

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137994.2 + phase: 0 /pseudo
(379 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
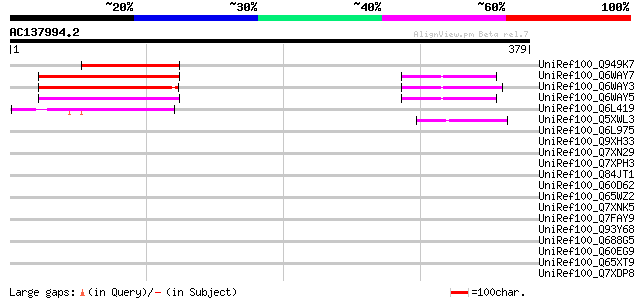
UniRef100_Q949K7 Putative gag polyprotein [Cicer arietinum] 89 2e-16
UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum] 79 3e-13
UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum] 74 5e-12
UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum] 74 9e-12
UniRef100_Q6L419 Putative gag polyprotein [Solanum demissum] 60 8e-08
UniRef100_Q5XWL3 Gag-pol polyprotein-like [Solanum tuberosum] 48 5e-04
UniRef100_Q6L975 GAG-POL [Vitis vinifera] 37 0.70
UniRef100_Q9XH33 Hypothetical protein [Nicotiana tabacum] 36 2.0
UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa] 35 2.6
UniRef100_Q7XPH3 OSJNBb0003B01.10 protein [Oryza sativa] 35 2.6
UniRef100_Q84JT1 Putative polyprotein [Oryza sativa] 35 3.5
UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solan... 35 3.5
UniRef100_Q65WZ2 Putative polyprotein [Oryza sativa] 35 3.5
UniRef100_Q7XNK5 OSJNBb0062B06.12 protein [Oryza sativa] 35 3.5
UniRef100_Q7FAY9 OSJNBb0005B05.9 protein [Oryza sativa] 35 3.5
UniRef100_Q93Y68 Putative gag-pol [Oryza sativa] 35 3.5
UniRef100_Q688G5 Hypothetical protein OSJNBa0039A21.9 [Oryza sat... 35 4.5
UniRef100_Q60EG9 Putative polyprotein [Oryza sativa] 35 4.5
UniRef100_Q65XT9 Putative polyprotein [Oryza sativa] 35 4.5
UniRef100_Q7XDP8 Putative polyprotein [Oryza sativa] 35 4.5
>UniRef100_Q949K7 Putative gag polyprotein [Cicer arietinum]
Length = 88
Score = 89.0 bits (219), Expect = 2e-16
Identities = 43/72 (59%), Positives = 54/72 (74%)
Query: 53 LCLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTA 112
+CLV +V IP KFKV +FEKYKG T PK HL MY RKM+ A ND+L IH+FQDSL+G +
Sbjct: 1 MCLVPDVTIPRKFKVLEFEKYKGATCPKNHLTMYGRKMASHAHNDKLLIHFFQDSLSGAS 60
Query: 113 LLGYMGLDKSDI 124
L YM L+++ I
Sbjct: 61 LSWYMHLERNRI 72
>UniRef100_Q6WAY7 Gag/pol polyprotein [Pisum sativum]
Length = 1814
Score = 78.6 bits (192), Expect = 3e-13
Identities = 36/103 (34%), Positives = 63/103 (60%)
Query: 22 RVDNLQEKYNEIQREMKALHRKELFGINAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKI 81
RVD K + + +++A+ + G + ++ LV + IP KFK P F+KY G + P+
Sbjct: 126 RVDERDRKVDALAEKIRAMECQNSLGFDVTNMGLVEGLRIPYKFKAPSFDKYNGTSCPRT 185
Query: 82 HLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYMGLDKSDI 124
H+ Y RK+S D++++++++FQDSL+G +L YM L + I
Sbjct: 186 HVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKRDSI 228
Score = 50.8 bits (120), Expect = 6e-05
Identities = 28/69 (40%), Positives = 37/69 (53%), Gaps = 1/69 (1%)
Query: 287 QIDLIPMKYADLFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFS 346
+ D +PM YA+L P LL +V + P P L Y N+ C FH GA H E C +
Sbjct: 413 RFDSLPMSYAELLPELLRLGMVELRTMAP-PTVLPPGYDANVRCDFHSGAPGHHTEKCRA 471
Query: 347 LKKIVQNLI 355
L+ VQ+LI
Sbjct: 472 LQHKVQDLI 480
>UniRef100_Q6WAY3 Gag/pol polyprotein [Pisum sativum]
Length = 2262
Score = 74.3 bits (181), Expect = 5e-12
Identities = 37/102 (36%), Positives = 63/102 (61%), Gaps = 1/102 (0%)
Query: 22 RVDNLQEKYNEIQREMKALHRKELFGINAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKI 81
R D K + + +++A+ + G + ++ LV + IP KFK P F+KY G + P+
Sbjct: 125 RNDARDRKVDALAEKIRAMECQNSLGFDVTNMGLVEGLRIPYKFKAPSFDKYNGTSCPRT 184
Query: 82 HLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYMGLDKSD 123
H+ Y RK+S D++++++++FQDSL+G +L YM L KSD
Sbjct: 185 HVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMEL-KSD 225
Score = 51.6 bits (122), Expect = 4e-05
Identities = 30/74 (40%), Positives = 38/74 (50%), Gaps = 1/74 (1%)
Query: 287 QIDLIPMKYADLFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFS 346
+ D +PM YA+L P LL LV P P L Y N+ C FH GA H E C +
Sbjct: 411 RFDSLPMSYAELLPELLRLGLVELCTMAP-PTVLPPGYDANVRCDFHSGAPGHHTEKCRA 469
Query: 347 LKKIVQNLIPENLL 360
L+ VQ+LI N +
Sbjct: 470 LQHKVQDLIDANAI 483
>UniRef100_Q6WAY5 Gag/pol polyprotein [Pisum sativum]
Length = 1105
Score = 73.6 bits (179), Expect = 9e-12
Identities = 34/103 (33%), Positives = 61/103 (59%)
Query: 22 RVDNLQEKYNEIQREMKALHRKELFGINAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKI 81
RVD K + + +++A+ + + ++ LV + IP KFK P F+KY + P+
Sbjct: 126 RVDERDRKVDALAEKIRAMECQNSLSFDVTNMGLVEGLRIPYKFKAPSFDKYNDTSCPRT 185
Query: 82 HLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYMGLDKSDI 124
H+ Y RK+S D++++++++FQDSL+G +L YM L + I
Sbjct: 186 HVQAYYRKISAYTDDEKMWMYFFQDSLSGASLDWYMELKRDSI 228
Score = 50.8 bits (120), Expect = 6e-05
Identities = 28/69 (40%), Positives = 37/69 (53%), Gaps = 1/69 (1%)
Query: 287 QIDLIPMKYADLFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFS 346
+ D +PM YA+L P LL +V + P P L Y N+ C FH GA H E C +
Sbjct: 413 RFDSLPMSYAELLPELLRLGMVELRTMAP-PTVLPPGYDANVRCDFHSGAPGHHTEKCRA 471
Query: 347 LKKIVQNLI 355
L+ VQ+LI
Sbjct: 472 LQHKVQDLI 480
>UniRef100_Q6L419 Putative gag polyprotein [Solanum demissum]
Length = 421
Score = 60.5 bits (145), Expect = 8e-08
Identities = 38/127 (29%), Positives = 64/127 (49%), Gaps = 16/127 (12%)
Query: 2 PTTQQNHGPIFHAESVKSYVRVDNLQEKYNEIQREMKALHR--KELFGINAY------DL 53
P T Q P ++VK+ E++ E+ ++MK+L +++ G+ + DL
Sbjct: 134 PQTHQYSSPFKIEKTVKN--------EEHEEMTKKMKSLEHSIRDMQGLGGHKGISFSDL 185
Query: 54 CLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTAL 113
C+ +V +P FK P+FEKY G+ P HL Y ++ +EL + YF +SL G A
Sbjct: 186 CMFPHVHLPTGFKTPNFEKYDGHGDPIAHLKRYYNQLRGAEGKEELLMAYFGESLVGIAS 245
Query: 114 LGYMGLD 120
++ D
Sbjct: 246 KWFIDQD 252
>UniRef100_Q5XWL3 Gag-pol polyprotein-like [Solanum tuberosum]
Length = 1716
Score = 47.8 bits (112), Expect = 5e-04
Identities = 25/66 (37%), Positives = 36/66 (53%), Gaps = 1/66 (1%)
Query: 298 LFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFSLKKIVQNLIPE 357
LF L +H P PV S YRP+ CA+H + HD E C +LK +Q+LI +
Sbjct: 433 LFERLTVAGYIHPVGPKPVDTS-SKFYRPDQRCAYHSNSVGHDTEDCINLKHKIQDLINQ 491
Query: 358 NLLPLE 363
N++ L+
Sbjct: 492 NVVSLQ 497
>UniRef100_Q6L975 GAG-POL [Vitis vinifera]
Length = 549
Score = 37.4 bits (85), Expect = 0.70
Identities = 22/64 (34%), Positives = 31/64 (48%), Gaps = 1/64 (1%)
Query: 62 PPK-FKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYMGLD 120
PP+ F VP F Y G + P H++ Y + M+ ND L F SL G AL + L
Sbjct: 15 PPRGFLVPKFSTYDGTSDPFDHIMHYRQLMTLDIGNDALLCKVFPASLQGQALSWFHRLP 74
Query: 121 KSDI 124
+ +
Sbjct: 75 PNSV 78
>UniRef100_Q9XH33 Hypothetical protein [Nicotiana tabacum]
Length = 152
Score = 35.8 bits (81), Expect = 2.0
Identities = 17/39 (43%), Positives = 20/39 (50%)
Query: 310 TKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFSLK 348
T P+ V LS PN CA+H G H IE C +LK
Sbjct: 109 TPIPVVVVKNLSQWINPNKTCAYHSGVKGHTIEECRTLK 147
>UniRef100_Q7XN29 OSJNBa0083I11.15 protein [Oryza sativa]
Length = 1563
Score = 35.4 bits (80), Expect = 2.6
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 6/71 (8%)
Query: 59 VVIPPKFKVPDFEKYKG----NTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALL 114
V +P +FK+PDF K+ G +T+ H+ Y+ + + D L + F+ SL+G+A
Sbjct: 467 VPLPNRFKIPDFSKFSGQEGVSTYE--HISRYLAQYGEASAVDALRVRMFRLSLSGSAFT 524
Query: 115 GYMGLDKSDIS 125
+ L ++
Sbjct: 525 WFSSLPYGSVN 535
>UniRef100_Q7XPH3 OSJNBb0003B01.10 protein [Oryza sativa]
Length = 1144
Score = 35.4 bits (80), Expect = 2.6
Identities = 20/55 (36%), Positives = 27/55 (48%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTA 112
NV PPKF+ EKY G+ P L +Y + +D + +YF SL G A
Sbjct: 194 NVRWPPKFRPNLTEKYDGSINPSEFLQIYTTVIEAAGGDDRVMANYFPMSLKGQA 248
>UniRef100_Q84JT1 Putative polyprotein [Oryza sativa]
Length = 1911
Score = 35.0 bits (79), Expect = 3.5
Identities = 20/67 (29%), Positives = 31/67 (45%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYM 117
NV PP+F+ EKY GN P L +Y + +D + ++F +L G A M
Sbjct: 230 NVRWPPRFRPTITEKYDGNVNPAEFLQIYTTGIEATGGDDRVMANFFPMALKGQARGWLM 289
Query: 118 GLDKSDI 124
L + +
Sbjct: 290 NLPPASV 296
>UniRef100_Q60D62 Putative gag/pol polyprotein, 3'-partial [Solanum demissum]
Length = 1784
Score = 35.0 bits (79), Expect = 3.5
Identities = 26/90 (28%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 277 ISTISTTG*TQIDLIPMKYADLFPILLERNL---VHTKAPLPVPARLSARYRPNLFCAFH 333
++T+ + I YA LF L N+ + K P P L CA+
Sbjct: 881 VATVGNGAKDEFTPIGESYASLFQKLRTLNVLSPIERKMSNPPPRNLDYSQH----CAYC 936
Query: 334 QGASCHDIEHCFSLKKIVQNLIPENLLPLE 363
A H+IE C+ LKK +Q+LI + +E
Sbjct: 937 SDAPGHNIERCWYLKKAIQDLIDTRRIIVE 966
>UniRef100_Q65WZ2 Putative polyprotein [Oryza sativa]
Length = 1490
Score = 35.0 bits (79), Expect = 3.5
Identities = 19/69 (27%), Positives = 35/69 (50%), Gaps = 2/69 (2%)
Query: 59 VVIPPKFKVPDFEKYKG--NTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGY 116
V +P ++KVPDF K+ G N H+ ++ + + D L + F SL+G+A +
Sbjct: 246 VPLPSRYKVPDFSKFSGQDNVSTYEHISRFLAQCGEASAIDTLRVRLFPLSLSGSAFTWF 305
Query: 117 MGLDKSDIS 125
L + ++
Sbjct: 306 SSLPYNSVN 314
>UniRef100_Q7XNK5 OSJNBb0062B06.12 protein [Oryza sativa]
Length = 1737
Score = 35.0 bits (79), Expect = 3.5
Identities = 20/67 (29%), Positives = 31/67 (45%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYM 117
NV PP+F+ EKY GN P L +Y + +D + ++F +L G A M
Sbjct: 222 NVRWPPRFRPTIAEKYDGNVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLM 281
Query: 118 GLDKSDI 124
L + +
Sbjct: 282 NLPPASV 288
>UniRef100_Q7FAY9 OSJNBb0005B05.9 protein [Oryza sativa]
Length = 847
Score = 35.0 bits (79), Expect = 3.5
Identities = 22/79 (27%), Positives = 36/79 (44%), Gaps = 7/79 (8%)
Query: 49 NAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKI--HLVMYVRKMSYQADNDELFIHYFQD 106
N YD+ V PP +K+P+F K+ G H+ ++ + + D + F
Sbjct: 154 NFYDV-----VPYPPDYKIPEFAKFSGKDSRTTCEHVGQFLAQCGRASSMDTCKLRLFSL 208
Query: 107 SLTGTALLGYMGLDKSDIS 125
SL+GTA + L + IS
Sbjct: 209 SLSGTAFTWFTSLPSNSIS 227
>UniRef100_Q93Y68 Putative gag-pol [Oryza sativa]
Length = 1829
Score = 35.0 bits (79), Expect = 3.5
Identities = 21/67 (31%), Positives = 27/67 (39%)
Query: 59 VVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYMG 118
V P + P F ++ G + HL + A N L +H F SLTG A Y
Sbjct: 1005 VPFPSGWHPPKFRQFDGTGDAREHLAYFEAACGDTASNPSLLLHQFSGSLTGPAFHWYSR 1064
Query: 119 LDKSDIS 125
L IS
Sbjct: 1065 LPIGSIS 1071
>UniRef100_Q688G5 Hypothetical protein OSJNBa0039A21.9 [Oryza sativa]
Length = 502
Score = 34.7 bits (78), Expect = 4.5
Identities = 20/67 (29%), Positives = 31/67 (45%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYM 117
NV PKF+ EKY GN P L +Y + +D + ++F +L G A + M
Sbjct: 228 NVRWTPKFRPSLTEKYDGNVNPSEFLQIYTAIIEAAGGDDRVMANFFPMTLKGQARVWLM 287
Query: 118 GLDKSDI 124
L + +
Sbjct: 288 NLPPASV 294
>UniRef100_Q60EG9 Putative polyprotein [Oryza sativa]
Length = 1626
Score = 34.7 bits (78), Expect = 4.5
Identities = 19/55 (34%), Positives = 28/55 (50%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTA 112
NV PPKF+ EKY G+ P L +Y + + +D + +YF +L G A
Sbjct: 241 NVRWPPKFRPNLTEKYDGSINPSEFLQIYTTVIVAEGGDDRVMANYFPIALKGQA 295
>UniRef100_Q65XT9 Putative polyprotein [Oryza sativa]
Length = 1823
Score = 34.7 bits (78), Expect = 4.5
Identities = 19/67 (28%), Positives = 32/67 (47%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYM 117
NV PP+F+ EKY G+ P L +Y + +D + +++F +L G A M
Sbjct: 230 NVRWPPRFRPTITEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMVNFFPMALKGQARGWLM 289
Query: 118 GLDKSDI 124
L + +
Sbjct: 290 NLPPASV 296
>UniRef100_Q7XDP8 Putative polyprotein [Oryza sativa]
Length = 465
Score = 34.7 bits (78), Expect = 4.5
Identities = 19/67 (28%), Positives = 32/67 (47%)
Query: 58 NVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQADNDELFIHYFQDSLTGTALLGYM 117
NV PP+F+ EKY G+ P L +Y + +D + +++F +L G A M
Sbjct: 230 NVRWPPRFRPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMVNFFPMALKGQARGWLM 289
Query: 118 GLDKSDI 124
L + +
Sbjct: 290 NLPPASV 296
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.340 0.147 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 577,409,530
Number of Sequences: 2790947
Number of extensions: 22034664
Number of successful extensions: 69849
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 22
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 69817
Number of HSP's gapped (non-prelim): 39
length of query: 379
length of database: 848,049,833
effective HSP length: 129
effective length of query: 250
effective length of database: 488,017,670
effective search space: 122004417500
effective search space used: 122004417500
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC137994.2


