

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139706.5 - phase: 0
(424 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
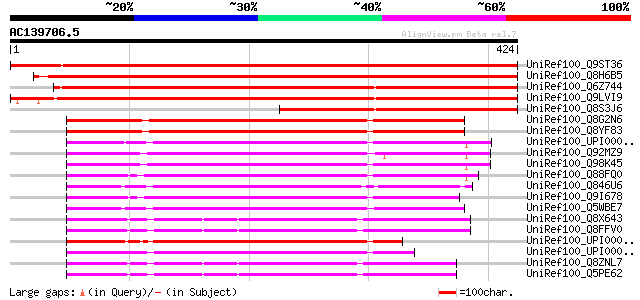
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ST36 JPR ORF1 protein [Pyrus pyrifolia] 726 0.0
UniRef100_Q8H6B5 Putative dehydrogenase [Lycopersicon esculentum] 702 0.0
UniRef100_Q6Z744 Putative dihydropyrimidine dehydrogenase [Oryza... 676 0.0
UniRef100_Q9LVI9 Senescencs-related protein; dihydroorotate dehy... 674 0.0
UniRef100_Q8S3J6 Putative dihydroorotate dehydrogenase [Oryza sa... 341 3e-92
UniRef100_Q8G2N6 Dihydroorotate dehydrogenase family protein [Br... 280 7e-74
UniRef100_Q8YF83 DIHYDROPYRIMIDINE DEHYDROGENASE [Brucella melit... 278 2e-73
UniRef100_UPI00002B95BA UPI00002B95BA UniRef100 entry 276 1e-72
UniRef100_Q92MZ9 PUTATIVE OXIDOREDUCTASE IRON-SULFUR PROTEIN [Rh... 273 7e-72
UniRef100_Q98K45 Probable oxidoreductase [Rhizobium loti] 266 8e-70
UniRef100_Q88FQ0 Dihydroorotate dehydrogenase family protein [Ps... 260 6e-68
UniRef100_Q846U6 Dihydropyrimidine dehydrogenase [Brevibacillus ... 258 3e-67
UniRef100_Q9I678 Probable oxidoreductase [Pseudomonas aeruginosa] 256 7e-67
UniRef100_Q5WBE7 Dihydropyrimidine dehydrogenase [Bacillus clausii] 252 1e-65
UniRef100_Q8X643 Hypothetical protein yeiA [Escherichia coli] 252 2e-65
UniRef100_Q8FFV0 Hypothetical protein yeiA [Escherichia coli O6] 252 2e-65
UniRef100_UPI00002FC63E UPI00002FC63E UniRef100 entry 248 2e-64
UniRef100_UPI00002C550A UPI00002C550A UniRef100 entry 247 5e-64
UniRef100_Q8ZNL7 Putative dihydropyrimidine dehydrogenase [Salmo... 246 7e-64
UniRef100_Q5PE62 Putative dihydropyrimidine dehydrogenase [Salmo... 246 9e-64
>UniRef100_Q9ST36 JPR ORF1 protein [Pyrus pyrifolia]
Length = 424
Score = 726 bits (1875), Expect = 0.0
Identities = 361/425 (84%), Positives = 391/425 (91%), Gaps = 2/425 (0%)
Query: 1 MANLSMTQLKTRNSASRFSFNFSKKV-YPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMP 59
MA++S TQ+ +N + FS +V RPSRV F+VFASE + +EPDLSV VNGLHMP
Sbjct: 1 MASISFTQIGGKNPVAEFSRRPRPEVRLTRPSRVGFRVFASESK-AEPDLSVTVNGLHMP 59
Query: 60 NPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKG 119
NPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSL+A KV NVTPRYARLR + NGSAKG
Sbjct: 60 NPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLEADKVKNVTPRYARLRVDGNGSAKG 119
Query: 120 EIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGI 179
+IIGW+NIELISDRPL+ MLKEFKQLK+EYPDRILIASIMEEYNKA WEELIDRVEQTG+
Sbjct: 120 QIIGWENIELISDRPLDIMLKEFKQLKQEYPDRILIASIMEEYNKAGWEELIDRVEQTGV 179
Query: 180 DAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPAR 239
DA+EINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDI+QPAR
Sbjct: 180 DALEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDITQPAR 239
Query: 240 VALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAK 299
VALSSGCEGV+AINTIMSVMGINL TLRPEPCVEGYSTPGGYSAKAVHPIAL KVM+IAK
Sbjct: 240 VALSSGCEGVSAINTIMSVMGINLTTLRPEPCVEGYSTPGGYSAKAVHPIALAKVMNIAK 299
Query: 300 MMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDF 359
+MKSEF +YSLS IGGVE+GGDAAEFILLGANTVQVCTGVMMHGYG+VKKL EL+DF
Sbjct: 300 LMKSEFGDTDYSLSGIGGVESGGDAAEFILLGANTVQVCTGVMMHGYGVVKKLCSELQDF 359
Query: 360 MKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESE 419
MK+HNF+SIEDFRGASL+YFTTHTDLV RQQEAIKQRKAI+KGLQSDKDWTGDGFVKE+E
Sbjct: 360 MKQHNFSSIEDFRGASLEYFTTHTDLVRRQQEAIKQRKAIRKGLQSDKDWTGDGFVKETE 419
Query: 420 SMVSN 424
SMVSN
Sbjct: 420 SMVSN 424
>UniRef100_Q8H6B5 Putative dehydrogenase [Lycopersicon esculentum]
Length = 429
Score = 702 bits (1813), Expect = 0.0
Identities = 347/404 (85%), Positives = 374/404 (91%), Gaps = 6/404 (1%)
Query: 21 NFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAF 80
NF +K RV F++ A EGQ EPDLSV VNGL MPNPFVIGSGPPGTNYTVMKRAF
Sbjct: 32 NFGRK------RVGFRIMALEGQSVEPDLSVTVNGLKMPNPFVIGSGPPGTNYTVMKRAF 85
Query: 81 DEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGEIIGWQNIELISDRPLETMLK 140
DEGWGGVIAKTVSL+A KV NVTPRYA+LRA+ANGSAKG+IIGWQNIELISDRPLETMLK
Sbjct: 86 DEGWGGVIAKTVSLEADKVKNVTPRYAKLRADANGSAKGQIIGWQNIELISDRPLETMLK 145
Query: 141 EFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGAA 200
EFKQLKEEYPDRILIASIMEEYNKAAWEELI R E+TGIDA EINFSCPHGMPER+MGAA
Sbjct: 146 EFKQLKEEYPDRILIASIMEEYNKAAWEELIYRCEETGIDAFEINFSCPHGMPERRMGAA 205
Query: 201 VGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMG 260
VGQDC LLEEVCGWINA ATVPVWAKMTPNITDI++PARVAL+ GCEGV+AINTIMSVMG
Sbjct: 206 VGQDCDLLEEVCGWINAVATVPVWAKMTPNITDITKPARVALNQGCEGVSAINTIMSVMG 265
Query: 261 INLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVET 320
INL+TLRPEPCVEGYSTPGGYS+KAVHPIAL KVM+IA+MMKSEF ++YSLSAIGGVET
Sbjct: 266 INLDTLRPEPCVEGYSTPGGYSSKAVHPIALAKVMNIARMMKSEFGDKDYSLSAIGGVET 325
Query: 321 GGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
GGDAAEFILLGA+TVQVCTGVMMHGYGLVK L ELKDFM+KHNF+SIEDFRG SL+YFT
Sbjct: 326 GGDAAEFILLGADTVQVCTGVMMHGYGLVKTLCSELKDFMRKHNFSSIEDFRGTSLEYFT 385
Query: 381 THTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESESMVSN 424
THTDLV RQQEAI+QRKA+KKGLQSDKDWTGDGFV+E+ESMVSN
Sbjct: 386 THTDLVRRQQEAIRQRKAVKKGLQSDKDWTGDGFVQETESMVSN 429
>UniRef100_Q6Z744 Putative dihydropyrimidine dehydrogenase [Oryza sativa]
Length = 414
Score = 676 bits (1744), Expect = 0.0
Identities = 333/388 (85%), Positives = 359/388 (91%), Gaps = 2/388 (0%)
Query: 37 VFASEGQVSEPDLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA 96
V AS G EPDLSV+VNGL MPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA
Sbjct: 29 VRASAG-AGEPDLSVRVNGLKMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDA 87
Query: 97 AKVINVTPRYARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIA 156
KVINVTPRYARLRA+ NGS K IIGWQNIELISDRPLETML EFKQLK+EYPDRILI
Sbjct: 88 EKVINVTPRYARLRADPNGSTKSPIIGWQNIELISDRPLETMLNEFKQLKKEYPDRILIG 147
Query: 157 SIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWIN 216
SIMEEYNKAAW ELI+RVE++G+DA+EINFSCPHGMPERKMGAAVGQDC LLEEVCGWIN
Sbjct: 148 SIMEEYNKAAWHELIERVEESGVDALEINFSCPHGMPERKMGAAVGQDCDLLEEVCGWIN 207
Query: 217 AKATVPVWAKMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYS 276
KATVPVWAKMTPNITDI++PAR++L SGCEGV+AINTIMSVMGINL TLRPEPCVEGYS
Sbjct: 208 EKATVPVWAKMTPNITDITKPARISLKSGCEGVSAINTIMSVMGINLKTLRPEPCVEGYS 267
Query: 277 TPGGYSAKAVHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQ 336
TPGGYSA+AVHPIAL KVM IA+MMK EF ++ SLSAIGGVETG DAAEFILLGA+TVQ
Sbjct: 268 TPGGYSARAVHPIALAKVMQIARMMKEEF-ADGQSLSAIGGVETGNDAAEFILLGADTVQ 326
Query: 337 VCTGVMMHGYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQR 396
VCTGVMMHGYGLVKKL EL+DFM++HNF+SIEDFRGASL YFTTHTDLVHRQ+EAI QR
Sbjct: 327 VCTGVMMHGYGLVKKLCAELQDFMRQHNFSSIEDFRGASLPYFTTHTDLVHRQREAINQR 386
Query: 397 KAIKKGLQSDKDWTGDGFVKESESMVSN 424
KAI+KGL+SDKDWTGDGFVKE+ESMVSN
Sbjct: 387 KAIRKGLESDKDWTGDGFVKETESMVSN 414
>UniRef100_Q9LVI9 Senescencs-related protein; dihydroorotate dehydrogenase-like
protein [Arabidopsis thaliana]
Length = 426
Score = 674 bits (1739), Expect = 0.0
Identities = 338/429 (78%), Positives = 381/429 (88%), Gaps = 8/429 (1%)
Query: 1 MANLS--MTQLKTRNSASRFSFNF---SKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNG 55
MA++S + + +S + S +F S++ + P+RV K+ S SEPDLSV VNG
Sbjct: 1 MASMSFALNRFSGLSSKTTLSADFDPSSRRSFLPPTRVGLKI--SSAAESEPDLSVTVNG 58
Query: 56 LHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANG 115
L MPNPFVIGSGPPGTNYTVMKRAFDEGWG VIAKTVSLDA+KVINVTPRYARLR +NG
Sbjct: 59 LKMPNPFVIGSGPPGTNYTVMKRAFDEGWGAVIAKTVSLDASKVINVTPRYARLRTGSNG 118
Query: 116 SAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVE 175
SAK ++IGWQNIELISDRPLETMLKEF++LK+EYPDRILIAS+MEEYNK AWEELIDRVE
Sbjct: 119 SAKTDVIGWQNIELISDRPLETMLKEFERLKKEYPDRILIASVMEEYNKTAWEELIDRVE 178
Query: 176 QTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDIS 235
QTG+DA+EINFSCPHGMPER+MGAAVGQDCALL+EVCGWINAKATVPVWAKMTPNITDI+
Sbjct: 179 QTGVDALEINFSCPHGMPERRMGAAVGQDCALLDEVCGWINAKATVPVWAKMTPNITDIT 238
Query: 236 QPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVM 295
+PARV+L SGCEG+AAINTIMSVMGI++ TLRPEPCVEGYSTPGGYS KAV PIAL KVM
Sbjct: 239 EPARVSLKSGCEGIAAINTIMSVMGIDMKTLRPEPCVEGYSTPGGYSYKAVRPIALAKVM 298
Query: 296 SIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLE 355
+IAKMMKSEF SE+ SLS IGGVETG DAAEFILLG+NTVQVCTGVMMHGYG VK L E
Sbjct: 299 NIAKMMKSEF-SEDRSLSGIGGVETGYDAAEFILLGSNTVQVCTGVMMHGYGHVKTLCAE 357
Query: 356 LKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFV 415
LKDFMK+HNF++IE+FRG SLQYFTTHTDLV RQ+EA++QRKA K+GL+SDKDWTGDGFV
Sbjct: 358 LKDFMKQHNFSTIEEFRGHSLQYFTTHTDLVKRQKEAVEQRKAEKRGLKSDKDWTGDGFV 417
Query: 416 KESESMVSN 424
KE+ESMVSN
Sbjct: 418 KETESMVSN 426
>UniRef100_Q8S3J6 Putative dihydroorotate dehydrogenase [Oryza sativa]
Length = 201
Score = 341 bits (874), Expect = 3e-92
Identities = 167/199 (83%), Positives = 186/199 (92%), Gaps = 1/199 (0%)
Query: 226 KMTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKA 285
+MTPNITDI++PAR++L SGCEGV+AINTIMSVMGINL TLRPEPCVEGYSTPGGYSA+A
Sbjct: 4 QMTPNITDITRPARISLKSGCEGVSAINTIMSVMGINLKTLRPEPCVEGYSTPGGYSARA 63
Query: 286 VHPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHG 345
VHPIAL KVM IA+MMK EF ++ SLSAIGGVETG DAAEFILLGA+TVQVCTGVMMHG
Sbjct: 64 VHPIALAKVMQIARMMKEEF-ADGQSLSAIGGVETGNDAAEFILLGADTVQVCTGVMMHG 122
Query: 346 YGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQS 405
YGLVKKL EL+DFM++HNF+SIEDFRGASL YFTTHTDLVHRQ+EAI QRKAI+KGL+S
Sbjct: 123 YGLVKKLCAELQDFMRQHNFSSIEDFRGASLPYFTTHTDLVHRQREAINQRKAIRKGLES 182
Query: 406 DKDWTGDGFVKESESMVSN 424
DKDWTGDGFVKE+ESMVSN
Sbjct: 183 DKDWTGDGFVKETESMVSN 201
>UniRef100_Q8G2N6 Dihydroorotate dehydrogenase family protein [Brucella suis]
Length = 436
Score = 280 bits (715), Expect = 7e-74
Identities = 146/334 (43%), Positives = 207/334 (61%), Gaps = 10/334 (2%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DLS G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLSTNFLGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGAEGPPVVNVNDPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLE L+E K++K +PDR LIASIM + A
Sbjct: 63 GAIHG-----ADRRLLGLNNIELITDRPLEINLREMKEVKMRWPDRALIASIMVPCEEEA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ + V W + +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYVGMVVKWCKQYSRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A ++G + V+ INTI S++ ++L+T+ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAFANGTDAVSLINTINSIVSVDLDTMSPVPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIAL V IA+ + ++ +S IGG+ T DAAEFI LG TVQVCT M +G+
Sbjct: 238 KPIALNMVAEIAR----DAETAGLPISGIGGITTWRDAAEFIALGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+++ L D+M + +++DF+G ++ + T
Sbjct: 294 KVVQEMIDGLSDWMDSKGYQTLDDFQGCAVSHVT 327
>UniRef100_Q8YF83 DIHYDROPYRIMIDINE DEHYDROGENASE [Brucella melitensis]
Length = 436
Score = 278 bits (712), Expect = 2e-73
Identities = 145/334 (43%), Positives = 207/334 (61%), Gaps = 10/334 (2%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DLS G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLSTNFLGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGAEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DRPLE L+E K++K +PDR LIASIM + A
Sbjct: 63 GAIHG-----ADRRLLGLNNIELITDRPLEINLREMKEVKMRWPDRALIASIMVPCEEEA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ + V W + +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYVGMVVKWCKQYSRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A ++G + ++ INTI S++ ++L+T+ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAFANGTDALSLINTINSIVSVDLDTMSPVPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIAL V IA+ + ++ +S IGG+ T DAAEFI LG TVQVCT M +G+
Sbjct: 238 KPIALNMVAEIAR----DAETAGLPISGIGGITTWQDAAEFIALGCGTVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+++ L D+M + +++DF+G ++ + T
Sbjct: 294 KVVQEMIDGLSDWMDSKGYQTLDDFQGCAVSHVT 327
>UniRef100_UPI00002B95BA UPI00002B95BA UniRef100 entry
Length = 442
Score = 276 bits (705), Expect = 1e-72
Identities = 149/358 (41%), Positives = 217/358 (59%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + RAF+ GWGGV+ KT+ LD V+NV+ RY
Sbjct: 3 DLRCTIAGITSPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGLDP-HVVNVSSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
++ N I G NIELI+DRPL+ LKE Q+K ++P+R +I S+M N+ W
Sbjct: 62 AVQWNGQ-----RIAGLNNIELITDRPLDVNLKEIAQVKRDWPERAMIVSLMVPCNERDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG DA+E+NF CPHGM ER MGAAVGQ +E V W+ + +P K+
Sbjct: 117 KWILPLVEDTGADAVELNFGCPHGMSERGMGAAVGQVPEYIEMVTRWVKEGSKLPCLVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI+DI +R A G +GV+ INTI S++ ++L+ + P P V+G T GGY AV
Sbjct: 177 TPNISDIRLGSRAAYKGGADGVSLINTINSIVAVDLDAMSPLPMVDGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + ++ N +S IGG+ T DAAEF++LGA +VQVCT M +G+
Sbjct: 237 PIALNMVAEIAR----DVETPNLPISGIGGISTWRDAAEFMVLGAGSVQVCTAAMHYGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKGL 403
+V L+ L ++M + + +++D RG ++ T + +L + + I Q K I+ GL
Sbjct: 293 IVSDLADGLSNWMDEKGYATLDDIRGRAVPNVTDWQYLNLKYDIKARIDQDKCIQCGL 350
>UniRef100_Q92MZ9 PUTATIVE OXIDOREDUCTASE IRON-SULFUR PROTEIN [Rhizobium meliloti]
Length = 437
Score = 273 bits (698), Expect = 7e-72
Identities = 156/360 (43%), Positives = 210/360 (58%), Gaps = 16/360 (4%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP ++RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLRNNFVGIKSPNPFWLASAPPTDKAYNVERAFKAGWGGVVWKTLGEEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G+ NIELI+DR L L+E KQ+K +PDR LIASIM + A
Sbjct: 63 GAI-----WGADRRLLGFNNIELITDRDLYVNLREMKQVKMNWPDRALIASIMVPCEENA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MG+AVGQ +E V W +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGSAVGQVPEYIEMVVRWCKQYTRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A + G + V+ INTI S+ G+NL+T PEP ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAKAGGTDAVSLINTINSITGVNLDTFSPEPSIDGRGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSL--SAIGGVETGGDAAEFILLGANTVQVCTGVMMH 344
PIAL V IA+ D E Y L S IGGV T DAAEF+ LGA VQVCT M +
Sbjct: 238 KPIALNMVAEIAR------DPETYGLPISGIGGVTTWRDAAEFMALGAGNVQVCTAAMTY 291
Query: 345 GYGLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
G+ +V+++ L D+M +++D G ++ T + +L + + I Q IK G
Sbjct: 292 GFKIVQEMITGLSDWMDAKGHRTLDDISGRAVPNVTDWQYLNLNYIAKAKIDQDACIKCG 351
>UniRef100_Q98K45 Probable oxidoreductase [Rhizobium loti]
Length = 437
Score = 266 bits (680), Expect = 8e-70
Identities = 149/358 (41%), Positives = 212/358 (58%), Gaps = 12/358 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP + RAF GWGGV+ KT+ + V+NV PRY
Sbjct: 3 DLRNNFVGIKSPNPFWLASAPPTDKAYNVIRAFKAGWGGVVWKTLGEEGPPVVNVNGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
+ A ++G NIELI+DR L+T L+E KQ+K ++PDR LIASIM +A+
Sbjct: 63 GAI-----WGADRRLLGLNNIELITDRDLQTNLREMKQVKMDWPDRALIASIMVPCEEAS 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ VE+TG D IE+NF CPHGM ER MGAAVGQ +E V W +PV K
Sbjct: 118 WKAILPLVEETGADGIELNFGCPHGMSERGMGAAVGQVPEYIEMVVRWCKQYTRMPVITK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPNITDI +PAR A + G + V+ INTI S+ G++L++ P P ++G + GGY AV
Sbjct: 178 LTPNITDIRKPARAAHAGGTDAVSLINTINSITGVDLDSFAPMPTIDGKGSHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGY 346
PIA+ V IA+ + ++ +S IGG+ T DAAEF+ LGA VQVCT M +G+
Sbjct: 238 KPIAMNMVAEIAR----DPETRGLPISGIGGITTWRDAAEFLALGAGNVQVCTAAMTYGF 293
Query: 347 GLVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT--THTDLVHRQQEAIKQRKAIKKG 402
+V+++ L+++M + S++D G + T + +L + + I Q IK G
Sbjct: 294 KIVQEMIAGLENWMDEKGHRSLDDIIGRATPNVTDWQYLNLNYVAKAHIDQDACIKCG 351
>UniRef100_Q88FQ0 Dihydroorotate dehydrogenase family protein [Pseudomonas putida]
Length = 424
Score = 260 bits (664), Expect = 6e-68
Identities = 147/352 (41%), Positives = 204/352 (57%), Gaps = 17/352 (4%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS+ G+ PNPF + S PP + RAF+ GWGGV+ KT+ D A V NV+ RY+
Sbjct: 3 DLSIVFAGIKAPNPFWLASAPPTDKAYNVVRAFEAGWGGVVWKTLGEDPAAV-NVSSRYS 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
A+ ++ G NIELI+DR LE L+E Q+K+++PDR LI S+M + +W
Sbjct: 62 -----AHYGPNRQVQGINNIELITDRSLEINLREITQVKKDWPDRALIVSLMVPCIEESW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG D IE+NF CPHGMPER MGAAVGQ +E V W +PV K+
Sbjct: 117 KFILPLVEATGADGIELNFGCPHGMPERGMGAAVGQVPEYVEMVTRWCKTYCALPVIVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNITDI Q AR A G + V+ INTI S+ ++L+ + P V ST GGY AV
Sbjct: 177 TPNITDIRQSARAAHRGGADAVSLINTINSITSVDLDRMVAHPIVGDQSTHGGYCGSAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + ++ + IGG+ DAAEF+ LG+ VQVCT M+HG+
Sbjct: 237 PIALNMVAEIAR----DPETRGLPICGIGGIGNWRDAAEFMALGSGAVQVCTAAMLHGFR 292
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT-------THTDLVHRQQEA 392
+V+ + L +M +H ++E FRG ++ + T + + H QEA
Sbjct: 293 IVEDMQDGLARWMDQHGHATVEAFRGQAVGHTTDWKYLDINYKSVAHIDQEA 344
>UniRef100_Q846U6 Dihydropyrimidine dehydrogenase [Brevibacillus agri]
Length = 425
Score = 258 bits (658), Expect = 3e-67
Identities = 139/340 (40%), Positives = 200/340 (57%), Gaps = 16/340 (4%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS+ + G+ PNPF + S PP + ++RAF+ GWGG + KT+ +INVT R+A
Sbjct: 3 DLSINLAGITSPNPFWLASAPPTNSGYQVQRAFEAGWGGAVWKTLG---EPIINVTSRFA 59
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L + G+ NIELI+D+PLE LKE + K+ +P LIAS+M E+ + AW
Sbjct: 60 GLTFGGQ-----RVFGFNNIELITDKPLEVNLKEMDETKKRFPKHTLIASLMVEHKQEAW 114
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
E++ +VE G+D +E+NF CPHGM ER MG+AVGQ L+ + W+ A PV K+
Sbjct: 115 HEIVKKVEAIGVDGLELNFGCPHGMAERGMGSAVGQHPDLIRQQVEWVKEVAQTPVIVKL 174
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNITDI AR A + G + ++ INTI S+MG++L+ P P V+G GGY AV
Sbjct: 175 TPNITDIRFTARAASAGGADAISMINTINSLMGVDLDKWLPIPHVDGKGAHGGYCGPAVK 234
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V A+ + F +S IGG+ DA EF+L+GA VQVCT M HG+
Sbjct: 235 PIALNMV---AECARDPF--VGVPISGIGGISNWRDAVEFLLMGATGVQVCTAAMHHGFR 289
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDLVH 387
+V+ + L +++ + S+ D G ++ T++D H
Sbjct: 290 IVEDMIDGLNNYLDQRKIASVMDIVGKAVH---TYSDWGH 326
>UniRef100_Q9I678 Probable oxidoreductase [Pseudomonas aeruginosa]
Length = 425
Score = 256 bits (655), Expect = 7e-67
Identities = 141/329 (42%), Positives = 194/329 (58%), Gaps = 10/329 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS++ G+ PNPF + S PP + RA+ GWGGV+ KT+ D A V NV+ RY+
Sbjct: 4 DLSIEFAGIKSPNPFWLASAPPTDKAYNVVRAYQAGWGGVVWKTLGEDPAAV-NVSSRYS 62
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
A+ ++ G NIELI+DR LE L+E Q+K+++PDR LI S+M + +W
Sbjct: 63 -----AHFGPNRQVQGINNIELITDRSLEINLREIAQVKKDWPDRALIVSLMVPCVEESW 117
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+ ++ VE TG D IE+NF CPHGMPER MGAAVGQ +E V W A ++PV K+
Sbjct: 118 KYILPLVEATGADGIELNFGCPHGMPERGMGAAVGQVPEYVEMVTRWCKAHCSLPVIVKL 177
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNITDI Q AR A G + V+ INTI S+ ++L + P V ST GGY AV
Sbjct: 178 TPNITDIRQSARAAHRGGADAVSLINTINSITSVDLERMVALPVVGTQSTHGGYCGSAVK 237
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL V IA+ + + + IGG+ + DAAEF+ LG VQVCT M+HG+
Sbjct: 238 PIALNMVAEIAR----DPQTRGLPICGIGGIGSWRDAAEFVALGCGAVQVCTAAMLHGFR 293
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASL 376
+V+ + L +M H + + DF G ++
Sbjct: 294 IVEDMQDGLARWMDSHGYQRLSDFSGGAV 322
>UniRef100_Q5WBE7 Dihydropyrimidine dehydrogenase [Bacillus clausii]
Length = 419
Score = 252 bits (644), Expect = 1e-65
Identities = 131/333 (39%), Positives = 197/333 (58%), Gaps = 13/333 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DL + G+ PNPF + S PP + ++RAF+ GWGG + KT+ ++N T R+A
Sbjct: 3 DLISNLAGITSPNPFWLASAPPTNSGYQVQRAFEAGWGGAVWKTLG---EPIMNSTSRFA 59
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L N + G+ NIELISDRPL+ L+E K+ +P+ ++AS+M E + W
Sbjct: 60 ALHFNGK-----RVAGFNNIELISDRPLDVNLREIADTKKRFPNHAIVASLMVEPKQEKW 114
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
E++ RVE G+D +E+NF CPHGM ER MG+A GQ AL+E+ W+ A PV K+
Sbjct: 115 HEIVKRVEAVGVDGLELNFGCPHGMAERGMGSASGQVPALVEQQTVWVKEVARTPVIVKL 174
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNITDI+ A A G + ++ INTI S+ G++++T + P V+G T GGY AV
Sbjct: 175 TPNITDITATAEAAAQGGADAISLINTINSLAGVDIDTWQTIPNVDGLGTHGGYCGPAVK 234
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL + A+ + N +S IGG+ T DA +F+L+GA+ VQVCT M HG+
Sbjct: 235 PIALHMLGDCAR-----HPNVNIPISGIGGIATWQDAVQFLLMGASNVQVCTAAMHHGFR 289
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFT 380
+V+ L L ++++ S+++ G S+ ++
Sbjct: 290 IVEDLLDGLSAYLEEKGLASVQELVGQSVHRYS 322
>UniRef100_Q8X643 Hypothetical protein yeiA [Escherichia coli]
Length = 413
Score = 252 bits (643), Expect = 2e-65
Identities = 141/338 (41%), Positives = 201/338 (58%), Gaps = 13/338 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS+ G+ PNPF + S P G Y + +A+D GWGGV+ KT+ A V+PR+
Sbjct: 7 DLSITFCGVKFPNPFCLSSSPVGNCYEMCAKAYDTGWGGVVFKTIGFFIAN--EVSPRFD 64
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L G IG++N+E I++ PLE L ++LKE+YPD++LIASIM E N+ W
Sbjct: 65 HLVKEDTG-----FIGFKNMEQIAEHPLEENLAALRRLKEDYPDKVLIASIMGE-NEQQW 118
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
EEL V++ G D IE NFSCP M MG+ VGQ L+E+ C + +T+P+ AKM
Sbjct: 119 EELARLVQEAGADMIECNFSCPQ-MTSHAMGSDVGQSPELVEKYCRAVKRGSTLPMLAKM 177
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI D+ + A A G +G+AAINT+ S+ I+LN P V G S+ GYS KAV
Sbjct: 178 TPNIGDMCEVALAAKRGGADGIAAINTVKSITNIDLNQKIGMPIVNGKSSISGYSGKAVK 237
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL + M++ + ++ +S IGG+ET DAAEF+LLGA T+QV TG+M +GY
Sbjct: 238 PIAL----RFIQQMRTHPELRDFPISGIGGIETWEDAAEFLLLGAATLQVTTGIMQYGYR 293
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDL 385
+V+ ++ L ++ F S+++ G + DL
Sbjct: 294 IVEDMASGLSHYLADQGFDSLQEMVGLANNNIVPAEDL 331
>UniRef100_Q8FFV0 Hypothetical protein yeiA [Escherichia coli O6]
Length = 413
Score = 252 bits (643), Expect = 2e-65
Identities = 141/338 (41%), Positives = 201/338 (58%), Gaps = 13/338 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS+ G+ PNPF + S P G Y + +A+D GWGGV+ KT+ A V+PR+
Sbjct: 7 DLSITFCGVKFPNPFCLSSSPVGNCYEMCAKAYDTGWGGVVFKTIGFFIAN--EVSPRFD 64
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L G IG++N+E I++ PLE L ++LKE+YPD++LIASIM E N+ W
Sbjct: 65 HLVKEDTG-----FIGFKNMEQIAEHPLEENLAALRRLKEDYPDKVLIASIMGE-NEQQW 118
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
EEL V++ G D IE NFSCP M MG+ VGQ L+E+ C + +T+P+ AKM
Sbjct: 119 EELARLVQEAGADMIECNFSCPQ-MTSHAMGSDVGQSPELVEKYCRAVKRGSTLPMLAKM 177
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI D+ + A A G +G+AAINT+ S+ I+LN P V G S+ GYS KAV
Sbjct: 178 TPNIGDMCEVALAAKRGGADGIAAINTVKSITNIDLNQKIGMPIVNGKSSISGYSGKAVK 237
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL + M++ + ++ +S IGG+ET DAAEF+LLGA T+QV TG+M +GY
Sbjct: 238 PIAL----RFIQQMRTHPELRDFPISGIGGIETWEDAAEFLLLGAATLQVTTGIMQYGYR 293
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDL 385
+V+ ++ L ++ F S+++ G + DL
Sbjct: 294 IVEDMASGLSHYLADQGFDSLQEMVGLANHNIVPAEDL 331
>UniRef100_UPI00002FC63E UPI00002FC63E UniRef100 entry
Length = 273
Score = 248 bits (634), Expect = 2e-64
Identities = 129/281 (45%), Positives = 178/281 (62%), Gaps = 10/281 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLS + G+ PNPF + S PP Y + RAF GWGGV+ KT+ +D ++NVT RY
Sbjct: 3 DLSCTIAGIPSPNPFWVASAPPSDKYYNVDRAFRAGWGGVVWKTLGIDPP-IVNVTSRYG 61
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
+ N NG +++G N ELI+DRPL+ L E KQ+KEE+PDR ++ S+M N+ W
Sbjct: 62 SM--NLNGQ---KMMGLNNNELITDRPLDVNLAEIKQIKEEWPDRAVVVSLMVPCNQNDW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
E ++ +VE TG D +E+NF CPHGM ER MGAAVGQ +E V W+ + +P K+
Sbjct: 117 EYVLRKVEDTGADGVELNFGCPHGMSERGMGAAVGQVPDYVEMVTRWVKENSDLPCIVKL 176
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNITDI PAR A + G +GV+ INTI S++ ++L+++ PEP +EG T GGY AV
Sbjct: 177 TPNITDIRYPARAAKAGGADGVSLINTISSIISVDLDSMSPEPSIEGKGTHGGYCGPAVK 236
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFI 328
PIAL IA+ + ++ +SAIGG+ DAAEF+
Sbjct: 237 PIALNMTSEIAR----DPETLGLPISAIGGISNWKDAAEFM 273
>UniRef100_UPI00002C550A UPI00002C550A UniRef100 entry
Length = 285
Score = 247 bits (630), Expect = 5e-64
Identities = 128/292 (43%), Positives = 177/292 (59%), Gaps = 10/292 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVT-PRY 106
DL G+ PNPF + S PP + RAF+ GWGGV+ KT+ + ++NV+ PRY
Sbjct: 3 DLKSNFVGIESPNPFWLASAPPTDKAYNVNRAFEAGWGGVVWKTLGEEGPPIVNVSGPRY 62
Query: 107 ARLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAA 166
L +IG+ NIELI+DRPL+ L+E Q+K+++PDR L+ S+M +AA
Sbjct: 63 EALHGKDRS-----VIGFNNIELITDRPLQLNLQEIAQVKKDWPDRALVVSLMVPCQEAA 117
Query: 167 WEELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAK 226
W+ ++ +VE TG D +E+NF CPHGM ER MGAAVGQ +E V W + +PV K
Sbjct: 118 WKSILSQVEDTGADGLELNFGCPHGMSERGMGAAVGQVPDYVEMVTAWCKEYSRLPVIVK 177
Query: 227 MTPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAV 286
+TPN+T I QPA+ A G + V+ INTI SVM ++L+ LR P ++ T GGY AV
Sbjct: 178 LTPNVTSIRQPAQAAKKGGADAVSLINTINSVMSVDLDALRINPTIDAMGTHGGYCGPAV 237
Query: 287 HPIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVC 338
PIAL V + + + + E +SAIGG+ DAAEF+ +GA VQVC
Sbjct: 238 KPIALHMVSDLGR----DPECEGLPISAIGGISNWRDAAEFLFVGAGNVQVC 285
>UniRef100_Q8ZNL7 Putative dihydropyrimidine dehydrogenase [Salmonella typhimurium]
Length = 411
Score = 246 bits (629), Expect = 7e-64
Identities = 135/326 (41%), Positives = 196/326 (59%), Gaps = 13/326 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLSV G+ PNPF + S P G Y + +A+D GWGG++ KT+ A V+PR+
Sbjct: 5 DLSVTFCGVKFPNPFCLSSSPVGNCYEMCAKAYDTGWGGIVFKTIGFFIAN--EVSPRFD 62
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L G IG++N+E I++ PLE L ++LK++YPD++LIASIM E N+ W
Sbjct: 63 HLTKEDTG-----FIGFKNMEQIAEHPLEENLAAIRRLKQDYPDKVLIASIMGE-NEQQW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+EL VE+ G D IE NFSCP M MG+ VGQ L+E+ C + +++P+ AKM
Sbjct: 117 QELARLVEEAGADMIECNFSCPQ-MTSHAMGSDVGQSPELVEKYCRAVKRGSSLPMLAKM 175
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI D+ + A A G +G+A INT+ S+ I+LN P V G S+ GYS KAV
Sbjct: 176 TPNIGDMCEVALAAKRGGADGIATINTVKSITNIDLNRKIGMPVVNGKSSISGYSGKAVK 235
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL + ++ + ++ +S IGG+ET DAAEF+LLGA T+QV TG+M +GY
Sbjct: 236 PIAL----RFIQQLRMHPELRDFPISGIGGIETWEDAAEFLLLGAATLQVTTGIMQYGYR 291
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRG 373
+V+ ++ L ++ F S+++ G
Sbjct: 292 IVEDMASGLSHYLADQGFASLQEMIG 317
>UniRef100_Q5PE62 Putative dihydropyrimidine dehydrogenase [Salmonella enterica
subsp. enterica serovar Paratypi A str. ATCC 9150]
Length = 411
Score = 246 bits (628), Expect = 9e-64
Identities = 135/326 (41%), Positives = 196/326 (59%), Gaps = 13/326 (3%)
Query: 48 DLSVKVNGLHMPNPFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYA 107
DLSV G+ PNPF + S P G Y + +A+D GWGG++ KT+ A V+PR+
Sbjct: 5 DLSVTFCGVKFPNPFCLSSSPVGNCYEMCAKAYDTGWGGIVFKTIGFFIAN--EVSPRFD 62
Query: 108 RLRANANGSAKGEIIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAW 167
L G IG++N+E I++ PLE L ++LK++YPD++LIASIM E N+ W
Sbjct: 63 HLTKEDTG-----FIGFKNMEQIAEHPLEENLAAIRRLKQDYPDKVLIASIMGE-NEQQW 116
Query: 168 EELIDRVEQTGIDAIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKM 227
+EL VE+ G D IE NFSCP M MG+ VGQ L+E+ C + +++P+ AKM
Sbjct: 117 QELARLVEEAGADMIECNFSCPQ-MTSHAMGSDVGQSPELVEKYCRAVKRGSSLPMLAKM 175
Query: 228 TPNITDISQPARVALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVH 287
TPNI D+ + A A G +G+A INT+ S+ I+LN P V G S+ GYS KAV
Sbjct: 176 TPNIGDMCEVALAAKRGGADGIATINTVKSITNIDLNRKIGMPVVNGKSSISGYSRKAVK 235
Query: 288 PIALGKVMSIAKMMKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYG 347
PIAL + ++ + ++ +S IGG+ET DAAEF+LLGA T+QV TG+M +GY
Sbjct: 236 PIAL----RFIQQLRMHPELRDFPISGIGGIETWEDAAEFLLLGAATLQVTTGIMQYGYR 291
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRG 373
+V+ ++ L ++ F S+++ G
Sbjct: 292 IVEDMASGLSHYLADQGFASLQEMIG 317
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 691,412,506
Number of Sequences: 2790947
Number of extensions: 28276909
Number of successful extensions: 83718
Number of sequences better than 10.0: 686
Number of HSP's better than 10.0 without gapping: 493
Number of HSP's successfully gapped in prelim test: 193
Number of HSP's that attempted gapping in prelim test: 82724
Number of HSP's gapped (non-prelim): 728
length of query: 424
length of database: 848,049,833
effective HSP length: 130
effective length of query: 294
effective length of database: 485,226,723
effective search space: 142656656562
effective search space used: 142656656562
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC139706.5


