJNBa0083D01.8 protein [Oryza sativa]
Length = 1161
Score = 91.3 bits (225), Expect = 5e-17
Identities = 91/375 (24%), Positives = 155/375 (41%), Gaps = 50/375 (13%)
Query: 17 GETKENFVENSGDVKPPNFEAKTNIDEASVHVEASKYVGGSGVLPLQPHEVDTAKFFSDD 76
G+ +N +NS ++ +A TN+D +V E + G+ +D D
Sbjct: 442 GQNVQNSADNSTEI---GNQATTNVD--TVEQEQEEATKGNNAA------IDAKYIPRVD 490
Query: 77 VKSKNRDELLEWVRRQANKAGFTIVT----------QRSSLINPMFRLVCERSGSH---- 122
V+ K+ E ++ A AGF++V + +I F+ C R G
Sbjct: 491 VQFKSIKEAHDFFNFYALLAGFSVVIAHNYHSTSKKRNGEVIRVTFK--CNRHGKAKSES 548
Query: 123 --------IVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALE 174
+V E+ S C C +IS ++ W + +N HN+ M P E
Sbjct: 549 QEEETEETVVAERNSNEIKATS--CNCALVIS----ERNLIWRITRVNLDHNYKMSPRDE 602
Query: 175 GHILAGR--LKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMT---NVKQVYNERQQIW 229
L + ++K ++R L + + R++++ L R + K V N R I
Sbjct: 603 VRFLKSHKNMTTEEKMLIRTLKECNIPTRHMIVILSTLRGGLTSLPYTKKDVSNVRTCIN 662
Query: 230 KANRGDK--KSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVM 287
K + + LQF K E+ +Y E+ + ++FW SI + + V+
Sbjct: 663 KETSSNDMMQVLQFFRKKKEKDPKFFYEFDLDENKKVTNLFWTDGRSIDWYEKYGDVVSF 722
Query: 288 DSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPK 347
D+TY TN Y +P VG++ T G F+ E E F W+ K +S K PK
Sbjct: 723 DTTYFTNRYNLPFAPFVGISGHGNTIVFGCAFLHDETAETFKWLFITFLKAMSKK--APK 780
Query: 348 VIVTDRDMSLMKAVA 362
I+TD+D ++ A++
Sbjct: 781 TIITDQDGAMSTALS 795
>UniRef100_Q8LH07 Far-red impaired response-like protein [Oryza sativa]
Length = 772
Score = 89.4 bits (220), Expect = 2e-16
Identities = 46/146 (31%), Positives = 72/146 (48%), Gaps = 2/146 (1%)
Query: 239 LQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRM 298
+++L K + YY T E+N ++ +FW S +L+ + + D+TYKTN Y M
Sbjct: 220 MEYLRQKEAKDPGYYYKLTLDENNKVKSMFWTDGRSTQLYEQYGEFVSFDTTYKTNRYNM 279
Query: 299 PMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLM 358
P +VGVT F+ E E F WV + K P+ I+TD+D+++
Sbjct: 280 PFAPIVGVTGHGNICIFACAFLGDETTETFKWVFETFLTAMGRK--HPETIITDQDLAMR 337
Query: 359 KAVAHVFPESYALNCFFHVQANVKQR 384
A+ VFP S NC FH+ ++R
Sbjct: 338 AAIRQVFPNSKHRNCLFHILKKYRER 363
>UniRef100_Q9SZL8 Hypothetical protein F20D10.300 [Arabidopsis thaliana]
Length = 788
Score = 87.0 bits (214), Expect = 1e-15
Identities = 79/324 (24%), Positives = 142/324 (43%), Gaps = 23/324 (7%)
Query: 113 RLVCERSGSHIVPEKKPKHANTGS----RKCGCLFMISGYQSKQTKEWGLNILNGVHNHP 168
+ VC + G + EK+ K + GC +S + + + +W ++ HNH
Sbjct: 118 QFVCAKEGFRNMNEKRTKDREIKRPRTITRVGCKASLS-VKMQDSGKWLVSGFVKDHNHE 176
Query: 169 MEPALEGHILAG--RLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHC----MTNVKQVY 222
+ P + H L ++ K ++ L + M PR I+ L + T V
Sbjct: 177 LVPPDQVHCLRSHRQISGPAKTLIDTLQAAGMGPRRIMSALIKEYGGISKVGFTEVDCRN 236
Query: 223 NERQQIWKANRGDKKSLQFLISKLEEHNYT----YYSRTQLESNTIEDIFWAHPTSIKLF 278
R K+ G+ +Q L+ L + N +YS E ++ ++FWA P +I F
Sbjct: 237 YMRNNRQKSIEGE---IQLLLDYLRQMNADNPNFFYSVQGSEDQSVGNVFWADPKAIMDF 293
Query: 279 NNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKL 338
+F + D+TY++N YR+P GV G F+ +E E +FVW+
Sbjct: 294 THFGDTVTFDTTYRSNRYRLPFAPFTGVNHHGQPILFGCAFIINETEASFVWLFNTWLAA 353
Query: 339 LSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCV-LNCKYPLGFKK 397
+S+ + P I TD D + A+ HVFP + C +H+ +++ + K+P F+
Sbjct: 354 MSA--HPPVSITTDHDAVIRAAIMHVFPGARHRFCKWHILKKCQEKLSHVFLKHP-SFES 410
Query: 398 D-GKEVSNRDVVKKIMNAWKAMVE 420
D K V+ + V+ W ++++
Sbjct: 411 DFHKCVNLTESVEDFERCWFSLLD 434
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 763,173,220
Number of Sequences: 2790947
Number of extensions: 31021773
Number of successful extensions: 78143
Number of sequences better than 10.0: 382
Number of HSP's better than 10.0 without gapping: 247
Number of HSP's successfully gapped in prelim test: 135
Number of HSP's that attempted gapping in prelim test: 77677
Number of HSP's gapped (non-prelim): 444
length of query: 461
length of database: 848,049,833
effective HSP length: 131
effective length of query: 330
effective length of database: 482,435,776
effective search space: 159203806080
effective search space used: 159203806080
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 77 (34.3 bits)
Medicago: description of AC139743.2


