

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138133.3 - phase: 0 /pseudo
(430 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
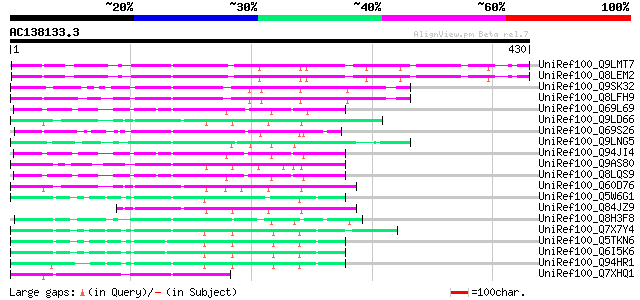
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana] 96 2e-18
UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana] 96 2e-18
UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana] 86 2e-15
UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana] 86 3e-15
UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [... 77 9e-13
UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein ... 77 1e-12
UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [... 77 1e-12
UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana] 76 2e-12
UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa] 75 5e-12
UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa] 75 5e-12
UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa] 75 5e-12
UniRef100_Q60D76 Putative polyprotein [Oryza sativa] 74 8e-12
UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa] 65 4e-09
UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa] 61 7e-08
UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like p... 59 3e-07
UniRef100_Q7X7Y4 OSJNBa0014F04.28 protein [Oryza sativa] 57 8e-07
UniRef100_Q5TKN6 Hypothetical protein OJ1058_F05.1 [Oryza sativa] 56 2e-06
UniRef100_Q6I5K6 Hypothetical protein OSJNBb0088F07.13 [Oryza sa... 56 2e-06
UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sat... 56 2e-06
UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sa... 55 3e-06
>UniRef100_Q9LMT7 F2H15.15 protein [Arabidopsis thaliana]
Length = 478
Score = 95.9 bits (237), Expect = 2e-18
Identities = 111/448 (24%), Positives = 187/448 (40%), Gaps = 75/448 (16%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLRLLG-------KLPKGELS 135
Query: 62 GY-ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFLA 180
+I + YL D + Y GA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLENYPVAARWK---HQKGHGEGITYRSLLDRIQLDDVCWRPYEKHR 237
H ++D +P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 Q-----MFVDFGTHTLKADARGEQAGEDTWRVAD--GYVLWYARVSHPQILPPIPRDLPR 346
+ + D T + R E E+ D Y+ W+ ++ +P + RD
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGVVAYADAQLGQEEVMSMTPQ----QWFEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ W+EA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
Query: 403 HMREQIAPILTRRRAQRPRRRHHQQDQD 430
T +R +RR QQ D
Sbjct: 462 ----------TNKR----KRREEQQQTD 475
>UniRef100_Q8LEM2 Hypothetical protein [Arabidopsis thaliana]
Length = 478
Score = 95.9 bits (237), Expect = 2e-18
Identities = 111/448 (24%), Positives = 187/448 (40%), Gaps = 75/448 (16%)
Query: 2 PMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYG 61
P GEMT+TLD+V+ + L ++G+ + G K + + + + LG K+ E
Sbjct: 84 PCGEMTITLDEVSLILGLAVDGKPVV-GVKEKDEDPSQVCLKLLG-------KLPKGELS 135
Query: 62 GY-ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
G ++ L++ + A + +E+E Y R YL+Y+VG +F
Sbjct: 136 GNRVTAKWLKESFAECPKGATM-------KEIE-------YHTRAYLIYIVGSTIFATTD 181
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFLA 180
+I + YL D + Y GA LA+LY ++ +A + +GG +TLL W
Sbjct: 182 PSKISVDYLILFED-FEKAGEYAWGAAALAFLYRQIGNASQRSQSIIGGCLTLLQCWSYF 240
Query: 181 HFPGFFSVDLNTDYLENYPVAARWK---HQKGHGEGITYRSLLDRIQLDDVCWRPYEKHR 237
H ++D +P+A WK + + YR LD + +V W P+E
Sbjct: 241 H----LNIDRPKRTTRQFPLALLWKGRQQSRSKNDLFKYRKALDDLDPSNVSWCPFEGDL 296
Query: 238 EI--QDFEE--VFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDLPPPSIV 293
+I Q F++ + S + G + V H P+R ++Q+G Q IP ++PP
Sbjct: 297 DIVPQSFKDNLLLGRSRTKLIGPKVVEWHFPDRCMKQFGLCQVIP------GEVPPRKNE 350
Query: 294 Q-----MFVDFGTHTLKADARGEQAGEDTWRVAD--GYVLWYARVSHPQILPPIPRDLPR 346
+ + D T + R E E+ D Y+ W+ ++ +P + RD
Sbjct: 351 KNHDEDLLEDMNTADEEWMRRRENIVENGGGNGDESEYMQWFNSIT----VPKLHRDTSL 406
Query: 347 PANEEQIIAEQWQRYEARSSPDTYDMVSGVVAYADAQLGQEEVMSMTPQ----QWFEAMT 402
A+ + A Q E S+ D+ + + +E VMSM+ W+EA T
Sbjct: 407 EADIMNVQAAILQFDEVASTLSLEDLHP-----EEREAIEEAVMSMSNALRVGDWYEAST 461
Query: 403 HMREQIAPILTRRRAQRPRRRHHQQDQD 430
T +R +RR QQ D
Sbjct: 462 ----------TNKR----KRREEQQQTD 475
>UniRef100_Q9SK32 Expressed protein [Arabidopsis thaliana]
Length = 509
Score = 85.9 bits (211), Expect = 2e-15
Identities = 90/351 (25%), Positives = 146/351 (40%), Gaps = 52/351 (14%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P+GEMT+TLD+VA + L I+G + K G + M G + N+E
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIVGSK-----VGDEVAMDMCGRLLGKLPSAANKE- 145
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRCYLLYLVGCLLFGDR 119
++ R++ ++L R +E PE+ ++ + T R YLLYL+G +F
Sbjct: 146 ---VNCSRVK---LNWLKR----TFSECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFL 179
++ + YL D + Y GA LA LY L +A + G +TLL W
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW-- 249
Query: 180 AHFPGFFSVDLNTDYLEN--YPVAARWKHQ--KGHGEGITYRSLLDRIQLDDVCWRPYEK 235
+F +D+ +P+A WK + + + YR LD + + W PYE+
Sbjct: 250 ----SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYER 305
Query: 236 HREI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP---------RHPT 282
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 306 FENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIPLKVKRRDRKNRRL 365
Query: 283 DVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVADG-YVLWYARVS 332
D D + + + G H + + G V DG Y+ WYAR+S
Sbjct: 366 DDLDTSMSLACEEWAERGDHIVDSPGGGNV-------VDDGAYMEWYARIS 409
>UniRef100_Q8LFH9 Hypothetical protein [Arabidopsis thaliana]
Length = 509
Score = 85.5 bits (210), Expect = 3e-15
Identities = 90/351 (25%), Positives = 143/351 (40%), Gaps = 52/351 (14%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P+GEMT+TLD+VA + L I+G + G K+ + LG + A K N
Sbjct: 92 LPLGEMTITLDEVALVLGLEIDGDPIV-GSKVDDEVAMDMCGRLLGKLPSAANKEVN--- 147
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELE-ELARVRTYCVRCYLLYLVGCLLFGDR 119
S +L ++ +E PE+ ++ + T R YLLYL+G +F
Sbjct: 148 ---CSRVKLNWLKRTF---------SECPEDASFDVVKCHT---RAYLLYLIGSTIFATT 192
Query: 120 SNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFL 179
++ + YL D + Y GA LA LY L +A + G +TLL W
Sbjct: 193 DGDKVSVKYLPLFED-FDQAGRYAWGAAALACLYRALGNASLKSQSNICGCLTLLQCW-- 249
Query: 180 AHFPGFFSVDLNTDYLEN--YPVAARWKHQ--KGHGEGITYRSLLDRIQLDDVCWRPYEK 235
+F +D+ +P+A WK + + + YR LD + + W PYE+
Sbjct: 250 ----SYFHLDIGRPEKSEACFPLALLWKGKGSRSKTDLSEYRRELDDLDPSKITWCPYER 305
Query: 236 HREI----QDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP---------RHPT 282
+ + + S + ++ H P+R LRQ+G Q IP
Sbjct: 306 FENLIPPHIKAKLILGRSKTTLVCFEKIELHFPDRCLRQFGKRQPIPLKVKRRDRKNRRL 365
Query: 283 DVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVADG-YVLWYARVS 332
D D + + + G H + + G V DG Y+ WYAR+S
Sbjct: 366 DDLDTSMSLACEEWAERGDHIVDSPGGGNV-------VDDGAYMEWYARIS 409
>UniRef100_Q69L69 Calcineurin-like phosphoesterase-like protein [Oryza sativa]
Length = 766
Score = 77.0 bits (188), Expect = 9e-13
Identities = 88/296 (29%), Positives = 126/296 (41%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYGGY 63
GEMTVTL DV+ L LPI+GR + H+ A +++ LG + +
Sbjct: 153 GEMTVTLQDVSMLLALPIDGRPVC---STTDHDYAQMVIDCLGHDPRGP----SMPGKSF 205
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + E PE ++ R VR Y+L L+ +LF D + R
Sbjct: 206 LHYKWLKKHF------------YELPEGTDDQTVERH--VRAYILSLLCGVLFPDGTG-R 250
Query: 124 IELIYLTTMADGYAGMRNYF*GAMTLAYLYGELAD-ACRPGHRALGGSVTLLTAWF---- 178
+ LIYL +AD + + Y G+ LA+LY L A + +GGS+ LL W
Sbjct: 251 MSLIYLPLIAD-LSLVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 309
Query: 179 -----LAHFPGFFSVDLNTDYLENYPVAARWKHQKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D L PV RW + E T YR L+ + D V
Sbjct: 310 HVGRPLVRSPVCPETDIPQDVL---PVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 366
Query: 229 CWRPYEKHREIQDFEEV------FWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 367 KWEPY-LHIESASLPLLCTKNADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 421
>UniRef100_Q9LD66 Similar to maize transposon MuDR mudrA protein [Oryza sativa]
Length = 1230
Score = 76.6 bits (187), Expect = 1e-12
Identities = 94/336 (27%), Positives = 136/336 (39%), Gaps = 49/336 (14%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRML----AHGKKMPRHEGAALLMTYLGVSQNEAEKIC 56
+P GE+TVTL+DVA + LPI G+ + A G R E YLG+ A
Sbjct: 592 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRERVE------EYLGLEPPVAPDGQ 645
Query: 57 NQEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLF 116
Q + LR + + P E +E A V+ YC R Y+LY+ G +LF
Sbjct: 646 RQTKTSGVPLSWLRANF------------GQCPAEADE-ATVQRYC-RAYVLYIFGSILF 691
Query: 117 GDRSNKRIELIYLTTMAD-GYAGMRNYF*GAMTLAYLYGELADACR--PGHRALGGSVTL 173
D ++L +AD AG ++ G+ LA+LY +L DACR G L G V L
Sbjct: 692 PDSGGDMASWMWLPLLADWDEAGTFSW--GSAALAWLYRQLCDACRRQGGDANLAGCVWL 749
Query: 174 LTAWFLAHFP------GFFSVDLNTDYLENYPVAARWKHQKGHGEG------ITYRSLLD 221
L W P + D VA W+ G Y + +D
Sbjct: 750 LQVWMWMRLPVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYYHYTNEMD 809
Query: 222 RIQLDDVCWRPYEKHREIQ-------DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYV 274
+Q + V W PY+ ++ E+ + V + RV+RQ+G +
Sbjct: 810 YLQPEHVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRL 869
Query: 275 QTIPRHPTDVRDLPPP-SIVQMFVDFGTHTLKADAR 309
QTIP + DL +I +F HT + D R
Sbjct: 870 QTIPHRFSTSIDLHNHFNIYYIFELDILHTCRVDRR 905
>UniRef100_Q69S26 Serine/threonine phosphatase PP7-related-like [Oryza sativa]
Length = 619
Score = 76.6 bits (187), Expect = 1e-12
Identities = 80/283 (28%), Positives = 126/283 (44%), Gaps = 37/283 (13%)
Query: 5 EMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYGGYI 64
EM +L DV+ + +P+ G ++ + + + + +LG S E++C G I
Sbjct: 96 EMAPSLRDVSYILGIPVTGHVVT-AEPIGDEAVRRMCLHFLGESPGNGEQLC-----GLI 149
Query: 65 SYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKRI 124
RL Y + E+P + E+A Y R YLLYLVG LF D +
Sbjct: 150 ---RLTWLYRKFHQLP------ENPT-INEIA----YSTRAYLLYLVGSTLFPDTMRGFV 195
Query: 125 ELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRA-LGGSVTLLTAWFLAHFP 183
YL +AD + +R Y G+ LA+LY L+ A P GS TLL AW + P
Sbjct: 196 SPRYLPLLAD-FRKIREYAWGSAALAHLYRGLSVAVTPNATTQFLGSATLLMAWIYEYLP 254
Query: 184 GFFSVDLNTDYLENYPVAARWKH---QKGHGEGI-TYRSLLDRIQLDDVCWRPYEKHRE- 238
N + L P A RW +G + + +R + D +Q DV W PY+
Sbjct: 255 LTQPQQKNQNTL--LPRACRWNFGGATRGQRKKVMEWRKVFDELQFSDVNWNPYKDMNPA 312
Query: 239 -IQDF----EEVFWYSGWIMC-GVRRVYRHLPERVLRQYGYVQ 275
I ++ + + + W++ ++ VY +P+R RQ+G Q
Sbjct: 313 IIPEYCIAADNICYSRTWLISFNIKEVY--VPDRFSRQFGREQ 353
>UniRef100_Q9LNG5 F21D18.16 [Arabidopsis thaliana]
Length = 1340
Score = 75.9 bits (185), Expect = 2e-12
Identities = 94/353 (26%), Positives = 142/353 (39%), Gaps = 62/353 (17%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GE+TVTL DV L L ++G + K + A L LG + +
Sbjct: 101 LPAGEITVTLQDVNILLGLRVDGPAVTGSTK---YNWADLCEDLLGHRPGPKDL-----H 152
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYC-VRCYLLYLVGCLLFGDR 119
G ++S LR+ + + DP+E V C R ++L L+ L+GD+
Sbjct: 153 GSHVSLAWLRENFRNL---------PADPDE------VTLKCHTRAFVLALMSGFLYGDK 197
Query: 120 SNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAWFL 179
S + L +L + D + + G+ TLA LY EL A + + G + LL W
Sbjct: 198 SKHDVALTFLPLLRD-FDEVAKLSWGSATLALLYRELCRASKRTVSTICGPLVLLQLWAW 256
Query: 180 AHF----PGFFSVDLNTDYLENY------PVAARWKHQKGHGEGIT-----YRSLLDRIQ 224
PG D+ Y++ P+ RW+ H E YR D+ +
Sbjct: 257 ERLHVGRPGRLK-DVGASYMDGIDGPLPDPLGCRWRASLSHKENPRGGLDFYRDQFDQQK 315
Query: 225 LDDVCWRPYE-----KHREIQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPR 279
+ V W+PY K I E W + + V H P+RVLRQ+G QTIP
Sbjct: 316 DEQVIWQPYTPDLLAKIPLICVSGENIWRTVAPLICFDVVEWHRPDRVLRQFGLHQTIPA 375
Query: 280 HPTDVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVADGYVLWYARVS 332
+ + L H + D RG+ + + R + LW ARVS
Sbjct: 376 PCDNEKAL--------------HAI--DKRGKSEYDWSARHSRHIGLWEARVS 412
>UniRef100_Q94JI4 P0638D12.5 protein [Oryza sativa]
Length = 761
Score = 74.7 bits (182), Expect = 5e-12
Identities = 87/296 (29%), Positives = 124/296 (41%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYGGY 63
GEM VTL DVA L LPI+GR + H+ A +++ LG + +
Sbjct: 170 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLGHDPRGP----SMPGKSF 222
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + E PE ++ R VR Y+L L+ +LF D + R
Sbjct: 223 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 267
Query: 124 IELIYLTTMADGYAGMRNYF*GAMTLAYLYGELAD-ACRPGHRALGGSVTLLTAWF---- 178
+ LIYL +AD + + Y G+ LA+LY L A + +GGS+ LL W
Sbjct: 268 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 326
Query: 179 -----LAHFPGFFSVDLNTDYLENYPVAARWKHQKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D PV RW + E T YR L+ + D V
Sbjct: 327 HVGRPLVRSPLCPETDIPQDL---PPVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 383
Query: 229 CWRPYEKHREIQDF------EEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 384 KWEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 438
>UniRef100_Q9AS80 P0028E10.12 protein [Oryza sativa]
Length = 792
Score = 74.7 bits (182), Expect = 5e-12
Identities = 85/302 (28%), Positives = 122/302 (40%), Gaps = 48/302 (15%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
MP GE+T+TL DVA + LPI G + P++E L+ YLG +A
Sbjct: 278 MPCGEITITLQDVAMILGLPIAGHAVTVNPTEPQNE---LVERYLG----KAPPPDRPRP 330
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
G +S+ R N D E +++ A R Y+L L+ LLF D S
Sbjct: 331 GLRVSWVR---------AEFNNCPEDADEETIKQHA-------RAYILSLISGLLFPDAS 374
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACR--PGHRALGGSVTLLTAWF 178
+AD + +Y G+ TLAYLY + DACR L G + LL W
Sbjct: 375 GDLYTFYPFPLIAD-LENIGSYSWGSATLAYLYRAMCDACRRQSDGSNLTGCLLLLQFWS 433
Query: 179 LAHFP---------GFFSVDLNTDYLENYPVAARW---KHQKGHGEGITYRSLLDRIQL- 225
HFP + +V+ D + V RW + Y + L
Sbjct: 434 WEHFPIGRPDLVKLKYPNVEELEDERDRPTVGLRWVVGVCARRAAPARCYEHFTNEFDLL 493
Query: 226 --DDVCWRPY--EKHREIQ-----DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQT 276
D V W PY E+ +E+Q + W + + V ++PERV+RQ+G Q
Sbjct: 494 TDDQVVWCPYREERMKELQLAPICTQDSHLWLTRAPLLYFFMVEIYMPERVMRQFGLHQV 553
Query: 277 IP 278
P
Sbjct: 554 CP 555
>UniRef100_Q8LQS9 P0702H08.15 protein [Oryza sativa]
Length = 718
Score = 74.7 bits (182), Expect = 5e-12
Identities = 87/296 (29%), Positives = 124/296 (41%), Gaps = 48/296 (16%)
Query: 4 GEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYGGY 63
GEM VTL DVA L LPI+GR + H+ A +++ LG + +
Sbjct: 184 GEMAVTLQDVAMLLALPIDGRPVC---STTDHDYAQMVIDCLGHDPRGP----SMPGKSF 236
Query: 64 ISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKR 123
+ Y L+ + E PE ++ R VR Y+L L+ +LF D + R
Sbjct: 237 LHYKWLKKHF------------YELPEGADDQTVERH--VRAYILSLLCGVLFPDGTG-R 281
Query: 124 IELIYLTTMADGYAGMRNYF*GAMTLAYLYGELAD-ACRPGHRALGGSVTLLTAWF---- 178
+ LIYL +AD + + Y G+ LA+LY L A + +GGS+ LL W
Sbjct: 282 MSLIYLPLIAD-LSRVGTYSWGSAVLAFLYRSLCSVASSHNIKNIGGSLLLLQLWSWERS 340
Query: 179 -----LAHFPGFFSVDLNTDYLENYPVAARWKHQKGHGEGIT-----YRSLLDRIQLDDV 228
L P D+ D PV RW + E T YR L+ + D V
Sbjct: 341 HVGRPLVRSPLCPETDIPQDV---PPVGFRWVGARTQSENATRCLKQYRDELNLQRADQV 397
Query: 229 CWRPYEKHREIQDF------EEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
W PY H E + W + + V +LPERV+RQ+G Q+IP
Sbjct: 398 KWEPY-LHIESSSLPLLCTKDADLWLTQAPLINFPIVEMYLPERVMRQFGLRQSIP 452
>UniRef100_Q60D76 Putative polyprotein [Oryza sativa]
Length = 1754
Score = 73.9 bits (180), Expect = 8e-12
Identities = 89/313 (28%), Positives = 130/313 (41%), Gaps = 48/313 (15%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRML----AHGKKMPRHEGAALLMTYLGVSQNEAEKIC 56
+P GE+TVTL+DVA + LPI G+ + A G R E YLG+ A
Sbjct: 974 LPCGELTVTLEDVAMILGLPIRGQAVTGDTASGNWRERVE------EYLGLEPPVAPDGQ 1027
Query: 57 NQEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLF 116
Q + LR + + P E +E A V+ YC R Y+LY+ G +LF
Sbjct: 1028 RQTKTSGVPLSWLRANF------------GQCPAEADE-ATVQRYC-RAYVLYIFGSILF 1073
Query: 117 GDRSNKRIELIYLTTMAD-GYAGMRNYF*GAMTLAYLYGELADACR--PGHRALGGSVTL 173
D ++L +AD AG ++ G+ LA+LY +L DACR G L G V L
Sbjct: 1074 PDSGGDMASWMWLPLLADWDEAGTFSW--GSAALAWLYRQLCDACRRQGGDANLAGCVWL 1131
Query: 174 LTAW----FLAHFPGFFSVDL--NTDYLENYPVAARWKHQKGHGEG------ITYRSLLD 221
L W L P + + + D VA W+ G Y + +D
Sbjct: 1132 LQVWMWMRLLVGRPMWRTHQAWPHQDADRRPTVAHLWESVPSPVVGRRNLAYCHYTNEMD 1191
Query: 222 RIQLDDVCWRPYEKHREIQ-------DFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYV 274
+Q + V W PY+ ++ E+ + V + RV+RQ+G +
Sbjct: 1192 YLQPEHVVWMPYQAQEVLELELNPMCHIEDALKTLRCPLICFYAVEFQMCHRVMRQFGRL 1251
Query: 275 QTIPRHPTDVRDL 287
QTIP + DL
Sbjct: 1252 QTIPHRFSTSIDL 1264
>UniRef100_Q5W6G1 Putative polyprotein [Oryza sativa]
Length = 1567
Score = 65.1 bits (157), Expect = 4e-09
Identities = 72/288 (25%), Positives = 114/288 (39%), Gaps = 31/288 (10%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GEMT+TL DVA + LP+ G + + P A + G+ N Q
Sbjct: 1042 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWH--AQVQQLFGIPLN-----IEQGQ 1094
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
GG + S+L + N +D E R Y R Y+L+L+G +LF D
Sbjct: 1095 GG---KKKQNGIPLSWLSQ-NFSHLDDDAEPW----RAECY-ARAYILHLLGGVLFPDAG 1145
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADAC--RPGHRALGGSVTLLTAWF 178
I++ +A+ + + G+ LA+ Y +L +AC + + G V L+ W
Sbjct: 1146 GDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQMWM 1204
Query: 179 LAHFPGFFSVDLNTDYLENYPVAARWKHQKGHGEGITYRSLLDRIQLDDVCWRPYEKHRE 238
P ++ YL A + Y + +D +Q + W PY +
Sbjct: 1205 WLRLPNEPDMEKTVAYLFESTATAHAHRDVAYKH---YVNEMDYLQPQHIEWLPYHTNEA 1261
Query: 239 --------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ F Y ++C V HLP RV+RQ+G Q P
Sbjct: 1262 SSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGKKQDWP 1308
>UniRef100_Q84JZ9 Mutator-like transposase [Oryza sativa]
Length = 1381
Score = 60.8 bits (146), Expect = 7e-08
Identities = 63/221 (28%), Positives = 92/221 (41%), Gaps = 26/221 (11%)
Query: 89 PEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKRIELIYLTTMAD-GYAGMRNYF*GAM 147
P E +E A V+ YC R Y+LY+ G +LF D ++L +AD AG ++ G+
Sbjct: 804 PAEADE-ATVQRYC-RAYVLYIFGSILFPDSGGDMASWMWLPLLADWDEAGTFSW--GSA 859
Query: 148 TLAYLYGELADACR--PGHRALGGSVTLLTAWFLAHFP------GFFSVDLNTDYLENYP 199
LA+LY +L DACR G L G V LL W P + D
Sbjct: 860 ALAWLYRQLCDACRRQGGDANLAGCVWLLQVWMWMRLPVGRPMWRTHQAWPHQDADRCPT 919
Query: 200 VAARWKHQKGHGEG------ITYRSLLDRIQLDDVCWRPYEKHREIQ-------DFEEVF 246
VA W+ G Y + +D +Q + V W PY+ ++ E+
Sbjct: 920 VAHLWESVPSPVVGRRNLAYYHYTNEMDYLQPEHVVWMPYQAQEVLELELNPMCHIEDAL 979
Query: 247 WYSGWIMCGVRRVYRHLPERVLRQYGYVQTIPRHPTDVRDL 287
+ V + RV+RQ+G +QTIP + DL
Sbjct: 980 KTLRCPLICFYAVEFQMCHRVMRQFGRLQTIPHRFSTSIDL 1020
>UniRef100_Q8H3F8 Serine/threonine phosphatase PP7-related-like protein [Oryza
sativa]
Length = 589
Score = 58.9 bits (141), Expect = 3e-07
Identities = 75/308 (24%), Positives = 121/308 (38%), Gaps = 51/308 (16%)
Query: 5 EMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEYGGYI 64
EM +L D A + +P+ GR++ G + + L YLG + C G ++
Sbjct: 83 EMAPSLRDAAYILGIPVTGRVVTTGAVLNKSV-EDLCFQYLGQVPD-----CRDCRGSHV 136
Query: 65 SYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRSNKRI 124
L+ ++ R T D Y R YLL+L+G L +R +
Sbjct: 137 KLSWLQSKFSRIPERP-----TNDQT---------MYGTRAYLLFLIGSALLPERDRGYV 182
Query: 125 ELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADA-CRPGHRALGGSVTLLTAWFLAHFP 183
YL ++D + ++ Y GA LA+LY L+ A + L GS LL W + P
Sbjct: 183 SPKYLPLLSD-FDKVQEYAWGAAALAHLYKALSIAVAHSARKRLFGSAALLMGWIYEYIP 241
Query: 184 GFFSVDLNTDYLENYPVAARWKHQKGHGEGIT--------YRSLLDRIQLDDVCWRPYE- 234
D+ +P +W G I+ R +Q+ DV W PY+
Sbjct: 242 A-LRPDMYDPPEHIFPRVLKWT-----GSTISQPAKNVSDIRKAFGLLQVSDVNWEPYKG 295
Query: 235 -------KHREIQDFEEVFWYSGWIMC-GVRRVYRHLPERVLRQYGYVQTIPRH--PTDV 284
KH D + + W++ ++ +Y P+R RQ+G Q P + P
Sbjct: 296 VDPASIPKHCAAPD--NLCFSRTWLVSFNLKEIY--APDRFARQFGQEQHRPLNDVPAFQ 351
Query: 285 RDLPPPSI 292
R L P++
Sbjct: 352 RQLWNPAV 359
>UniRef100_Q7X7Y4 OSJNBa0014F04.28 protein [Oryza sativa]
Length = 1575
Score = 57.4 bits (137), Expect = 8e-07
Identities = 82/343 (23%), Positives = 128/343 (36%), Gaps = 45/343 (13%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GEMT+TL DVA + LP+ G + P A + G+ N Q
Sbjct: 984 LPSGEMTITLQDVAMILALPLRGHAVTGRTDTPGWH--AQVQQLFGIPLN-----IEQGQ 1036
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
GG + S+L + N +D E RV Y R Y+L+L+G +LF D
Sbjct: 1037 GG---KKKQNGIPLSWLSQ-NFSHLDDDAEPW----RVECY-ARAYILHLLGEVLFPDAG 1087
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADAC--RPGHRALGGSVTLLTAWF 178
I++ +A+ + + G+ LA+ Y +L +AC + + G V L+ W
Sbjct: 1088 GDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQMWM 1146
Query: 179 LAHFP--------GFFSVDLNTDYLENYPVAARWKHQKGHG-EGITYR---SLLDRIQLD 226
P F N +E H + Y+ + +D +Q
Sbjct: 1147 WLRLPVGRPKWRQSFTPWPYNEPDMEKTVAYLFESTATAHAHRDVAYKHYVNEMDCLQPQ 1206
Query: 227 DVCWRPYEKHRE--------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ W PY + + F Y ++C V HLP RV+RQ+G Q P
Sbjct: 1207 HIEWLPYHTNEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGKKQDWP 1265
Query: 279 RHPTDVRDLPPPSIVQMFVDFGTHTLKADARGEQAGEDTWRVA 321
V D+ + + T +K D WR+A
Sbjct: 1266 -----VEDISTGVELHKYDRVRTKKVKDWGLEHNRYIDEWRIA 1303
>UniRef100_Q5TKN6 Hypothetical protein OJ1058_F05.1 [Oryza sativa]
Length = 725
Score = 56.2 bits (134), Expect = 2e-06
Identities = 73/300 (24%), Positives = 114/300 (37%), Gaps = 40/300 (13%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GEMT+TL DVA + LP+ G + + P A + G+ N Q
Sbjct: 140 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWH--AQVQQLFGIPLN-----IEQGQ 192
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
GG + S+L + N +D E RV Y R Y+L+L+G +LF D
Sbjct: 193 GG---KKKQNGIPLSWLSQ-NFSHLDDDAEPW----RVECY-ARAYILHLLGGVLFPDAG 243
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADAC--RPGHRALGGSVTLLTAWF 178
I++ + + + G+ LA+ Y +L +AC + + G V L+ W
Sbjct: 244 GDIASAIWI-PLVTNLGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQMWM 302
Query: 179 LAHFP--------GFFSVDLNTDYLENYPVAARWKHQKGHG-EGITYR---SLLDRIQLD 226
P F N +E H + Y+ + +D +Q
Sbjct: 303 WLRLPIGRPKWRQSFTPWPYNEPDMEKTVAYLFESTATAHAHRDVAYKHYVNEMDCLQPQ 362
Query: 227 DVCWRPYEKHRE--------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ W PY + + F Y ++C V HLP RV+RQ+G Q P
Sbjct: 363 HIEWLPYHTNEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGKKQDWP 421
>UniRef100_Q6I5K6 Hypothetical protein OSJNBb0088F07.13 [Oryza sativa]
Length = 1564
Score = 56.2 bits (134), Expect = 2e-06
Identities = 73/300 (24%), Positives = 114/300 (37%), Gaps = 40/300 (13%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMPRHEGAALLMTYLGVSQNEAEKICNQEY 60
+P GEMT+TL DVA + LP+ G + + P A + G+ N Q
Sbjct: 979 LPSGEMTITLQDVAMILALPLRGHAVTGRTETPGWH--AQVQQLFGIPLN-----IEQGQ 1031
Query: 61 GGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGDRS 120
GG + S+L + N +D E RV Y R Y+L+L+G +LF D
Sbjct: 1032 GG---KKKQNGIPLSWLSQ-NFSHLDDDAEPW----RVECY-ARAYILHLLGGVLFPDAG 1082
Query: 121 NKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADAC--RPGHRALGGSVTLLTAWF 178
I++ + + + G+ LA+ Y +L +AC + + G V L+ W
Sbjct: 1083 GDIASAIWI-PLVTNLGDLGRFSWGSAVLAWTYRQLCEACCRQAPSSNMSGCVLLIQMWM 1141
Query: 179 LAHFP--------GFFSVDLNTDYLENYPVAARWKHQKGHG-EGITYR---SLLDRIQLD 226
P F N +E H + Y+ + +D +Q
Sbjct: 1142 WLRLPIGRPKWRQSFTPWPYNEPDMEKTVAYLFESTATAHAHRDVAYKHYVNEMDCLQPQ 1201
Query: 227 DVCWRPYEKHRE--------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQTIP 278
+ W PY + + F Y ++C V HLP RV+RQ+G Q P
Sbjct: 1202 HIEWLPYHTNEASSLTLNSMCNRDSDYFMYQCPLIC-FWAVEYHLPHRVMRQFGKKQDWP 1260
>UniRef100_Q94HR1 Hypothetical protein OSJNBa0065C16.8 [Oryza sativa]
Length = 681
Score = 56.2 bits (134), Expect = 2e-06
Identities = 72/302 (23%), Positives = 118/302 (38%), Gaps = 44/302 (14%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRMLAHGKKMP--RHEGAALLMTYLGVSQNEAEKICNQ 58
+P GEMT+TL DVA + LP++G ++ + P R + L L + Q + K
Sbjct: 188 LPSGEMTITLQDVAMILALPLQGHVVTGRTETPGWRAQVEQLFGIPLNIEQGQGGK---- 243
Query: 59 EYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFGD 118
+ S+L + N +D E RV Y R Y+L+L+G +LF +
Sbjct: 244 --------KKQNGIPLSWLSQ-NFSHLDDDAEPW----RVECY-ARAYILHLLGGVLFPN 289
Query: 119 RSNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADAC--RPGHRALGGSVTLLTA 176
I++ +A+ + + G+ LA+ Y +L +AC + + G V L+
Sbjct: 290 AGGDIASAIWIPLVAN-LGDLGRFSWGSAVLAWTYRQLCEACCRQALSSNMSGCVLLIQL 348
Query: 177 WFLAHFP--------GFFSVDLNTDYLENYPVAARWKHQKGHG-EGITYR---SLLDRIQ 224
W P F N +E H + Y+ + +D +Q
Sbjct: 349 WMGLRLPVGRPKWRQSFTPWPYNEPDMEKTVAYLFESTATAHAHRDVAYKHYVNEMDCLQ 408
Query: 225 LDDVCWRPYEKHRE--------IQDFEEVFWYSGWIMCGVRRVYRHLPERVLRQYGYVQT 276
+ W PY + + F Y ++C V HLP RV+RQ+G Q
Sbjct: 409 PQHIEWLPYHTNEASSLSLNSMCNRDSDYFMYQCPLVC-FWAVEYHLPHRVMRQFGKKQD 467
Query: 277 IP 278
P
Sbjct: 468 WP 469
>UniRef100_Q7XHQ1 Calcineurin-like phosphoesterase-like [Oryza sativa]
Length = 327
Score = 55.5 bits (132), Expect = 3e-06
Identities = 49/186 (26%), Positives = 80/186 (42%), Gaps = 13/186 (6%)
Query: 1 MPMGEMTVTLDDVACLTHLPIEGRML---AHGKKMPRHEGAALLMTYLGVSQNEAEKICN 57
+P+GEMT+TL DV+CL LPI GR + A G + E LL + + +K
Sbjct: 87 LPVGEMTITLQDVSCLWGLPIHGRPITGQADGSWVDMIE--RLLGIPMEEQHMKQKKRKK 144
Query: 58 QEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRCYLLYLVGCLLFG 117
++ +SY R + R V+ E+ + R +L ++G ++F
Sbjct: 145 EDDMTMVSYSRYSISLSKLRDRFRVMPKNATEREI-------NWYTRALVLDIIGSMVFT 197
Query: 118 DRSNKRIELIYLTTMADGYAGMRNYF*GAMTLAYLYGELADACRPGHRALGGSVTLLTAW 177
D S + +YL M + + Y GA L+ LY +L+ A + + LL W
Sbjct: 198 DTSGDGVPAMYLQFMMN-LSEQTEYNWGAAALSMLYRQLSIASEKERAEISRPLLLLQLW 256
Query: 178 FLAHFP 183
+ P
Sbjct: 257 SWSRLP 262
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 765,920,331
Number of Sequences: 2790947
Number of extensions: 33134294
Number of successful extensions: 86536
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 91
Number of HSP's that attempted gapping in prelim test: 86379
Number of HSP's gapped (non-prelim): 177
length of query: 430
length of database: 848,049,833
effective HSP length: 130
effective length of query: 300
effective length of database: 485,226,723
effective search space: 145568016900
effective search space used: 145568016900
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC138133.3


