

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
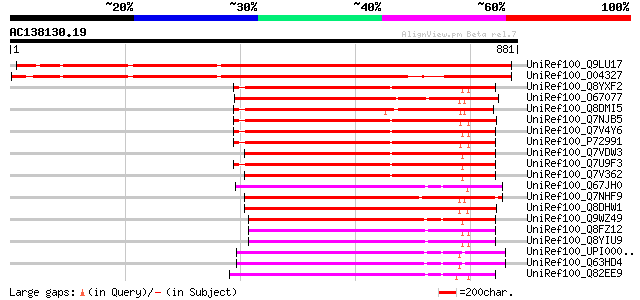
UniRef100_Q9LU17 Cell division protein FtsH-like [Arabidopsis th... 1082 0.0
UniRef100_O04327 Cell division protein FtsH isolog [Arabidopsis ... 967 0.0
UniRef100_Q8YXF2 Cell division protein [Anabaena sp.] 341 5e-92
UniRef100_O67077 Cell division protein ftsH homolog [Aquifex aeo... 341 5e-92
UniRef100_Q8DMI5 Cell division protein [Synechococcus elongatus] 341 6e-92
UniRef100_Q7NJB5 Cell division protein [Gloeobacter violaceus] 340 8e-92
UniRef100_Q7V4Y6 Cell division protein FtsH2 [Prochlorococcus ma... 340 1e-91
UniRef100_P72991 Cell division protein ftsH homolog 4 [Synechocy... 338 4e-91
UniRef100_Q7VDW3 Cell division protein FtsH [Prochlorococcus mar... 337 1e-90
UniRef100_Q7U9F3 Cell division protein FtsH2 precursor [Synechoc... 336 2e-90
UniRef100_Q7V362 Cell division protein FtsH2 [Prochlorococcus ma... 334 8e-90
UniRef100_Q67JH0 Cell division protein [Symbiobacterium thermoph... 329 3e-88
UniRef100_Q7NHF9 Cell division protein [Gloeobacter violaceus] 327 1e-87
UniRef100_Q8DHW1 Cell division protein [Synechococcus elongatus] 323 1e-86
UniRef100_Q9WZ49 Cell division protein FtsH [Thermotoga maritima] 322 4e-86
UniRef100_Q8FZ12 Cell division protein FtsH [Brucella suis] 319 3e-85
UniRef100_Q8YIU9 CELL DIVISION PROTEIN FTSH [Brucella melitensis] 319 3e-85
UniRef100_UPI00003CC442 UPI00003CC442 UniRef100 entry 318 6e-85
UniRef100_Q63HD4 Cell division protein [Bacillus cereus] 317 1e-84
UniRef100_Q82EE9 Putative cell division protein FtsH [Streptomyc... 316 2e-84
>UniRef100_Q9LU17 Cell division protein FtsH-like [Arabidopsis thaliana]
Length = 976
Score = 1082 bits (2797), Expect = 0.0
Identities = 552/862 (64%), Positives = 687/862 (79%), Gaps = 22/862 (2%)
Query: 13 SSSSKLKKFPKPSKFRSISSQIQTPESENDEKNQKNLNFNNLNFLKFTVTLTVISASLPQ 72
++S L + ++ R+ ++ + +ND+ N +N L +TLT+ISASL +
Sbjct: 121 NTSMVLLRCRSDNRIRNATNVVHEDGDDNDKAKT-----NQVNLLAIPITLTIISASLAK 175
Query: 73 AATAVAAAGKKRAPRKASTKKVEALSIEEVKTWIEGLPIVSERIPYTEIAELKNLGMLKH 132
+ A A +++ +K K EAL++E++K W + LP+VS RIPYT+I LK G LKH
Sbjct: 176 PSFAAAKVTERKRTQK---KPQEALTLEQLKAWSKDLPVVSNRIPYTDILSLKAEGKLKH 232
Query: 133 IVKPSAVELRERAVAVLVVLEDSRVLRTVLPNVESDRKFWGLWDELKIENLCVNAYSPPV 192
++KP + LR++A VLVVLEDSRVLRTVLP++E +++FW WDEL I+ CVNAY+PPV
Sbjct: 233 VIKPPNLSLRQKAEPVLVVLEDSRVLRTVLPSLEGNKRFWEQWDELGIDVQCVNAYTPPV 292
Query: 193 KVPEIPLSVLARIWLSLPFHKPLVEFVNRFQPKKKSKKELALREARMQLQRQKKEEVVKT 252
K P +P L +W K + +PKK+SK+ L+ R +RQ+KEE+
Sbjct: 293 KRPPVPSPYLGFLW------KVPAYMLTWVKPKKESKRAAELKRMREDFKRQRKEEIETM 346
Query: 253 MKEREMIERNERNKKREAENEKRMR-RRKEYKEKMVEVKANEFFNTTIWTRMAKDKMAIN 311
+ER M+E+ + +K++ E +KR R+K+Y+E + E + N +W R+A+D
Sbjct: 347 KEERVMMEKTMKAQKKQQERKKRKAVRKKKYEESLREARKNYRDMADMWARLAQDPNVAT 406
Query: 312 GIGVLFFVIFYRTVVVSYKKQKKDYEDRIKIQKADAEERRKMREMEAEMGWSEAGGDEDE 371
+G++FF IFYR VV++Y+KQKKDYEDR+KI+KA+A+ER+KMRE+E EM G E+E
Sbjct: 407 ALGLVFFYIFYRVVVLNYRKQKKDYEDRLKIEKAEADERKKMRELEREME-----GIEEE 461
Query: 372 SELVKEG--EENPYLKMTKEFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLE 429
E V+EG E+NPYL+M +FMKSGARVRRA N+RLP+YLERGVDVKFTDVAGLGKIRLE
Sbjct: 462 DEEVEEGTGEKNPYLQMAMQFMKSGARVRRASNKRLPEYLERGVDVKFTDVAGLGKIRLE 521
Query: 430 LEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVE 489
LEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVE
Sbjct: 522 LEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVE 581
Query: 490 IYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLD 549
IYVGVGASRVR+LYQEA+ENAPSVVFIDELDAVGR+RGLIKGSGGQERDATLNQLLV LD
Sbjct: 582 IYVGVGASRVRALYQEARENAPSVVFIDELDAVGRERGLIKGSGGQERDATLNQLLVSLD 641
Query: 550 GFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAED 609
GFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG IGR+EIL+VHARKKP+AED
Sbjct: 642 GFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRMEILQVHARKKPMAED 701
Query: 610 VDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERS 669
+DY VASMTDGMVGAELANIVE+AAINMMRD RTE++TDDLLQAAQ+EERGMLDRK+RS
Sbjct: 702 LDYMAVASMTDGMVGAELANIVEIAAINMMRDGRTELTTDDLLQAAQIEERGMLDRKDRS 761
Query: 670 KEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQ 729
E W QVAINEAAMAV A+N P+ NIE++TI PRAGRELGYVR ++ I F +GML+RQ
Sbjct: 762 LETWRQVAINEAAMAVVAVNFPDMKNIEFLTINPRAGRELGYVRVKMDHIKFKEGMLSRQ 821
Query: 730 SLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMIGGLSDKYRGVSNFWV 789
S+ DHITVQLAPRAADE+W+G+DQLSTIWAET+DNAR AAR ++GGLSDK+ G++NFWV
Sbjct: 822 SILDHITVQLAPRAADELWYGEDQLSTIWAETSDNARSAARSLVLGGLSDKHHGLNNFWV 881
Query: 790 TDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGH 849
DRIN+ID+EA++ILN+CYERAKEIL +N+TLMD +V +LV KK+LTK++ LV+L+G
Sbjct: 882 ADRINDIDVEALRILNMCYERAKEILGRNRTLMDEVVEKLVQKKSLTKQEFFTLVELYGS 941
Query: 850 AKPIPISVLDIRDAKHKELQEI 871
+KP+P S+L++R K EL+E+
Sbjct: 942 SKPMPPSILELRKIKRLELEEM 963
>UniRef100_O04327 Cell division protein FtsH isolog [Arabidopsis thaliana]
Length = 983
Score = 967 bits (2501), Expect = 0.0
Identities = 516/871 (59%), Positives = 639/871 (73%), Gaps = 84/871 (9%)
Query: 4 HCFVHFPPSSSSSKLKKFPKPSKFRSISSQIQTPESENDEKNQKNLNFNNLNFLKFTVTL 63
H F+ +P + S K P+ E+ + N K N +N L +TL
Sbjct: 181 HFFIDYPEITLQSHAKTIQLPNVVH-----------EDGDDNDK-AKTNQVNLLAIPITL 228
Query: 64 TVISASLPQAATAVAAAGKKRAPRKASTKKVEALSIEEVKTWIEGLPIVSERIPYTEIAE 123
T+ISASL + + A A +++ +K K EAL++E++K W + LP+VS RIPYT+I
Sbjct: 229 TIISASLAKPSFAAAKVTERKRTQK---KPQEALTLEQLKAWSKDLPVVSNRIPYTDILS 285
Query: 124 LKNLGMLKHIVKPSAVELRERAVAVLVVLEDSRVLRTVLPNVESDRKFWGLWDELKIENL 183
LK G LKH++KP + LR++A VLVVLEDSRVLRTVLP++E +++FW WDEL I+
Sbjct: 286 LKAEGKLKHVIKPPNLSLRQKAEPVLVVLEDSRVLRTVLPSLEGNKRFWEQWDELGIDVQ 345
Query: 184 CVNAYSPPVKVPEIPLSVLARIWLSLPFHKPLVEFVNRFQPKKKSKKELALREARMQLQR 243
CVNAY+PPVK P +P L +W K + +PKK+SK+ L+ R +R
Sbjct: 346 CVNAYTPPVKRPPVPSPYLGFLW------KVPAYMLTWVKPKKESKRAAELKRMREDFKR 399
Query: 244 QKKEEVVKTMKEREMIERNERNKKREAENEKRMR-RRKEYKEKMVEVKANEFFNTTIWTR 302
Q+KEE+ +ER M+E+ + +K++ E +KR R+K+Y+E + E + N +W R
Sbjct: 400 QRKEEIETMKEERVMMEKTMKAQKKQQERKKRKAVRKKKYEESLREARKNYRDMADMWAR 459
Query: 303 MAKDKMAINGIGVLFFVIFYRTVVVSYKKQKKDYEDRIKIQKADAEERRKMREMEAEMGW 362
+A+D +G++FF IFYR VV++Y+KQKKDYEDR+KI+KA+A+ER+KMRE+E EM
Sbjct: 460 LAQDPNVATALGLVFFYIFYRVVVLNYRKQKKDYEDRLKIEKAEADERKKMRELEREME- 518
Query: 363 SEAGGDEDESELVKEG--EENPYLKMTKEFMKSGARVRRAQNRRLPQYLERGVDVKFTDV 420
G E+E E V+EG E+NPYL+M +FMKSGARVRRA N+RLP+YLERGVDVKFTDV
Sbjct: 519 ----GIEEEDEEVEEGTGEKNPYLQMAMQFMKSGARVRRASNKRLPEYLERGVDVKFTDV 574
Query: 421 AGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFF 480
AGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFF
Sbjct: 575 AGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFF 634
Query: 481 SISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDAT 540
SISASQFVEIYVGVGASRVR+LYQEA+ENAPSVVFIDELDAVGR+RGLIKGSGGQERDAT
Sbjct: 635 SISASQFVEIYVGVGASRVRALYQEARENAPSVVFIDELDAVGRERGLIKGSGGQERDAT 694
Query: 541 LNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVH 600
LNQLLV LDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG IGR+EIL+VH
Sbjct: 695 LNQLLVSLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRMEILQVH 754
Query: 601 ARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEER 660
ARKKP+AED+DY VASMTDGMVGAELANIVE+AAINMMRD RTE++TDDLLQAAQ+EER
Sbjct: 755 ARKKPMAEDLDYMAVASMTDGMVGAELANIVEIAAINMMRDGRTELTTDDLLQAAQIEER 814
Query: 661 GMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESIN 720
GMLDRK+RS E W QVAINEAAMAV A+N P+ +++I
Sbjct: 815 GMLDRKDRSLETWRQVAINEAAMAVVAVNFPD-----------------------MKNIE 851
Query: 721 FNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMIGGLSDK 780
F LSTIWAET+DNAR AAR ++GGLSDK
Sbjct: 852 F--------------------------------LSTIWAETSDNARSAARSLVLGGLSDK 879
Query: 781 YRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDI 840
+ G++NFWV DRIN+ID+EA++ILN+CYERAKEIL +N+TLMD +V +LV KK+LTK++
Sbjct: 880 HHGLNNFWVADRINDIDVEALRILNMCYERAKEILGRNRTLMDEVVEKLVQKKSLTKQEF 939
Query: 841 VRLVQLHGHAKPIPISVLDIRDAKHKELQEI 871
LV+L+G +KP+P S+L++R K EL+E+
Sbjct: 940 FTLVELYGSSKPMPPSILELRKIKRLELEEM 970
>UniRef100_Q8YXF2 Cell division protein [Anabaena sp.]
Length = 613
Score = 341 bits (875), Expect = 5e-92
Identities = 204/475 (42%), Positives = 293/475 (60%), Gaps = 32/475 (6%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E V F DVAG+ + +LEL E+V F + + + G K
Sbjct: 140 FGKSKARVQ----------MEPQTQVTFGDVAGIDQAKLELNEVVDFLKNADRFTAVGAK 189
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLA+AVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK N
Sbjct: 190 IPKGVLLVGPPGTGKTLLARAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKSN 249
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I IA+TNRPD+L
Sbjct: 250 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIIIAATNRPDVL 309
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P + GR EILKVHAR K +A+DVD + +A T G GA+L+N
Sbjct: 310 DAALLRPGRFDRQVVVDRPDYAGRSEILKVHARGKTLAKDVDLDKIARRTPGFTGADLSN 369
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAA 687
++ AAI R + TE+S D++ A G ++K+R S+++ VA +EA A+
Sbjct: 370 LLNEAAILAARRNLTEISMDEINDAIDRVLAGP-EKKDRVMSEKRKVLVAYHEAGHALVG 428
Query: 688 MNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEM 747
+P++D ++ I+I PR GR G + G+ +R L + + V L R A+E+
Sbjct: 429 ALMPDYDPVQKISIIPR-GRAGGLTWFTPSEDRMDTGLYSRAYLENQMAVALGGRIAEEI 487
Query: 748 WFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKYRGV-------SNFWVTDRINE---- 795
FG+++++T + ARVA +M G+SDK V + F D ++E
Sbjct: 488 IFGEEEVTTGASNDLQQVARVARQMITRFGMSDKLGPVALGRQQGNMFLGRDIMSERDFS 547
Query: 796 ------IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
ID E K++ Y RAKE+L N+ ++D + LV K+T+ +++ ++
Sbjct: 548 EETAAAIDEEVHKLVETAYTRAKEVLVNNRHILDQIAQMLVDKETVDADELQEIL 602
>UniRef100_O67077 Cell division protein ftsH homolog [Aquifex aeolicus]
Length = 634
Score = 341 bits (875), Expect = 5e-92
Identities = 210/482 (43%), Positives = 303/482 (62%), Gaps = 31/482 (6%)
Query: 391 MKSGARVRRAQN---RRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRG 447
M G V RA N R Y+E V F DVAG+ +++ E++EI+++ +++ G
Sbjct: 125 MSGGGNVNRAFNFGKSRAKVYIEEKPKVTFKDVAGIEEVKEEVKEIIEYLKDPVKFQKLG 184
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
+ P G+LL G PGVGKTLLAKA+AGEA V F S+S S FVE++VGVGA+RVR L++ AK
Sbjct: 185 GRPPKGVLLYGEPGVGKTLLAKAIAGEAHVPFISVSGSDFVEMFVGVGAARVRDLFETAK 244
Query: 508 ENAPSVVFIDELDAVGRKRGLIK-GSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRP 566
++AP ++FIDE+DAVGR RG I G G ER+ TLNQLLV +DGF+ +I IA+TNRP
Sbjct: 245 KHAPCIIFIDEIDAVGRARGAIPVGGGHDEREQTLNQLLVEMDGFDTSDGIIVIAATNRP 304
Query: 567 DILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAE 626
DILDPAL+RPGRFDR+IFIPKP GR EILKVHAR K +A+DVD E VA T G GA+
Sbjct: 305 DILDPALLRPGRFDRQIFIPKPDVRGRYEILKVHARNKKLAKDVDLEFVARATPGFTGAD 364
Query: 627 LANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERS---KEKWEQVAINEAAM 683
L N++ AA+ R + E++ +++ +A G L+RK + KEK E++AI+EA
Sbjct: 365 LENLLNEAALLAARKGKEEITMEEIEEALDRITMG-LERKGMTISPKEK-EKIAIHEAGH 422
Query: 684 AVAAMNLPNFDNIEYITIAPRAGRELGYVRTM-LESINFNDGMLTRQSLFDHITVQLAPR 742
A+ + + D + I+I PR G LG + + +E + D ++ L++ I V L R
Sbjct: 423 ALMGLVSDDDDKVHKISIIPR-GMALGVTQQLPIEDKHIYD----KKDLYNKILVLLGGR 477
Query: 743 AADEMWFGKDQLSTIWAETADNAR-VAARMYMIGGLSDK-----YRGVSNFWV------- 789
AA+E++FGKD ++T A +A RM + G+SDK R V+N ++
Sbjct: 478 AAEEVFFGKDGITTGAENDLQRATDLAYRMVSMWGMSDKVGPIAIRRVANPFLGGMTTAV 537
Query: 790 ---TDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQL 846
D + EID E +I+ YE+AK I+++ K + +V +L+ K+T+T E+ V + +L
Sbjct: 538 DTSPDLLREIDEEVKRIITEQYEKAKAIVEEYKEPLKAVVKKLLEKETITCEEFVEVFKL 597
Query: 847 HG 848
+G
Sbjct: 598 YG 599
>UniRef100_Q8DMI5 Cell division protein [Synechococcus elongatus]
Length = 612
Score = 341 bits (874), Expect = 6e-92
Identities = 199/476 (41%), Positives = 290/476 (60%), Gaps = 42/476 (8%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E V F DVAG+ + +LEL E+V+F + + + G K
Sbjct: 139 FGKSRARVQ----------MEPQTQVTFNDVAGIDQAKLELGEVVEFLKYADRFTEVGAK 188
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLA+AVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK N
Sbjct: 189 IPKGVLLVGPPGTGKTLLARAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKAN 248
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I IA+TNRPD+L
Sbjct: 249 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIVIAATNRPDVL 308
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P + GR++ILKVHAR K +A+DVD + +A T G GA+L+N
Sbjct: 309 DAALLRPGRFDRQVVVDRPDYKGRLDILKVHARGKTLAKDVDLDKIARRTPGFTGADLSN 368
Query: 630 IVEVAAINMMRDSRTEVSTDDL-------LQAAQMEERGMLDRKERSKEKWEQVAINEAA 682
++ AAI R + TE+S D++ L + ++R M DR+++ VA +EA
Sbjct: 369 LLNEAAILAARRNLTEISMDEINDAIDRVLAGPEKKDRVMSDRRKK------LVAYHEAG 422
Query: 683 MAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPR 742
A+ +P++D ++ ++I PR GR G + G+ +R L + + V L R
Sbjct: 423 HALVGALMPDYDPVQKVSIIPR-GRAGGLTWFTPNEDQMDSGLYSRAYLQNQMAVALGGR 481
Query: 743 AADEMWFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKY------RGVSNFWV------ 789
A+E+ FG+D+++T + ARVA +M G+SD+ R N ++
Sbjct: 482 IAEEIVFGEDEVTTGASNDLQQVARVARQMVTRFGMSDRLGPVALGRQTGNVFLGRDIMA 541
Query: 790 -----TDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDI 840
+ ID E ++ Y RAKE+L N+ ++D + L+ K+T+ E++
Sbjct: 542 ERDFSEETAATIDDEVRNLVEQAYRRAKEVLVNNRHVLDQIAQVLIEKETIDAEEL 597
>UniRef100_Q7NJB5 Cell division protein [Gloeobacter violaceus]
Length = 611
Score = 340 bits (873), Expect = 8e-92
Identities = 206/477 (43%), Positives = 291/477 (60%), Gaps = 34/477 (7%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E FTDVAG+ + +LEL+E+V F + E + G K
Sbjct: 140 FGKSKARVQ----------MEPQTKTTFTDVAGVEEAKLELQEVVDFLKNSERFTAVGAK 189
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLAKAVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK+N
Sbjct: 190 IPKGVLLVGPPGTGKTLLAKAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKKN 249
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLLV +DGFEG VI IA+TNRPD+L
Sbjct: 250 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLVEMDGFEGNTGVIIIAATNRPDVL 309
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P F GR+EILKVHAR K + +D+D E +A T G GA+LAN
Sbjct: 310 DAALLRPGRFDRQVVVDRPDFKGRLEILKVHARGKTLGKDIDLEKIARRTPGFTGADLAN 369
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER---SKEKWEQVAINEAAMAVA 686
++ AAI R S TE+S D++ A G ++K R K KW VA +E A+
Sbjct: 370 LLNEAAILAARRSLTEISMDEVNDAVDRVLAGP-EKKNRLMTEKRKW-LVAYHEVGHALV 427
Query: 687 AMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADE 746
LP +D ++ I+I PR G G + + + G+ +R + + + V L R A+E
Sbjct: 428 GALLPEYDPVQKISIIPR-GMAGGLTWFVPDEERADSGLYSRVYMTNMMAVALGGRIAEE 486
Query: 747 MWFGKDQLST-IWAETADNARVAARMYMIGGLSDKY-------RGVSNFWVTDRINE--- 795
+ +G+ +++T + A++A M G+S+K +G S F D + E
Sbjct: 487 IVYGEAEVTTGATNDLQQVAQIARNMVTRYGMSEKLGPVALGRQGGSMFLGRDIMTERDF 546
Query: 796 -------IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
ID E +++ Y +K +L ++ LMD + LV K+T+ E++ +L++
Sbjct: 547 SEHTASVIDEEIRELIEKAYALSKSVLLSHRNLMDRVTEVLVQKETVDAEELEQLIE 603
>UniRef100_Q7V4Y6 Cell division protein FtsH2 [Prochlorococcus marinus]
Length = 615
Score = 340 bits (871), Expect = 1e-91
Identities = 201/475 (42%), Positives = 295/475 (61%), Gaps = 32/475 (6%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E V F+DVAG+ +LEL E+V F + + + G K
Sbjct: 142 FGKSKARVQ----------MEPSTQVTFSDVAGIEGAKLELTEVVDFLKNPDRFTAVGAK 191
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLAKAVAGEA V FFSIS S+FVE++VGVGASRVR L+++AK+N
Sbjct: 192 IPKGVLLVGPPGTGKTLLAKAVAGEAAVPFFSISGSEFVEMFVGVGASRVRDLFEQAKKN 251
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I +A+TNRPD+L
Sbjct: 252 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIIVAATNRPDVL 311
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P + GR++IL VHAR K +++DVD + VA T G GA+LAN
Sbjct: 312 DSALMRPGRFDRQVVVERPDYTGRLQILNVHARDKTLSKDVDLDKVARRTPGFTGADLAN 371
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAA 687
++ AAI R TEVS D++ A + G ++K+R S+ + VA +E+ A+
Sbjct: 372 LLNEAAILAARRELTEVSNDEISDAIERVMVGP-EKKDRVMSERRKRLVAYHESGHALVG 430
Query: 688 MNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEM 747
+P++D+++ I+I PR G+ G G+ +R L + + V L R A+E+
Sbjct: 431 ALMPDYDSVQKISIIPR-GQAGGLTFFTPSEERMESGLYSRAYLQNQMAVALGGRVAEEI 489
Query: 748 WFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKYRGVS-------NFWVTDRINE---- 795
+G+D+++T + A+VA +M G+SDK V+ F D +E
Sbjct: 490 VYGEDEVTTGASNDLQQVAQVARQMVTRFGMSDKLGPVALGRSQGGMFLGRDIASERDFS 549
Query: 796 ------IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
ID E ++++ Y+RA ++L +N++++D L + LV K+TL +D+ L+
Sbjct: 550 EDTAAIIDAEVSDLVDVAYKRATKVLIENRSVLDELADLLVEKETLDAQDLQELL 604
>UniRef100_P72991 Cell division protein ftsH homolog 4 [Synechocystis sp.]
Length = 616
Score = 338 bits (867), Expect = 4e-91
Identities = 201/475 (42%), Positives = 292/475 (61%), Gaps = 32/475 (6%)
Query: 390 FMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVK 449
F KS ARV+ +E V F DVAG+ + +LEL E+V F + + + G K
Sbjct: 143 FGKSKARVQ----------MEPQTQVTFGDVAGIEQAKLELTEVVDFLKNADRFTELGAK 192
Query: 450 IPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKEN 509
IP G+LL GPPG GKTLLAKAVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK N
Sbjct: 193 IPKGVLLVGPPGTGKTLLAKAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKAN 252
Query: 510 APSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDIL 569
AP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I +A+TNRPD+L
Sbjct: 253 APCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIIVAATNRPDVL 312
Query: 570 DPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELAN 629
D AL+RPGRFDR++ + +P + GR EIL VHAR K +++DVD + +A T G GA+L+N
Sbjct: 313 DSALMRPGRFDRQVVVDRPDYAGRREILNVHARGKTLSQDVDLDKIARRTPGFTGADLSN 372
Query: 630 IVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAA 687
++ AAI R + TE+S D++ A G ++K R S+++ VA +EA A+
Sbjct: 373 LLNEAAILAARRNLTEISMDEVNDAIDRVLAGP-EKKNRVMSEKRKTLVAYHEAGHALVG 431
Query: 688 MNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEM 747
+P++D ++ I+I PR GR G G+ +R L + + V L R A+E+
Sbjct: 432 ALMPDYDPVQKISIIPR-GRAGGLTWFTPSEDRMESGLYSRSYLQNQMAVALGGRIAEEI 490
Query: 748 WFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKY-------RGVSNFWVTDRINE---- 795
FG+++++T + ARVA +M G+SD+ +G F D ++
Sbjct: 491 IFGEEEVTTGASNDLQQVARVARQMVTRFGMSDRLGPVALGRQGGGVFLGRDIASDRDFS 550
Query: 796 ------IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
ID E ++++ Y+RAK++L +N+ ++D L LV K+T+ E++ L+
Sbjct: 551 DETAAAIDEEVSQLVDQAYQRAKQVLVENRGILDQLAEILVEKETVDSEELQTLL 605
>UniRef100_Q7VDW3 Cell division protein FtsH [Prochlorococcus marinus]
Length = 599
Score = 337 bits (863), Expect = 1e-90
Identities = 192/456 (42%), Positives = 286/456 (62%), Gaps = 22/456 (4%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ +LEL E+V F + + + G KIP G+LL GPPG GKTLLA
Sbjct: 135 MEPSTQVTFGDVAGIEGAKLELAEVVDFLKNPDRFTAVGAKIPKGVLLVGPPGTGKTLLA 194
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KAVAGEA V FFSIS S+FVE++VGVGASRVR L+++AK+NAP +VFIDE+DAVGR+RG
Sbjct: 195 KAVAGEAAVPFFSISGSEFVEMFVGVGASRVRDLFEQAKKNAPCIVFIDEIDAVGRQRGA 254
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I +A+TNRPD+LD AL+RPGRFDR++ + +P
Sbjct: 255 GLGGGNDEREQTLNQLLTEMDGFEGNTGIIIVAATNRPDVLDSALLRPGRFDRQVVVDRP 314
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
++GR++ILKVHAR+K +++DVD + VA T G GA+LAN++ +AI R TEVS
Sbjct: 315 DYLGRLQILKVHAREKTLSKDVDLDQVARRTPGFTGADLANLLNESAILAARREHTEVSN 374
Query: 649 DDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAG 706
++ A + G ++K+R S ++ E VA +EA A+ +P++D ++ I+I PR G
Sbjct: 375 IEISDAIERVMAGP-EKKDRVMSNKRKELVAYHEAGHALVGAVMPDYDPVQKISIIPR-G 432
Query: 707 RELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNA 765
+ G G+ +R L + + V L R A+E+ +G+D+++T + A
Sbjct: 433 QAGGLTFFTPSEERMESGLYSRSYLQNQMAVALGGRVAEEIVYGEDEVTTGASNDLKQVA 492
Query: 766 RVAARMYMIGGLSDKYRGVS-----------------NFWVTDRINEIDLEAMKILNLCY 808
+VA +M G+S+K V+ + D ID E ++++ Y
Sbjct: 493 QVARQMVTRFGMSEKLGPVALGRSQGGMFLGRDIAAERDFSEDTAATIDDEVSCLVDIAY 552
Query: 809 ERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+RA + L +N++++D L L+ K+T+ ED+ +L+
Sbjct: 553 KRATKALLENRSVLDELAEMLIEKETVDSEDLQQLL 588
>UniRef100_Q7U9F3 Cell division protein FtsH2 precursor [Synechococcus sp.]
Length = 615
Score = 336 bits (862), Expect = 2e-90
Identities = 198/476 (41%), Positives = 289/476 (60%), Gaps = 32/476 (6%)
Query: 389 EFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGV 448
+F KS ARV+ +E V FTDVAG+ +LEL E+V F + + + G
Sbjct: 141 QFGKSKARVQ----------MEPSTQVTFTDVAGIEGAKLELTEVVDFLKNPDRFTAVGA 190
Query: 449 KIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKE 508
KIP G+LL GPPG GKTLLAKAVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK+
Sbjct: 191 KIPKGVLLVGPPGTGKTLLAKAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKK 250
Query: 509 NAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDI 568
NAP +VFIDE+DAVGR+RG G G ER+ TLNQLL +DGFEG +I +A+TNRPD+
Sbjct: 251 NAPCIVFIDEIDAVGRQRGAGLGGGNDEREQTLNQLLTEMDGFEGNTGIIIVAATNRPDV 310
Query: 569 LDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELA 628
LD AL+RPGRFDR++ + +P + GR++IL VHAR K +++DVD + VA T G GA+LA
Sbjct: 311 LDAALMRPGRFDRQVTVDRPDYAGRLQILNVHARGKTLSKDVDLDKVARRTPGYTGADLA 370
Query: 629 NIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVA 686
N++ AAI R TEVS D++ A + G ++K+R S+ + VA +EA A+
Sbjct: 371 NLLNEAAILAARRELTEVSNDEISDAIERVMAGP-EKKDRVMSERRKRLVAYHEAGHALV 429
Query: 687 AMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADE 746
+P++D ++ I+I PR G G G+ +R L + + V L R A+E
Sbjct: 430 GALMPDYDPVQKISIIPR-GNAGGLTFFTPSEERMESGLYSRAYLQNQMAVALGGRVAEE 488
Query: 747 MWFGKDQLSTIWAETADN-ARVAARMYMIGGLSDKYRGVS-----------------NFW 788
+ +G+D+++T + A A +M G+SD V+ +
Sbjct: 489 IVYGEDEVTTGASNDLQQVASTARQMITRFGMSDTLGPVALGRAQGGMFLGRDIAAERDF 548
Query: 789 VTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
D ID E +++++ Y+RA ++L N+ ++D L + LV ++T+ E++ L+
Sbjct: 549 SEDTAATIDQEVSELVDVAYKRATKVLVDNRAVLDELADMLVEQETVDAEELQELL 604
>UniRef100_Q7V362 Cell division protein FtsH2 [Prochlorococcus marinus subsp.
pastoris]
Length = 618
Score = 334 bits (856), Expect = 8e-90
Identities = 190/456 (41%), Positives = 281/456 (60%), Gaps = 22/456 (4%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F+DVAG+ +LEL E+V F + + G KIP G+LL GPPG GKTLLA
Sbjct: 154 MEPSTQVTFSDVAGVEGAKLELTEVVDFLKSPDRFTAVGAKIPKGVLLVGPPGTGKTLLA 213
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KAVAGEAGV FFSIS S+FVE++VGVGASRVR L+++AK+NAP +VFIDE+DAVGR+RG
Sbjct: 214 KAVAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFEQAKKNAPCIVFIDEIDAVGRQRGA 273
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I +A+TNRPD+LD AL+RPGRFDR++ + +P
Sbjct: 274 GMGGGNDEREQTLNQLLTEMDGFEGNSGIIIVAATNRPDVLDSALMRPGRFDRQVTVDRP 333
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
+ GR++IL VHA+ K +++DVD + VA T G GA+LAN++ AAI R VS
Sbjct: 334 DYAGRLQILNVHAKDKTLSKDVDLDKVARRTPGFTGADLANLLNEAAILAARKDLDTVSN 393
Query: 649 DDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAG 706
D++ A + G ++K+R S K E VA +EA A+ +P++D + ++I PR G
Sbjct: 394 DEVGDAIERVMAGP-EKKDRVISDRKKELVAYHEAGHALVGACMPDYDAVAKVSIIPR-G 451
Query: 707 RELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADN-A 765
+ G G+ +R L + + V L R A+E+ +G+++++T + A
Sbjct: 452 QAGGLTFFTPSEERMESGLYSRSYLQNQMAVALGGRVAEEIVYGEEEVTTGASNDLQQVA 511
Query: 766 RVAARMYMIGGLSDKYRGV-----------------SNFWVTDRINEIDLEAMKILNLCY 808
VA +M G+SDK V + + D ID+E ++++ Y
Sbjct: 512 NVARQMITKFGMSDKIGPVALGQSQGGMFLGRDMSATRDFSEDTAATIDVEVSELVDTAY 571
Query: 809 ERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+RA ++L N++++D + + L+ ++T+ EDI L+
Sbjct: 572 KRATKVLSDNRSVLDEMASMLIERETIDTEDIQDLL 607
>UniRef100_Q67JH0 Cell division protein [Symbiobacterium thermophilum]
Length = 626
Score = 329 bits (843), Expect = 3e-88
Identities = 198/484 (40%), Positives = 286/484 (58%), Gaps = 26/484 (5%)
Query: 393 SGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPG 452
SG RV + R + V F DVAG+ +++ EL EIV F H + Y G +IP
Sbjct: 132 SGNRVMQFGKSRARLVTDDRKRVTFDDVAGIDEVKEELAEIVDFLKHPKRYLELGARIPK 191
Query: 453 GILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPS 512
G+LL GPPG GKTLLAKAVAGEAGV FFSIS S FVE++VGVGASRVR L+++AK+N+P
Sbjct: 192 GVLLYGPPGTGKTLLAKAVAGEAGVPFFSISGSDFVEMFVGVGASRVRDLFEQAKKNSPC 251
Query: 513 VVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPA 572
+VFIDE+DAVGR+RG G G ER+ TLNQLLV +DGF +I IA+TNRPD+LDPA
Sbjct: 252 IVFIDEIDAVGRQRGAGYGGGHDEREQTLNQLLVEMDGFSANEGIIIIAATNRPDVLDPA 311
Query: 573 LVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVE 632
L+RPGRFDR+I I +P GR+ I +VHA+ KP+ DVD E++A T G GA++AN++
Sbjct: 312 LLRPGRFDRQIVIDRPDLKGRLAIFQVHAKGKPLEPDVDLEVLAKRTPGFTGADIANLMN 371
Query: 633 VAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNL 690
AA+ R + ++S D+ A G ++K R S+++ A +EA AV L
Sbjct: 372 EAALLAARRRKKKISMQDVEDAIDRVLAGGPEKKSRVISEKEKRVTAYHEAGHAVVGHML 431
Query: 691 PNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFG 750
P+ D + ITI PR GR +GY + +N +++ + D +T+ L RAA+E+ FG
Sbjct: 432 PHMDPLHKITIIPR-GRAMGYTLFLPVEDRYN---ISKSEILDRMTMALGGRAAEEITFG 487
Query: 751 KDQLSTIWAETADNARVAARMYMIGGLSDKYRGVSNFWVTDRI----------------- 793
+ S + + A RM G+S+K ++ D +
Sbjct: 488 -EITSGAQDDIERTTQWARRMVTEWGMSEKLGPLTYGMKQDEVFLARDMTRLRNYSEEVA 546
Query: 794 NEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQ--LHGHAK 851
ID E K +++ Y+RA +IL +++ ++ + L+ K+TL +++ L++ L K
Sbjct: 547 GLIDEEVRKFVHMAYQRAIDILTEHRDALEKVSEVLLEKETLEGKELQDLLEQLLPPRPK 606
Query: 852 PIPI 855
P P+
Sbjct: 607 PEPL 610
>UniRef100_Q7NHF9 Cell division protein [Gloeobacter violaceus]
Length = 630
Score = 327 bits (838), Expect = 1e-87
Identities = 190/463 (41%), Positives = 282/463 (60%), Gaps = 23/463 (4%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E VKF DVAG+ + + EL+E+V+F E + G KIP G+LL GPPG GKTLLA
Sbjct: 165 MEAKTGVKFDDVAGIDEAKEELQEVVQFLKRPERFTAVGAKIPKGVLLVGPPGTGKTLLA 224
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VFIDE+DAVGR+RG
Sbjct: 225 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENAPCIVFIDEIDAVGRQRGA 284
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLLV +DGFEG +I IA+TNRPD+LD A++RPGRFDR+I + +P
Sbjct: 285 GIGGGNDEREQTLNQLLVEMDGFEGNTGIIIIAATNRPDVLDAAILRPGRFDRQITVDRP 344
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
GR+EILKVH+R K +A D+D +++A T G GA+L+N++ AAI R +TE++
Sbjct: 345 DMAGRLEILKVHSRNKKLAPDIDLDVIARRTPGFAGADLSNLLNEAAILAARRRQTEITM 404
Query: 649 DDLLQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRE 708
++ A G+ +K +A +E A+ L D ++ +TI PR GR
Sbjct: 405 REIDDATDRVIAGLEKPPLVDSKKKRLIAYHEVGHALVGTLLAEHDPVQKVTIIPR-GRA 463
Query: 709 LGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARV 767
G + + ++TR L IT L RAA+E+ FG+D+++T ++ + +
Sbjct: 464 GGLT---WFTPSEEQMLITRNQLLARITGALGGRAAEEVVFGEDEVTTGASSDLQQVSNL 520
Query: 768 AARMYMIGGLSD----KYRGVSNFWV-----------TDRINEIDLEAMKILNLCYERAK 812
A +M G+S+ G ++ D + +D + I+ C+ +A
Sbjct: 521 ARQMVTRFGMSELGLLSLTGGGEVFLGRDLMQRSDMSEDVASMVDEQVRAIVKQCHRQAV 580
Query: 813 EILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAKPIPI 855
+L +++ LMD +V+ L+ K+T+ E++ R+V P+P+
Sbjct: 581 SMLTEHRALMDRIVDVLLEKETVDGEELRRIV---SEVVPVPM 620
>UniRef100_Q8DHW1 Cell division protein [Synechococcus elongatus]
Length = 644
Score = 323 bits (828), Expect = 1e-86
Identities = 192/455 (42%), Positives = 277/455 (60%), Gaps = 19/455 (4%)
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + EL+E+V F + E + G +IP G+LL GPPG GKTLLA
Sbjct: 163 MEAQTGVTFGDVAGIEEAKEELQEVVTFLKNSEKFTSIGARIPKGVLLIGPPGTGKTLLA 222
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VGVGASRVR L+++AKENAP +VFIDE+DAVGR+RG
Sbjct: 223 KAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFRKAKENAPCLVFIDEIDAVGRQRGA 282
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG +I IA+TNRPD+LD AL+RPGRFDR+I + P
Sbjct: 283 GIGGGNDEREQTLNQLLTEMDGFEGNTGIIVIAATNRPDVLDAALLRPGRFDRQITVDLP 342
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVST 648
+ GR++IL+VHAR K IA +V E +A T G GAELAN++ AAI R + ++
Sbjct: 343 SYKGRLQILQVHARNKKIAPEVSLEAIARRTPGFSGAELANLLNEAAILTARRRKPAITN 402
Query: 649 DDLLQAAQMEERGM-LDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGR 707
++ A GM L SK+KW +A +E A+ L + D + +TI PR+G
Sbjct: 403 AEIDDAIDRVTIGMTLTPLLDSKKKW-LIAYHEVGHALLMTLLKHADPLNKVTIIPRSGG 461
Query: 708 ELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNAR 766
G+ + + + + G+ TR L D IT+ L RAA+ FG +++ ++ A
Sbjct: 462 VGGFAQQIFDEERVDSGLYTRAWLLDEITILLGGRAAEVEIFGDAEVTVGASSDLRAVAN 521
Query: 767 VAARMYMIGGLSD------KYRGVSNFWVTDRIN----------EIDLEAMKILNLCYER 810
+A M G+SD + G F D + +ID + +I+ CYE
Sbjct: 522 LAREMVTRYGMSDLGHLALETTGNEVFLGRDLMPRAEYSEAVAVQIDHQVREIVMHCYEI 581
Query: 811 AKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQ 845
A++++++++ +D LV L+ K+T+ ++ LV+
Sbjct: 582 ARKLIREHRVAIDKLVELLLEKETIDGDEFRALVR 616
>UniRef100_Q9WZ49 Cell division protein FtsH [Thermotoga maritima]
Length = 610
Score = 322 bits (824), Expect = 4e-86
Identities = 185/448 (41%), Positives = 272/448 (60%), Gaps = 22/448 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DV G + EL+E+V+F + R G ++P GILL GPPG GKTLLA+AVAGE
Sbjct: 158 VTFKDVGGAEEAIEELKEVVEFLKDPSKFNRIGARMPKGILLVGPPGTGKTLLARAVAGE 217
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
A V FF IS S FVE++VGVGA+RVR L+ +AK +AP +VFIDE+DAVGR RG G G
Sbjct: 218 ANVPFFHISGSDFVELFVGVGAARVRDLFAQAKAHAPCIVFIDEIDAVGRHRGAGLGGGH 277
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ TLNQLLV +DGF+ + +I +A+TNRPDILDPAL+RPGRFD+KI + P +GR
Sbjct: 278 DEREQTLNQLLVEMDGFDSKEGIIVMAATNRPDILDPALLRPGRFDKKIVVDPPDMLGRK 337
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL++H R KP+AEDV+ EI+A T G VGA+L N+V AA+ R+ R +++ D +A
Sbjct: 338 KILEIHTRNKPLAEDVNLEIIAKRTPGFVGADLENLVNEAALLAAREGRDKITMKDFEEA 397
Query: 655 AQMEERGMLDRKERSKEKWEQ-VAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVR 713
G + + K ++ +A +EA AV + +PN + + I+I PR + LGY
Sbjct: 398 IDRVIAGPARKSKLISPKEKRIIAYHEAGHAVVSTVVPNGEPVHRISIIPRGYKALGYTL 457
Query: 714 TMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYM 773
+ E + +++R L D +T L RAA+E+ FG D S + +A M
Sbjct: 458 HLPEEDKY---LVSRNELLDKLTALLGGRAAEEVVFG-DVTSGAANDIERATEIARNMVC 513
Query: 774 IGGLSDKYRGVS-----------------NFWVTDRINEIDLEAMKILNLCYERAKEILQ 816
G+S++ ++ + + ++ID E KI+ CYERAKEI++
Sbjct: 514 QLGMSEELGPLAWGKEEQEVFLGKEITRLRNYSEEVASKIDEEVKKIVTNCYERAKEIIR 573
Query: 817 QNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+ + +D +V L+ K+T+ +++ R++
Sbjct: 574 KYRKQLDNIVEILLEKETIEGDELRRIL 601
>UniRef100_Q8FZ12 Cell division protein FtsH [Brucella suis]
Length = 644
Score = 319 bits (817), Expect = 3e-85
Identities = 186/448 (41%), Positives = 267/448 (59%), Gaps = 22/448 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG+ + + +LEEIV+F + ++R G KIP G+LL GPPG GKTLLA++VAGE
Sbjct: 154 VTFQDVAGVDEAKQDLEEIVEFLRDPQKFQRLGGKIPRGVLLVGPPGTGKTLLARSVAGE 213
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
A V FF+IS S FVE++VGVGASRVR ++++AK+NAP ++FIDE+DAVGR RG G G
Sbjct: 214 ANVPFFTISGSDFVEMFVGVGASRVRDMFEQAKKNAPCIIFIDEIDAVGRHRGAGLGGGN 273
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ TLNQLLV +DGFE +I IA+TNRPD+LDPAL+RPGRFDR++ +P P +GR
Sbjct: 274 DEREQTLNQLLVEMDGFEANESIILIAATNRPDVLDPALLRPGRFDRQVVVPNPDIVGRE 333
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+ILKVH R P+A +VD ++VA T G GA+LAN+V AA+ R ++ V+ + +
Sbjct: 334 QILKVHVRNVPLAPNVDLKVVARGTPGFSGADLANLVNEAALMTARRNKRLVTMQEFEDS 393
Query: 655 AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
G R + E+ A +EA A+ A+N+P D + TI PR GR LG V
Sbjct: 394 KDKIMMGAERRSAMTPEEKTNTAYHEAGHAIVALNVPKADPVHKATIIPR-GRALGMVMQ 452
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNA-RVAARMYM 773
+ E ++ T + + + + R A+E+ FGK+ +++ + A ++A M
Sbjct: 453 LPEGDRYS---ATYTWMVSRLAIMMGGRVAEELKFGKENITSGASSDIQQATKLARSMVT 509
Query: 774 IGGLSDKYRGVS------------NFWVTDRINE-----IDLEAMKILNLCYERAKEILQ 816
G SDK V+ + T I+E ID E ++++ Y A IL
Sbjct: 510 QWGYSDKLGRVAYGDNQEEVFLGHSVSRTQNISEETAQIIDAEVRRLIDEAYAEATRILT 569
Query: 817 QNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+ K L L+ +TLT ++I L+
Sbjct: 570 KKKKDWIALAEGLLEYETLTGDEINELI 597
>UniRef100_Q8YIU9 CELL DIVISION PROTEIN FTSH [Brucella melitensis]
Length = 651
Score = 319 bits (817), Expect = 3e-85
Identities = 186/448 (41%), Positives = 267/448 (59%), Gaps = 22/448 (4%)
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG+ + + +LEEIV+F + ++R G KIP G+LL GPPG GKTLLA++VAGE
Sbjct: 161 VTFQDVAGVDEAKQDLEEIVEFLRDPQKFQRLGGKIPRGVLLVGPPGTGKTLLARSVAGE 220
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
A V FF+IS S FVE++VGVGASRVR ++++AK+NAP ++FIDE+DAVGR RG G G
Sbjct: 221 ANVPFFTISGSDFVEMFVGVGASRVRDMFEQAKKNAPCIIFIDEIDAVGRHRGAGLGGGN 280
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ TLNQLLV +DGFE +I IA+TNRPD+LDPAL+RPGRFDR++ +P P +GR
Sbjct: 281 DEREQTLNQLLVEMDGFEANESIILIAATNRPDVLDPALLRPGRFDRQVVVPNPDIVGRE 340
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+ILKVH R P+A +VD ++VA T G GA+LAN+V AA+ R ++ V+ + +
Sbjct: 341 QILKVHVRNVPLAPNVDLKVVARGTPGFSGADLANLVNEAALMAARRNKRLVTMQEFEDS 400
Query: 655 AQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRT 714
G R + E+ A +EA A+ A+N+P D + TI PR GR LG V
Sbjct: 401 KDKIMMGAERRSAMTPEEKTNTAYHEAGHAIVALNVPKADPVHKATIIPR-GRALGMVMQ 459
Query: 715 MLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNA-RVAARMYM 773
+ E ++ T + + + + R A+E+ FGK+ +++ + A ++A M
Sbjct: 460 LPEGDRYS---ATYTWMVSRLAIMMGGRVAEELKFGKENITSGASSDIQQATKLARSMVT 516
Query: 774 IGGLSDKYRGVS------------NFWVTDRINE-----IDLEAMKILNLCYERAKEILQ 816
G SDK V+ + T I+E ID E ++++ Y A IL
Sbjct: 517 QWGYSDKLGRVAYGDNQEEVFLGHSVSRTQNISEETAQIIDAEVRRLIDEAYAEATRILT 576
Query: 817 QNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+ K L L+ +TLT ++I L+
Sbjct: 577 KKKKDWIALAEGLLEYETLTGDEINELI 604
>UniRef100_UPI00003CC442 UPI00003CC442 UniRef100 entry
Length = 633
Score = 318 bits (814), Expect = 6e-85
Identities = 199/490 (40%), Positives = 277/490 (55%), Gaps = 31/490 (6%)
Query: 394 GARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGG 453
G+RV + Y + V+F DVAG + + EL E+V+F + G +IP G
Sbjct: 138 GSRVMNFGKSKAKLYNDEKKKVRFRDVAGADEEKQELVEVVEFLKDPRKFAEVGARIPKG 197
Query: 454 ILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSV 513
+LL GPPG GKTLLA+AVAGEAGV FFSIS S FVE++VGVGASRVR L++ AK+NAP +
Sbjct: 198 VLLVGPPGTGKTLLARAVAGEAGVPFFSISGSDFVEMFVGVGASRVRDLFENAKKNAPCI 257
Query: 514 VFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPAL 573
+FIDE+DAVGR+RG G G ER+ TLNQLLV +DGF +I IA+TNRPDILDPAL
Sbjct: 258 IFIDEIDAVGRQRGAGLGGGHDEREQTLNQLLVEMDGFGANEGIIIIAATNRPDILDPAL 317
Query: 574 VRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEV 633
+RPGRFDR+I + +P GR +LKVHAR KP+ E+++ +A+ T G GA+L N++
Sbjct: 318 LRPGRFDRQITVDRPDVNGREAVLKVHARNKPLDENINLRAIATRTPGFSGADLENLLNE 377
Query: 634 AAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERSKEKWEQ-VAINEAAMAVAAMNLPN 692
AA+ R + ++ D+ +A G + EK VA +EA V + L
Sbjct: 378 AALVAARQDKKKIDMSDIDEATDRVIAGPAKKSRVISEKERNIVAFHEAGHTVIGVVLDE 437
Query: 693 FDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKD 752
D + +TI PR G+ GY + + + +T+ L D IT L R A+E+ FG
Sbjct: 438 ADVVHKVTIVPR-GQAGGYAVMLPKEDRY---FMTKPELLDKITGLLGGRVAEEIVFG-- 491
Query: 753 QLSTIWAETADNAR-VAARMYMIGGLSDK-------------------YRGVSNFWVTDR 792
++ST A +A RM G+SDK + N+ +D
Sbjct: 492 EVSTGAHNDFQRATGIARRMVTEFGMSDKLGPMQFGSSQGGQVFLGRDFHSEQNY--SDA 549
Query: 793 I-NEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAK 851
I +EID+E I+ CY RAK+IL +N+ +D + L+ +TL E I L +G
Sbjct: 550 IAHEIDMEMQTIMKECYARAKQILTENRDKLDLIAKTLLEVETLDAEQINHLCD-YGRLP 608
Query: 852 PIPISVLDIR 861
P S D++
Sbjct: 609 ERPTSSADVK 618
>UniRef100_Q63HD4 Cell division protein [Bacillus cereus]
Length = 633
Score = 317 bits (812), Expect = 1e-84
Identities = 199/490 (40%), Positives = 276/490 (55%), Gaps = 31/490 (6%)
Query: 394 GARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGG 453
G+RV + Y + V+F DVAG + + EL E+V+F + G +IP G
Sbjct: 138 GSRVMNFGKSKAKLYNDEKKKVRFRDVAGADEEKQELVEVVEFLKDPRKFAEVGARIPKG 197
Query: 454 ILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSV 513
+LL GPPG GKTLLA+AVAGEAGV FFSIS S FVE++VGVGASRVR L++ AK+NAP +
Sbjct: 198 VLLVGPPGTGKTLLARAVAGEAGVPFFSISGSDFVEMFVGVGASRVRDLFENAKKNAPCI 257
Query: 514 VFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPAL 573
+FIDE+DAVGR+RG G G ER+ TLNQLLV +DGF +I IA+TNRPDILDPAL
Sbjct: 258 IFIDEIDAVGRQRGAGLGGGHDEREQTLNQLLVEMDGFGANEGIIIIAATNRPDILDPAL 317
Query: 574 VRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEV 633
+RPGRFDR+I + +P GR +LKVHAR KP+ E+++ +A+ T G GA+L N++
Sbjct: 318 LRPGRFDRQITVDRPDVNGREAVLKVHARNKPLDENINLRAIATRTPGFSGADLENLLNE 377
Query: 634 AAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERSKEKWEQ-VAINEAAMAVAAMNLPN 692
AA+ R + ++ D+ +A G + EK VA +EA V + L
Sbjct: 378 AALVAARQDKKKIDMSDIDEATDRVIAGPAKKSRVISEKERNIVAFHEAGHTVIGVVLDE 437
Query: 693 FDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKD 752
D + +TI PR G+ GY + + + +T+ L D IT L R A+E+ FG
Sbjct: 438 ADVVHKVTIVPR-GQAGGYAVMLPKEDRY---FMTKPELLDKITGLLGGRVAEEIVFG-- 491
Query: 753 QLSTIWAETADNAR-VAARMYMIGGLSDK-------------------YRGVSNFWVTDR 792
++ST A +A RM G+SDK + N+ +D
Sbjct: 492 EVSTGAHNDFQRATGIARRMVTEFGMSDKLGPMQFGSSQGGQVFLGRDFHSEQNY--SDA 549
Query: 793 I-NEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLVQLHGHAK 851
I +EID+E I+ CY RAK+IL N+ +D + L+ +TL E I L +G
Sbjct: 550 IAHEIDMEMQTIMKECYARAKQILTDNRDKLDLIAKTLLEVETLDAEQINHLCD-YGRLP 608
Query: 852 PIPISVLDIR 861
P S D++
Sbjct: 609 ERPTSSADVK 618
>UniRef100_Q82EE9 Putative cell division protein FtsH [Streptomyces avermitilis]
Length = 664
Score = 316 bits (810), Expect = 2e-84
Identities = 196/480 (40%), Positives = 271/480 (55%), Gaps = 25/480 (5%)
Query: 383 YLKMTKEFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEM 442
+L + + G+RV + + F+DVAG + EL EI +F
Sbjct: 125 FLFLMNQMQGGGSRVMNFGKSKAKLITKDTPKTTFSDVAGSDEAVEELHEIKEFLQEPAK 184
Query: 443 YRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSL 502
++ G KIP G+LL GPPG GKTLLA+AVAGEAGV F+SIS S FVE++VGVGASRVR L
Sbjct: 185 FQAVGAKIPKGVLLYGPPGTGKTLLARAVAGEAGVPFYSISGSDFVEMFVGVGASRVRDL 244
Query: 503 YQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIAS 562
+++AK NAP++VF+DE+DAVGR RG G G ER+ TLNQLLV +DGF+ +G VI IA+
Sbjct: 245 FEQAKANAPAIVFVDEIDAVGRHRGAGLGGGHDEREQTLNQLLVEMDGFDVKGGVILIAA 304
Query: 563 TNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGM 622
TNRPDILDPAL+RPGRFDR+I + +P GR+EILKVH + KP+A DVD VA T G
Sbjct: 305 TNRPDILDPALLRPGRFDRQIAVDRPDMQGRLEILKVHQKGKPVAPDVDLSAVARRTPGF 364
Query: 623 VGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKERSKEKWEQV-AINEA 681
GA+LAN++ AA+ R + V L +A G R +K +++ A +E
Sbjct: 365 TGADLANVLNEAALLTARSDKKLVDNSMLDEAIDRVVAGPQKRTRIMSDKEKKITAYHEG 424
Query: 682 AMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAP 741
A+ A PN D + ITI R GR LGY + + ++ TR + D + L
Sbjct: 425 GHALVAAASPNSDPVHKITILSR-GRALGYTMVLPDEDKYS---TTRNEMLDQLAYMLGG 480
Query: 742 RAADEMWFGKDQLSTIWAETADNARVAARMYMI--------------GGLSDKYRGVSNF 787
RAA+E+ F +T A + A AR + G ++ + G
Sbjct: 481 RAAEELVF--HDPTTGAANDIEKATATARAMVTQYGMTERLGAIKFGGDNTEPFLGREMS 538
Query: 788 WVTDRINE----IDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
D E +D E K++ + A EIL +N+ ++D LV +L+ K+TL+KE I +
Sbjct: 539 HPRDYSEEVAALVDEEVKKLIENAHNEAWEILVENRDVLDALVLQLLEKETLSKEQIAEV 598
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,389,734,344
Number of Sequences: 2790947
Number of extensions: 58708981
Number of successful extensions: 375491
Number of sequences better than 10.0: 7652
Number of HSP's better than 10.0 without gapping: 4058
Number of HSP's successfully gapped in prelim test: 3751
Number of HSP's that attempted gapping in prelim test: 323623
Number of HSP's gapped (non-prelim): 27424
length of query: 881
length of database: 848,049,833
effective HSP length: 136
effective length of query: 745
effective length of database: 468,481,041
effective search space: 349018375545
effective search space used: 349018375545
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 79 (35.0 bits)
Medicago: description of AC138130.19


