

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.6 - phase: 0
(342 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
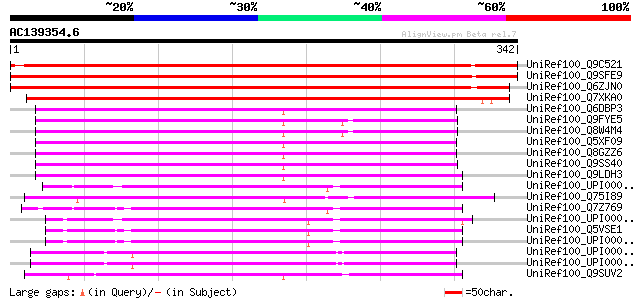
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C521 Hypothetical protein At1g77610 [Arabidopsis tha... 618 e-176
UniRef100_Q9SFE9 T26F17.9 [Arabidopsis thaliana] 611 e-173
UniRef100_Q6ZJN0 Putative glucose-6-phosphate/phosphate transloc... 557 e-157
UniRef100_Q7XKA0 OSJNBb0020J19.10 protein [Oryza sativa] 555 e-157
UniRef100_Q6DBP3 At5g05820 [Arabidopsis thaliana] 184 3e-45
UniRef100_Q9FYE5 Phosphate/phosphoenolpyruvate translocator-like... 183 5e-45
UniRef100_Q8W4M4 Phosphate/phosphoenolpyruvate translocator-like... 183 7e-45
UniRef100_Q5XF09 At3g11320 [Arabidopsis thaliana] 182 2e-44
UniRef100_Q8GZZ6 Putative phosphate/phosphoenolpyruvate transloc... 177 3e-43
UniRef100_Q9SS40 F14P13.11 protein [Arabidopsis thaliana] 175 2e-42
UniRef100_Q9LDH3 T12C24.5 [Arabidopsis thaliana] 164 2e-39
UniRef100_UPI00003A96AC UPI00003A96AC UniRef100 entry 139 9e-32
UniRef100_Q75I89 Expressed protein [Oryza sativa] 136 7e-31
UniRef100_Q7Z769 SLC35E3 protein [Homo sapiens] 136 9e-31
UniRef100_UPI0000365B60 UPI0000365B60 UniRef100 entry 135 2e-30
UniRef100_Q5VSE1 Novel protein [Brachydanio rerio] 134 4e-30
UniRef100_UPI000043840D UPI000043840D UniRef100 entry 134 5e-30
UniRef100_UPI00002BACDF UPI00002BACDF UniRef100 entry 133 8e-30
UniRef100_UPI000036626E UPI000036626E UniRef100 entry 130 4e-29
UniRef100_Q9SUV2 Hypothetical protein F8B4.90 [Arabidopsis thali... 127 6e-28
>UniRef100_Q9C521 Hypothetical protein At1g77610 [Arabidopsis thaliana]
Length = 336
Score = 618 bits (1593), Expect = e-176
Identities = 304/343 (88%), Positives = 322/343 (93%), Gaps = 8/343 (2%)
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEEG S+ RSLLAILQWW FNVTVIIMNKWIFQKLDFKFPLSVSC+HFICS+IGAY
Sbjct: 1 MEEG-----SMFRSLLAILQWWGFNVTVIIMNKWIFQKLDFKFPLSVSCVHFICSSIGAY 55
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
+VIKVLKLKPLI VDP+DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT
Sbjct: 56 IVIKVLKLKPLIVVDPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 115
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAE 180
VVLQWLVWRKYFDWRIWASLVPIVGGILLTS+TELSFNMFGFCAALFGCLATSTKTILAE
Sbjct: 116 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSVTELSFNMFGFCAALFGCLATSTKTILAE 175
Query: 181 ALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVL 240
+LLHGYKFDSINTVY+MAPFAT+I+ PALLLEG+GIL WF HP PW+A+III SSGVL
Sbjct: 176 SLLHGYKFDSINTVYYMAPFATMILGIPALLLEGSGILSWFEAHPAPWSALIIILSSGVL 235
Query: 241 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTF 300
AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAV++SWLIFRNPISYMNAVGC ITLVGCTF
Sbjct: 236 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVMVSWLIFRNPISYMNAVGCGITLVGCTF 295
Query: 301 YGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLV-NDKLDDKV 342
YGYVR+M+SQQ PGTPRTPRTPRSKMELLPLV NDKL+ KV
Sbjct: 296 YGYVRHMLSQQ--TPGTPRTPRTPRSKMELLPLVNNDKLEGKV 336
>UniRef100_Q9SFE9 T26F17.9 [Arabidopsis thaliana]
Length = 341
Score = 611 bits (1575), Expect = e-173
Identities = 295/343 (86%), Positives = 321/343 (93%), Gaps = 3/343 (0%)
Query: 1 MEEG-LVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGA 59
MEEG L QW++ RSLL+ILQWW FNVTVIIMNKWIFQKLDFKFPLSVSC+HFICS+IGA
Sbjct: 1 MEEGSLWRQWTMFRSLLSILQWWGFNVTVIIMNKWIFQKLDFKFPLSVSCVHFICSSIGA 60
Query: 60 YVVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPAT 119
Y+VIKVLKLKPLI VDP+DRWRRIFPMSFVFCINIVLGN+SLRYIPVSFMQTIKS TPAT
Sbjct: 61 YIVIKVLKLKPLIVVDPEDRWRRIFPMSFVFCINIVLGNISLRYIPVSFMQTIKSLTPAT 120
Query: 120 TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILA 179
TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFN+FGFCAALFGCLATSTKTILA
Sbjct: 121 TVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNVFGFCAALFGCLATSTKTILA 180
Query: 180 EALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGV 239
E+LLHGYKFDSINTVY+MAPFAT+I+ PA LLE NGIL+WF HP PW+A+II+F+SGV
Sbjct: 181 ESLLHGYKFDSINTVYYMAPFATMILGLPAFLLERNGILDWFEAHPSPWSALIILFNSGV 240
Query: 240 LAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCT 299
LAFCLNFSIFYVI STTAVTFNVAGNLKVAVAV +SW+IFRNPIS MNAVGC ITLVGCT
Sbjct: 241 LAFCLNFSIFYVIQSTTAVTFNVAGNLKVAVAVFVSWMIFRNPISPMNAVGCGITLVGCT 300
Query: 300 FYGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLVNDKLDDKV 342
FYGYVR+M+SQQ PGTPRTPRTPR+KMEL+PLVNDKL+ K+
Sbjct: 301 FYGYVRHMLSQQQ--PGTPRTPRTPRNKMELIPLVNDKLESKI 341
>UniRef100_Q6ZJN0 Putative glucose-6-phosphate/phosphate translocator [Oryza sativa]
Length = 337
Score = 557 bits (1436), Expect = e-157
Identities = 268/337 (79%), Positives = 304/337 (89%), Gaps = 4/337 (1%)
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEE + + +R++LAILQWW FNVTVIIMNKWIFQKL+FKFPL+VSC+HFICS+IGAY
Sbjct: 1 MEEAKMGDVATIRAVLAILQWWGFNVTVIIMNKWIFQKLEFKFPLTVSCVHFICSSIGAY 60
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
+ IK+LK+KPLI V P+DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT
Sbjct: 61 IAIKILKMKPLIEVAPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAE 180
V+LQWLVWRKYF+WRIWASLVPIVGGI+LTSITELSFNMFGFCAA+ GCLATSTKTILAE
Sbjct: 121 VILQWLVWRKYFEWRIWASLVPIVGGIMLTSITELSFNMFGFCAAMVGCLATSTKTILAE 180
Query: 181 ALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVL 240
+LLHGYKFDSINTVY+MAPFAT+I+ PA++LEG+G++ W + A+III +SGVL
Sbjct: 181 SLLHGYKFDSINTVYYMAPFATMILSVPAIVLEGSGVINWLYTYDSIVPALIIITTSGVL 240
Query: 241 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTF 300
AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVL+SW+IFRNPIS MNAVGCAITLVGCTF
Sbjct: 241 AFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLVSWMIFRNPISAMNAVGCAITLVGCTF 300
Query: 301 YGYVRNMISQQPAVPGTPRTPRTPRSKMELLPLVNDK 337
YGYVR++ISQQ +PRTPRS+ME+LPLV DK
Sbjct: 301 YGYVRHLISQQ----SVNSSPRTPRSRMEMLPLVGDK 333
>UniRef100_Q7XKA0 OSJNBb0020J19.10 protein [Oryza sativa]
Length = 350
Score = 555 bits (1429), Expect = e-157
Identities = 271/331 (81%), Positives = 301/331 (90%), Gaps = 5/331 (1%)
Query: 12 VRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPL 71
+R+LLAILQWW FNVTVII+NKWIFQKLDFKFPL+VSC+HFICS+IGAY+ I VLK KPL
Sbjct: 16 LRALLAILQWWGFNVTVIIINKWIFQKLDFKFPLTVSCVHFICSSIGAYIAIHVLKAKPL 75
Query: 72 ISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKY 131
I V+P+DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTV+LQWLVW K+
Sbjct: 76 IQVEPEDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVILQWLVWSKH 135
Query: 132 FDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSI 191
F+WRIWASLVPIVGGILLTSITELSFNMFGFCAA+ GCLATSTKTILAE+LLHGYKFDSI
Sbjct: 136 FEWRIWASLVPIVGGILLTSITELSFNMFGFCAAMVGCLATSTKTILAESLLHGYKFDSI 195
Query: 192 NTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYV 251
NTVY+MAPFAT+I+ PA+LLEG G++ WF H +A++II SGVLAFCLNFSIFYV
Sbjct: 196 NTVYYMAPFATMILALPAVLLEGGGVVTWFYTHDSIASALVIIIGSGVLAFCLNFSIFYV 255
Query: 252 IHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQ 311
IHSTTAVTFNVAGNLKVAVAVL+SWLIFRNPIS MNA+GCAITLVGCTFYGYVR++I QQ
Sbjct: 256 IHSTTAVTFNVAGNLKVAVAVLVSWLIFRNPISPMNAIGCAITLVGCTFYGYVRHLIPQQ 315
Query: 312 PAV-PGT--PRTPRT--PRSKMELLPLVNDK 337
AV PGT P T +T PRS+ME+LPLV DK
Sbjct: 316 QAVAPGTGSPTTSQTNSPRSRMEMLPLVGDK 346
>UniRef100_Q6DBP3 At5g05820 [Arabidopsis thaliana]
Length = 309
Score = 184 bits (467), Expect = 3e-45
Identities = 95/286 (33%), Positives = 172/286 (59%), Gaps = 2/286 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I LK+ P+ ++ +
Sbjct: 15 VASWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLLSYVAIAWLKMVPMQTIRSR 74
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S VFC+++V GN+SLR++PVSF Q I + TP T V +L+ RK W +
Sbjct: 75 VQFFKIAALSLVFCVSVVFGNISLRFLPVSFNQAIGATTPFFTAVFAYLMTRKKEAWLTY 134
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E SF++FGF + A + K++L LL G K +S+N +
Sbjct: 135 FTLVPVVTGVVIASGGEPSFHLFGFLMCIAATAARALKSVLQGILLSSEGEKLNSMNLLL 194
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A ++++ L++E N + ++ + + + + LA+ +N + F V + T
Sbjct: 195 YMAPIAVVLLLPATLIMEKNVVGITIALARDDFRIVWYLLFNSALAYLVNLTNFLVTNHT 254
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+A+T V GN K AVAV++S LIF+NP+S +G ++T+ G Y
Sbjct: 255 SALTLQVLGNAKGAVAVVVSILIFKNPVSVTGMLGYSLTVCGVILY 300
>UniRef100_Q9FYE5 Phosphate/phosphoenolpyruvate translocator-like protein
[Arabidopsis thaliana]
Length = 309
Score = 183 bits (465), Expect = 5e-45
Identities = 102/290 (35%), Positives = 169/290 (58%), Gaps = 8/290 (2%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I+ W++ N+ V+++NK++ FKFP+ ++ H AI +Y+ I LKL PL + +
Sbjct: 16 IISWYSSNIGVLLLNKFLLSNYGFKFPIFLTMCHMSACAILSYISIVFLKLVPLQHLKSR 75
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ ++ +S VFC ++V GN+SLRY+PVSF Q + + TP T + +L+ K W +
Sbjct: 76 SQFLKVATLSIVFCASVVGGNISLRYLPVSFNQAVGATTPFFTALFAYLMTFKREAWVTY 135
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E F+ FGF + A + K++L LL G K +S+N +
Sbjct: 136 GALVPVVAGVVIASGGEPGFHWFGFIMCISATAARAFKSVLQGILLSSEGEKLNSMNLML 195
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSI---HPYPWAAMIIIFSSGVLAFCLNFSIFYVI 252
+M+P A + ++ L +E + I ++ H Y W I++ + V+A+ N F V
Sbjct: 196 YMSPIAVIALLPVTLFMEPDVISVTLTLAKQHQYMW---ILLLVNSVMAYSANLLNFLVT 252
Query: 253 HSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
T+A+T V GN K AVAV+IS LIF+NP++ M G +IT++G YG
Sbjct: 253 KHTSALTLQVLGNAKGAVAVVISILIFQNPVTVMGIGGYSITVLGVVAYG 302
>UniRef100_Q8W4M4 Phosphate/phosphoenolpyruvate translocator-like protein
[Arabidopsis thaliana]
Length = 309
Score = 183 bits (464), Expect = 7e-45
Identities = 102/290 (35%), Positives = 169/290 (58%), Gaps = 8/290 (2%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I+ W++ N+ V+++NK++ FKFP+ ++ H AI +Y+ I LKL PL + +
Sbjct: 16 IISWYSSNIGVLLLNKFLLSNYGFKFPIFLTMCHMSACAILSYISIVFLKLVPLQHLKSR 75
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ ++ +S VFC ++V GN+SLRY+PVSF Q + + TP T + +L+ K W +
Sbjct: 76 SQFLKVATLSIVFCASVVGGNISLRYLPVSFNQAVGATTPFFTALFAYLMTFKREAWVTY 135
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E F+ FGF + A + K++L LL G K +S+N +
Sbjct: 136 GALVPVVAGVVIASGGEPGFHWFGFIMCISATAARAFKSVLQGILLSSEGEKLNSMNLML 195
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSI---HPYPWAAMIIIFSSGVLAFCLNFSIFYVI 252
+M+P A + ++ L +E + I ++ H Y W I++ + V+A+ N F V
Sbjct: 196 YMSPVAVIALLPVTLFMEPDVISVTLTLAKQHQYMW---ILLLVNSVMAYSANLLNFLVT 252
Query: 253 HSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
T+A+T V GN K AVAV+IS LIF+NP++ M G +IT++G YG
Sbjct: 253 KHTSALTLQVLGNAKGAVAVVISILIFQNPVTVMGIGGYSITVLGVVAYG 302
>UniRef100_Q5XF09 At3g11320 [Arabidopsis thaliana]
Length = 308
Score = 182 bits (461), Expect = 2e-44
Identities = 94/286 (32%), Positives = 170/286 (58%), Gaps = 2/286 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I +K+ P+ ++ +
Sbjct: 15 VASWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLLSYVAIAWMKMVPMQTIRSR 74
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S VFC+++V GN+SLR++PVSF Q I + TP T V +L+ K W +
Sbjct: 75 VQFLKIAALSLVFCVSVVFGNISLRFLPVSFNQAIGATTPFFTAVFAYLITFKREAWLTY 134
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S +E SF++FGF + A + K++L LL G K +S+N +
Sbjct: 135 FTLVPVVTGVVIASGSEPSFHLFGFIMCIAATAARALKSVLQGILLSSEGEKLNSMNLLL 194
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A + ++ L++E N + ++ + + + + LA+ +N + F V T
Sbjct: 195 YMAPIAVVFLLPATLIMEKNVVGITIALARDDFRIVWYLLFNSALAYFVNLTNFLVTKHT 254
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+A+T V GN K AVAV++S LIFRNP+S +G ++T+ G Y
Sbjct: 255 SALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMLGYSLTVCGVILY 300
>UniRef100_Q8GZZ6 Putative phosphate/phosphoenolpyruvate translocator protein [Oryza
sativa]
Length = 322
Score = 177 bits (450), Expect = 3e-43
Identities = 95/286 (33%), Positives = 164/286 (57%), Gaps = 2/286 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H A+ +Y I L++ P+ V +
Sbjct: 28 VTAWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMSACALLSYAAIAWLRVVPMQLVRSR 87
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
+ +I +S VFC ++V GNVSLRY+PVSF Q + + TP T V +++ K W +
Sbjct: 88 VQLAKIAALSLVFCGSVVSGNVSLRYLPVSFNQAVGATTPFFTAVFAYIMTVKRESWVTY 147
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E SF++FGF + A + KT+L LL G K +S+N +
Sbjct: 148 LTLVPVVTGVMIASGGEPSFHLFGFIMCIGATAARALKTVLQGILLSSEGEKLNSMNLLL 207
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A ++++ + +E N + + + ++ + LA+ +N + F V T
Sbjct: 208 YMAPIAVILLLPATIFMEDNVVGITIELAKKDTTIVWLLLFNSCLAYFVNLTNFLVTKHT 267
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
+A+T V GN K AVAV++S LIFRNP+S +G +T++G Y
Sbjct: 268 SALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMLGYTLTVIGVILY 313
>UniRef100_Q9SS40 F14P13.11 protein [Arabidopsis thaliana]
Length = 355
Score = 175 bits (443), Expect = 2e-42
Identities = 96/287 (33%), Positives = 165/287 (57%), Gaps = 2/287 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I+ W+ N+ V+++NK++ FKFP+ ++ H AI +YV I LKL PL + +
Sbjct: 62 IILWYTSNIGVLLLNKFLLSNYGFKFPIFLTMCHMSACAILSYVSIVFLKLVPLQYLKSR 121
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ ++ +S VFC ++V GN+SLRY+PVSF Q + + TP T + +++ K W +
Sbjct: 122 SQFLKVATLSIVFCASVVGGNISLRYLPVSFNQAVGATTPFFTALFAYIMTFKREAWVTY 181
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+LVP+V G+++ S E F+ FGF + A + K++L LL G + +S+N +
Sbjct: 182 GALVPVVTGVVIASGGEPGFHWFGFIMCISATAARAFKSVLQGILLSSEGERLNSMNLML 241
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+M+P A + ++ + +E + + ++ I++ + V+A+ N F V T
Sbjct: 242 YMSPIAVIALLPVTIFMEPDVMSVTLTLGRQHKYMYILLLVNSVMAYSANLLNFLVTKHT 301
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
+A+T V GN K AVAV+IS L+FRNP++ M G +IT++G YG
Sbjct: 302 SALTLQVLGNAKGAVAVVISILLFRNPVTVMGIGGYSITVLGVVAYG 348
>UniRef100_Q9LDH3 T12C24.5 [Arabidopsis thaliana]
Length = 361
Score = 164 bits (416), Expect = 2e-39
Identities = 91/290 (31%), Positives = 158/290 (54%), Gaps = 2/290 (0%)
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
I W+ N+ V+++NK++ F++P+ ++ H + A + VI + + P + +
Sbjct: 63 IAAWFGSNIGVLLLNKYLLFYYGFRYPIFLTMTHMLSCAAYSSAVINIAGIVPRQHILSR 122
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
++ +I +S +FC+++V GN SLRYIPVSF Q I + TP T V +L+ K ++
Sbjct: 123 RQFLKILSLSAIFCLSVVCGNTSLRYIPVSFNQAIGATTPFFTAVFSFLITCKTESTEVY 182
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTVY 195
+L+P+V GI+L S +E SF++FGF + + K+++ +L K S+N +
Sbjct: 183 LALLPVVSGIVLASNSEPSFHLFGFLICVASTAGRALKSVVQGIILTSESEKLHSMNLLL 242
Query: 196 HMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHST 255
+MAP A I++ L +EGN + + ++ + +A+ +N + F V T
Sbjct: 243 YMAPMAACILLPFTLYIEGNVLRVLIEKARTDPLIIFLLAGNATVAYLVNLTNFLVTKHT 302
Query: 256 TAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+A+T V GN K AVA +S LIFRNP++ M G +T++G Y R
Sbjct: 303 SALTLQVLGNGKAAVAAGVSVLIFRNPVTVMGIAGFGVTIMGVVLYSEAR 352
>UniRef100_UPI00003A96AC UPI00003A96AC UniRef100 entry
Length = 302
Score = 139 bits (351), Expect = 9e-32
Identities = 84/286 (29%), Positives = 152/286 (52%), Gaps = 13/286 (4%)
Query: 23 AFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRR 82
A ++ ++ +NKW++ +L F LS++ +HF + +G Y+ + P R +
Sbjct: 12 AASICIVFLNKWLYVRLGFP-NLSLTLVHFAITWLGLYLCQALGAFAP-----KSLRAAQ 65
Query: 83 IFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVP 142
+ P++ FC +V N+SL+ + Q K+ T V++Q L + K F RI +LVP
Sbjct: 66 VLPLALSFCGFVVFTNLSLQSNTIGTYQLAKAMTTPVIVLIQSLAYGKSFPLRIKLTLVP 125
Query: 143 IVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFAT 202
I G+ L S ++ FN+ G A G L TS + A H + +S+ +Y+ AP ++
Sbjct: 126 ITLGVFLNSYYDVKFNVLGTVFATLGVLVTSLYQVWVGAKQHELQVNSMQLLYYQAPMSS 185
Query: 203 LIMVFPALLLE---GNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVT 259
+++F E G G + P+ +A+I++ SGV+AF +N SI+++I +T+ VT
Sbjct: 186 AMLLFIIPFFEPVFGEGGI----FGPWTLSAVIMVLLSGVIAFMVNLSIYWIIGNTSPVT 241
Query: 260 FNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
+N+ G+ K + +L L+F++P+S +G TL G Y + +
Sbjct: 242 YNMFGHFKFCITLLGGCLLFKDPLSVNQGLGILCTLFGILAYTHFK 287
>UniRef100_Q75I89 Expressed protein [Oryza sativa]
Length = 379
Score = 136 bits (343), Expect = 7e-31
Identities = 91/322 (28%), Positives = 162/322 (50%), Gaps = 10/322 (3%)
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFP--LSVSCIHFICSAIGAYVVIKVLKL 68
+V + L +L + + VI+ NKW+ FKFP ++++ IH S + + +++V K+
Sbjct: 7 LVLTYLYLLIYVCLSSGVILFNKWVLSPKYFKFPFPITLTMIHMAFSGVVTFFLVRVFKV 66
Query: 69 KPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVW 128
+ + Q + P+S F ++ GN + YI V+F+Q +K+ P T ++ L
Sbjct: 67 VAPVKMTFQIYATCVIPISAFFASSLWFGNTAYLYISVAFIQMLKALMPVATFIMAVLCG 126
Query: 129 RKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--GY 186
W ++ ++V + G++++S E+ FN+ G + G A + + +L + LL G
Sbjct: 127 TDKLRWDLFLNMVLVSVGVVVSSYGEIHFNIIGTLYQVTGIFAEALRLVLTQVLLQKKGL 186
Query: 187 KFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNF 246
+ I ++Y++AP + + + P LLE ++ I W I F + V AF LN
Sbjct: 187 TLNPITSLYYIAPCSFIFLFVPWFLLE-KPEMDVSQIQFNYW----IFFFNAVAAFALNI 241
Query: 247 SIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIF-RNPISYMNAVGCAITLVGCTFYGYVR 305
SIF VI T AVT VAG LK + + +S +IF + I+ +N +G A+ L G Y Y++
Sbjct: 242 SIFLVIGRTGAVTIRVAGVLKDWILIALSTIIFPESIITSLNIIGYAVALSGVVMYNYLK 301
Query: 306 NMISQQPAVPGTPRTPRTPRSK 327
+ +P R + K
Sbjct: 302 MKDVRANQLPADIAPDRATKDK 323
>UniRef100_Q7Z769 SLC35E3 protein [Homo sapiens]
Length = 313
Score = 136 bits (342), Expect = 9e-31
Identities = 82/300 (27%), Positives = 156/300 (51%), Gaps = 16/300 (5%)
Query: 9 WSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKL 68
W + LL L ++ ++ +NKWI+ F +S++ +HF+ + +G Y+ K+
Sbjct: 12 WRIAAGLLFNL---LVSICIVFLNKWIYVYHGFP-NMSLTLVHFVVTWLGLYICQKLDIF 67
Query: 69 KPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVW 128
P S+ P R+ ++ FC +V N+SL+ + Q K+ T + +Q +
Sbjct: 68 APK-SLPPS----RLLLLALSFCGFVVFTNLSLQNNTIGTYQLAKAMTTPVIIAIQTFCY 122
Query: 129 RKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKF 188
+K F RI +L+PI G++L S ++ FN G A G L TS + A H +
Sbjct: 123 QKTFSTRIQLTLIPITLGVILNSYYDVKFNFLGMVFAALGVLVTSLYQVWVGAKQHELQV 182
Query: 189 DSINTVYHMAPFATLIMVFPALLLE---GNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+S+ +Y+ AP ++ +++ E G G + P+ +A++++ SGV+AF +N
Sbjct: 183 NSMQLLYYQAPMSSAMLLVAVPFFEPVFGEGGI----FGPWSVSALLMVLLSGVIAFMVN 238
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
SI+++I +T+ VT+N+ G+ K + + +++F++P+S A+G TL G Y + +
Sbjct: 239 LSIYWIIGNTSPVTYNMFGHFKFCITLFGGYVLFKDPLSINQALGILCTLFGILAYTHFK 298
>UniRef100_UPI0000365B60 UPI0000365B60 UniRef100 entry
Length = 306
Score = 135 bits (340), Expect = 2e-30
Identities = 85/300 (28%), Positives = 160/300 (53%), Gaps = 23/300 (7%)
Query: 25 NVTVIIMNKWIFQKLDFKFP-LSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRI 83
++ ++ +NKWI+ + + FP ++++ IHF+ + +G Y+ K+ P + R+I
Sbjct: 18 SICIVFINKWIY--MHYGFPNMTLTLIHFVVTWLGLYICQKMDIFSP-----KRLPIRKI 70
Query: 84 FPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPI 143
++ FC + N+SL+ + Q K+ T +++Q ++K F +I +LVPI
Sbjct: 71 VLLALSFCGFVAFTNLSLQNNSIGTYQLAKTMTTPVIIIIQTTYYKKTFSTKIKLTLVPI 130
Query: 144 VGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF--A 201
G++L S ++ FN+ G A G L TS + A H + +S+ +Y+ AP A
Sbjct: 131 TLGVILNSYYDVRFNLLGTVFATLGVLVTSLYQVWVGAKQHELQVNSMQLLYYQAPLSSA 190
Query: 202 TLIMVFP-ALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTF 260
L+ + P + L G+G + P+ AA+ + SGV+AF +N SI+++I +T+ VT+
Sbjct: 191 FLLAIIPFSEPLSGDGGI----FGPWSLAALATVLFSGVIAFLVNLSIYWIIGNTSPVTY 246
Query: 261 NVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYV--------RNMISQQP 312
N+ G+ K + ++ +L+F +P+S A+G TL G Y + +N ++Q+P
Sbjct: 247 NMFGHFKFCITLVGGYLLFHDPLSLNQALGILCTLAGILSYTHFKLVEPEDGKNRLAQRP 306
>UniRef100_Q5VSE1 Novel protein [Brachydanio rerio]
Length = 317
Score = 134 bits (337), Expect = 4e-30
Identities = 83/285 (29%), Positives = 156/285 (54%), Gaps = 15/285 (5%)
Query: 25 NVTVIIMNKWIFQKLDFKFP-LSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRI 83
+V ++ +NKWI+ + + FP ++++ IHF+ + +G ++ K+ P S+ P +I
Sbjct: 29 SVCIVFINKWIY--VHYGFPNMTLTLIHFVMTWLGLFICQKMDIFAPK-SLRPS----KI 81
Query: 84 FPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPI 143
++ FC +V N+SL+ + Q K T + +Q + +RK F +I +LVPI
Sbjct: 82 LLLALSFCGFVVFTNLSLQSNTIGTYQLAKVMTTPVIIAIQTMYYRKTFSTKIKLTLVPI 141
Query: 144 VGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF--A 201
G++L S ++ FN+ G A G L TS + A H + +S+ +Y+ AP A
Sbjct: 142 TLGVILNSYYDVRFNLMGMIFATLGVLVTSLYQVWVGAKQHELQVNSMQLLYYQAPMSSA 201
Query: 202 TLIMVFPAL-LLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTF 260
L+++ P L G+G + P+ + A+ ++ SGV+AF +N SI+++I +T+ VT+
Sbjct: 202 FLLVLVPFFEPLTGDGGI----FGPWSFLALFMVLLSGVIAFLVNLSIYWIIGNTSPVTY 257
Query: 261 NVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
N+ G+ K + +L +++F++P+S +G TL G Y + +
Sbjct: 258 NMFGHFKFCITLLGGYVLFQDPLSLNQGLGILCTLTGILAYTHFK 302
>UniRef100_UPI000043840D UPI000043840D UniRef100 entry
Length = 313
Score = 134 bits (336), Expect = 5e-30
Identities = 82/285 (28%), Positives = 156/285 (53%), Gaps = 15/285 (5%)
Query: 25 NVTVIIMNKWIFQKLDFKFP-LSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRI 83
++ ++ +NKWI+ + + FP ++++ IHF+ + +G ++ K+ P S+ P +I
Sbjct: 25 SICIVFINKWIY--VHYGFPNMTLTLIHFVMTWLGLFICQKMDIFAPK-SLRPS----KI 77
Query: 84 FPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPI 143
++ FC +V N+SL+ + Q K T + +Q + +RK F +I +LVPI
Sbjct: 78 LLLALSFCGFVVFTNLSLQSNTIGTYQLAKVMTTPVIIAIQTMYYRKTFSTKIKLTLVPI 137
Query: 144 VGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF--A 201
G++L S ++ FN+ G A G L TS + A H + +S+ +Y+ AP A
Sbjct: 138 TLGVILNSYYDVRFNLMGMIFATLGVLVTSLYQVWVGAKQHELQVNSMQLLYYQAPMSSA 197
Query: 202 TLIMVFPAL-LLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTTAVTF 260
L+++ P L G+G + P+ + A+ ++ SGV+AF +N SI+++I +T+ VT+
Sbjct: 198 FLLVLVPFFEPLTGDGGI----FGPWSFLALFMVLLSGVIAFLVNLSIYWIIGNTSPVTY 253
Query: 261 NVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
N+ G+ K + +L +++F++P+S +G TL G Y + +
Sbjct: 254 NMFGHFKFCITLLGGYVLFQDPLSLNQGLGILCTLTGILAYTHFK 298
>UniRef100_UPI00002BACDF UPI00002BACDF UniRef100 entry
Length = 344
Score = 133 bits (334), Expect = 8e-30
Identities = 89/291 (30%), Positives = 146/291 (49%), Gaps = 7/291 (2%)
Query: 15 LLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISV 74
L A++ W T+ +NKWIF +F++PL +S +H + + + Y +IK L+L + V
Sbjct: 43 LSAVMVWLVTGSTISSLNKWIFAVFNFRYPLLLSALHMLTAMVVDYGLIK-LRLIRHVGV 101
Query: 75 DPQDRWR----RIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRK 130
QD ++F +S FC +I GNV L Y+ +SF Q I + TP T+ + LV K
Sbjct: 102 RQQDLTPGAKCKVFMLSLTFCASIAFGNVGLNYVQLSFAQMIYTTTPIFTLAISTLVLGK 161
Query: 131 YFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDS 190
+ +++PI G + + E+ F+ G + K+I LL K +S
Sbjct: 162 QHHILKYTAMMPICLGASFSIMGEVQFDQTGCFFVFAATMLRGVKSIQQSILLQEEKINS 221
Query: 191 INTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFY 250
+ +Y M+ + I+ AL LE +LEW +H Y + I S + + N +
Sbjct: 222 VFLLYLMSIPSFCILAVAALALENWALLEW-PLH-YDRRLWVFILLSCLGSVLYNLASCC 279
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
VI T+AVT ++ GNL V +L+S L+F + +S ++ G +TL G Y
Sbjct: 280 VISLTSAVTLHILGNLNVVGNLLLSQLLFGSELSTLSCAGAVLTLSGMLIY 330
>UniRef100_UPI000036626E UPI000036626E UniRef100 entry
Length = 307
Score = 130 bits (328), Expect = 4e-29
Identities = 88/291 (30%), Positives = 146/291 (49%), Gaps = 7/291 (2%)
Query: 15 LLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISV 74
L A++ W T+ +NKWIF +F++PL +S +H + + + Y +IK L++ I V
Sbjct: 19 LSAVVVWLVTGSTISSLNKWIFAVFNFRYPLLLSALHMLTAIVVDYGLIK-LRVVRHIGV 77
Query: 75 DPQDRWR----RIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRK 130
QD ++F +S FC +I GN+ L Y+ +SF Q I + TP T+ + LV K
Sbjct: 78 REQDLTPSAKCKVFMLSLTFCASIAFGNMGLNYVQLSFAQMIYTTTPIFTLAISTLVLGK 137
Query: 131 YFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDS 190
+ +++PI G + + E+ F+ G + K+I LL K +S
Sbjct: 138 QHHILKYTAMMPICLGASFSIMGEVQFDQTGCLFVFAATMLRGVKSIQQSILLQEEKINS 197
Query: 191 INTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFY 250
+ +Y M+ + I+ AL LE +LEW +H Y + I S + + N +
Sbjct: 198 VFLLYLMSIPSFCILAVAALALENWALLEW-PLH-YDRRLWLFILLSCLGSVLYNLASCC 255
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
VI T+AVT ++ GNL V +L+S L+F + +S ++ G +TL G Y
Sbjct: 256 VISLTSAVTLHILGNLNVVGNLLLSQLLFGSELSALSCAGAVLTLSGMFIY 306
>UniRef100_Q9SUV2 Hypothetical protein F8B4.90 [Arabidopsis thaliana]
Length = 350
Score = 127 bits (318), Expect = 6e-28
Identities = 76/300 (25%), Positives = 156/300 (51%), Gaps = 10/300 (3%)
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHF-ICSAIGAYVVIKVLK 67
++ S + W + TVI+ NK+I K ++ FP++++ IH CS++ A ++IKV K
Sbjct: 15 ILLSYTYVAIWIFLSFTVIVYNKYILDKKMYNWPFPITLTMIHMAFCSSL-AVILIKVFK 73
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
+ +S+ R + P+ ++ +++ L N + Y+ VSF+Q +K+ P + L+
Sbjct: 74 IVEPVSMSRDTYIRSVVPIGALYSLSLWLSNSAYIYLSVSFIQMLKALMPVAVYSIGVLL 133
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HG 185
++ F +++ I G+ + + E F+ +G L +T+ +L + LL G
Sbjct: 134 KKESFKSETMTNMLSISFGVAIAAYGEAKFDTWGVMLQLGAVAFEATRLVLIQILLTSKG 193
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ I ++Y++AP + + FP + +E + E S H +I ++ V AF LN
Sbjct: 194 INLNPITSLYYVAPCCLVFLFFPWIFVELPILRETSSFH----FDFVIFGTNSVCAFALN 249
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVR 305
++F ++ T+A+T NVAG +K + + SW + ++ ++ +N G + +G +Y + +
Sbjct: 250 LAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPLNLFGYGLAFLGVAYYNHCK 309
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.331 0.142 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 578,475,230
Number of Sequences: 2790947
Number of extensions: 22845685
Number of successful extensions: 77745
Number of sequences better than 10.0: 656
Number of HSP's better than 10.0 without gapping: 333
Number of HSP's successfully gapped in prelim test: 324
Number of HSP's that attempted gapping in prelim test: 76829
Number of HSP's gapped (non-prelim): 728
length of query: 342
length of database: 848,049,833
effective HSP length: 128
effective length of query: 214
effective length of database: 490,808,617
effective search space: 105033044038
effective search space used: 105033044038
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 75 (33.5 bits)
Medicago: description of AC139354.6


