

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
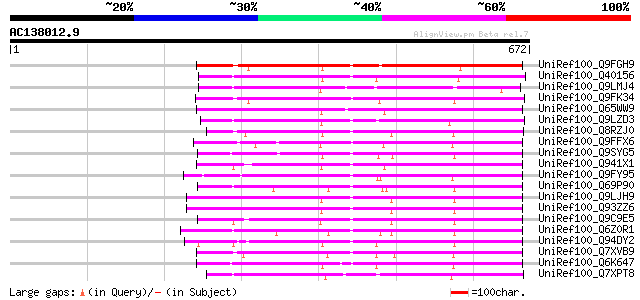
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FGH9 Leucine zipper protein [Arabidopsis thaliana] 302 2e-80
UniRef100_Q40156 L.esculentum protein with leucine zipper [Lycop... 276 1e-72
UniRef100_Q9LMJ4 F10K1.28 protein [Arabidopsis thaliana] 233 1e-59
UniRef100_Q9FK34 Gb|AAD25781.1 [Arabidopsis thaliana] 197 8e-49
UniRef100_Q65WW9 Putative leucine zipper protein [Oryza sativa] 188 4e-46
UniRef100_Q9LZD3 Hypothetical protein F12E4_320 [Arabidopsis tha... 186 3e-45
UniRef100_Q8RZJ0 Putative leucine zipper protein [Oryza sativa] 181 8e-44
UniRef100_Q9FFX6 Putative leucine zipper protein [Arabidopsis th... 179 2e-43
UniRef100_Q9SYG5 F15I1.17 protein [Arabidopsis thaliana] 177 1e-42
UniRef100_Q941X1 Leucine zipper protein-like [Oryza sativa] 176 2e-42
UniRef100_Q9FY95 Hypothetical protein T19L5_110 [Arabidopsis tha... 175 4e-42
UniRef100_Q69P90 Putative leucine zipper-containing protein [Ory... 173 1e-41
UniRef100_Q9LJH9 Gb|AAD25781.1 [Arabidopsis thaliana] 173 2e-41
UniRef100_Q93ZZ6 Hypothetical protein At3g14090 [Arabidopsis tha... 173 2e-41
UniRef100_Q9C9E5 Hypothetical protein T10D10.6 [Arabidopsis thal... 169 2e-40
UniRef100_Q6Z0R1 Putative leucine zipper-containing protein [Ory... 166 3e-39
UniRef100_Q94DY2 P0403C05.2 protein [Oryza sativa] 163 2e-38
UniRef100_Q7XVB9 OSJNBa0072D21.9 protein [Oryza sativa] 159 2e-37
UniRef100_Q6K647 Putative leucine zipper-containing protein [Ory... 155 5e-36
UniRef100_Q7XPT8 OSJNBa0088H09.18 protein [Oryza sativa] 154 8e-36
>UniRef100_Q9FGH9 Leucine zipper protein [Arabidopsis thaliana]
Length = 624
Score = 302 bits (774), Expect = 2e-80
Identities = 174/434 (40%), Positives = 264/434 (60%), Gaps = 20/434 (4%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKIN 301
D +D LPS +++LHE+ K ML AG+ K CS VY S R+ L++ + L K++
Sbjct: 193 DLIIDALPSATINDLHEMAKRMLGAGFGKACSHVYSSCRREFLEESMSR----LGLQKLS 248
Query: 302 TERERE---RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEA 358
E + + L+ RW+ A+++A +LFP E++ CD VF GFSSA F+E+C+ +
Sbjct: 249 IEEVHKMPWQELEDEIDRWIKAANVALRILFPSERRLCDRVFFGFSSAADLSFMEVCRGS 308
Query: 359 TFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNR 414
T QLLNFAD IA GS S RLFK++D+F + +L+P+FES+F + L NEA+ + R
Sbjct: 309 TIQLLNFADAIAIGSRSPERLFKVLDVFETMRDLMPEFESVFSDQFCSVLRNEAVTIWKR 368
Query: 415 LGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEY 474
LG+A R +FM++ N I R PA V G H +T VM+Y+ +ACR R+ LEQ+ EE
Sbjct: 369 LGEAIRGIFMELENLIRRDPAKAAVPG--GGLHPITRYVMNYLRAACRSRQTLEQVFEES 426
Query: 475 PEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSR 534
V + + S+ QM IM +L+ L VKS+ KD AL ++F++NN +I K
Sbjct: 427 NGVPS--KDSTLLTVQMSWIMELLESNLEVKSKVYKDPALCYVFLMNNGRYIVQKVKDGD 484
Query: 535 LETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKEL---LKEKIHLFN 591
L + G+DW + + K++Q Y+RS+W++++ LK+DN + L +KEK+ FN
Sbjct: 485 LGLLLGDDWIRKHNVKVKQYHMNYQRSSWNKMLGLLKVDNTAAGMNGLGKTMKEKLKQFN 544
Query: 592 NRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLH--GILGNQAYKYIKYG 649
+F+ IC+V S W ++ QL+ E+ S+ +L+PAYG F+GR G +G A KYIKYG
Sbjct: 545 IQFDEICKVHSTWVVFDEQLKEELKISLARLLVPAYGSFIGRFQNLGDIGKNADKYIKYG 604
Query: 650 MIEIQDLLNHLFLG 663
+ +I+ +N LF G
Sbjct: 605 VEDIEARINELFKG 618
>UniRef100_Q40156 L.esculentum protein with leucine zipper [Lycopersicon esculentum]
Length = 631
Score = 276 bits (706), Expect = 1e-72
Identities = 163/444 (36%), Positives = 255/444 (56%), Gaps = 23/444 (5%)
Query: 245 MDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTER 304
++ LP+ + +LHEI K M+ AGY+KECS Y R+ L++ L +++ + + ++
Sbjct: 187 IEALPAGIISDLHEIAKRMVAAGYDKECSHAYSVSRREFLEESL-SRLGLQKLSMDQVQK 245
Query: 305 ERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLN 364
+ L+ ++W+ A ++A +LFP E++ CD VF GF+S + F+E+ + +T QLLN
Sbjct: 246 MQWNELEDEIEKWVKAVNVALRILFPSERRLCDRVFFGFNSVSDLSFMEVSRGSTIQLLN 305
Query: 365 FADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNRLGDASR 420
FAD +A S + RLFK++D++ L +L+PKFE +F + L NEA+ + RLG+A R
Sbjct: 306 FADAVAISSRAPERLFKVLDVYEALRDLMPKFEFMFSDQYCVLLRNEALTIWRRLGEAIR 365
Query: 421 VLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE------- 473
+FM++ N I R PA V G H +T VM+Y+ +ACR R LEQ+ +E
Sbjct: 366 GIFMELENLIRRDPAKTPVPG--GGLHPITRYVMNYIRAACRSRITLEQVFKEIIVPSAS 423
Query: 474 ---YPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMN 530
Y E + +SS QM IM +L+ L KS+ KD AL +FM+NN +I
Sbjct: 424 AVDYREGDDRALSSSSLAVQMAWIMELLESNLETKSKIYKDSALLAVFMMNNERYIVQKV 483
Query: 531 KFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESIT----KELLKEK 586
K S L + G+DW + + AK++Q Y RS+W +V LK+DNN + LKEK
Sbjct: 484 KDSELGLLLGDDWVRKHAAKVKQYHVNYHRSSWSKVSGVLKIDNNAMSSPTGASRSLKEK 543
Query: 587 IHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAY--K 644
+ LFN+ FE IC+ QS W I+ QL+ E+ SV L PAY F+GRL + + +
Sbjct: 544 LKLFNSYFEEICKTQSTWIIFDEQLKEELRISVAGALSPAYRNFIGRLQSNNDSSRHTER 603
Query: 645 YIKYGMIEIQDLLNHLFLGNKMYG 668
+IK+ + +++ ++ LF G+ G
Sbjct: 604 HIKFSVEDLEARISELFQGSSGSG 627
>UniRef100_Q9LMJ4 F10K1.28 protein [Arabidopsis thaliana]
Length = 599
Score = 233 bits (594), Expect = 1e-59
Identities = 147/427 (34%), Positives = 236/427 (54%), Gaps = 20/427 (4%)
Query: 245 MDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTER 304
++ L S + +L+ I M+ G+ KECS VY S R+ L++ L +++ + + +
Sbjct: 178 IEALQSSVIGDLNAIAVRMVAGGFAKECSRVYSSRRREFLEESL-SRLHLRGLSMEEVQE 236
Query: 305 ERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGF--SSATSHCFIEICQEATFQL 362
+ L+ RW+ A + V FP E+ CD VFS SS T F+E+C+ T QL
Sbjct: 237 SPWQDLEDEIDRWIKAVTLIFHVFFPSERLLCDRVFSDLPVSSVTDLSFMEVCRGTTTQL 296
Query: 363 LNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAVRNRLGDA 418
LNFAD IA GS RLFK+VD++ + +L+PK E+LF + L +EA+A+ RLG+A
Sbjct: 297 LNFADAIALGSRLPERLFKVVDLYEAMQDLIPKMETLFSDRYCSPLRHEALAIHKRLGEA 356
Query: 419 SRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVH 478
R +FM++ N I R P + G H +T VM+Y+ +AC+ R+ LEQIL+ +
Sbjct: 357 IRGIFMELENLIRRDP--PKTAFPGGGIHPITRYVMNYLRAACKSRQSLEQILD---QTG 411
Query: 479 NEVEASSFFLK-QMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLET 537
NE + + L Q+ ++ +L+ L K +D +L +FM+NN +I K + L
Sbjct: 412 NETGSDTRPLSVQIIWVLELLESNLEGKKRTYRDPSLCFLFMMNNDKYILDKAKDNELGL 471
Query: 538 IFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAI 597
+ G DW + AK++Q Y+RS+W++V+ L+ D L E + LF ++F+ +
Sbjct: 472 VLGEDWIVKHAAKLRQYHSNYRRSSWNQVVGLLRTDG----PYPKLVENLRLFKSQFDEV 527
Query: 598 CRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLH---GILGNQAYKYIKYGMIEIQ 654
C+VQS W + QLR E+ SSV I+ PAY F+ RL I G + +I Y + +++
Sbjct: 528 CKVQSQWVVSDGQLREELRSSVAGIVSPAYSNFIRRLKESPEINGRRGEPFIPYTVEDVE 587
Query: 655 DLLNHLF 661
++ LF
Sbjct: 588 FIIKRLF 594
>UniRef100_Q9FK34 Gb|AAD25781.1 [Arabidopsis thaliana]
Length = 637
Score = 197 bits (501), Expect = 8e-49
Identities = 139/450 (30%), Positives = 226/450 (49%), Gaps = 30/450 (6%)
Query: 241 CDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKI 300
CD +D + + +L EI + M+ AGYEKEC VY S R+ L L+ +L K+
Sbjct: 185 CDLWVDLINPTAVEDLKEIAERMIRAGYEKECVQVYSSVRRDALDDCLM----ILGVEKL 240
Query: 301 NTERERE---RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQE 357
+ E ++ + +D ++W+ A I VL E+K CD +FS S+ CF E +
Sbjct: 241 SIEEVQKIDWKSMDEKMKKWIQAVKITVRVLLVGEKKICDEIFSSSESSKEVCFNETTKS 300
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLV-NEAIAVRNRLG 416
QLLNF + +A G S +LF+++D++ L N++ E + + V NE V LG
Sbjct: 301 CVMQLLNFGEAVAIGRRSSEKLFRILDMYDALANVLQTLEVMVTDCFVCNETKGVLEALG 360
Query: 417 DASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPE 476
DA+R F++ N + R +K+ ++ G+ H M VM+Y+ L +LE
Sbjct: 361 DAARGTFVEFENNV-RNETSKRPTTN-GEVHPMIRYVMNYMKLIVDYAVTLNSLLESNES 418
Query: 477 V----HNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKF 532
+ E S K++ ++ L+ L KS+ +D L+H+FM+NN +I K
Sbjct: 419 SGVSGDDSTEEMSPLAKRILGLITSLESNLEDKSKLYEDGGLQHVFMMNNIYYIVQKVKD 478
Query: 533 SRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN--------------NESI 578
S L + G+DW + + +I+Q Y R++W V+ L+ ++ + +
Sbjct: 479 SELGKLLGDDWVRKRRGQIRQYATGYLRASWSRVLSALRDESMGGSSSGSPSYGQRSNNS 538
Query: 579 TKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL 638
+K LKE+ FN FE + R+Q+AW + QLR E+ S+ ++PAY F GR L
Sbjct: 539 SKMALKERFRGFNASFEELYRLQTAWKVPDPQLREELRISISEKVIPAYRAFFGRNRSQL 598
Query: 639 --GNQAYKYIKYGMIEIQDLLNHLFLGNKM 666
G A KYIKY +++ L LF GN++
Sbjct: 599 EGGRHAGKYIKYTPDDLESYLPDLFEGNQL 628
>UniRef100_Q65WW9 Putative leucine zipper protein [Oryza sativa]
Length = 589
Score = 188 bits (478), Expect = 4e-46
Identities = 121/435 (27%), Positives = 220/435 (49%), Gaps = 16/435 (3%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKIN 301
D +D L + N+H+I + M+ AG+ +EC+ VY + R+ + + + ++ V P
Sbjct: 155 DVVIDALSPGSVANVHQIARRMVDAGFGRECAEVYAAARRGFVDESVA-RLGVRPRTAEE 213
Query: 302 TERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQ 361
L+ RW+ A ++ +L P E++ CD VF G + F+ + Q
Sbjct: 214 VHASSWEELEFDIARWIPAFNMVFRILIPSERRLCDRVFDGLAPFGDLAFVAAVRTQALQ 273
Query: 362 LLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEAIAVRNRLGD 417
L++F D I+ S + RLF++VD++ + +L+P + +F + +L E AV N LG
Sbjct: 274 LISFGDAISSSSRAPERLFRVVDMYEAVRDLLPDLDPVFADPYSAALRAEVTAVCNTLGS 333
Query: 418 ASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILE-EYPE 476
+ + +FM++ N I R PA +V + G H +T VM+Y+ +AC R+ LE+++E ++
Sbjct: 334 SIKGIFMELENLIRRDPA--RVAAQGGGIHPITRYVMNYLRAACGSRQTLEEVMEGDFGA 391
Query: 477 VHNEVEA------SSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMN 530
V A +S + IM +L + L +KS+ +D +L +F++NN +I
Sbjct: 392 VGGAAAAVDPDRPTSSLAVHIAWIMDVLHKNLDIKSKIYRDPSLACVFLMNNGKYIIQKV 451
Query: 531 KFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNN--ESITKELLKEKIH 588
S L + G++W + ++++ Y+R W +V L+ + +K+K+
Sbjct: 452 NDSELGVLLGDEWIKQMTNRVRRWSMDYQRVTWGKVTTVLQTGGPGVGGLPATAMKQKLR 511
Query: 589 LFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAYKYIKY 648
+FN F+ I VQS W I QLR ++ ++V ++P Y + RL + YIKY
Sbjct: 512 MFNTYFQEIYEVQSEWVIADEQLRVDVRAAVAEAVMPVYTALISRLKSSPEARHDLYIKY 571
Query: 649 GMIEIQDLLNHLFLG 663
+++ + HLF G
Sbjct: 572 TPEDVEACIQHLFEG 586
>UniRef100_Q9LZD3 Hypothetical protein F12E4_320 [Arabidopsis thaliana]
Length = 638
Score = 186 bits (471), Expect = 3e-45
Identities = 122/438 (27%), Positives = 220/438 (49%), Gaps = 25/438 (5%)
Query: 248 LPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERE 307
+PS L LH++ + M+ AG++++ +Y R +L++ L K+ V +K + +R +
Sbjct: 202 IPSRVLPLLHDLAQQMVQAGHQQQLLQIYRDTRSFVLEESL-KKLGVEKLSKEDVQRMQW 260
Query: 308 RYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFAD 367
L+ W+ IA +LF E++ CD +F GF S + CF E+ + LL+F D
Sbjct: 261 EVLEAKIGNWIHFMRIAVKLLFAGERQVCDQIFRGFDSLSDQCFAEVTVSSVSMLLSFGD 320
Query: 368 VIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEAIAVRNRLGDASRVLF 423
IA S +LF ++D++ + L + E++F + + A + RL ++ F
Sbjct: 321 AIARSKRSPEKLFVLLDMYEIMRELHTEIETIFKGKACLEIRDSATGLTKRLAQTAQETF 380
Query: 424 MKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEA 483
+ + V+ G H +T V++YV + L+Q+ E+ N ++
Sbjct: 381 GDFEEAVEKDATKTAVLD--GTVHPLTSYVINYVKFLFDYQTTLKQLFLEF---GNGDDS 435
Query: 484 SSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDW 543
+S +IM+ LQ L KS+ KD AL H+F++NN ++ + S + + G+DW
Sbjct: 436 NSQLASVTMRIMQALQNNLDGKSKQYKDPALTHLFLMNNIHYMVRSVRRSEAKDLLGDDW 495
Query: 544 FQNNKAKIQQNLDLYKRSAWDEVMD-------------FLKLDNNESITKELLKEKIHLF 590
Q ++ +QQ+ + YKR AW +++ L+ N+ +++ LLKE+ +F
Sbjct: 496 VQRHRRIVQQHANQYKRVAWTKILQSSSAQGLTSSGGGSLEGGNSSGVSRGLLKERFKMF 555
Query: 591 NNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKY 648
N +F+ + + QS W + ++LR + +V +LLPAY F+ R ++ G KYIKY
Sbjct: 556 NMQFDELHQRQSQWTVPDTELRESLRLAVAEVLLPAYRSFLKRFGPLVESGKNPQKYIKY 615
Query: 649 GMIEIQDLLNHLFLGNKM 666
+++ LL LF G M
Sbjct: 616 TAEDLERLLGELFEGKSM 633
>UniRef100_Q8RZJ0 Putative leucine zipper protein [Oryza sativa]
Length = 646
Score = 181 bits (458), Expect = 8e-44
Identities = 130/429 (30%), Positives = 214/429 (49%), Gaps = 26/429 (6%)
Query: 256 LHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE---RERERYLDT 312
L I +ML AGY E VY R+ L + L VL K++ E R LD
Sbjct: 222 LRGIADVMLRAGYGPELCQVYGEMRRDTLMECLA----VLGVDKMSLEEVQRVEWGVLDG 277
Query: 313 MFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAYG 372
++W+ A + L E++ C+ +F+ + A CF E + QLLNF D IA G
Sbjct: 278 KMKKWIQALKVVVRGLVAEERRICNQIFAADAEAEEDCFTEAAKGCVLQLLNFGDAIAIG 337
Query: 373 SLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNRLGDASRVLFMKMHN 428
S +LF+++ ++ L+ ++P+ E LF + EA+ + RLGDA R + N
Sbjct: 338 KRSSEKLFRILGMYEALDEVLPELEGLFSGDARDFIKEEAVGILMRLGDAVRGTVAEFAN 397
Query: 429 FIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFL 488
I + + + G+ H +T VM+YV R L Q+L+++ + E + +
Sbjct: 398 AIQGETSRRALPG--GEIHPLTRYVMNYVRLLADYSRSLNQLLKDW-DTELENGGDNVNM 454
Query: 489 KQMEQ----IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWF 544
+ Q ++ LQ K+ KS+ +D AL++IF++NN +I K S L+T+ G++W
Sbjct: 455 TPLGQCVLILITHLQAKIEEKSKLYEDEALQNIFLMNNLLYIVQKVKDSELKTLLGDNWI 514
Query: 545 QNNKAKIQQNLDLYKRSAWDEVMDFLKLD------NNESITKELLKEKIHLFNNRFEAIC 598
+ + +I++ Y RS+W V+ L+ D + S K LKE+ FN FE +
Sbjct: 515 RQRRGQIRRYSTGYLRSSWTRVLACLRDDGLPQTMGSSSALKASLKERFKNFNLAFEELY 574
Query: 599 RVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKYGMIEIQDL 656
+ Q+ W + QLR E+ S+ +LPAY FVGR G L G + +YIKY ++++
Sbjct: 575 KTQTTWKVVDPQLREELKISISEKVLPAYRSFVGRFRGQLEGGRNSARYIKYNPEDLENQ 634
Query: 657 LNHLFLGNK 665
++ F G +
Sbjct: 635 VSDFFEGRR 643
>UniRef100_Q9FFX6 Putative leucine zipper protein [Arabidopsis thaliana]
Length = 695
Score = 179 bits (454), Expect = 2e-43
Identities = 128/444 (28%), Positives = 216/444 (47%), Gaps = 26/444 (5%)
Query: 239 GKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEA 298
G + E P + + L +I + M GY EC VY+ R+ +L + L
Sbjct: 247 GDQEIEYPGYPEDVVVVLRKIAEKMKAGGYGWECREVYLVGRRNILMRTLKQDCEF---E 303
Query: 299 KINTERERERYLDTMFQR---WMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEIC 355
K++ + ++ DT+ + W +++ FP E K + +F G + F +
Sbjct: 304 KVSIDEVQKMSWDTLEREIPIWNKTFKDCSSLFFPGELKLAERIFPGDEG---NLFCIVT 360
Query: 356 QEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAV 411
Q L FA+ +A S +LFK++DI+ L + P E LFP + L NE +
Sbjct: 361 HGLAIQFLGFAEAVAMTRRSTEKLFKILDIYETLRDSFPAMEELFPEELRSELRNEVTSA 420
Query: 412 RNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQIL 471
R+RLG+ + +F + + I + ++K V G H +T M+Y+ +C + LEQ+
Sbjct: 421 RSRLGETAIHIFCDLEHSI-KSDSSKTPVPG-GAVHPLTRYTMNYLKYSCEYKDTLEQVF 478
Query: 472 EEYPEVHNEVE-----ASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI 526
+ + ++ E E +S F Q+ +IM +L L KS+ KD L IFM+NN +I
Sbjct: 479 KSHSKMEREEEEPVESGNSAFASQLMRIMELLDGNLETKSKQYKDIPLSCIFMMNNGRYI 538
Query: 527 EAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLK---LDNNESITKEL 582
K S + + G+ W + ++++ Y+R W +++ FL L +N I K
Sbjct: 539 VQKIKGSAEIHEVMGDTWCRRRSSELRNYHKNYQRETWGKLLGFLGHEGLMHNGKIVKPN 598
Query: 583 LKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GN 640
LKE+ FN F+ I + Q+ W + QL+ E+ S+ +++PAY F+ R L G
Sbjct: 599 LKERFKSFNATFDEIHKTQTTWVVNDEQLQSELRVSITAVMIPAYRAFMARFGQYLDPGR 658
Query: 641 QAYKYIKYGMIEIQDLLNHLFLGN 664
Q KY+KY +I+DL++ LF GN
Sbjct: 659 QTEKYVKYQPEDIEDLIDQLFEGN 682
>UniRef100_Q9SYG5 F15I1.17 protein [Arabidopsis thaliana]
Length = 622
Score = 177 bits (448), Expect = 1e-42
Identities = 122/448 (27%), Positives = 222/448 (49%), Gaps = 33/448 (7%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE 303
E+D + E + +L I + M+ AGY +EC VY + RK ++ + ++ ++ + +
Sbjct: 169 EIDLVTPEAVSDLRSIAQRMIGAGYSRECVQVYGNVRKSAMEM-IFKQLGIVKLGIGDVQ 227
Query: 304 RERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLL 363
R ++ +RW+ A+ + V+F E++ C+ +F G T CF+EI + + QL
Sbjct: 228 RLDWEAVEVKIRRWIRAAKVCVRVVFASEKRLCEQIFEGTMEET--CFMEIVKTSALQLF 285
Query: 364 NFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNRLGDAS 419
NF + I+ S +LFK++D+ + +L+P E +F +S ++ +A +++RL +A+
Sbjct: 286 NFPEAISISRRSPEKLFKILDLHDAITDLLPDMEEIFDSSSSESILVQATEIQSRLAEAA 345
Query: 420 RVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYP---- 475
R + + N +FR P+ V G H +T VM+Y++ R L ++ P
Sbjct: 346 RGILTEFENAVFREPSVVPVPG--GTIHPLTRYVMNYLNLISDYRETLIDLVMTKPCRGL 403
Query: 476 EVHNEVEASSFFLKQMEQI----------MRMLQRKLIVKSENCKDRALRHIFMLNNRSH 525
+ N+ + ++E I M MLQ L KS + +D L HIF++NN +
Sbjct: 404 KCTNDRNDPDMDISELEGISPLALHMIWTMVMLQFNLEEKSLHYRDEPLSHIFVMNNVHY 463
Query: 526 IEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN-------NES 577
I K S L + G+ + + +Q Y+R+ W V++ L+ + +
Sbjct: 464 IVQKVKSSPELMELIGDKYLRKLTGIFRQAATKYQRATWVRVLNSLRDEGLHVSGSFSSG 523
Query: 578 ITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI 637
++K L+E+ FN FE + R+QS W + +QLR E+ S+ L+PAY F+GR G
Sbjct: 524 VSKSALRERFKAFNTMFEEVHRIQSTWSVPDTQLREELRISLSEHLIPAYRSFLGRFRGH 583
Query: 638 L--GNQAYKYIKYGMIEIQDLLNHLFLG 663
+ G Y+KY + ++ + F G
Sbjct: 584 IESGRHPENYLKYSVENLETAVLDFFEG 611
>UniRef100_Q941X1 Leucine zipper protein-like [Oryza sativa]
Length = 652
Score = 176 bits (446), Expect = 2e-42
Identities = 118/444 (26%), Positives = 220/444 (48%), Gaps = 34/444 (7%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL---------LNKI 292
D +D LP + ++H+I + M+ AG+ +EC+ Y + R+ + + + ++++
Sbjct: 218 DVVIDALPPGSVSDVHQIARRMVDAGFGRECAEAYAAARRGFIDESVARLGIRARTIDEV 277
Query: 293 FVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFI 352
LP ++ + RW+ A + +L P E++ CD VF G + F+
Sbjct: 278 HSLPWEELEFD----------IARWIPAFKMVFRILIPSERRLCDRVFDGLAPYGDLAFV 327
Query: 353 EICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEA 408
+ QL++F D ++ S + RLF+++D++ + +L+P + +F + +L E
Sbjct: 328 AAVRTQVLQLISFGDAVSAASRAPERLFRVIDMYEAVRDLLPDLDPVFADPYSAALRAEV 387
Query: 409 IAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLE 468
AV N LG + + +FM++ N I R PA V G H +T VM+Y+ +AC R+ LE
Sbjct: 388 SAVCNTLGSSIKGIFMELENLIRRDPARVSVPG--GGIHPITRYVMNYLRAACGSRQTLE 445
Query: 469 QILE-EYPEVHNEVEA------SSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLN 521
+++E + V A +S + IM +L + L KS+ +D L IF++N
Sbjct: 446 EVMEGDLGAVGGAAIAVDPDRPTSSLAVHIAWIMDVLHKNLETKSKIYRDPPLASIFLMN 505
Query: 522 NRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNN--ESIT 579
N +I S L + G++W + +++++ Y+R AW +VM L+ S+
Sbjct: 506 NGKYIIHKVNDSELGVLLGDEWMKQMMSRVRRWSLEYQRGAWAKVMSVLQTGGPGIGSLP 565
Query: 580 KELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILG 639
+ L +K+ +FN E IC +QS W I QLR ++ +++ + + AY + RL
Sbjct: 566 AKALLQKLRMFNGYLEEICAIQSEWVIADEQLREDVRAAITDSVKSAYMGLISRLKSSPE 625
Query: 640 NQAYKYIKYGMIEIQDLLNHLFLG 663
+IK+ +++ + HLF G
Sbjct: 626 AAQDLFIKHSPEDVEARIQHLFEG 649
>UniRef100_Q9FY95 Hypothetical protein T19L5_110 [Arabidopsis thaliana]
Length = 653
Score = 175 bits (443), Expect = 4e-42
Identities = 130/467 (27%), Positives = 220/467 (46%), Gaps = 32/467 (6%)
Query: 225 SLNIIDR-LMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVL 283
S N DR +++++ + + + D P E + L +I M+ AGYE EC Y R+
Sbjct: 184 STNDSDRCVLQDHEEAEEESFHDFSP-ESISTLKKIAGAMISAGYEAECCMSYEMSRRHA 242
Query: 284 LQKGLLNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVF--S 341
++ L F + + +R L+ W++ +TVLFP E C+ VF
Sbjct: 243 FKEELTEVGFEGINVE-DVQRIGWESLEGEIASWISIVRRCSTVLFPGELSLCNAVFPDQ 301
Query: 342 GFSSATSHCFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP 401
SS F + T + L+F+ + S +LFK +D++ L +L+P E
Sbjct: 302 DHSSVRKRLFTGLVSAVTIRFLDFSGAVVLTKRSSEKLFKFLDMYETLRDLIPAVEQS-D 360
Query: 402 NSLVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSAC 461
+ L+ E + RLG+A+ +F ++ I V S G H +T M+Y+ AC
Sbjct: 361 SDLIQEIKLAQTRLGEAAVTIFGELEKSIKSDNGRTPVPS--GAVHPLTRYTMNYLKYAC 418
Query: 462 RKRRKLEQILEEY----------PEVH--------NEVEASSFFLKQMEQIMRMLQRKLI 503
+ L+Q+ + Y PE +E S F +QM ++M +L L
Sbjct: 419 EYKETLDQVFQHYEANQTDNKPEPETKPRQQQREDDEEYKVSAFARQMIRVMELLDANLE 478
Query: 504 VKSENCKDRALRHIFMLNNRSHIEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSA 562
+KS +D +LR IF++NN +I K S + + G W + +++Q Y+R
Sbjct: 479 IKSRLYRDPSLRFIFLMNNGRYILQKIKGSIEIRDLMGQSWTRKRSTELRQYHKSYQRET 538
Query: 563 WDEVMDFLK---LDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSV 619
W +V+ + L N ++K +LKE+ +FN F+ I + QS W + Q++ E+ S+
Sbjct: 539 WGKVLQCMNQEGLQVNGKVSKPVLKERFKIFNAMFDEIHKTQSTWIVSDEQMQSELRVSI 598
Query: 620 GNILLPAYGIFVGRL--HGILGNQAYKYIKYGMIEIQDLLNHLFLGN 664
++++PAY F GR H G Q KY+KY +I+ ++ LF GN
Sbjct: 599 SSLVIPAYRSFFGRYKQHLDSGKQTDKYVKYQPEDIESFIDDLFDGN 645
>UniRef100_Q69P90 Putative leucine zipper-containing protein [Oryza sativa]
Length = 638
Score = 173 bits (439), Expect = 1e-41
Identities = 134/462 (29%), Positives = 221/462 (47%), Gaps = 45/462 (9%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTE 303
E+D LP++ + +LH I M AGY +EC VY S RK + L ++ V + + +
Sbjct: 167 EIDLLPADAISDLHAIASRMAVAGYGRECVQVYASVRKPAVDSAL-RRLGVEKLSIGDVQ 225
Query: 304 RERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVF-------SGFSSATSHC-FIEIC 355
R L+ +RW+ A+ A +F E++ C L+F S ++AT F E
Sbjct: 226 RLEWEVLEAKIRRWIRAARAAVRGVFASERRLCFLIFHDLPLSSSTITTATHDAPFAEAV 285
Query: 356 QEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAV---- 411
+ A QL FA+ I+ G S +LFK++D+ + +L+P +F S E+I V
Sbjct: 286 KGAALQLFGFAEAISIGRRSPEKLFKIIDLHDAIADLLPDVSDIFAASKAGESIYVQAAE 345
Query: 412 -RNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQI 470
R+RL DA R + + N + R P+ V G H +T VM+Y S + L ++
Sbjct: 346 IRSRLADAVRGILSEFENAVLRDPSKTPVPG--GTIHPLTRYVMNYSSLISDYKTTLSEL 403
Query: 471 LEEYPEVHNEV------EASSF-------------FLKQMEQIMRMLQRKLIVKSENCKD 511
+ P + + A SF + I+ +L+ L K+ KD
Sbjct: 404 IVSRPLACSRIAPEGNENAPSFPDLDLADPDSQLPLAAHLIWIIVVLEHNLESKASLYKD 463
Query: 512 RALRHIFMLNNRSHIEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFL 570
AL H+F++NN +I K S L + G+++ + K + Y+R+AW ++++ L
Sbjct: 464 AALSHLFVMNNVHYIAHKIKDSPELRGLIGDEYLKQLTGKFRLAATRYQRTAWLKILNCL 523
Query: 571 KLDN-------NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNIL 623
+ + + ++K L+E+ FN FE RVQSAW++ +QLR E+ S+ L
Sbjct: 524 RDEGLHVSGGFSSGVSKSALRERFKSFNAAFEEAHRVQSAWYVPDTQLREELRISIAEKL 583
Query: 624 LPAYGIFVGRL--HGILGNQAYKYIKYGMIEIQDLLNHLFLG 663
LPAY F+GR H G YIKY + +++ + + F G
Sbjct: 584 LPAYRSFLGRFRHHIENGRHPELYIKYSVEDLETSVTNFFEG 625
>UniRef100_Q9LJH9 Gb|AAD25781.1 [Arabidopsis thaliana]
Length = 623
Score = 173 bits (438), Expect = 2e-41
Identities = 110/460 (23%), Positives = 227/460 (48%), Gaps = 28/460 (6%)
Query: 229 IDRLMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL 288
+ R Y EMD + E + +L IV+ M+ AGY +EC VY + RK ++ +
Sbjct: 157 LTRRRSSYRSTSSIREMDLISPEAVSDLRSIVQRMVAAGYSRECIQVYGTVRKSAME-AI 215
Query: 289 LNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATS 348
++ ++ + + +R ++ ++W+ A+ + V+F E++ C+ +F G +A
Sbjct: 216 FKQLGIVKISIGDVQRLEWEVVEGKIRKWIRAAKVCIRVVFSSEKRLCEQLFDGICTAMD 275
Query: 349 H-CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----S 403
CF+E + + +L F + I+ S +LFK++D+ L +++P E++F + +
Sbjct: 276 ETCFMETVKASALRLFTFPEAISISRRSPEKLFKILDLHDALTDMLPDIEAIFDSDSSDA 335
Query: 404 LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRK 463
+ +A+ +++RL +A+R + + N + R P+ V G H +T VM+Y+
Sbjct: 336 IRAQAVEIQSRLAEAARGILSEFENAVLREPSIVPVPG--GTIHPLTRYVMNYIVMISDY 393
Query: 464 RRKLEQILEEYPEVHNEVEASSFFLKQMEQ----------IMRMLQRKLIVKSENCKDRA 513
++ L+ ++ P ++ +++ ++ +L L KS++ +D +
Sbjct: 394 KQTLDDLIMSNPSTGSDPNTPDMDFTELDSKSPLDLHLIWLIVVLHFNLEEKSKHYRDTS 453
Query: 514 LRHIFMLNNRSHI-EAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL 572
L HIF++NN +I + + + L + G+ + + + Y+R+ W V++ L+
Sbjct: 454 LAHIFIMNNIHYIVQKVKRSPELREMIGDHYLRKLTGIFRHAATNYQRATWVRVLNSLRD 513
Query: 573 DN-------NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
+ + +++ L+E+ FN FE + R QS W + +QLR E+ S+ L+P
Sbjct: 514 EGLHVSGSFSSGVSRSALRERFKAFNTMFEEVHRTQSTWSVPDAQLREELRISLSEHLIP 573
Query: 626 AYGIFVGRLHGIL--GNQAYKYIKYGMIEIQDLLNHLFLG 663
AY F+GR G + G Y+KY + +I+ ++ F G
Sbjct: 574 AYRSFLGRFRGHIESGRHPENYLKYSVEDIETIVLDFFEG 613
>UniRef100_Q93ZZ6 Hypothetical protein At3g14090 [Arabidopsis thaliana]
Length = 503
Score = 173 bits (438), Expect = 2e-41
Identities = 110/460 (23%), Positives = 227/460 (48%), Gaps = 28/460 (6%)
Query: 229 IDRLMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL 288
+ R Y EMD + E + +L IV+ M+ AGY +EC VY + RK ++ +
Sbjct: 37 LTRRRSSYRSTSSIREMDLISPEAVSDLRSIVQRMVAAGYSRECIQVYGTVRKSAME-AI 95
Query: 289 LNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATS 348
++ ++ + + +R ++ ++W+ A+ + V+F E++ C+ +F G +A
Sbjct: 96 FKQLGIVKISIGDVQRLEWEVVEGKIRKWIRAAKVCIRVVFSSEKRLCEQLFDGICTAMD 155
Query: 349 H-CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----S 403
CF+E + + +L F + I+ S +LFK++D+ L +++P E++F + +
Sbjct: 156 ETCFMETVKASALRLFTFPEAISISRRSPEKLFKILDLHDALTDMLPDIEAIFDSDSSDA 215
Query: 404 LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRK 463
+ +A+ +++RL +A+R + + N + R P+ V G H +T VM+Y+
Sbjct: 216 IRAQAVEIQSRLAEAARGILSEFENAVLREPSIVPVPG--GTIHPLTRYVMNYIVMISDY 273
Query: 464 RRKLEQILEEYPEVHNEVEASSFFLKQMEQ----------IMRMLQRKLIVKSENCKDRA 513
++ L+ ++ P ++ +++ ++ +L L KS++ +D +
Sbjct: 274 KQTLDDLIMSNPSTGSDPNTPDMDFTELDSKSPLDLHLIWLIVVLHFNLEEKSKHYRDTS 333
Query: 514 LRHIFMLNNRSHI-EAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL 572
L HIF++NN +I + + + L + G+ + + + Y+R+ W V++ L+
Sbjct: 334 LAHIFIMNNIHYIVQKVKRSPELREMIGDHYLRKLTGIFRHAATNYQRATWVRVLNSLRD 393
Query: 573 DN-------NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLP 625
+ + +++ L+E+ FN FE + R QS W + +QLR E+ S+ L+P
Sbjct: 394 EGLHVSGSFSSGVSRSALRERFKAFNTMFEEVHRTQSTWSVPDAQLREELRISLSEHLIP 453
Query: 626 AYGIFVGRLHGIL--GNQAYKYIKYGMIEIQDLLNHLFLG 663
AY F+GR G + G Y+KY + +I+ ++ F G
Sbjct: 454 AYRSFLGRFRGHIESGRHPENYLKYSVEDIETIVLDFFEG 493
>UniRef100_Q9C9E5 Hypothetical protein T10D10.6 [Arabidopsis thaliana]
Length = 633
Score = 169 bits (428), Expect = 2e-40
Identities = 131/452 (28%), Positives = 218/452 (47%), Gaps = 39/452 (8%)
Query: 244 EMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL----LNKIFVLPEAK 299
E++ +P E + +L I + M+ AGY +EC VY S RK + + K+ + +
Sbjct: 177 EIELIPIESVIHLSWIARRMVSAGYLRECIQVYGSVRKSAVDSSFRRLGIEKLSIGDVQR 236
Query: 300 INTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSH--CFIEICQE 357
+N E L+ +RW+ A+ I V+F E+ C+ VF + H CF+E +
Sbjct: 237 LNWEA-----LEQKIRRWIRAAKICVRVVFASEKLLCEHVFESVGAVNIHEACFMETVKG 291
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFP----NSLVNEAIAVRN 413
QL NFA+ I+ S +LFK++D+ L L+P ES+F +S+ +A + +
Sbjct: 292 PAIQLFNFAEAISISRRSPEKLFKILDLHDALIELLPDIESVFDLKSSDSIRVQAAEILS 351
Query: 414 RLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE 473
RL +A+R + + N + R P+ V G H +T VM+Y+S R L ++
Sbjct: 352 RLAEAARGILSEFENAVLREPSRVPVPG--GTIHPLTRYVMNYISLISEYRPTLIDLIMS 409
Query: 474 YPEVH-NEVEASSFFLKQMEQ-----------IMRMLQRKLIVKSENCKDRALRHIFMLN 521
P + + F ++E I+ +LQ L KS+ K+ AL H+F++N
Sbjct: 410 KPSRNATDSNTPDFDFSELENNKGPLALHLIWIIVILQFNLEGKSKYYKNAALSHLFIMN 469
Query: 522 NRSHIEAMNKFS-RLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDN------ 574
N +I K S L + G+ + + K +Q Y+R+AW +V+ L+ +
Sbjct: 470 NAHYIVQKIKGSPELREMIGDLYLRKLTGKFRQAATYYQRAAWVKVLYCLRDEGLHTKGS 529
Query: 575 -NESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGR 633
+ +++ L+E+ FN FE + RVQS W + SQLR E+ S+ L PAY F+GR
Sbjct: 530 FSSGVSRSALRERFKSFNALFEEVHRVQSQWLVPDSQLREELKISILEKLSPAYRSFLGR 589
Query: 634 L--HGILGNQAYKYIKYGMIEIQDLLNHLFLG 663
H G YIK + +++ + LF G
Sbjct: 590 FRSHIESGKHPENYIKISVEDLETEVLDLFEG 621
>UniRef100_Q6Z0R1 Putative leucine zipper-containing protein [Oryza sativa]
Length = 632
Score = 166 bits (419), Expect = 3e-39
Identities = 139/487 (28%), Positives = 224/487 (45%), Gaps = 48/487 (9%)
Query: 222 QQDSLNIIDRLMEEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRK 281
+ DS++ + Y E+D LP + + +L I M AGY +EC+ VY S RK
Sbjct: 136 EDDSVSSLVGRRSSYRSLPSIREIDLLPDDAVSDLRAIASRMAAAGYGRECAQVYASVRK 195
Query: 282 VLLQKGLLNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFS 341
+ L ++ V + + +R + L+ +RW+ A+ A +F E++ C L+F
Sbjct: 196 PAVDASL-RRLGVERLSIGDVQRLEWKALEAKIRRWIRAARAAVRGVFASERRLCFLIFH 254
Query: 342 GF---------SSATSHC--FIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLN 390
++ +H F E + A QL FA+ I+ G S +LFK++D+ L+
Sbjct: 255 DLPISNITVTAAAPATHDTPFAEAVKGAALQLFGFAEAISIGRRSPEKLFKIIDLHDALS 314
Query: 391 NLVPKFESLFPNSLVNEAIAV-----RNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQ 445
+L+P +F S V E+I V R+RL DA R + + N + R P V G
Sbjct: 315 DLLPDVSDIFAASKVAESIYVQAAEIRSRLADAVRGILSEFENAVLRDPPKTAVPG--GT 372
Query: 446 HHQMTIQVMSYVSSACRKRRKLEQILEEYPEVH-------NEVEASSFFLKQMEQ----- 493
H +T VM+Y S + L +++ P NE+ S L E
Sbjct: 373 VHPLTRYVMNYSSLISDYKVTLSELIVSRPSASARLAAEGNELAPSLAELDLPEPDNQTP 432
Query: 494 -------IMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSR-LETIFGNDWFQ 545
I+ +L+ L K+ +D AL H+F++NN +I K S L + G+D+ +
Sbjct: 433 LAAHIIWIIVVLEHNLEGKASLYRDTALSHLFLMNNVYYIVHKVKDSPDLWNLIGDDYLK 492
Query: 546 NNKAKIQQNLDLYKRSAWDEVMDFLKLDN-------NESITKELLKEKIHLFNNRFEAIC 598
K Y+RSAW ++++ L+ + + I+K L+E+ FN FE
Sbjct: 493 RLTGKFTMAATNYQRSAWLKILNCLRDEGLHVSGGFSSGISKSALRERFRSFNAAFEEAH 552
Query: 599 RVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRL--HGILGNQAYKYIKYGMIEIQDL 656
RVQS W + +QLR E+ S+ L+PAY F+GR H G YIKY +++
Sbjct: 553 RVQSGWCVPDTQLREELRISISEKLVPAYRSFLGRFRHHIENGKHPELYIKYSAEDLEIA 612
Query: 657 LNHLFLG 663
+N F G
Sbjct: 613 VNDFFEG 619
>UniRef100_Q94DY2 P0403C05.2 protein [Oryza sativa]
Length = 601
Score = 163 bits (412), Expect = 2e-38
Identities = 123/477 (25%), Positives = 222/477 (45%), Gaps = 47/477 (9%)
Query: 227 NIIDRLMEEYNQGKCD--------------FEMDTLPSEKLHNLHEIVKLMLCAGYEKEC 272
++ D +N+ +CD F D + L + I M + Y+KEC
Sbjct: 127 SVDDFSSSSFNEEQCDGKTTQTETTGGSEYFATDLIQHGALSAVKSIANFMFLSEYDKEC 186
Query: 273 SAVYISWRKVLLQKGL----LNKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVL 328
S YIS R+ + + L ++K+ + E ++T + L ++ +RW A + V
Sbjct: 187 SQAYISTRQSAVDENLGSLRIDKLSM--EELLSTNWTK---LSSLIKRWNRAMKVFVQVY 241
Query: 329 FPIEQKFCDLVFSGFSSATSH-CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFV 387
E++ + VF S +T+ CF EI + QLL F + +A G +LF+++D++
Sbjct: 242 LTSEKRLSNHVFGELSESTADLCFYEISLSSVMQLLTFYESVAIGPPKPEKLFRLLDMYE 301
Query: 388 KLNNLVPKFESLFPNS----LVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSY 443
LN+L+P+ E LF ++ E V +LG++ R + + ++ +
Sbjct: 302 VLNDLLPEVEFLFQEGCDDIVLTEYNEVLLQLGESVRKTITEFKYAVQSYTSSNAMAR-- 359
Query: 444 GQHHQMTIQVMSYVSSACRKRRKLEQILEE-----------YPEVHNEVEASSFFLKQME 492
G+ H +T VM+Y+ + + L+ +L++ + N+ + ++
Sbjct: 360 GEVHPLTKYVMNYIKALTAYSKTLDSLLKDTDRRCQHFSTDIQSMANQCPHFTVSALHLQ 419
Query: 493 QIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQ 552
+ +L+ L S +D LR+IFM+NN ++ K S L+ G+DW + + K Q
Sbjct: 420 SVTAILEENLEAGSRLYRDDRLRNIFMMNNIYYMVQKVKNSELKIFLGDDWIRVHNRKFQ 479
Query: 553 QNLDLYKRSAWDEVMDFLKLDN----NESITKELLKEKIHLFNNRFEAICRVQSAWFIYG 608
Q Y+R++W V+ FL D + +++++KEK FN FE R Q+ W I
Sbjct: 480 QQAMSYERASWSHVLSFLSDDGLCAAGDGASRKIIKEKFKNFNLSFEDAYRTQTGWSIPD 539
Query: 609 SQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAY--KYIKYGMIEIQDLLNHLFLG 663
QLR ++ S+ ++ AY F GR + L + +YIKY +++ LL LF G
Sbjct: 540 DQLREDVRISISLKIIQAYRTFTGRYYSRLDGTRHLERYIKYKPEDLEKLLLDLFEG 596
>UniRef100_Q7XVB9 OSJNBa0072D21.9 protein [Oryza sativa]
Length = 688
Score = 159 bits (403), Expect = 2e-37
Identities = 124/454 (27%), Positives = 215/454 (47%), Gaps = 37/454 (8%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKIN 301
D D + E + +L I + M AGY E VY R+ LL + L VL +++
Sbjct: 231 DQVFDLVRPEAIDDLRSIAQRMDRAGYASELEQVYCGVRRDLLDECLA----VLGVERLS 286
Query: 302 ---TERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEA 358
+R + L+ ++W+ L E++ CD V + CF+E +
Sbjct: 287 IDEVQRMEWKLLNDKMKKWVHGVKTVVRSLLTGERRICDQVLAVSDELRDECFVESTKGC 346
Query: 359 TFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIA----VRNR 414
Q+LNF D +A S S +L +++D++ L ++P+ + LF + N+ I V R
Sbjct: 347 IMQILNFGDAVAVCSRSPEKLSRILDMYEALAEVIPELKELFFGNSGNDVICDLEGVLER 406
Query: 415 LGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE- 473
LGDA + ++ + + + + +++ G+ H MT VM+Y+ L+++L +
Sbjct: 407 LGDAVKGTLLEFGKVLQQESSRRPMMA--GEIHPMTRYVMNYLRLLVVYSDTLDKLLGDD 464
Query: 474 ------YPEVHNEVEASSFFLKQMEQIMR-------MLQRKLIVKSENCKDRALRHIFML 520
+ + H + +L+ + + R L+ L KS+ +D AL+ IF +
Sbjct: 465 SAGDVDHSDTHRGGDDEEEYLESLSPLGRHLVKLISYLEANLEEKSKLYEDGALQCIFSM 524
Query: 521 NNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLD------- 573
NN +I K S L I G+ W + + KI+QN Y R +W +V+ FLK D
Sbjct: 525 NNILYIVQKVKDSELGRILGDHWIRRRRGKIRQNSKNYLRISWTKVLSFLKDDAHGGRSG 584
Query: 574 -NNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVG 632
+ S +KEK FN F+ I R Q+ W + QLR E+ S+ ++PAY F+G
Sbjct: 585 SGSGSGNSSRIKEKFKNFNLAFDEIYRSQTLWKVPDPQLREELKISISENVIPAYRAFLG 644
Query: 633 RLHGIL--GNQAYKYIKYGMIEIQDLLNHLFLGN 664
R ++ G + +YIKY ++++ L+ LF G+
Sbjct: 645 RYGSLVDSGRNSGRYIKYTPEDLENQLSDLFEGS 678
>UniRef100_Q6K647 Putative leucine zipper-containing protein [Oryza sativa]
Length = 689
Score = 155 bits (391), Expect = 5e-36
Identities = 122/452 (26%), Positives = 216/452 (46%), Gaps = 33/452 (7%)
Query: 242 DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKIN 301
D D + E + +L I M AGY +E + Y R+ LL + L+ + V +
Sbjct: 229 DQVFDPVRPEAVDDLRAIADRMARAGYSRELADAYCGIRRDLLDE-YLSALGVERLSIDE 287
Query: 302 TERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQ 361
+R ++L+ ++W+ A VL E++ CD V S CFIE + Q
Sbjct: 288 VQRIEWKHLNDKMKKWVQAVKTVVRVLLAGERRLCDQVLSVSDELREECFIESTKGCIMQ 347
Query: 362 LLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRNRLGD 417
+L+F D +A S +L +++D++ L ++P+ + L S ++++ A +RLGD
Sbjct: 348 ILSFGDAVAVCPRSPEKLSRILDMYEALAEVIPEMKDLCLGSSGDGVISDVQANLDRLGD 407
Query: 418 ASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEE---- 473
A R + + ++ ++++ +++ G+ H MT VM+Y+ L+ +L++
Sbjct: 408 AIRGTLFEFGK-VLQLESSRRAMTA-GEIHPMTRYVMNYLRLLVVYSDTLDALLDDNADD 465
Query: 474 -------YPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI 526
+ +E+ + K++ +++ L+ L KS+ +D AL IF +NN +I
Sbjct: 466 QIDLARAEDQDQEHLESMTPLGKRLLKLISYLEANLEEKSKLYEDSALECIFSMNNLLYI 525
Query: 527 EAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLD------------- 573
+ S L I G+ W + KI+Q Y R +W +V+ FLK D
Sbjct: 526 VQKVRDSELGKILGDHWVKRRNGKIRQYSKSYLRISWMKVLSFLKDDGHGSGSGSSSGSG 585
Query: 574 NNESITKELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGR 633
+ S ++ +KEK FN FE I R Q+ W + QLR E+ S+ ++PAY F+GR
Sbjct: 586 SGHSSSRMSIKEKFKNFNLAFEEIYRNQTTWKVPDPQLREELKISISENVIPAYRAFLGR 645
Query: 634 LHGIL--GNQAYKYIKYGMIEIQDLLNHLFLG 663
+ G + KYIKY +++ L+ LF G
Sbjct: 646 YGSQVDGGRNSGKYIKYTPEDLESQLSDLFEG 677
>UniRef100_Q7XPT8 OSJNBa0088H09.18 protein [Oryza sativa]
Length = 634
Score = 154 bits (389), Expect = 8e-36
Identities = 112/432 (25%), Positives = 204/432 (46%), Gaps = 28/432 (6%)
Query: 256 LHEIVKLMLCAGYEKECSAVYISWRKVLLQKGLLNKIFVLPEAKINTERERERYLDTMFQ 315
L ++ + ++ AG +++CS +Y R L+ L + V +K ++ L++
Sbjct: 206 LAKLAQQLVQAGCQQQCSEIYSEARASALESSL-KSLGVEKLSKDEVQKMPWEILESKIG 264
Query: 316 RWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVIAYGSLS 375
W+ IA +LF E++ CD VF S CF +I + + LL+F + IA S
Sbjct: 265 NWIHFMRIAVKLLFAAERQLCDQVFECSQSLRDKCFAQITRNSLATLLSFGEAIAMSKRS 324
Query: 376 KWRLFKMVDIFVKLNNLVPKFESLFPNSLVNE----AIAVRNRLGDASRVLFMKMHNFIF 431
+LF ++D++ + L +++F ++ A+++ L ++ F +
Sbjct: 325 PEKLFVLLDMYEIMCELQADIDTIFVGESCSQMRESALSLTKCLAQTAQKTFSDFEEAVE 384
Query: 432 RVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQM 491
+ A + + G H +T V++YV + L+Q+ +E+ E S
Sbjct: 385 K--DATKNIHIDGTVHPLTSYVINYVKFLFDYQSTLKQLFQEFK---GEDGTGSELATVT 439
Query: 492 EQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKI 551
IM+ LQ L K++ KD AL HIF++NN +I + S + + G+DW Q ++ +
Sbjct: 440 MSIMQALQNNLDAKAKQYKDPALMHIFLMNNIHYIVKSVRRSEAKDLLGDDWIQRHRRIV 499
Query: 552 QQNLDLYKRSAWDEVMDFL----------------KLDNNESITKELLKEKIHLFNNRFE 595
QQN + Y+R AW +V+ L + N+ ++ +KE+ FN FE
Sbjct: 500 QQNANHYRRIAWSKVLQCLSGQGLTSSGGSGQVGSEGGNSSGASRAAVKERFRSFNVLFE 559
Query: 596 AICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAY--KYIKYGMIEI 653
I + Q W + ++LR + +V ILLPAY F+ R ++ N KY+K+ ++
Sbjct: 560 EIYQKQCGWSVPDTELRESLRLAVAEILLPAYRSFLKRFGPLIENSKAPGKYVKHTPEQV 619
Query: 654 QDLLNHLFLGNK 665
+ LL +LF G +
Sbjct: 620 ELLLANLFEGKQ 631
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,033,332,208
Number of Sequences: 2790947
Number of extensions: 41616834
Number of successful extensions: 153093
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 65
Number of HSP's successfully gapped in prelim test: 72
Number of HSP's that attempted gapping in prelim test: 152802
Number of HSP's gapped (non-prelim): 171
length of query: 672
length of database: 848,049,833
effective HSP length: 134
effective length of query: 538
effective length of database: 474,062,935
effective search space: 255045859030
effective search space used: 255045859030
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC138012.9


