

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
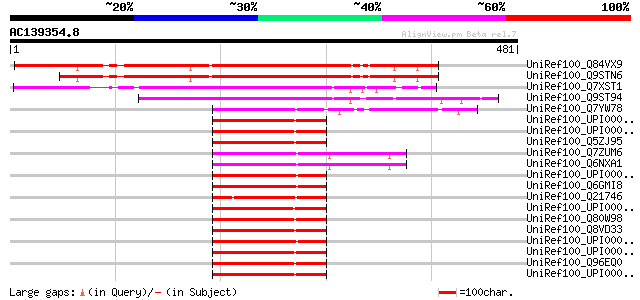
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84VX9 At4g08320 [Arabidopsis thaliana] 334 3e-90
UniRef100_Q9STN6 Hypothetical protein T28D5.10 [Arabidopsis thal... 275 2e-72
UniRef100_Q7XST1 OSJNBa0039K24.25 protein [Oryza sativa] 228 3e-58
UniRef100_Q9ST94 CAA30373.1 protein [Oryza sativa] 184 4e-45
UniRef100_Q7YW78 Small glutamine-rich tetratricopeptide [Schisto... 109 2e-22
UniRef100_UPI00003AFD68 UPI00003AFD68 UniRef100 entry 109 2e-22
UniRef100_UPI00003AFD67 UPI00003AFD67 UniRef100 entry 109 2e-22
UniRef100_Q5ZJ95 Hypothetical protein [Gallus gallus] 109 2e-22
UniRef100_Q7ZUM6 Small glutamine-rich tetratricopeptide repeat (... 108 4e-22
UniRef100_Q6NXA1 Small glutamine-rich tetratricopeptide repeat (... 108 4e-22
UniRef100_UPI0000360266 UPI0000360266 UniRef100 entry 105 2e-21
UniRef100_Q6GMI8 Zgc:92462 [Brachydanio rerio] 105 2e-21
UniRef100_Q21746 Hypothetical protein R05F9.10 [Caenorhabditis e... 105 4e-21
UniRef100_UPI0000432F49 UPI0000432F49 UniRef100 entry 104 6e-21
UniRef100_Q80W98 Small glutamine rich protein with tetratricopep... 104 6e-21
UniRef100_Q8VD33 Small glutamine-rich tetratricopeptide repeat-c... 104 6e-21
UniRef100_UPI00002DA2F7 UPI00002DA2F7 UniRef100 entry 103 1e-20
UniRef100_UPI000036D10D UPI000036D10D UniRef100 entry 103 1e-20
UniRef100_Q96EQ0 Small glutamine-rich tetratricopeptide repeat-c... 103 1e-20
UniRef100_UPI00003AFB3D UPI00003AFB3D UniRef100 entry 102 2e-20
>UniRef100_Q84VX9 At4g08320 [Arabidopsis thaliana]
Length = 427
Score = 334 bits (857), Expect = 3e-90
Identities = 201/418 (48%), Positives = 268/418 (64%), Gaps = 30/418 (7%)
Query: 5 RITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPDS- 63
++TTDSPL +IVRSFL+FL+ VE +PGVD EG+EVARECL EAFK+N+ +S + DS
Sbjct: 3 KLTTDSPLCRKIVRSFLNFLDSVEVAPGVDEEGLEVARECLREAFKLNSDSSRDDDDDSF 62
Query: 64 ----LIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHT 119
L+++F S D N+ + A++ Q+ + + +H + D + P
Sbjct: 63 KPISLVNLFTSLDNNETIPPPPPPPVAATT----QDPSSSGSH------VSVDTCKEPSF 112
Query: 120 SAVSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRR--WKDLGVS 177
+ S+DEL GQ F LEK YFR DG DD QLEKA+R+F D + + V+
Sbjct: 113 TGTSRDELFGQLFGALEKVRYFRNTPDGDDDPAQLEKATRIFHDCVNEMEKSGCQTFDVN 172
Query: 178 SLI*RIWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQ 237
SL + K AMQS Y +A+ELY+ AIA+ +K+AV+YCNRAAAYTQIN +EAI+
Sbjct: 173 SLAETLKCQGNK--AMQSNLYLEAVELYSFAIALTDKNAVFYCNRAAAYTQINMCSEAIK 230
Query: 238 DSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHK 297
D L+SIEIDPNYSKAYSRLGLAYYAQG Y +AI+KGFKKAL LDP+NESVKENIRVAE K
Sbjct: 231 DCLKSIEIDPNYSKAYSRLGLAYYAQGKYAEAIEKGFKKALLLDPHNESVKENIRVAEQK 290
Query: 298 LMEERHRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFM-AAANGGQG 356
+ EE+ R +Q + S + G P+ F SMP NP +L SMFM A N G
Sbjct: 291 IREEQQRQRRSQQNTSTFYTQEPEMGG-VQGIPSQF-SMPVNP-DLMSMFMNMAGNTFPG 347
Query: 357 SHSQEGQ----EDANSSGANEPEIRFGGNVNVN---QDQIPQELRGAFQSVMHMFSGN 407
+HS+ + D +GA+EPEI GGN+N++ +Q+P++L GA +S+M MF G+
Sbjct: 348 NHSRNNEGGAGGDGTRNGADEPEIHVGGNINIDLGGTEQMPEDLSGALRSMMQMFGGS 405
>UniRef100_Q9STN6 Hypothetical protein T28D5.10 [Arabidopsis thaliana]
Length = 382
Score = 275 bits (703), Expect = 2e-72
Identities = 171/375 (45%), Positives = 231/375 (61%), Gaps = 30/375 (8%)
Query: 48 AFKINNSASVTGEPDS-----LIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTH 102
AFK+N+ +S + DS L+++F S D N+ + A++ Q+ + + +H
Sbjct: 1 AFKLNSDSSRDDDDDSFKPISLVNLFTSLDNNETIPPPPPPPVAATT----QDPSSSGSH 56
Query: 103 PDESKPMDEDWTQGPHTSAVSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFD 162
+ D + P + S+DEL GQ F LEK YFR DG DD QLEKA+R+F
Sbjct: 57 ------VSVDTCKEPSFTGTSRDELFGQLFGALEKVRYFRNTPDGDDDPAQLEKATRIFH 110
Query: 163 DGFTLYRR--WKDLGVSSLI*RIWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYC 220
D + + V+SL + K AMQS Y +A+ELY+ AIA+ +K+AV+YC
Sbjct: 111 DCVNEMEKSGCQTFDVNSLAETLKCQGNK--AMQSNLYLEAVELYSFAIALTDKNAVFYC 168
Query: 221 NRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQL 280
NRAAAYTQIN +EAI+D L+SIEIDPNYSKAYSRLGLAYYAQG Y +AI+KGFKKAL L
Sbjct: 169 NRAAAYTQINMCSEAIKDCLKSIEIDPNYSKAYSRLGLAYYAQGKYAEAIEKGFKKALLL 228
Query: 281 DPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNP 340
DP+NESVKENIRVAE K+ EE+ R +Q + S + G P+ F SMP NP
Sbjct: 229 DPHNESVKENIRVAEQKIREEQQRQRRSQQNTSTFYTQEPEMGG-GQGIPSQF-SMPVNP 286
Query: 341 SNLASMFM-AAANGGQGSHSQEGQ----EDANSSGANEPEIRFGGNVNVN---QDQIPQE 392
+L SMFM A N G+HS+ + D +GA+EPEI GGN+N++ +Q+P++
Sbjct: 287 -DLMSMFMNMAGNTFPGNHSRNNEGGAGGDGTRNGADEPEIHVGGNINIDLGGTEQMPED 345
Query: 393 LRGAFQSVMHMFSGN 407
L GA +S+M MF G+
Sbjct: 346 LSGALRSMMQMFGGS 360
>UniRef100_Q7XST1 OSJNBa0039K24.25 protein [Oryza sativa]
Length = 410
Score = 228 bits (581), Expect = 3e-58
Identities = 153/416 (36%), Positives = 224/416 (53%), Gaps = 47/416 (11%)
Query: 4 NRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTG-EPD 62
N +DSP+S IV SFL FLN VE +PG D E +EVARECL F IN+S+ V P
Sbjct: 3 NMTRSDSPISRLIVLSFLDFLNSVELAPGADPEALEVARECLESIFSINSSSVVERVHPG 62
Query: 63 SLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHTSAV 122
L+++F S +A +Q N+A P S+ +ED H
Sbjct: 63 LLLELFSSMEAAQQ-----------------DNSA-----PGPSEGQNEDTFDLDH---- 96
Query: 123 SKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRRWKDLGVSSLI*R 182
S DEL +F+ L++ ++F+T+ G +D QL KA++ FDD R+ S
Sbjct: 97 SGDELFAKFYTSLDEINFFKTSSAGAEDPGQLSKATQFFDDALLGMRKSGRKRASLGDLA 156
Query: 183 IWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRS 242
+ + M+SKQ+ A+ELY CAIA+ +A+YYCNRAAAYT +N + EA++D L+S
Sbjct: 157 EFFKSKGNEFMRSKQHLKAVELYTCAIALSRNNAIYYCNRAAAYTLLNMFNEAVEDCLKS 216
Query: 243 IEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
IEIDPNYSKAYSRLG AY+A G Y DA+ KG+ KA +LDP+NE+V +NI V + KL E+R
Sbjct: 217 IEIDPNYSKAYSRLGSAYFALGKYHDALYKGYLKASELDPSNENVWQNIEVTKKKLAEQR 276
Query: 303 HRADHNQNSRSQEFQNHYAR--GSRSHAAPASF---GSMPFNPSNLASM--------FMA 349
+ QN+ + + Q + + G S P +F G+ P P A++ F
Sbjct: 277 GPPE-EQNTYAPQSQASHGQFPGQSSSGVPFTFFPPGNSP-TPEFFANIINLPPPFSFTG 334
Query: 350 AANGGQGSHSQEGQEDANSSGANEPEIRFGGNVNVNQDQIPQELRGAFQSVMHMFS 405
+ G + + G E + +P + + +N P++ A ++VM M +
Sbjct: 335 STEGNRPQQTSSGHEGEH----GQPGMHRDAGIQINLAG-PEQAADAMRAVMEMLA 385
>UniRef100_Q9ST94 CAA30373.1 protein [Oryza sativa]
Length = 431
Score = 184 bits (468), Expect = 4e-45
Identities = 127/351 (36%), Positives = 185/351 (52%), Gaps = 18/351 (5%)
Query: 123 SKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRRWKDLGVSSLI*R 182
S DEL +F+ L++ ++F+T+ G +D QL KA++ FDD R+ S
Sbjct: 72 SGDELFAKFYTSLDEINFFKTSSAGAEDPGQLSKATQFFDDALLGMRKSGRKRASLGDLA 131
Query: 183 IWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRS 242
+ + M+SKQ+ A+ELY CAIA+ +A+YYCNRAAAYT +N + EA++D L+S
Sbjct: 132 EFFKSKGNEFMRSKQHLKAVELYTCAIALSRNNAIYYCNRAAAYTLLNMFNEAVEDCLKS 191
Query: 243 IEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
IEIDPNYSKAYSRLG AY+A G Y DA+ KG+ KA +LDP+NE+V++NI V + KL E+R
Sbjct: 192 IEIDPNYSKAYSRLGSAYFALGKYHDALYKGYLKASELDPSNENVRQNIEVTKKKLAEQR 251
Query: 303 HRADHNQNSRSQEFQNHYAR--GSRSHAAPASFGSMPFNPSNLAS-MFMAAANGGQGSHS 359
+ QN+ + + Q + + G S P +F F P N + F A S
Sbjct: 252 GPPE-EQNTYAPQSQASHGQFPGQSSSGVPFTF----FPPGNSPTPEFFANIINRVSDIS 306
Query: 360 QEGQEDANSSGANEPEIRFGGNVNVNQDQIPQELRGAFQSVMHMFSGNA---PPGQPHDQ 416
Q+ E +S N +I NVN N PQ + + F N PP
Sbjct: 307 QQSSE--HSININLNDIFNHANVNGNSQGTPQTETSSNHTPPPSFPTNTAVPPPFSFTGS 364
Query: 417 MNGSHEDANSS----EDNEPDIQFGGNVNLNSDQIPQELSGVFQNVMEMLS 463
G+ SS E +P + + +N P++ + + VMEML+
Sbjct: 365 TEGNRPQQTSSGHEGEHGQPGMHRDAGIQINLAG-PEQAADAMRAVMEMLA 414
>UniRef100_Q7YW78 Small glutamine-rich tetratricopeptide [Schistosoma japonicum]
Length = 348
Score = 109 bits (273), Expect = 2e-22
Identities = 76/266 (28%), Positives = 137/266 (50%), Gaps = 26/266 (9%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ +++ +A+ Y+ AI + +AV+YCNRAAA+++++ + +AI D L+++EIDP YSKA
Sbjct: 94 MKQEKFEEAVACYSKAIELSPYNAVFYCNRAAAHSRLDHHQDAINDCLKALEIDPYYSKA 153
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS- 311
Y R+G+AY + GN+ A++ ++K L+LDPNNE+ ++N+ +AE KL ++ +D +Q+S
Sbjct: 154 YGRMGIAYSSIGNHAKAVE-CYRKGLELDPNNENCQQNLSIAEEKL---KNSSDASQSSG 209
Query: 312 ----------RSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANGGQGSHSQE 361
S + AR S P + M +NL ++ ++
Sbjct: 210 LFGGFDLNSILSNPMMQNMARQFMSD--PNAQNMM----TNLLRNTFGMSDNPTANNPTS 263
Query: 362 GQEDANSSGANEPEIRFGGNVNVNQDQIPQELRGAFQSVMHMFSGNAPPGQPHDQMNGSH 421
E NS+G+ E ++ V + + LR Q + + N P + H
Sbjct: 264 NTEATNSNGSTESNLQSDNGTGVGGQSMDEFLRLGQQLAQQLQASN--PQLVNHLRQTFH 321
Query: 422 EDA---NSSEDNEPDIQFGGNVNLNS 444
+ NS+ +N P+ N + +S
Sbjct: 322 TNVAGQNSNPENNPNFNDEHNNSTDS 347
>UniRef100_UPI00003AFD68 UPI00003AFD68 UniRef100 entry
Length = 252
Score = 109 bits (272), Expect = 2e-22
Identities = 53/108 (49%), Positives = 76/108 (70%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++N+Y+EAI+D R+I IDP YSKA
Sbjct: 74 MKEENYGAAVDCYTRAIELDPNNAVYYCNRAAAQSKLNKYSEAIKDCERAIAIDPKYSKA 133
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + Y +AI ++KAL LDP N+S K N+++AE KL +
Sbjct: 134 YGRMGLALTSVNKYEEAI-TSYQKALDLDPENDSYKSNLKIAEQKLRD 180
Score = 45.1 bits (105), Expect = 0.005
Identities = 25/60 (41%), Positives = 35/60 (57%), Gaps = 3/60 (5%)
Query: 15 RIVRSFLHFL---NQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPDSLIDIFKSF 71
R+V +F+HFL +Q+E + E +EVA +CL FKIN + P LI+IFK F
Sbjct: 8 RLVYAFIHFLREQSQMETFTADEQESLEVAIQCLETVFKINPEDTHLAPPQHLIEIFKFF 67
>UniRef100_UPI00003AFD67 UPI00003AFD67 UniRef100 entry
Length = 304
Score = 109 bits (272), Expect = 2e-22
Identities = 53/108 (49%), Positives = 76/108 (70%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++N+Y+EAI+D R+I IDP YSKA
Sbjct: 96 MKEENYGAAVDCYTRAIELDPNNAVYYCNRAAAQSKLNKYSEAIKDCERAIAIDPKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + Y +AI ++KAL LDP N+S K N+++AE KL +
Sbjct: 156 YGRMGLALTSVNKYEEAI-TSYQKALDLDPENDSYKSNLKIAEQKLRD 202
Score = 42.4 bits (98), Expect = 0.030
Identities = 26/64 (40%), Positives = 36/64 (55%), Gaps = 4/64 (6%)
Query: 15 RIVRSFLHFL---NQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPDSLIDIF-KS 70
R+V +F+HFL +Q+E + E +EVA +CL FKIN + P LI+IF S
Sbjct: 6 RLVYAFIHFLREQSQMETFTADEQESLEVAIQCLETVFKINPEDTHLAPPQHLIEIFANS 65
Query: 71 FDAN 74
F N
Sbjct: 66 FHKN 69
>UniRef100_Q5ZJ95 Hypothetical protein [Gallus gallus]
Length = 304
Score = 109 bits (272), Expect = 2e-22
Identities = 53/108 (49%), Positives = 76/108 (70%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++N+Y+EAI+D R+I IDP YSKA
Sbjct: 96 MKEENYGAAVDCYTRAIELDPNNAVYYCNRAAAQSKLNKYSEAIKDCERAIAIDPKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + Y +AI ++KAL LDP N+S K N+++AE KL +
Sbjct: 156 YGRMGLALTSVNKYEEAI-TSYQKALDLDPENDSYKSNLKIAEQKLRD 202
Score = 41.6 bits (96), Expect = 0.051
Identities = 26/64 (40%), Positives = 36/64 (55%), Gaps = 4/64 (6%)
Query: 15 RIVRSFLHFL---NQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPDSLIDIF-KS 70
R+V +F+HFL +Q+E + E +EVA +CL FKIN + P LI+IF S
Sbjct: 6 RLVYAFIHFLREQSQMETFTPDEQESLEVAIQCLETVFKINPEDTHLAPPQHLIEIFANS 65
Query: 71 FDAN 74
F N
Sbjct: 66 FHKN 69
>UniRef100_Q7ZUM6 Small glutamine-rich tetratricopeptide repeat (TPR)-containing
[Brachydanio rerio]
Length = 320
Score = 108 bits (269), Expect = 4e-22
Identities = 67/199 (33%), Positives = 106/199 (52%), Gaps = 16/199 (8%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + + A+E Y+ AI + ++AVY+CNRAAAY+++ Y A+QD R+I ID NYSKA
Sbjct: 102 MKVENFSAAVEFYSKAIQLNPQNAVYFCNRAAAYSKLGNYAGAVQDCERAIGIDANYSKA 161
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER---------- 302
Y R+GLA + Y +A+ +KKAL+LDP+N++ K N++VAE K+ E +
Sbjct: 162 YGRMGLALASLNKYSEAVSY-YKKALELDPDNDTYKVNLQVAEQKVKETQPSTAGGLGGV 220
Query: 303 HRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANGGQGSHS--- 359
A N ++ + + S ++P + A+ AA GG G++
Sbjct: 221 DLAGLLSNPGFMNMASNLMNNPQVQQLVSGMMSGSYSPVSSATSPTAAGAGGGGANDLSS 280
Query: 360 --QEGQEDANSSGANEPEI 376
Q GQ+ A PE+
Sbjct: 281 LIQAGQQFAQQMQQQNPEL 299
>UniRef100_Q6NXA1 Small glutamine-rich tetratricopeptide repeat (TPR)-containing
[Brachydanio rerio]
Length = 320
Score = 108 bits (269), Expect = 4e-22
Identities = 67/199 (33%), Positives = 106/199 (52%), Gaps = 16/199 (8%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + + A+E Y+ AI + ++AVY+CNRAAAY+++ Y A+QD R+I ID NYSKA
Sbjct: 102 MKVENFSAAVEFYSKAIQLNPQNAVYFCNRAAAYSKLGNYAGAVQDCERAIGIDANYSKA 161
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER---------- 302
Y R+GLA + Y +A+ +KKAL+LDP+N++ K N++VAE K+ E +
Sbjct: 162 YGRMGLALASLNKYSEAVSY-YKKALELDPDNDTYKVNLQVAEQKVKETQPSTAGGLGGV 220
Query: 303 HRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANGGQGSHS--- 359
A N ++ + + S ++P + A+ AA GG G++
Sbjct: 221 DLAGLLSNPGFMNMASNLMNNPQVQQLVSGMMSGSYSPVSSATSPTAAGAGGGGANDLSS 280
Query: 360 --QEGQEDANSSGANEPEI 376
Q GQ+ A PE+
Sbjct: 281 LIQAGQQFAQQMQQQNPEL 299
>UniRef100_UPI0000360266 UPI0000360266 UniRef100 entry
Length = 306
Score = 105 bits (263), Expect = 2e-21
Identities = 55/108 (50%), Positives = 73/108 (66%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A+E Y AI + ++AVYYCNRAAA++++ YTEA D R+I IDP YSKA
Sbjct: 98 MKEENYRCAVECYTKAIELDLRNAVYYCNRAAAHSKLGNYTEATCDCERAIGIDPTYSKA 157
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A Y +AI FKKAL LDP N++ K N+++AE K E
Sbjct: 158 YGRMGLALTAMSKYPEAISY-FKKALVLDPENDTYKSNLKIAEQKHKE 204
>UniRef100_Q6GMI8 Zgc:92462 [Brachydanio rerio]
Length = 306
Score = 105 bits (263), Expect = 2e-21
Identities = 51/108 (47%), Positives = 74/108 (68%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +++AVYYCNRAAA++++ YTEA+ D R+I IDP+YSKA
Sbjct: 98 MKEENYSSAVDCYTKAIELDQRNAVYYCNRAAAHSKLENYTEAMGDCERAIAIDPSYSKA 157
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + Y +AI F KAL LDP N++ K N+++ E K E
Sbjct: 158 YGRMGLALTSMSKYPEAISY-FNKALVLDPENDTYKSNLKIVEQKQKE 204
>UniRef100_Q21746 Hypothetical protein R05F9.10 [Caenorhabditis elegans]
Length = 337
Score = 105 bits (261), Expect = 4e-21
Identities = 51/108 (47%), Positives = 75/108 (69%), Gaps = 2/108 (1%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M++ Q+ A++ YN AI + + VY+CNRAAAY ++ +Y AIQD ++ +DP+YSKA
Sbjct: 116 MKASQFEAAVQKYNAAIKL-NRDPVYFCNRAAAYCRLEQYDLAIQDCRTALALDPSYSKA 174
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
+ R+GLAY Q Y A + +KKAL+L+PN ES K N+++AE KL E
Sbjct: 175 WGRMGLAYSCQNRYEHAAE-AYKKALELEPNQESYKNNLKIAEDKLKE 221
>UniRef100_UPI0000432F49 UPI0000432F49 UniRef100 entry
Length = 294
Score = 104 bits (259), Expect = 6e-21
Identities = 49/108 (45%), Positives = 80/108 (73%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+++++ +A+ Y +I + ++AVYYCNRAAAY++I Y +AI D ++ IDP+YSKA
Sbjct: 90 MKAEKHHEALANYTKSIQLDGRNAVYYCNRAAAYSKIGNYQQAINDCHTALSIDPSYSKA 149
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y RLGLAY + +++A + ++KAL+++P+NES K N++VAE KL +
Sbjct: 150 YGRLGLAYSSLQRHKEA-KESYQKALEMEPDNESYKNNLQVAEEKLAQ 196
>UniRef100_Q80W98 Small glutamine rich protein with tetratricopeptide repeats 2
[Rattus norvegicus]
Length = 304
Score = 104 bits (259), Expect = 6e-21
Identities = 50/108 (46%), Positives = 74/108 (68%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA ++++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLSHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTAMNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>UniRef100_Q8VD33 Small glutamine-rich tetratricopeptide repeat-containing protein B
[Mus musculus]
Length = 304
Score = 104 bits (259), Expect = 6e-21
Identities = 50/108 (46%), Positives = 74/108 (68%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA ++++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLSHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTAMNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>UniRef100_UPI00002DA2F7 UPI00002DA2F7 UniRef100 entry
Length = 337
Score = 103 bits (257), Expect = 1e-20
Identities = 54/108 (50%), Positives = 72/108 (66%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A+E Y AI + ++AVYYCNRAAA++++ Y EA D R+I IDP YSKA
Sbjct: 91 MKEENYRCAVECYTKAIELDLRNAVYYCNRAAAHSKLGNYMEATCDCERAIGIDPTYSKA 150
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A Y +AI FKKAL LDP N++ K N+++AE K E
Sbjct: 151 YGRMGLALTAMSKYPEAISY-FKKALVLDPENDTYKSNLKIAEQKHKE 197
>UniRef100_UPI000036D10D UPI000036D10D UniRef100 entry
Length = 304
Score = 103 bits (256), Expect = 1e-20
Identities = 50/108 (46%), Positives = 73/108 (67%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLGHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTALNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>UniRef100_Q96EQ0 Small glutamine-rich tetratricopeptide repeat-containing protein B
[Homo sapiens]
Length = 304
Score = 103 bits (256), Expect = 1e-20
Identities = 50/108 (46%), Positives = 73/108 (67%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLGHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTALNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>UniRef100_UPI00003AFB3D UPI00003AFB3D UniRef100 entry
Length = 313
Score = 102 bits (255), Expect = 2e-20
Identities = 48/108 (44%), Positives = 76/108 (69%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+++ + A+ Y AI + +AVY+CNRAAAY+++ Y A++D R+I IDPNYSKA
Sbjct: 102 MKAENFEAAVSFYGKAIELNPSNAVYFCNRAAAYSKLGNYAGAVRDCERAIGIDPNYSKA 161
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + + +A+ +KKAL+LDP+N++ K N+++AE K+ E
Sbjct: 162 YGRMGLALSSLNKHTEAV-VYYKKALELDPDNDTYKSNLKIAEQKMKE 208
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 805,999,362
Number of Sequences: 2790947
Number of extensions: 33742282
Number of successful extensions: 112285
Number of sequences better than 10.0: 2337
Number of HSP's better than 10.0 without gapping: 1289
Number of HSP's successfully gapped in prelim test: 1060
Number of HSP's that attempted gapping in prelim test: 106259
Number of HSP's gapped (non-prelim): 5912
length of query: 481
length of database: 848,049,833
effective HSP length: 131
effective length of query: 350
effective length of database: 482,435,776
effective search space: 168852521600
effective search space used: 168852521600
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Medicago: description of AC139354.8


