

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.3 + phase: 0
(282 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
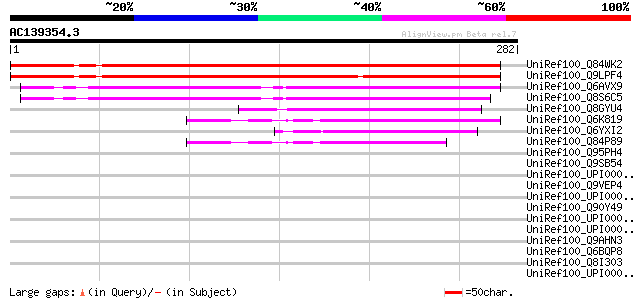
Sequences producing significant alignments: (bits) Value
UniRef100_Q84WK2 At1g44770 [Arabidopsis thaliana] 305 7e-82
UniRef100_Q9LPF4 T12C22.4 protein [Arabidopsis thaliana] 283 4e-75
UniRef100_Q6AVX9 Expressed protein [Oryza sativa] 163 5e-39
UniRef100_Q8S6C5 Putative growth factor [Oryza sativa] 161 2e-38
UniRef100_Q8GYU4 Hypothetical protein At4g24590/F22K18_210 [Arab... 78 3e-13
UniRef100_Q6K819 MADS box interactor-like [Oryza sativa] 74 3e-12
UniRef100_Q6YXI2 Hypothetical protein OJ1005_D12.16 [Oryza sativa] 58 2e-07
UniRef100_Q84P89 Hypothetical protein [Oryza sativa] 48 3e-04
UniRef100_Q95PH4 Histidine kinase DhkM [Dictyostelium discoideum] 42 0.023
UniRef100_Q9SB54 Hypothetical protein F22K18.210 [Arabidopsis th... 41 0.031
UniRef100_UPI00002FEF9E UPI00002FEF9E UniRef100 entry 40 0.089
UniRef100_Q9VEP4 CG5225-PA [Drosophila melanogaster] 40 0.089
UniRef100_UPI000042C67D UPI000042C67D UniRef100 entry 38 0.26
UniRef100_Q90Y49 POU domain transcription factor brn-1 [Ambystom... 38 0.26
UniRef100_UPI000036390A UPI000036390A UniRef100 entry 38 0.34
UniRef100_UPI0000363909 UPI0000363909 UniRef100 entry 38 0.34
UniRef100_Q9AHN3 DcbE [Pasteurella multocida] 38 0.34
UniRef100_Q6BQP8 Similar to sp|P22035 Saccharomyces cerevisiae M... 38 0.34
UniRef100_Q8I303 Hypothetical protein PFI0730w [Plasmodium falci... 37 0.44
UniRef100_UPI000042CEFB UPI000042CEFB UniRef100 entry 37 0.75
>UniRef100_Q84WK2 At1g44770 [Arabidopsis thaliana]
Length = 271
Score = 305 bits (782), Expect = 7e-82
Identities = 150/274 (54%), Positives = 198/274 (71%), Gaps = 6/274 (2%)
Query: 1 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
M +H+A+E+ KTVLEVADVAWTAVE +HHHH H D H+ D P D++LEAL
Sbjct: 1 MTSHHAIEVTKTVLEVADVAWTAVETYHHHHHHQDE--NHESTNPISD---PRDRELEAL 55
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 120
+ ENRRLR LL+ NLKL L+ES +F +CP DL+ RL V S ++L RL+ L++ +
Sbjct: 56 RQENRRLRTLLESNLKLFETLAESAAFSHDCPSDLYARLVTMVTSRDFLARLENLRQALS 115
Query: 121 SGG-NQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLII 179
+G NQFPFKE TE D ++ ++LI +D +EPSWWV V D+ P N EE S ID+E Y+++
Sbjct: 116 NGTQNQFPFKEPTEDDVKTVEVLIEMDHQEPSWWVLVTDDMVPSNVEEQSAIDNEHYIVV 175
Query: 180 SEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYA 239
+EEHV+D VA+F+A+CIMSNPKA+N+ PEELQ L + SK+ KV+DIW AGK+FY
Sbjct: 176 NEEHVIDAVAHFLAKCIMSNPKAKNLKPEELQKLLVQEVTALSKVGKVVDIWHAGKMFYT 235
Query: 240 LSTWGLALAGLYQTRSLLKVAAKGVHSGGKLALK 273
LSTWGLA GLYQ R +LK+AAKGVH+ K+ L+
Sbjct: 236 LSTWGLAFGGLYQARGVLKIAAKGVHATSKVVLR 269
>UniRef100_Q9LPF4 T12C22.4 protein [Arabidopsis thaliana]
Length = 269
Score = 283 bits (724), Expect = 4e-75
Identities = 143/274 (52%), Positives = 192/274 (69%), Gaps = 8/274 (2%)
Query: 1 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
M +H+A+E+ KTVLEVADVAWTAVE +HHHH H D H+ D P D++LEAL
Sbjct: 1 MTSHHAIEVTKTVLEVADVAWTAVETYHHHHHHQDE--NHESTNPISD---PRDRELEAL 55
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 120
+ ENRRLR LL+ NLKL L+ES +F +CP DL+ RL V S ++L RL+ L++ +
Sbjct: 56 RQENRRLRTLLESNLKLFETLAESAAFSHDCPSDLYARLVTMVTSRDFLARLENLRQALS 115
Query: 121 SGG-NQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLII 179
+G NQFPFKE TE D ++ ++LI +D +EPSWWV V D+ P N EE S ID+E Y+++
Sbjct: 116 NGTQNQFPFKEPTEDDVKTVEVLIEMDHQEPSWWVLVTDDMVPSNVEEQSAIDNEHYIVV 175
Query: 180 SEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYA 239
+EEHV+D VA+F+A+ + K +N+ PEELQ L + SK+ KV+DIW AGK+FY
Sbjct: 176 NEEHVIDAVAHFLAK--STGFKLQNLKPEELQKLLVQEVTALSKVGKVVDIWHAGKMFYT 233
Query: 240 LSTWGLALAGLYQTRSLLKVAAKGVHSGGKLALK 273
LSTWGLA GLYQ R +LK+AAKGVH+ K+ L+
Sbjct: 234 LSTWGLAFGGLYQARGVLKIAAKGVHATSKVVLR 267
>UniRef100_Q6AVX9 Expressed protein [Oryza sativa]
Length = 258
Score = 163 bits (412), Expect = 5e-39
Identities = 91/267 (34%), Positives = 149/267 (55%), Gaps = 17/267 (6%)
Query: 7 VEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRR 66
+E+A T++E+A+VA A+E H R H P +D ++ E L+ EN
Sbjct: 7 LEVAHTLVEIAEVARYAIE----HRRGHGPA------HDGVSPPAVDGEEAERLRAENVI 56
Query: 67 LRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETASGGNQF 126
LR L +L +L L CP DLH RL A V + +L +L+ +++E+ +
Sbjct: 57 LRARLADDLSILRELQGEPCVSQECPADLHNRLVAAVNNASFLAQLEKIRDESRHQQTEL 116
Query: 127 PFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVD 186
T++ Y K SW V VA + N EE SGID+E+Y++++++ ++D
Sbjct: 117 SPDNMTDIPYTEGG------GKNGSW-VLVACDKPGANMEEISGIDNENYVLVNDDDIID 169
Query: 187 GVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLA 246
G+ +F+ARCI+ +PK++++SP ELQ ++ A + + K + IW AGK+ Y L+TWG+
Sbjct: 170 GMTSFIARCILEDPKSKSISPVELQKAVAMALSTLNDKWKWMSIWEAGKVLYILATWGIT 229
Query: 247 LAGLYQTRSLLKVAAKGVHSGGKLALK 273
+ GLY++R +LK+AAKG K +K
Sbjct: 230 IVGLYRSRHVLKIAAKGAVVSAKFVMK 256
>UniRef100_Q8S6C5 Putative growth factor [Oryza sativa]
Length = 479
Score = 161 bits (407), Expect = 2e-38
Identities = 90/261 (34%), Positives = 147/261 (55%), Gaps = 17/261 (6%)
Query: 7 VEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRR 66
+E+A T++E+A+VA A+E H R H P +D ++ E L+ EN
Sbjct: 7 LEVAHTLVEIAEVARYAIE----HRRGHGPA------HDGVSPPAVDGEEAERLRAENVI 56
Query: 67 LRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETASGGNQF 126
LR L +L +L L CP DLH RL A V + +L +L+ +++E+ +
Sbjct: 57 LRARLADDLSILRELQGEPCVSQECPADLHNRLVAAVNNASFLAQLEKIRDESRHQQTEL 116
Query: 127 PFKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVD 186
T++ Y K SW V VA + N EE SGID+E+Y++++++ ++D
Sbjct: 117 SPDNMTDIPYTEGG------GKNGSW-VLVACDKPGANMEEISGIDNENYVLVNDDDIID 169
Query: 187 GVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLA 246
G+ +F+ARCI+ +PK++++SP ELQ ++ A + + K + IW AGK+ Y L+TWG+
Sbjct: 170 GMTSFIARCILEDPKSKSISPVELQKAVAMALSTLNDKWKWMSIWEAGKVLYILATWGIT 229
Query: 247 LAGLYQTRSLLKVAAKGVHSG 267
+ GLY++R +LK+AAKG G
Sbjct: 230 IVGLYRSRHVLKIAAKGAVMG 250
>UniRef100_Q8GYU4 Hypothetical protein At4g24590/F22K18_210 [Arabidopsis thaliana]
Length = 241
Score = 77.8 bits (190), Expect = 3e-13
Identities = 40/135 (29%), Positives = 70/135 (51%), Gaps = 5/135 (3%)
Query: 128 FKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVDG 187
F+E E+ S ++ DS W D + EEC G ++ Y+++ EE + DG
Sbjct: 99 FREKIELAQVSVPQVLAEDSSS-----WDVVSEDDLWDEECVGQTEDDYVVVREEDIADG 153
Query: 188 VANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLAL 247
+A FMA + S + +++SP++LQ LS F+ + K+ W K+ Y +++W
Sbjct: 154 IACFMATYLSSLKQTKDISPDQLQKALSTMFSVKKRKGKLRKAWEGSKVIYNVASWSATA 213
Query: 248 AGLYQTRSLLKVAAK 262
G+YQ +L +A+K
Sbjct: 214 IGIYQNPMILSIASK 228
>UniRef100_Q6K819 MADS box interactor-like [Oryza sativa]
Length = 265
Score = 74.3 bits (181), Expect = 3e-12
Identities = 50/177 (28%), Positives = 89/177 (50%), Gaps = 23/177 (12%)
Query: 99 LAATVRSDEYLTRLKVLQEETASGGNQFPFKEATEVDYQ-SADILINVDSKEPSWWVWVA 157
L + V S E+L +L +Q+ G VD S DI+ D +W
Sbjct: 108 LKSRVASPEFLEKLDNIQKSVYQNG---------AVDETISWDIISAAD-------IW-- 149
Query: 158 DETDP-INFEECSGIDDESYLIISEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSK 216
D+ D +N + S ++ Y++I +E +VDG+A+FMA ++S + ++++P +LQ LSK
Sbjct: 150 DDIDKGMNISDDS---EDGYVLIKQEDIVDGIASFMAAYLLSLKQTKDLTPNQLQQALSK 206
Query: 217 AFAGTSKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSLLKVAAKGVHSGGKLALK 273
F+ + K+ W K+ Y +++W G+YQ ++LK A + ++A K
Sbjct: 207 TFSAKKRKSKLQKAWDGTKVIYNIASWSATAIGIYQNPAILKAATAAFWTSCRVASK 263
>UniRef100_Q6YXI2 Hypothetical protein OJ1005_D12.16 [Oryza sativa]
Length = 257
Score = 58.2 bits (139), Expect = 2e-07
Identities = 30/113 (26%), Positives = 56/113 (49%), Gaps = 6/113 (5%)
Query: 148 KEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVDGVANFMARCIMSNPKARNMSP 207
++PSW D ++ E + D Y+++ + +G+A F+A I S A SP
Sbjct: 136 QDPSW-----DIVKAVDLWEDDDLGD-GYVLVKNDDATEGMAFFIATYISSLKTANECSP 189
Query: 208 EELQNNLSKAFAGTSKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSLLKVA 260
++++ L K F+ + K+ W K+ Y + +W G+Y +++LKVA
Sbjct: 190 DQIRKALKKTFSSRKRKGKLRKAWNGTKVIYNVDSWSATAIGIYHNQAILKVA 242
>UniRef100_Q84P89 Hypothetical protein [Oryza sativa]
Length = 377
Score = 48.1 bits (113), Expect = 3e-04
Identities = 39/147 (26%), Positives = 68/147 (45%), Gaps = 23/147 (15%)
Query: 99 LAATVRSDEYLTRLKVLQEETASGGNQFPFKEATEVDYQ-SADILINVDSKEPSWWVWVA 157
L + V S E+L +L +Q+ G VD S DI+ D +W
Sbjct: 164 LKSRVASPEFLEKLDNIQKSVYQNG---------AVDETISWDIISAAD-------IW-- 205
Query: 158 DETDP-INFEECSGIDDESYLIISEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSK 216
D+ D +N + S ++ Y++I E +VDG+A+ MA ++S + +++ P +LQ LSK
Sbjct: 206 DDIDKGMNISDDS---EDGYVLIKXEDIVDGIASXMAAYLLSLKQTKDLXPNQLQQALSK 262
Query: 217 AFAGTSKLEKVLDIWAAGKLFYALSTW 243
F + + K+ Y + +W
Sbjct: 263 TFXAKKRKXXLQKXXDGTKVIYNIXSW 289
>UniRef100_Q95PH4 Histidine kinase DhkM [Dictyostelium discoideum]
Length = 1318
Score = 41.6 bits (96), Expect = 0.023
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 3/55 (5%)
Query: 26 YHHHHHRHHDPVVTHDDKYD---DKDNNCPSDQDLEALKLENRRLRNLLDQNLKL 77
+HHHHH HH + DD YD D DNN + + ++++ D+NL+L
Sbjct: 176 HHHHHHHHHHHHIIDDDDYDDDNDDDNNTEDSSSCCNIDELSDKIKDNQDENLEL 230
>UniRef100_Q9SB54 Hypothetical protein F22K18.210 [Arabidopsis thaliana]
Length = 204
Score = 41.2 bits (95), Expect = 0.031
Identities = 23/79 (29%), Positives = 40/79 (50%), Gaps = 5/79 (6%)
Query: 128 FKEATEVDYQSADILINVDSKEPSWWVWVADETDPINFEECSGIDDESYLIISEEHVVDG 187
F+E E+ S ++ DS W D + EEC G ++ Y+++ EE + DG
Sbjct: 99 FREKIELAQVSVPQVLAEDSSS-----WDVVSEDDLWDEECVGQTEDDYVVVREEDIADG 153
Query: 188 VANFMARCIMSNPKARNMS 206
+A FMA + S + +++S
Sbjct: 154 IACFMATYLSSLKQTKDIS 172
>UniRef100_UPI00002FEF9E UPI00002FEF9E UniRef100 entry
Length = 197
Score = 39.7 bits (91), Expect = 0.089
Identities = 16/31 (51%), Positives = 18/31 (57%)
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSDQD 56
+HHHHH HHD DD DD N+ SD D
Sbjct: 3 HHHHHHHHHDGDDDDDDDDDDTGNDNDSDND 33
Score = 35.8 bits (81), Expect = 1.3
Identities = 14/31 (45%), Positives = 16/31 (51%)
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSDQD 56
+HHHHH HH DD DD D +D D
Sbjct: 1 HHHHHHHHHHHDGDDDDDDDDDDTGNDNDSD 31
>UniRef100_Q9VEP4 CG5225-PA [Drosophila melanogaster]
Length = 594
Score = 39.7 bits (91), Expect = 0.089
Identities = 38/141 (26%), Positives = 54/141 (37%), Gaps = 14/141 (9%)
Query: 27 HHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSESNS 86
HH H HH P T K + P D DL+ L + N++D+N NN E
Sbjct: 428 HHQHPHHHQPKPT---KPEGSGEQTPRD-DLDLLDENVEIVENIIDEN-DNENNFMEGEE 482
Query: 87 FLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETASGGNQFPFKEATEVDYQSADILINVD 146
+SN P DL EY V E+ SG A E+D + + ++
Sbjct: 483 EMSNEPNDLETEPEDADYGLEYADN-TVENPESPSG--------AEEIDDEDSSYMVPEG 533
Query: 147 SKEPSWWVWVADETDPINFEE 167
+ D+ DP+N E
Sbjct: 534 DDAMEEDHMMEDQMDPVNMME 554
>UniRef100_UPI000042C67D UPI000042C67D UniRef100 entry
Length = 1373
Score = 38.1 bits (87), Expect = 0.26
Identities = 12/21 (57%), Positives = 16/21 (76%)
Query: 21 WTAVEYHHHHHRHHDPVVTHD 41
+++V HHHHH HHDP+ HD
Sbjct: 1070 FSSVSGHHHHHHHHDPMAKHD 1090
>UniRef100_Q90Y49 POU domain transcription factor brn-1 [Ambystoma mexicanum]
Length = 403
Score = 38.1 bits (87), Expect = 0.26
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 8/52 (15%)
Query: 25 EYHHHHHRHH--------DPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLR 68
E+HHHHH+HH P H D + D+D D + A + + RR++
Sbjct: 187 EHHHHHHQHHHPHQHHPPPPQHHHQDPHSDEDTPTSDDLEQFAKQFKQRRIK 238
>UniRef100_UPI000036390A UPI000036390A UniRef100 entry
Length = 763
Score = 37.7 bits (86), Expect = 0.34
Identities = 21/71 (29%), Positives = 33/71 (45%)
Query: 5 NAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLEN 64
++V M + +L A++ HHHHH HH V+T D D ++ + +L +
Sbjct: 89 HSVIMPENILGTEVAIDEALDSHHHHHHHHHHVLTSDLIQDANHHHHDMTDQVFVAELLS 148
Query: 65 RRLRNLLDQNL 75
N LDQ L
Sbjct: 149 EHHDNTLDQQL 159
>UniRef100_UPI0000363909 UPI0000363909 UniRef100 entry
Length = 378
Score = 37.7 bits (86), Expect = 0.34
Identities = 21/71 (29%), Positives = 33/71 (45%)
Query: 5 NAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLEN 64
++V M + +L A++ HHHHH HH V+T D D ++ + +L +
Sbjct: 86 HSVIMPENILGTEVAIDEALDSHHHHHHHHHHVLTSDLIQDANHHHHDMTDQVFVAELLS 145
Query: 65 RRLRNLLDQNL 75
N LDQ L
Sbjct: 146 EHHDNTLDQQL 156
>UniRef100_Q9AHN3 DcbE [Pasteurella multocida]
Length = 603
Score = 37.7 bits (86), Expect = 0.34
Identities = 44/216 (20%), Positives = 94/216 (43%), Gaps = 20/216 (9%)
Query: 46 DKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRS 105
+K+ N +DLE +++E +L+ L++ N L E N LS DL+++ ++S
Sbjct: 324 EKEKNIKDKKDLE-IQIEALQLK--LEKYFIESNALKEDNKKLSIEIDDLNIKKEENIKS 380
Query: 106 --DEYLTRLKVLQEETASGGNQFPFKEATEVDYQSADILINVDSKEPSWWVWVADETDPI 163
E T +++ N +++ ++ + + D+K+ + + + D I
Sbjct: 381 LKKEIETLKAEIEKNHIENKNLKETNKSSLIEIEKLNREKEDDNKQKEKTITLLKKEDSI 440
Query: 164 NFEECSGIDDE--SYLIISEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGT 221
E ++E +Y II E+++ + K+ E LQ L K F
Sbjct: 441 KSNELDRKNEEIKNYKIIIEDNIKE-------------KKSLENQVEALQFELEKYFIEN 487
Query: 222 SKLEKVLDIWAAGKLFYALSTWGLALAGLYQTRSLL 257
K+++ +W A K+ + + L + ++RS++
Sbjct: 488 KKIKEKPPLWGASKVIRSTLEYQLGRTMIEKSRSII 523
>UniRef100_Q6BQP8 Similar to sp|P22035 Saccharomyces cerevisiae MYB-like DNA-binding
protein BAS1 [Debaryomyces hansenii]
Length = 744
Score = 37.7 bits (86), Expect = 0.34
Identities = 17/40 (42%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Query: 26 YHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKLENR 65
+HHHHH HH ++K D KD + + DL +KL NR
Sbjct: 481 HHHHHHHHHHDNENENEKQDRKDPSNSKESDL--IKLLNR 518
>UniRef100_Q8I303 Hypothetical protein PFI0730w [Plasmodium falciparum]
Length = 709
Score = 37.4 bits (85), Expect = 0.44
Identities = 17/48 (35%), Positives = 29/48 (60%), Gaps = 5/48 (10%)
Query: 43 KYDDKDNNCPSD-----QDLEALKLENRRLRNLLDQNLKLLNNLSESN 85
K D++ N CPSD +D+ +K+EN + + + N K L+ +SE+N
Sbjct: 512 KEDNEQNICPSDSNIYLKDISKIKIENNDFQKISEYNTKFLDEISEAN 559
>UniRef100_UPI000042CEFB UPI000042CEFB UniRef100 entry
Length = 437
Score = 36.6 bits (83), Expect = 0.75
Identities = 32/127 (25%), Positives = 55/127 (43%), Gaps = 11/127 (8%)
Query: 28 HHHHRHHDP---VVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKLLNNLSES 84
HHHH H DP V++ D Y + S N N + N NN S S
Sbjct: 121 HHHHHHKDPGCVGVSNSDVYSIVNGEKKSHYHHHHRNDNNNNNNNNNNNNNNNNNNNSNS 180
Query: 85 NSFLSNCP----PDLHVRL---AATVRSDEYLTRLKVLQEETASGGNQFPFKEATEVDYQ 137
NSF CP D ++ + + T S+ Y + + ++ + G + P E++E+ +
Sbjct: 181 NSFSFTCPSEGNSDSNIIIPINSQTCESNNY-NNMHYEESDSKNLGIKSPLIESSELYNE 239
Query: 138 SADILIN 144
++++ N
Sbjct: 240 KSNMINN 246
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.315 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 462,293,802
Number of Sequences: 2790947
Number of extensions: 18925565
Number of successful extensions: 85478
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 84856
Number of HSP's gapped (non-prelim): 264
length of query: 282
length of database: 848,049,833
effective HSP length: 126
effective length of query: 156
effective length of database: 496,390,511
effective search space: 77436919716
effective search space used: 77436919716
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 74 (33.1 bits)
Medicago: description of AC139354.3


