

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.10 - phase: 0
(381 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
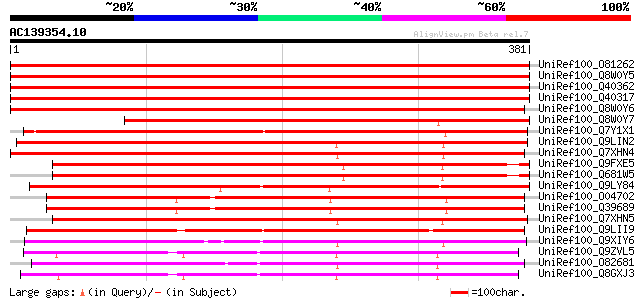
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O81262 Early nodule-specific protein [Medicago truncat... 779 0.0
UniRef100_Q8W0Y5 Enod8.1 [Medicago truncatula] 777 0.0
UniRef100_Q40362 Early nodulin [Medicago sativa] 714 0.0
UniRef100_Q40317 Nodulin [Medicago sativa] 711 0.0
UniRef100_Q8W0Y6 Enod8.2 [Medicago truncatula] 656 0.0
UniRef100_Q8W0Y7 Enod8.3 [Medicago truncatula] 500 e-140
UniRef100_Q7Y1X1 Esterase precursor [Hevea brasiliensis] 474 e-132
UniRef100_Q9LIN2 Nodulin-like protein protein [Arabidopsis thali... 392 e-107
UniRef100_Q7XHN4 Putative early nodulin 8 [Oryza sativa] 381 e-104
UniRef100_Q9FXE5 F12A21.4 [Arabidopsis thaliana] 368 e-100
UniRef100_Q681W5 ENOD8-like protein [Arabidopsis thaliana] 368 e-100
UniRef100_Q9LY84 Putative early nodule-specific protein [Arabido... 362 1e-98
UniRef100_O04702 IEP4 [Daucus carota] 359 6e-98
UniRef100_Q39689 Secreted glycoprotein precursor [Daucus carota] 359 8e-98
UniRef100_Q7XHN5 Putative early nodulin 8 [Oryza sativa] 350 3e-95
UniRef100_Q9LII9 Arabidopsis thaliana genomic DNA, chromosome 3,... 321 2e-86
UniRef100_Q9XIY6 Putative lanatoside 15'-O-acetylesterase [Oryza... 294 3e-78
UniRef100_Q9ZVL5 T22H22.20 protein [Arabidopsis thaliana] 285 1e-75
UniRef100_O82681 Lanatoside 15'-O-acetylesterase precursor [Digi... 282 1e-74
UniRef100_Q8GXJ3 Hypothetical protein At1g54790/T22H22_20 [Arabi... 272 1e-71
>UniRef100_O81262 Early nodule-specific protein [Medicago truncatula]
Length = 381
Score = 779 bits (2011), Expect = 0.0
Identities = 381/381 (100%), Positives = 381/381 (100%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP
Sbjct: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK
Sbjct: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND
Sbjct: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS
Sbjct: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF
Sbjct: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
Query: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS
Sbjct: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
Query: 361 QILTGVFNDPPISLDRACYRK 381
QILTGVFNDPPISLDRACYRK
Sbjct: 361 QILTGVFNDPPISLDRACYRK 381
>UniRef100_Q8W0Y5 Enod8.1 [Medicago truncatula]
Length = 381
Score = 777 bits (2007), Expect = 0.0
Identities = 380/381 (99%), Positives = 381/381 (99%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP
Sbjct: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK
Sbjct: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND
Sbjct: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
LTIGFFGNKTIQQVNATVPDIVNNYI+NIKNIYNLGARSFWIHGTGPKGCAPVILANFPS
Sbjct: 181 LTIGFFGNKTIQQVNATVPDIVNNYIKNIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF
Sbjct: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
Query: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS
Sbjct: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
Query: 361 QILTGVFNDPPISLDRACYRK 381
QILTGVFNDPPISLDRACYRK
Sbjct: 361 QILTGVFNDPPISLDRACYRK 381
>UniRef100_Q40362 Early nodulin [Medicago sativa]
Length = 381
Score = 714 bits (1843), Expect = 0.0
Identities = 342/381 (89%), Positives = 363/381 (94%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MA+IELSRH+PLVTLIVLVLC TPPIFA+ +CDFPAIFSFGASNVDTGGLAAAFRAPPSP
Sbjct: 1 MARIELSRHMPLVTLIVLVLCTTPPIFASTHCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
YG+TYF+RSTGRFSDGRII+DFIA+SFRLPY SPYLNSLGSNFTHGANFA+ GSTINIP
Sbjct: 61 YGQTYFNRSTGRFSDGRIIIDFIAQSFRLPYPSPYLNSLGSNFTHGANFATAGSTINIPT 120
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
SILP G LSPFSLQIQYIQFK+FISKTKLIRDQGGVFATL+PKEDYFSKALY+FDIGQND
Sbjct: 121 SILPKGILSPFSLQIQYIQFKDFISKTKLIRDQGGVFATLVPKEDYFSKALYVFDIGQND 180
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
LTIGFFGNKTIQQVNATVPD+VNNYIENIKNIYNLGARSFWIH TGPKGCAPVILANFPS
Sbjct: 181 LTIGFFGNKTIQQVNATVPDLVNNYIENIKNIYNLGARSFWIHSTGPKGCAPVILANFPS 240
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
AIKDSYGCAKQYNEVSQYFN KLKEALA+LRS+L AAITYVDIY+PKYSLFTNP+KYGF
Sbjct: 241 AIKDSYGCAKQYNEVSQYFNLKLKEALAQLRSDLPLAAITYVDIYSPKYSLFTNPKKYGF 300
Query: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
ELP+VACCGYGGEYNIG GCGA+IN+NGTKIVAGSCKNPSTRI WDG HYTEAAN+IVF
Sbjct: 301 ELPYVACCGYGGEYNIGAGCGATINVNGTKIVAGSCKNPSTRITWDGTHYTEAANKIVFD 360
Query: 361 QILTGVFNDPPISLDRACYRK 381
QI TG FNDPPISLD ACYRK
Sbjct: 361 QISTGAFNDPPISLDMACYRK 381
>UniRef100_Q40317 Nodulin [Medicago sativa]
Length = 381
Score = 711 bits (1836), Expect = 0.0
Identities = 341/381 (89%), Positives = 362/381 (94%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MA+IELSRH+PLVTLIVLVLC TPPIFA+ +CDFPAIFSFGASNVDTGGLAAAFRAPPSP
Sbjct: 1 MARIELSRHMPLVTLIVLVLCTTPPIFASTHCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
YG+TYF+RSTGRFSDGRII+DFIA+SFRLPY SPYLNSLGSNFTHGANFA+ GSTINIP
Sbjct: 61 YGQTYFNRSTGRFSDGRIIIDFIAQSFRLPYPSPYLNSLGSNFTHGANFATAGSTINIPT 120
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
SILP G LSPFSLQIQYIQFK+FISKTKLIRDQGGVFATL+PKEDYFSKALY+FDIGQND
Sbjct: 121 SILPKGILSPFSLQIQYIQFKDFISKTKLIRDQGGVFATLVPKEDYFSKALYVFDIGQND 180
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
LTIGFFGNKTIQQVNATVPD+VNNYIENIKNIYNLGARSFWIH TGPKGCAPVILANFPS
Sbjct: 181 LTIGFFGNKTIQQVNATVPDLVNNYIENIKNIYNLGARSFWIHSTGPKGCAPVILANFPS 240
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
AIKDSYGCAKQYNEVSQYFN KLKEALA+LRS+L AAITYVDIY+PKYSLFTNP+KYGF
Sbjct: 241 AIKDSYGCAKQYNEVSQYFNLKLKEALAQLRSDLPLAAITYVDIYSPKYSLFTNPKKYGF 300
Query: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
ELP+VACCGYGGEYNIG GCGA+IN+NGTKIVAGSCKNPSTRI WDG HYTE AN+ VF
Sbjct: 301 ELPYVACCGYGGEYNIGAGCGATINVNGTKIVAGSCKNPSTRITWDGTHYTEEANKFVFY 360
Query: 361 QILTGVFNDPPISLDRACYRK 381
QI TGVFNDPPISLD ACYRK
Sbjct: 361 QISTGVFNDPPISLDMACYRK 381
>UniRef100_Q8W0Y6 Enod8.2 [Medicago truncatula]
Length = 385
Score = 656 bits (1693), Expect = 0.0
Identities = 319/379 (84%), Positives = 343/379 (90%), Gaps = 1/379 (0%)
Query: 1 MAKIELSRHIPLVTLIVLVLCI-TPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
M K EL RH+ LV+LIVL+LCI TPPIFAT+NCDFPAIFSFGASNVDTGGLAAAF+APPS
Sbjct: 1 MEKFELLRHMSLVSLIVLILCIITPPIFATRNCDFPAIFSFGASNVDTGGLAAAFQAPPS 60
Query: 60 PYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIP 119
PYGETYFHRSTGRFSDGRIILDFIA+SF LPYLSPYLNSLGSNFTHGANFA+GGSTINIP
Sbjct: 61 PYGETYFHRSTGRFSDGRIILDFIAQSFGLPYLSPYLNSLGSNFTHGANFATGGSTINIP 120
Query: 120 KSILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQN 179
SI+PNG SPFSLQIQYIQFK+FISKT LIRDQGGVFATLIPKEDYFSKALY FDIGQN
Sbjct: 121 NSIIPNGIFSPFSLQIQYIQFKDFISKTNLIRDQGGVFATLIPKEDYFSKALYTFDIGQN 180
Query: 180 DLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP 239
DL G+FGNKTI+QVNATVPDIVNN+I NIKNIYNLGARSFWIH T P GC P ILANFP
Sbjct: 181 DLIGGYFGNKTIKQVNATVPDIVNNFIVNIKNIYNLGARSFWIHSTVPSGCTPTILANFP 240
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
SAIKDSYGCAKQYNEVSQYFN KLK+ALA+LR +L AAITYVDIY+P YSLF NP+KYG
Sbjct: 241 SAIKDSYGCAKQYNEVSQYFNLKLKKALAQLRVDLPLAAITYVDIYSPNYSLFQNPKKYG 300
Query: 300 FELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVF 359
FELP VACCGYGG+YNI VGCG ++NINGTKI AGSCKNPSTRIIWDG H+TE +IVF
Sbjct: 301 FELPHVACCGYGGKYNIRVGCGETLNINGTKIEAGSCKNPSTRIIWDGSHFTERRYKIVF 360
Query: 360 SQILTGVFNDPPISLDRAC 378
QI TG F+DPPISL+RAC
Sbjct: 361 DQISTGAFSDPPISLNRAC 379
>UniRef100_Q8W0Y7 Enod8.3 [Medicago truncatula]
Length = 299
Score = 500 bits (1288), Expect = e-140
Identities = 241/299 (80%), Positives = 259/299 (86%), Gaps = 2/299 (0%)
Query: 85 RSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFI 144
+SF LPYLSPYLNSLGSNFTHGANFA+ GSTI IP SI+PNG SPFSLQIQ IQFK+FI
Sbjct: 1 QSFGLPYLSPYLNSLGSNFTHGANFATAGSTIKIPNSIIPNGMFSPFSLQIQSIQFKDFI 60
Query: 145 SKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNN 204
K K IRDQGGVFATLIPKEDY+SKALY FDIGQNDLT GFFGNKTIQQVN TVPDIV +
Sbjct: 61 PKAKFIRDQGGVFATLIPKEDYYSKALYTFDIGQNDLTAGFFGNKTIQQVNTTVPDIVKS 120
Query: 205 YIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQYNEVSQYFNFKLK 264
+I+NIKNIYNLGARSFWIH TGP GC P+ILANFPSAIKD YGCAKQYNEVSQYFN KLK
Sbjct: 121 FIDNIKNIYNLGARSFWIHNTGPIGCVPLILANFPSAIKDRYGCAKQYNEVSQYFNLKLK 180
Query: 265 EALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGE--YNIGVGCGA 322
EALA+LR +L AAITYVDIY+PKYSLF NP+KYGFELP VACCG GG+ YNI GCGA
Sbjct: 181 EALAQLRKDLPLAAITYVDIYSPKYSLFQNPKKYGFELPLVACCGNGGKYNYNIRAGCGA 240
Query: 323 SININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYRK 381
+ININGT V GSCK PSTRIIWDG HYTEAAN+IVF QI G F DPPI L+RACY+K
Sbjct: 241 TININGTNTVVGSCKKPSTRIIWDGTHYTEAANKIVFDQISNGAFTDPPIPLNRACYKK 299
>UniRef100_Q7Y1X1 Esterase precursor [Hevea brasiliensis]
Length = 391
Score = 474 bits (1219), Expect = e-132
Identities = 229/374 (61%), Positives = 286/374 (76%), Gaps = 6/374 (1%)
Query: 11 PLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRST 70
P++TL L LC+ +A++ CDFPAIF+FG SN DTGG AAAF PYGET+FHRST
Sbjct: 10 PIITLSFL-LCMLSLAYASETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRST 68
Query: 71 GRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILP-NGKLS 129
GR+SDGR+I+DFIA SF LPYLSPYL+SLGSNF HGA+FA+ GSTI +P +I+P +G S
Sbjct: 69 GRYSDGRLIIDFIAESFNLPYLSPYLSSLGSNFKHGADFATAGSTIKLPTTIIPAHGGFS 128
Query: 130 PFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNK 189
PF L +QY QF++FI +++ IR+ GG+FA L+P+E YF KALY FDIGQNDLT GF N
Sbjct: 129 PFYLDVQYSQFRQFIPRSQFIRETGGIFAELVPEEYYFEKALYTFDIGQNDLTEGFL-NL 187
Query: 190 TIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCA 249
T+++VNATVPD+VN++ N+K IY+LGAR+FWIH TGP GC IL FP A KDS GCA
Sbjct: 188 TVEEVNATVPDLVNSFSANVKKIYDLGARTFWIHNTGPIGCLSFILTYFPWAEKDSAGCA 247
Query: 250 KQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCG 309
K YNEV+Q+FN KLKE +A+LR +L A +VDIY+ KYSLF+ PEK+GFE P + CCG
Sbjct: 248 KAYNEVAQHFNHKLKEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEFPLITCCG 307
Query: 310 YGGEYNIGV--GCGASINI-NGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGV 366
YGG+YN V CG ++ +GTKIV GSC PS R+ WDG HYTEAANE F QI TG
Sbjct: 308 YGGKYNFSVTAPCGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAANEYFFDQISTGA 367
Query: 367 FNDPPISLDRACYR 380
F+DPP+ L+ AC++
Sbjct: 368 FSDPPVPLNMACHK 381
>UniRef100_Q9LIN2 Nodulin-like protein protein [Arabidopsis thaliana]
Length = 380
Score = 392 bits (1006), Expect = e-107
Identities = 186/380 (48%), Positives = 260/380 (67%), Gaps = 5/380 (1%)
Query: 6 LSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETY 65
+ ++ LV ++L C+ P + +C+FPAIF+FG SN DTGGL+A+F P P G+T+
Sbjct: 1 METNLLLVKCVLLASCLIHPRACSPSCNFPAIFNFGDSNSDTGGLSASFGQAPYPNGQTF 60
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FH +GRFSDGR+I+DFIA LPYL+ +L+S+GSNF+HGANFA+ GST+ P + +
Sbjct: 61 FHSPSGRFSDGRLIIDFIAEELGLPYLNAFLDSIGSNFSHGANFATAGSTVRPPNATIAQ 120
Query: 126 GKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
+SP SL +Q +QF +FI++++LIR++GGVF L+PK++YFS+ALY FDIGQNDLT G
Sbjct: 121 SGVSPISLDVQLVQFSDFITRSQLIRNRGGVFKKLLPKKEYFSQALYTFDIGQNDLTAGL 180
Query: 186 FGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--PSAIK 243
N T Q+ A +PD+ + I+ +Y+ G R FWIH T P GC P +L F P++
Sbjct: 181 KLNMTSDQIKAYIPDVHDQLSNVIRKVYSKGGRRFWIHNTAPLGCLPYVLDRFPVPASQI 240
Query: 244 DSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELP 303
D++GCA NE+++Y+N +LK + ELR LS AA TYVDIY+ K +L T +K GF P
Sbjct: 241 DNHGCAIPRNEIARYYNSELKRRVIELRKELSEAAFTYVDIYSIKLTLITQAKKLGFRYP 300
Query: 304 FVACCGYGGEYNIG--VGCGASININGTKIV-AGSCKNPSTRIIWDGVHYTEAANEIVFS 360
VACCG+GG+YN + CGA + I G +IV A SC + S R+ WDG+H+TE N +F
Sbjct: 301 LVACCGHGGKYNFNKLIKCGAKVMIKGKEIVLAKSCNDVSFRVSWDGIHFTETTNSWIFQ 360
Query: 361 QILTGVFNDPPISLDRACYR 380
QI G F+DPP+ + AC R
Sbjct: 361 QINDGAFSDPPLPVKSACTR 380
>UniRef100_Q7XHN4 Putative early nodulin 8 [Oryza sativa]
Length = 391
Score = 381 bits (979), Expect = e-104
Identities = 189/385 (49%), Positives = 256/385 (66%), Gaps = 7/385 (1%)
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MA + + + T+ VLVL + A +C FPA+F+FG SN DTGGL+A F A P P
Sbjct: 1 MAPPPYAAAVVVATVAVLVLVSQVSVAAGADCRFPAVFNFGDSNSDTGGLSATFGAAPPP 60
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
G T+F GR+ DGR+++DFIA S LPYLS YLNS+GSNFT GANFA+ GS+I
Sbjct: 61 NGRTFFGMPVGRYCDGRLVIDFIAESLGLPYLSAYLNSIGSNFTQGANFATAGSSIRRQN 120
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLI-RDQGGVFATLIPKEDYFSKALYIFDIGQN 179
+ L SP SL +Q +F++FI++++ + ++GG++ L+PK +YFS+ALY FDIGQN
Sbjct: 121 TSLFLSGFSPISLDVQSWEFEQFINRSQFVYNNKGGIYRELLPKAEYFSQALYTFDIGQN 180
Query: 180 DLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP 239
D+T GFF N T +QV A +PD++ I+N+Y LG R FWIH TGP GC P + + P
Sbjct: 181 DITTGFFINMTSEQVIAYIPDLMERLTNIIQNVYGLGGRYFWIHNTGPIGCLPYAMVHRP 240
Query: 240 --SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEK 297
+ +KD GC+ YNEV+Q FN +LKE + LR + AA TYVD+Y+ KY L ++ +K
Sbjct: 241 DLAVVKDGSGCSVAYNEVAQLFNQRLKETVGHLRKTHADAAFTYVDVYSAKYKLISDAKK 300
Query: 298 YGFELPFVACCGY-GGEYNIG--VGCGASININGTKIVAG-SCKNPSTRIIWDGVHYTEA 353
G + P + CCGY GG YN VGCG + +NGT +VAG SC +P R+ WDGVH+TEA
Sbjct: 301 LGMDDPMLTCCGYGGGRYNFDDRVGCGGKVKVNGTWVVAGKSCDDPLKRVSWDGVHFTEA 360
Query: 354 ANEIVFSQILTGVFNDPPISLDRAC 378
AN+ VF QI G +DPP+ L +AC
Sbjct: 361 ANKFVFDQIAGGKLSDPPVPLRQAC 385
>UniRef100_Q9FXE5 F12A21.4 [Arabidopsis thaliana]
Length = 372
Score = 368 bits (944), Expect = e-100
Identities = 174/355 (49%), Positives = 241/355 (67%), Gaps = 13/355 (3%)
Query: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
C FPAIF+FG SN DTGGL+AAF P+G ++F GR+ DGR+++DFIA S LPY
Sbjct: 26 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 85
Query: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
LS +L+S+GSNF+HGANFA+ GS I S L SPFSL +Q++QF F ++++ +R
Sbjct: 86 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTVR 145
Query: 152 DQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKN 211
+GGV+ T++P+ D FSKALY FDIGQNDLT G+F NKT++QV VP+I++ ++ IKN
Sbjct: 146 SRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAIKN 205
Query: 212 IYNLGARSFWIHGTGPKGCAPVILANFPSAIK--DSYGCAKQYNEVSQYFNFKLKEALAE 269
IY G R FWIH TGP GC ++ FP+ DS+GC N ++Q FN LK+A+ E
Sbjct: 206 IYGQGGRYFWIHNTGPIGCLAYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQAVIE 265
Query: 270 LRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNI--GVGCGASININ 327
LRS+LS AAITYVD+Y+ K+ LF + + +GF+ V+CCG+GG+YN G+GCG +
Sbjct: 266 LRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKKIVK 325
Query: 328 GTKIVAGS-CKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYRK 381
G ++ G C P ++WDGVH+T+AAN+ +F +I G L +AC R+
Sbjct: 326 GKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKRQ 372
>UniRef100_Q681W5 ENOD8-like protein [Arabidopsis thaliana]
Length = 364
Score = 368 bits (944), Expect = e-100
Identities = 174/355 (49%), Positives = 241/355 (67%), Gaps = 13/355 (3%)
Query: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
C FPAIF+FG SN DTGGL+AAF P+G ++F GR+ DGR+++DFIA S LPY
Sbjct: 18 CHFPAIFNFGDSNSDTGGLSAAFGQAGPPHGSSFFGSPAGRYCDGRLVIDFIAESLGLPY 77
Query: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
LS +L+S+GSNF+HGANFA+ GS I S L SPFSL +Q++QF F ++++ +R
Sbjct: 78 LSAFLDSVGSNFSHGANFATAGSPIRALNSTLRQSGFSPFSLDVQFVQFYNFHNRSQTVR 137
Query: 152 DQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKN 211
+GGV+ T++P+ D FSKALY FDIGQNDLT G+F NKT++QV VP+I++ ++ IKN
Sbjct: 138 SRGGVYKTMLPESDSFSKALYTFDIGQNDLTAGYFANKTVEQVETEVPEIISQFMNAIKN 197
Query: 212 IYNLGARSFWIHGTGPKGCAPVILANFPSAIK--DSYGCAKQYNEVSQYFNFKLKEALAE 269
IY G R FWIH TGP GC ++ FP+ DS+GC N ++Q FN LK+A+ E
Sbjct: 198 IYGQGGRYFWIHNTGPIGCLAYVIERFPNKASDFDSHGCVSPLNHLAQQFNHALKQAVIE 257
Query: 270 LRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNI--GVGCGASININ 327
LRS+LS AAITYVD+Y+ K+ LF + + +GF+ V+CCG+GG+YN G+GCG +
Sbjct: 258 LRSSLSEAAITYVDVYSLKHELFVHAQGHGFKGSLVSCCGHGGKYNYNKGIGCGMKKIVK 317
Query: 328 GTKIVAGS-CKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYRK 381
G ++ G C P ++WDGVH+T+AAN+ +F +I G L +AC R+
Sbjct: 318 GKEVYIGKPCDEPDKAVVWDGVHFTQAANKFIFDKIAPG--------LSKACKRQ 364
>UniRef100_Q9LY84 Putative early nodule-specific protein [Arabidopsis thaliana]
Length = 389
Score = 362 bits (929), Expect = 1e-98
Identities = 179/372 (48%), Positives = 238/372 (63%), Gaps = 7/372 (1%)
Query: 15 LIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFS 74
L++ + +T + C FPAI++FG SN DTGG++AAF PYG+ +FHR TGR S
Sbjct: 20 LLLCLFAVTTSVSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDPYGQGFFHRPTGRDS 79
Query: 75 DGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQ 134
DGR+ +DFIA LPYLS YLNSLGSNF HGANFA+GGSTI + +SPFSL
Sbjct: 80 DGRLTIDFIAERLGLPYLSAYLNSLGSNFRHGANFATGGSTIRRQNETIFQYGISPFSLD 139
Query: 135 IQYIQFKEFISKTKLIRDQ--GGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQ 192
+Q QF +F +++ L+ Q +P+++ F+KALY FDIGQNDL++G F ++
Sbjct: 140 MQIAQFDQFKARSALLFTQIKSRYDREKLPRQEEFAKALYTFDIGQNDLSVG-FRTMSVD 198
Query: 193 QVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPV---ILANFPSAIKDSYGCA 249
Q+ AT+PDIVN+ ++NIY G R+FW+H TGP GC PV + D GC
Sbjct: 199 QLKATIPDIVNHLASAVRNIYQQGGRTFWVHNTGPFGCLPVNMFYMGTPAPGYLDKSGCV 258
Query: 250 KQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCG 309
K NE++ FN KLKE + LR L+ AAITYVD+YT KY + +NP+K GF P CCG
Sbjct: 259 KAQNEMAMEFNRKLKETVINLRKELTQAAITYVDVYTAKYEMMSNPKKLGFANPLKVCCG 318
Query: 310 YGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFND 369
Y +Y+ + CG +N T+I GSC NP + WDGVHYTEAAN+ V + L G+ D
Sbjct: 319 YHEKYD-HIWCGNKGKVNNTEIYGGSCPNPVMAVSWDGVHYTEAANKHVADRTLNGLLTD 377
Query: 370 PPISLDRACYRK 381
PP+ + RACYR+
Sbjct: 378 PPVPITRACYRQ 389
>UniRef100_O04702 IEP4 [Daucus carota]
Length = 391
Score = 359 bits (922), Expect = 6e-98
Identities = 175/360 (48%), Positives = 248/360 (68%), Gaps = 12/360 (3%)
Query: 28 ATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSF 87
+++ CDFPAIF+FG +N DTG AA F P +G++YF+ S GR SDGR+++DF+A
Sbjct: 24 SSQTCDFPAIFNFGDANSDTGAFAAWFFGNPPFFGQSYFNGSAGRVSDGRLLIDFMATDL 83
Query: 88 RLPYLSPYLNSLGSNFTHGANFASGGSTINIPKS-----ILPNGKLSPFSLQIQYIQFKE 142
LP+L PY++SLG+NF+HGANFA+ STI +P S + P L+P +L IQ QF +
Sbjct: 84 GLPFLHPYMDSLGANFSHGANFANILSTIALPTSNIIPGVRPPRGLNPVNLDIQVAQFAQ 143
Query: 143 FISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIV 202
F++++ + QG F +PK+DYFS+ALY DIGQ D+T F NKT ++ A VP ++
Sbjct: 144 FVNRS---QTQGEAFDNFMPKQDYFSQALYTLDIGQIDITQEFLTNKTDDEIKAVVPGLI 200
Query: 203 NNYIENIKNIYNLGARSFWIHGTGPKGCAPVI--LANFPSAIKDSYGCAKQYNEVSQYFN 260
++ NI+ +Y+LG RSFWIH GP GC P++ LA P DS GCAK+YN+++QYFN
Sbjct: 201 SSLSSNIQILYSLGGRSFWIHNLGPNGCLPILLTLAPVPDDQLDSAGCAKRYNDLTQYFN 260
Query: 261 FKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVG- 319
+LK+ + +LR++L AA+TYVD+YT KYSL+ P KYGF P CCG+GG YN G
Sbjct: 261 SELKKGVDQLRTDLPLAAVTYVDVYTAKYSLYQEPAKYGFTHPLETCCGFGGRYNYGEFS 320
Query: 320 -CGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRAC 378
CG++I +NGT++ G C+NP+ I ++G YT+AA++I F++I TG +DPP SL AC
Sbjct: 321 LCGSTITVNGTQLTVGPCENPAEYINYEGQTYTQAADQITFNKISTGELSDPPNSLKTAC 380
>UniRef100_Q39689 Secreted glycoprotein precursor [Daucus carota]
Length = 383
Score = 359 bits (921), Expect = 8e-98
Identities = 175/360 (48%), Positives = 248/360 (68%), Gaps = 12/360 (3%)
Query: 28 ATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSF 87
+++ CDFPAIF+FG +N DTG AA F P +G++YF+ S GR SDGR+++DF+A
Sbjct: 16 SSQTCDFPAIFNFGDANSDTGAFAAWFFGNPPFFGQSYFNGSAGRVSDGRLLIDFMATDL 75
Query: 88 RLPYLSPYLNSLGSNFTHGANFASGGSTINIPKS-----ILPNGKLSPFSLQIQYIQFKE 142
LP+L PY++SLG+NF+HGANFA+ STI +P S + P L+P +L IQ QF +
Sbjct: 76 GLPFLHPYMDSLGANFSHGANFANILSTIALPTSNIIPGVRPPRGLNPVNLDIQVAQFAQ 135
Query: 143 FISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIV 202
F++++ + QG F +PK+DYFS+ALY DIGQ D+T F NKT ++ A VP ++
Sbjct: 136 FVNRS---QTQGEAFDNFMPKQDYFSQALYTLDIGQIDITQEFLTNKTDDEIKAVVPGLI 192
Query: 203 NNYIENIKNIYNLGARSFWIHGTGPKGCAPVI--LANFPSAIKDSYGCAKQYNEVSQYFN 260
++ NI+ +Y+LG RSFWIH GP GC P++ LA P DS GCAK+YN+++QYFN
Sbjct: 193 SSLSSNIQILYSLGGRSFWIHNLGPNGCLPILWTLAPVPDDQIDSAGCAKRYNDLTQYFN 252
Query: 261 FKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVG- 319
+LK+ + +LR++L AA+TYVD+YT KYSL+ P KYGF P CCG+GG YN G
Sbjct: 253 SELKKGVDQLRTDLPLAAVTYVDVYTAKYSLYQEPAKYGFTHPLETCCGFGGRYNYGEFS 312
Query: 320 -CGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRAC 378
CG++I +NGT++ G C+NP+ I ++G YT+AA++I F++I TG +DPP SL AC
Sbjct: 313 LCGSTITVNGTQLAVGPCENPAEYINYEGQTYTQAADQITFNKISTGELSDPPNSLKTAC 372
>UniRef100_Q7XHN5 Putative early nodulin 8 [Oryza sativa]
Length = 405
Score = 350 bits (899), Expect = 3e-95
Identities = 167/356 (46%), Positives = 238/356 (65%), Gaps = 7/356 (1%)
Query: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
CDFPAIF+FG SN DTGGL+A P P+G TYF GRFSDGR+ +DF+A+S + Y
Sbjct: 45 CDFPAIFNFGDSNSDTGGLSALIAVVPPPFGRTYFGMPAGRFSDGRLTIDFMAQSLGIRY 104
Query: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
LS YL+S+GSNF+ GANFA+ ++I + +SP SL +Q QF++FI++++ +
Sbjct: 105 LSAYLDSVGSNFSQGANFATAAASIRPANGSIFVSGISPISLDVQTSQFEQFINRSQFVY 164
Query: 152 DQ-GGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIK 210
GG++ ++PK +YFS+ALY FDIGQNDLT+G+F N + +QV A VPD++ + I+
Sbjct: 165 SNIGGIYREILPKAEYFSRALYTFDIGQNDLTMGYFDNMSTEQVEAYVPDLMERFSAAIQ 224
Query: 211 NIYNLGARSFWIHGTGPKGCAPVILANFP--SAIKDSYGCAKQYNEVSQYFNFKLKEALA 268
+Y+LG R FW+H T P GC + P +A +D GC+ YN +++FN +L+E +
Sbjct: 225 KVYSLGGRYFWVHNTAPLGCLTYAVVLLPKLAAPRDDAGCSVAYNAAARFFNARLRETVD 284
Query: 269 ELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGY-GGEYNI--GVGCGASIN 325
LR+ L AA+TYVD+Y+ KY L + ++ GF P + CCGY GGEYN + CG +
Sbjct: 285 RLRAALPDAALTYVDVYSAKYRLISQAKQLGFGDPLLVCCGYGGGEYNFDRDIRCGGKVE 344
Query: 326 INGTKIVAG-SCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYR 380
+NGT ++AG SC +PS + WDGVH+TEAAN VF I+ G +DPP+ L +AC R
Sbjct: 345 VNGTSVLAGKSCDDPSRSVSWDGVHFTEAANRFVFELIVGGKLSDPPVPLRQACRR 400
>UniRef100_Q9LII9 Arabidopsis thaliana genomic DNA, chromosome 3, TAC clone:K24A2
[Arabidopsis thaliana]
Length = 371
Score = 321 bits (822), Expect = 2e-86
Identities = 168/367 (45%), Positives = 230/367 (61%), Gaps = 9/367 (2%)
Query: 13 VTLIVLVLCITPPIFA-TKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTG 71
+ +IVL+L T + A + +C+FPA+F+FG SN DTG ++AA P P G +F RS G
Sbjct: 8 LAIIVLLLGFTEKLSALSSSCNFPAVFNFGDSNSDTGAISAAIGEVPPPNGVAFFGRSAG 67
Query: 72 RFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPF 131
R SDGR+I+DFI + LPYL+PYL+S+G+N+ HGANFA+GGS I + SPF
Sbjct: 68 RHSDGRLIIDFITENLTLPYLTPYLDSVGANYRHGANFATGGSCIRPTLACF-----SPF 122
Query: 132 SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTI 191
L Q QF F ++T + +Q + +YFSKALY DIGQNDL IGF N T
Sbjct: 123 HLGTQVSQFIHFKTRTLSLYNQTNGKFNRLSHTNYFSKALYTLDIGQNDLAIGF-QNMTE 181
Query: 192 QQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQ 251
+Q+ AT+P I+ N+ +K +Y GAR F IH TGP GC P +L FP+ +D YGC K
Sbjct: 182 EQLKATIPLIIENFTIALKLLYKEGARFFSIHNTGPTGCLPYLLKAFPAIPRDPYGCLKP 241
Query: 252 YNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYG 311
N V+ FN +LK + +L+ L S+ TYVD+Y+ KY+L T + GF PF CC
Sbjct: 242 LNNVAIEFNKQLKNKITQLKKELPSSFFTYVDVYSAKYNLITKAKALGFIDPFDYCC--V 299
Query: 312 GEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPP 371
G G+GCG +I +NGT++ + SC+N I WDG+HYTE AN +V ++IL G +DPP
Sbjct: 300 GAIGRGMGCGKTIFLNGTELYSSSCQNRKNFISWDGIHYTETANMLVANRILDGSISDPP 359
Query: 372 ISLDRAC 378
+ +AC
Sbjct: 360 LPTQKAC 366
>UniRef100_Q9XIY6 Putative lanatoside 15'-O-acetylesterase [Oryza sativa]
Length = 379
Score = 294 bits (752), Expect = 3e-78
Identities = 154/373 (41%), Positives = 216/373 (57%), Gaps = 10/373 (2%)
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTG 71
L T + C+ AT C FPA+F+FG SN DTGG AAF A +P+G TYF R G
Sbjct: 7 LATAAAALCCLVAAASATAQCRFPAVFNFGDSNSDTGGFWAAFPAQQAPFGMTYFRRPAG 66
Query: 72 RFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPF 131
R SDGR+++DF+ ++ LP LSPYL S+GS + HGANFA+ ST P + L +SPF
Sbjct: 67 RASDGRLVVDFLVQAMGLPLLSPYLQSVGSGYRHGANFATLASTALQPNTSLFVTGISPF 126
Query: 132 SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTI 191
L +Q Q KE +TK++ G +P D ALY DIGQNDLT G+++I
Sbjct: 127 FLAVQLNQMKEL--RTKVLTSNGN--NDQLPAPDVLHNALYTIDIGQNDLTSN-LGSQSI 181
Query: 192 QQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP--SAIKDSYGCA 249
+ V ++P +V+ ++ +YN+GAR+ + P GC P L P S D YGC
Sbjct: 182 ETVKQSLPSVVSKISSTVQELYNIGARNIMVFNMAPIGCYPAFLTKLPHTSNDMDGYGCM 241
Query: 250 KQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCG 309
K YN Y+N L +LAE+R L A+I Y+D + LF +P+ +G + ACCG
Sbjct: 242 KTYNSAVTYYNELLNNSLAEVRKKLQDASIVYLDKHAVTLELFRHPKAHGLKYGTKACCG 301
Query: 310 YG-GEYNIG--VGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGV 366
YG G YN V CG+S +NG + A +C +P + WDG+H TEAAN+I+ S +++G
Sbjct: 302 YGDGAYNFNPDVYCGSSKLLNGQTVTAKACADPQNYVSWDGIHATEAANKIIASSLMSGS 361
Query: 367 FNDPPISLDRACY 379
++ PP L + C+
Sbjct: 362 YSYPPFDLSKLCH 374
>UniRef100_Q9ZVL5 T22H22.20 protein [Arabidopsis thaliana]
Length = 383
Score = 285 bits (729), Expect = 1e-75
Identities = 157/377 (41%), Positives = 223/377 (58%), Gaps = 21/377 (5%)
Query: 11 PLVTLIVLVLCITPPIFATKNCD----FPAIFSFGASNVDTGGL-AAAFRAPPSPYGETY 65
P+V + +VL I +F + +PAI +FG SN DTG L +A PYG+TY
Sbjct: 3 PIVNNLSIVLFILISLFLPSSFSIILKYPAIINFGDSNSDTGNLISAGIENVNPPYGQTY 62
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLG-SNFTHGANFASGGSTINIPKSILP 124
F+ +GR+ DGR+I+DF+ LP+L+PYL+SLG NF G NFA+ GSTI LP
Sbjct: 63 FNLPSGRYCDGRLIVDFLLDEMDLPFLNPYLDSLGLPNFKKGCNFAAAGSTI------LP 116
Query: 125 NG--KLSPFSLQIQYIQFKEFISKT-KLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDL 181
+SPFS +Q QF F S+ +L+ G + +P DY+SK LY+ DIGQND+
Sbjct: 117 ANPTSVSPFSFDLQISQFIRFKSRAIELLSKTGRKYEKYLPPIDYYSKGLYMIDIGQNDI 176
Query: 182 TIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--P 239
G F +KT+ QV A++P I+ + +K +Y G R+ WIH TGP GC +A F
Sbjct: 177 A-GAFYSKTLDQVLASIPSILETFEAGLKRLYEEGGRNIWIHNTGPLGCLAQNIAKFGTD 235
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
S D +GC +N+ ++ FN +L + ++ A +TYVDI++ K +L N ++G
Sbjct: 236 STKLDEFGCVSSHNQAAKLFNLQLHAMSNKFQAQYPDANVTYVDIFSIKSNLIANYSRFG 295
Query: 300 FELPFVACCGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANE 356
FE P +ACCG GG Y+ + CG + ++G + A +C + S I WDG+HYTEAANE
Sbjct: 296 FEKPLMACCGVGGAPLNYDSRITCGQTKVLDGISVTAKACNDSSEYINWDGIHYTEAANE 355
Query: 357 IVFSQILTGVFNDPPIS 373
V SQILTG ++DPP S
Sbjct: 356 FVSSQILTGKYSDPPFS 372
>UniRef100_O82681 Lanatoside 15'-O-acetylesterase precursor [Digitalis lanata]
Length = 386
Score = 282 bits (721), Expect = 1e-74
Identities = 158/368 (42%), Positives = 211/368 (56%), Gaps = 9/368 (2%)
Query: 17 VLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDG 76
V+VL + CDF AIF+FG SN DTGG AAF A P G TYF R GR +DG
Sbjct: 16 VVVLMEVSVRSSESKCDFKAIFNFGDSNSDTGGFWAAFPAENPPNGMTYFKRPAGRVTDG 75
Query: 77 RIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQ 136
R+I+DF+A+ +P+LSPYL S+GS+F HGANFA+ ST+ +P++ L +SPFSL IQ
Sbjct: 76 RLIIDFLAQGIGIPFLSPYLLSIGSDFRHGANFATAASTVLLPRTSLFVTGVSPFSLGIQ 135
Query: 137 YIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQV-N 195
Q K+F + + G +P D F K+LY IGQND T G G+ I V
Sbjct: 136 LNQMKQFKLQVDRLHHSPGKLN--LPAPDIFRKSLYTLYIGQNDFT-GNLGSLGISGVKK 192
Query: 196 ATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP--SAIKDSYGCAKQYN 253
+P +V+ IK +Y LG R+F + P GC P+ L + P S+ DS+GC YN
Sbjct: 193 KIIPQVVSQISSTIKKLYELGGRTFLVLNLAPIGCYPLFLVDLPHNSSDIDSFGCMISYN 252
Query: 254 EVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGG- 312
+ +N+ LKEALA+ R ++ A + Y DI++ LF +P G + ACCGYGG
Sbjct: 253 KAVVEYNYMLKEALAQTRKDIQDADVIYTDIHSVMLQLFQHPTSNGLKYGTKACCGYGGG 312
Query: 313 --EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDP 370
+N V C S ING + A +CK+P + WDG+H TEAAN+ V IL G DP
Sbjct: 313 AFNFNQQVFCSYSKLINGKNVTANACKDPQNYVSWDGIHATEAANKHVAHAILEGSHFDP 372
Query: 371 PISLDRAC 378
P SL + C
Sbjct: 373 PFSLHKLC 380
>UniRef100_Q8GXJ3 Hypothetical protein At1g54790/T22H22_20 [Arabidopsis thaliana]
Length = 382
Score = 272 bits (696), Expect = 1e-71
Identities = 152/377 (40%), Positives = 218/377 (57%), Gaps = 19/377 (5%)
Query: 9 HIPLVTLIVLVLCITPPIFATKNCDF--PAIFSFGASNVDTGGLAAAFRAPPS-PYGETY 65
+I + L ++L + + + DF P+ F+FG SN DTG L A P G+
Sbjct: 2 NITKMKLFYVILFFISSLQISNSIDFNYPSAFNFGDSNSDTGDLVAGLGIRLDLPNGQNS 61
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLG-SNFTHGANFASGGSTINIPKSILP 124
F S+ RF DGR+++DF+ LP+L+PYL+SLG NF G NFA+ GSTI LP
Sbjct: 62 FKTSSQRFCDGRLVIDFLMDEMDLPFLNPYLDSLGLPNFKKGCNFAAAGSTI------LP 115
Query: 125 NG--KLSPFSLQIQYIQFKEFISKT-KLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDL 181
+SPFS +Q QF F S+ +L+ G + +P DY+SK LY+ DIGQND+
Sbjct: 116 ANPTSVSPFSFDLQISQFIRFKSRAIELLSKTGRKYEKYLPPIDYYSKGLYMIDIGQNDI 175
Query: 182 TIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF--P 239
G F +KT+ QV A++P I+ + +K +Y G R+ WIH TGP GC +A F
Sbjct: 176 A-GAFYSKTLDQVLASIPSILETFEAGLKRLYEEGGRNIWIHNTGPLGCLAQNIAKFGTD 234
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
S D +GC +N+ ++ FN +L + ++ A +TYVDI++ K +L N ++G
Sbjct: 235 STKLDEFGCVSSHNQAAKLFNLQLHAMSNKFQAQYPDANVTYVDIFSIKSNLIANYSRFG 294
Query: 300 FELPFVACCGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANE 356
FE P +ACCG GG Y+ + CG + ++G + A +C + S I WDG+HYTEAANE
Sbjct: 295 FEKPLMACCGVGGAPLNYDSRITCGQTKVLDGISVTAKACNDSSEYINWDGIHYTEAANE 354
Query: 357 IVFSQILTGVFNDPPIS 373
V SQILTG ++DPP S
Sbjct: 355 FVSSQILTGKYSDPPFS 371
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 673,730,149
Number of Sequences: 2790947
Number of extensions: 29661973
Number of successful extensions: 67122
Number of sequences better than 10.0: 337
Number of HSP's better than 10.0 without gapping: 276
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 65683
Number of HSP's gapped (non-prelim): 388
length of query: 381
length of database: 848,049,833
effective HSP length: 129
effective length of query: 252
effective length of database: 488,017,670
effective search space: 122980452840
effective search space used: 122980452840
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC139354.10


