

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
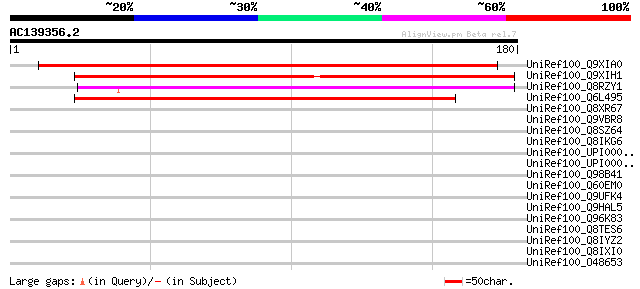
Sequences producing significant alignments: (bits) Value
UniRef100_Q9XIA0 F13F21.24 protein [Arabidopsis thaliana] 172 4e-42
UniRef100_Q9XIH1 Hypothetical protein At2g16190 [Arabidopsis tha... 163 2e-39
UniRef100_Q8RZY1 P0034C09.31 protein [Oryza sativa] 152 5e-36
UniRef100_Q6L495 Hypothetical protein P0554F08.11 [Oryza sativa] 142 4e-33
UniRef100_Q8XR67 PROBABLE EXCINUCLEASE ABC SUBUNIT A (DNA REPAIR... 35 0.68
UniRef100_Q9VBR8 CG11902-PA [Drosophila melanogaster] 34 2.0
UniRef100_Q8SZ64 RE15630p [Drosophila melanogaster] 34 2.0
UniRef100_Q8IKG6 DNA-3-methyladenine glycosylase, putative [Plas... 33 2.6
UniRef100_UPI00002F0EF3 UPI00002F0EF3 UniRef100 entry 32 5.8
UniRef100_UPI0000266B44 UPI0000266B44 UniRef100 entry 32 9.9
UniRef100_Q98B41 Ribose ABC transporter [Rhizobium loti] 32 9.9
UniRef100_Q60EM0 Unknow protein [Oryza sativa] 32 9.9
UniRef100_Q9UFK4 Hypothetical protein DKFZp564D0764 [Homo sapiens] 32 9.9
UniRef100_Q9HAL5 Hypothetical protein FLJ11390 [Homo sapiens] 32 9.9
UniRef100_Q96K83 Hypothetical protein FLJ14448 [Homo sapiens] 32 9.9
UniRef100_Q8TES6 FLJ00107 protein [Homo sapiens] 32 9.9
UniRef100_Q8IYZ2 EHZF protein [Homo sapiens] 32 9.9
UniRef100_Q8IXI0 Early hematopoietic zinc finger [Homo sapiens] 32 9.9
UniRef100_O48653 DNA polymerase alpha catalytic subunit [Oryza s... 32 9.9
>UniRef100_Q9XIA0 F13F21.24 protein [Arabidopsis thaliana]
Length = 331
Score = 172 bits (436), Expect = 4e-42
Identities = 83/163 (50%), Positives = 110/163 (66%)
Query: 11 RRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRT 70
R + +S I PF WATN R +I +L +L N+I +ITG+VQC+ C+ +Q+ ++LR
Sbjct: 157 RSTVSKKSDTISPPFPWATNRRGEIQSLEYLESNQITTITGEVQCRHCEKVYQVSYNLRE 216
Query: 71 KFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLG 130
+F + ++ +T M DRA K P RC C +E +VKPVIAE+K INWLFLLLG
Sbjct: 217 RFAEVVKFYLTEKRKMRDRAHKDWAYPEQRRCELCGREKAVKPVIAERKSQINWLFLLLG 276
Query: 131 QMLGCCKLEQLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLDP 173
Q LG C LEQLK FCK++ +HRTGAK+RVLYLTY LC+ L P
Sbjct: 277 QTLGFCTLEQLKNFCKHSKNHRTGAKDRVLYLTYMGLCKMLQP 319
>UniRef100_Q9XIH1 Hypothetical protein At2g16190 [Arabidopsis thaliana]
Length = 303
Score = 163 bits (413), Expect = 2e-39
Identities = 76/156 (48%), Positives = 101/156 (64%), Gaps = 2/156 (1%)
Query: 24 PFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNM 83
P+ WAT KI + L N I I+G V CK+C + ++L KF + Y+ N
Sbjct: 150 PYPWATKKPGKIQSFRDLSSNNINVISGQVHCKTCDRTDTVEYNLEEKFSELYGYIKVNK 209
Query: 84 NTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 143
M RAP P+L C C E +KPV++E+K+ INWLFLLLGQMLGCC L+QL+Y
Sbjct: 210 EEMRHRAPGSWSTPKLIPCRTCKSE--MKPVMSERKEEINWLFLLLGQMLGCCTLDQLRY 267
Query: 144 FCKYNNHHRTGAKNRVLYLTYFELCQQLDPSGPFNV 179
FC+ N+ HRTG+K+RV+Y+TY LC+QLDP GPFN+
Sbjct: 268 FCQLNSKHRTGSKDRVVYITYLSLCKQLDPEGPFNL 303
>UniRef100_Q8RZY1 P0034C09.31 protein [Oryza sativa]
Length = 316
Score = 152 bits (383), Expect = 5e-36
Identities = 72/157 (45%), Positives = 95/157 (59%), Gaps = 2/157 (1%)
Query: 25 FIWATNHRAKIHN--LNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTN 82
F W T+ + + L +L I S+ G C C + + +DL KF + Y+ N
Sbjct: 160 FPWVTSADLPVLHCTLESMLLKGITSVEGKATCNRCSAEVPIAYDLDAKFREVRDYVAAN 219
Query: 83 MNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLK 142
++ M DRAP+ M PRLP C C ++ + P I +K+ INWLFL LGQMLGCC LE LK
Sbjct: 220 IHIMDDRAPEHWMCPRLPDCGSCGKKACMWPQIPNEKREINWLFLFLGQMLGCCTLEGLK 279
Query: 143 YFCKYNNHHRTGAKNRVLYLTYFELCQQLDPSGPFNV 179
+FCK +H TGAK RVLY Y E+C+QLDP GPFN+
Sbjct: 280 FFCKNTKNHCTGAKTRVLYYAYIEMCRQLDPQGPFNI 316
>UniRef100_Q6L495 Hypothetical protein P0554F08.11 [Oryza sativa]
Length = 315
Score = 142 bits (358), Expect = 4e-33
Identities = 65/135 (48%), Positives = 88/135 (65%)
Query: 24 PFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNM 83
P+ WATN AK H+L L + I +I G+ +C+ C T+ + +++ TKF +S Y N
Sbjct: 172 PYPWATNEVAKHHSLVELARRDIININGEARCRRCDTRKMIVYNIATKFREVSDYFRQNY 231
Query: 84 NTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKY 143
M+DRA MNP +P C C E ++PVIA +K+ INWLFLLLG+ LG C L+QLKY
Sbjct: 232 QHMNDRAQARWMNPVVPNCDSCGHERCMRPVIAAEKERINWLFLLLGETLGLCTLDQLKY 291
Query: 144 FCKYNNHHRTGAKNR 158
FC + N HRTGAK+R
Sbjct: 292 FCAHTNRHRTGAKDR 306
>UniRef100_Q8XR67 PROBABLE EXCINUCLEASE ABC SUBUNIT A (DNA REPAIR ATP-BINDING) ABC
TRANSPORTER PROTEIN [Ralstonia solanacearum]
Length = 1945
Score = 35.4 bits (80), Expect = 0.68
Identities = 19/51 (37%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 5 HTHVPPRRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQN----KIFSITG 51
H P R +APRSG P W T H A +HNL +L ++ ++TG
Sbjct: 1545 HPLQPRREVAAPRSGAAASPERWLTVHGASLHNLRNLTATLPLARLVAVTG 1595
>UniRef100_Q9VBR8 CG11902-PA [Drosophila melanogaster]
Length = 1266
Score = 33.9 bits (76), Expect = 2.0
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 22 PIPFIWATN---HRAKIHNLNHL-LQNKIFSITGDVQCKSCQTKFQMPFDL 68
P+ F W N H H L+ L +Q+ + + GD+QC C F+M DL
Sbjct: 770 PLRFRWRENLKLHMEVFHMLDELSIQDDLPKVDGDLQCGKCNRSFRMQKDL 820
>UniRef100_Q8SZ64 RE15630p [Drosophila melanogaster]
Length = 1426
Score = 33.9 bits (76), Expect = 2.0
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 4/51 (7%)
Query: 22 PIPFIWATN---HRAKIHNLNHL-LQNKIFSITGDVQCKSCQTKFQMPFDL 68
P+ F W N H H L+ L +Q+ + + GD+QC C F+M DL
Sbjct: 930 PLRFRWRENLKLHMEVFHMLDELSIQDDLPKVDGDLQCGKCNRSFRMQKDL 980
>UniRef100_Q8IKG6 DNA-3-methyladenine glycosylase, putative [Plasmodium falciparum]
Length = 501
Score = 33.5 bits (75), Expect = 2.6
Identities = 25/88 (28%), Positives = 42/88 (47%), Gaps = 6/88 (6%)
Query: 35 IHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKIL 94
IHN +++ N I +I + +S + + + + L KF+ ISQYL++N +H +
Sbjct: 172 IHNCLNIVTN-IQNIPDAILVRSTEPLYNIEYFLSNKFEKISQYLLSNSIFIHKNIKEQQ 230
Query: 95 MNPRLPRCAHCNQENSVKPVIAEKKKNI 122
N P+ Q N I+ K NI
Sbjct: 231 KNNSTPKNKRNQQGN-----ISNKDGNI 253
>UniRef100_UPI00002F0EF3 UPI00002F0EF3 UniRef100 entry
Length = 221
Score = 32.3 bits (72), Expect = 5.8
Identities = 19/46 (41%), Positives = 25/46 (54%)
Query: 104 HCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYFCKYNN 149
+ N ENSVK + K KN+N +LL Q C K ++L K NN
Sbjct: 124 YLNSENSVKYKVINKIKNLNRDNVLLTQTPQCFKFKELFNLYKTNN 169
>UniRef100_UPI0000266B44 UPI0000266B44 UniRef100 entry
Length = 118
Score = 31.6 bits (70), Expect = 9.9
Identities = 22/67 (32%), Positives = 30/67 (43%), Gaps = 5/67 (7%)
Query: 18 SGPIPIPFIWATN--HRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVI 75
+ PI F W HR K+ N N L++ F+ + + C K M DL D
Sbjct: 40 TNPIASIFAWTRGLAHRGKLDNNNELIK---FASSLEKVCVETVEKGSMTKDLAILIDKS 96
Query: 76 SQYLVTN 82
S+YL TN
Sbjct: 97 SKYLTTN 103
>UniRef100_Q98B41 Ribose ABC transporter [Rhizobium loti]
Length = 380
Score = 31.6 bits (70), Expect = 9.9
Identities = 16/60 (26%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Query: 66 FDLRTKFDVISQYLVTNMNTMHDRAPKI----LMNPRLPRCAHCNQENSVKPVIAEKKKN 121
F +R + + L TN+ +M + +P I ++ P +P C++ +KP++A K N
Sbjct: 8 FAMRQLLPIAALLLATNVASMAEGSPGITVPTVLQPFVPTAPSCSKPTDLKPILAFAKDN 67
>UniRef100_Q60EM0 Unknow protein [Oryza sativa]
Length = 874
Score = 31.6 bits (70), Expect = 9.9
Identities = 22/85 (25%), Positives = 35/85 (40%), Gaps = 4/85 (4%)
Query: 89 RAPKILMNPRLPRCAHCNQENSVKPVI--AEKKKNINWLFLLLGQMLGCCKLEQLKYFCK 146
R+ + N + A C K V E + ++W L+ G L C+LE L+ F
Sbjct: 233 RSSVFVCNSLMNMYAKCGLVEDAKSVFNWMETRDMVSWNTLMAGLQLNECELEALQLF-- 290
Query: 147 YNNHHRTGAKNRVLYLTYFELCQQL 171
+ + G + Y T +LC L
Sbjct: 291 HESRATMGKMTQSTYATVIKLCANL 315
>UniRef100_Q9UFK4 Hypothetical protein DKFZp564D0764 [Homo sapiens]
Length = 583
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 411 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 456
>UniRef100_Q9HAL5 Hypothetical protein FLJ11390 [Homo sapiens]
Length = 693
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 521 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 566
>UniRef100_Q96K83 Hypothetical protein FLJ14448 [Homo sapiens]
Length = 1311
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 1139 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 1184
>UniRef100_Q8TES6 FLJ00107 protein [Homo sapiens]
Length = 1234
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 1062 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 1107
>UniRef100_Q8IYZ2 EHZF protein [Homo sapiens]
Length = 1346
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 1174 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 1219
>UniRef100_Q8IXI0 Early hematopoietic zinc finger [Homo sapiens]
Length = 1311
Score = 31.6 bits (70), Expect = 9.9
Identities = 13/46 (28%), Positives = 24/46 (51%)
Query: 54 QCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRL 99
+C SC KF+ +L+ I + LV + N+ + P++ PR+
Sbjct: 1139 RCSSCNVKFESESELQNHIQTIHRELVPDSNSTQLKTPQVSPMPRI 1184
>UniRef100_O48653 DNA polymerase alpha catalytic subunit [Oryza sativa]
Length = 1243
Score = 31.6 bits (70), Expect = 9.9
Identities = 27/100 (27%), Positives = 40/100 (40%), Gaps = 7/100 (7%)
Query: 53 VQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVK 112
+ C SC T F P + + S V+N N +D + R PRC E+ V
Sbjct: 1047 LSCPSCSTTFDCP-PVSSLIIGSSSGNVSNPNEGNDASINFWRRMRCPRCPDDTDESRVS 1105
Query: 113 PVIA--EKKKNINWLFLLLGQMLGCCKLEQLKYFCKYNNH 150
P + + K+ + L + L C E CKY+ H
Sbjct: 1106 PAVLANQMKRQADSFINLYYKGLLMCDDEG----CKYSTH 1141
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 314,970,820
Number of Sequences: 2790947
Number of extensions: 12131579
Number of successful extensions: 32484
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 32471
Number of HSP's gapped (non-prelim): 19
length of query: 180
length of database: 848,049,833
effective HSP length: 119
effective length of query: 61
effective length of database: 515,927,140
effective search space: 31471555540
effective search space used: 31471555540
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 70 (31.6 bits)
Medicago: description of AC139356.2


