

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
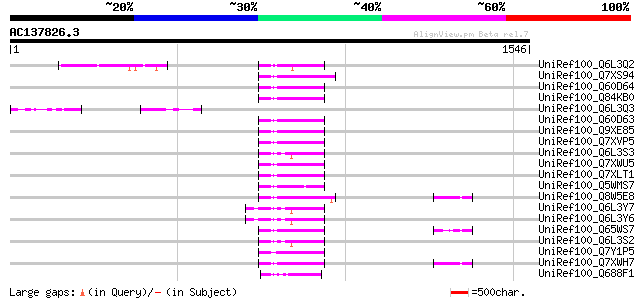
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6L3Q2 Putative polyprotein, 3'-partial [Solanum demis... 99 8e-19
UniRef100_Q7XS94 OSJNBa0074B10.6 protein [Oryza sativa] 94 3e-17
UniRef100_Q60D64 Putative polyprotein [Oryza sativa] 92 1e-16
UniRef100_Q84KB0 Pol protein [Cucumis melo] 92 1e-16
UniRef100_Q6L3Q3 Hypothetical protein [Solanum demissum] 92 2e-16
UniRef100_Q60D63 Putative polyprotein [Oryza sativa] 91 3e-16
UniRef100_Q9XE85 Polyprotein [Sorghum bicolor] 91 3e-16
UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa] 91 3e-16
UniRef100_Q6L3S3 Putative gag-pol polyprotein [Solanum demissum] 91 3e-16
UniRef100_Q7XWU5 OSJNBa0065B15.6 protein [Oryza sativa] 91 4e-16
UniRef100_Q7XLT1 OSJNBa0057M08.3 protein [Oryza sativa] 90 5e-16
UniRef100_Q5WMS7 Putative polyprotein [Oryza sativa] 90 6e-16
UniRef100_Q8W5E8 Putative polyprotein [Oryza sativa] 90 6e-16
UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum] 89 8e-16
UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum] 89 8e-16
UniRef100_Q65WS7 Putative polyprotein [Oryza sativa] 89 8e-16
UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demis... 89 8e-16
UniRef100_Q7Y1P5 Putative polyprotein [Oryza sativa] 89 1e-15
UniRef100_Q7XWH7 OSJNBa0085C10.17 protein [Oryza sativa] 89 1e-15
UniRef100_Q688F1 Hypothetical protein OSJNBa0035I01.9 [Oryza sat... 69 1e-09
>UniRef100_Q6L3Q2 Putative polyprotein, 3'-partial [Solanum demissum]
Length = 1622
Score = 99.4 bits (246), Expect = 8e-19
Identities = 78/207 (37%), Positives = 104/207 (49%), Gaps = 14/207 (6%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D +M MC DYRQLN+VT +N+Y LP IDDLF+QL AS +SK+ L +
Sbjct: 858 DGTMRMCIDYRQLNKVTVKNRYPLPRIDDLFDQLQGASV---FSKIDLRFDYH---QLRI 911
Query: 800 KGIQDPLWAL*VSSDIVWVDKCSFGCHEFD*-LCDQMTLGF------ILIVCVDDILLFL 852
+ P A + SFG D MT F +IV +DDIL++
Sbjct: 912 RAADIPKTAFRTRYGHYELLVMSFGLTNAPAAFMDLMTRVFRPYLDSFVIVFIDDILIYS 971
Query: 853 ESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEPVMN 911
S+ D HL +V Q+LYA F F + +G+VV E IMVD IE + +
Sbjct: 972 RSRGDHEQHLRVVLQTLRDQRLYAKFSKCQFWLDSVAFLGHVVSKEGIMVDPAKIEAIRD 1031
Query: 912 WV*PSFVIESRGFMGLASYYRGFEDAF 938
W P+ V E R F+GLA YYR F ++F
Sbjct: 1032 WARPTSVTEIRSFVGLAGYYRRFVESF 1058
Score = 63.9 bits (154), Expect = 4e-08
Identities = 82/347 (23%), Positives = 141/347 (40%), Gaps = 32/347 (9%)
Query: 146 NRPVMTGNEHKMLTKFLKLMSYMFFGSESEDTFKFILHYYERLNKLVIVKQYGVEIVTLQ 205
+RP M + KML FL+L F G+ ED +F+ +ERL L +V+ G + + Q
Sbjct: 282 DRP-MPLEDQKMLGVFLRLTPPRFSGAVGEDAHEFLTTCHERLRTLGLVESRGADFIAYQ 340
Query: 206 FRGEAKQW*RAYVKCRSSTFPPLSWMQFYALF*KDMCSR-LSEIVRWVISLL*SMVVSL* 264
G A+QW R +++ R PP+SW +F F R + + +R L +S
Sbjct: 341 LDGPARQWWRTFIQTRPVGSPPVSWDEFSEAFLAKFIPRSVRDRLRDQFCRLEQGSMS-- 398
Query: 265 LIMRLSSMICLDMIHNWCLL--KSTGFAYLLRELNFEL*VFRVHMTCARKSSNKAIDFVK 322
+ + H +L + +R L +L + + A +S +D
Sbjct: 399 -VAEYETRFHELSRHAEMILPTEEERVRCFVRGLRLQLRLETQSLVSAGRSFLDVVDHAS 457
Query: 323 KVEGVKRDRDAKAFSKKDKSKGNF*VSYSRGKG---------RMTLVDQPIQSTMPSS-- 371
+E ++R+ + K+ + +G++ S R + R +P Q+++ +S
Sbjct: 458 TMEHLRREAQGGS-EKRARHEGSYSDSRPRSRDYHDRVGRRFRQGQSSRPTQASLQASDG 516
Query: 372 ----TSNYRGTPHHNFIQYS*GVVSSASVRPSFYRTFYNCGDFRHISRYCPKPRILDLMQ 427
S+ +G G SS S RP + Y CG+ H +R CP+ +
Sbjct: 517 GRGEASSSQGREVFQSGSSGHGSRSSGS-RPR--QGCYECGEMGHWARDCPQHSRIAPAP 573
Query: 428 K*TRAVVPA------SNDKNSRGHPRGW*GGNHRGCEGRENGNVGRG 468
+PA ++ RG +G GG G G + GRG
Sbjct: 574 HAAPLALPAPPVRGRGQGQDRRGGQQGIRGGPRGGRSGGRSDGQGRG 620
Score = 48.5 bits (114), Expect = 0.002
Identities = 35/118 (29%), Positives = 56/118 (46%), Gaps = 10/118 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLW-----ENA* 1318
RLTK++HF+ V+ ++A++L+++Y+R P+ + W E
Sbjct: 1422 RLTKSAHFLPVQCSFSAERLARIYIREVVRLHGVPVSIISDRGSQFTSNFWRTFQDELGT 1481
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
V T F + TI+ +R C +DF G WD+FL L EF+YNN+Y
Sbjct: 1482 RVDLSTAFHPQTDGQSERTIQVLEDMLRA-----CVMDFGGQWDQFLPLAEFAYNNSY 1534
>UniRef100_Q7XS94 OSJNBa0074B10.6 protein [Oryza sativa]
Length = 1447
Score = 94.4 bits (233), Expect = 3e-17
Identities = 81/247 (32%), Positives = 124/247 (49%), Gaps = 24/247 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 587 DHTQRMCVDYRALNEVTIKNKYPLPRIDDLFDQLEGATV---FSKIDLRS------GYHQ 637
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 638 LRIREEDIPKTAFTTRYGLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDIL 697
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ ++K + +HL + + +LYA F F ++ +G+V+ + + VD N+E
Sbjct: 698 IYSKTKEEHEEHLRLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISSGGVAVDPSNVES 757
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAFDH----NTYLGTTSVT*SLIINYYASQLIL 964
V++W P V E R F+GLA YYR F + F T L V + AS+L L
Sbjct: 758 VLSWKQPKTVSEIRSFLGLAGYYRRFIENFSKIARPMTRLLQKEVKKGFQVYCDASRLGL 817
Query: 965 GVVLMQD 971
G VLMQ+
Sbjct: 818 GCVLMQE 824
Score = 47.4 bits (111), Expect = 0.004
Identities = 39/118 (33%), Positives = 54/118 (45%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*-EVGH 1322
RLTK +HFI VK Y +L+++YM R P + W+ E+G
Sbjct: 1113 RLTKVAHFIPVKTTYTGNKLAELYMARVVCLHGVPKKIVSDRGSQFTSKFWQKLQVEMGT 1172
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C + F G WDK L EFSYNN+Y S
Sbjct: 1173 RLNFSTAYHPQTDGQTERVNQ--ILEDMLRACVLAFGGSWDKNLPYAEFSYNNSYQAS 1228
>UniRef100_Q60D64 Putative polyprotein [Oryza sativa]
Length = 1777
Score = 92.0 bits (227), Expect = 1e-16
Identities = 71/210 (33%), Positives = 107/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A +SK+ L G +
Sbjct: 887 DKTKRMCVDYRALNEVTIKNKYPLPRIDDLFDQLKGAKV---FSKIDLRS------GYHQ 937
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A ++ SFG F L +++ + F+ ++V +DDIL
Sbjct: 938 LRIREEDIPKTAFTTRYELYECTVMSFGLTNAPAFFMNLMNKVFMEFLDKFVVVFIDDIL 997
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ ES+ + HL +V + +LYA F F ++ +G+V+ + + VD N+E
Sbjct: 998 IYSESEEEHEQHLRLVLEKLKEHQLYAKFSKCDFWLTEVKFLGHVITAQGVAVDPSNVES 1057
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V W P V + R F+GLA YYR F + F
Sbjct: 1058 VTKWTPPKTVSQIRSFLGLAGYYRRFIENF 1087
Score = 47.8 bits (112), Expect = 0.003
Identities = 38/118 (32%), Positives = 55/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI V Y ++L+++Y+ R P + W+ E +G
Sbjct: 1443 RLTKVAHFIPVHTTYTGKRLAELYLSRIMCLHGVPKKIVSDRGSQFTSKFWQKLQEELGT 1502
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1503 RLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGAWDKSLPYAEFSYNNSYQAS 1558
>UniRef100_Q84KB0 Pol protein [Cucumis melo]
Length = 923
Score = 92.0 bits (227), Expect = 1e-16
Identities = 76/210 (36%), Positives = 107/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D SM +C DYR+LN+VT +N+Y LP IDDLF+QL A+ +SK+ L G +
Sbjct: 40 DGSMRLCIDYRELNKVTVKNRYPLPRIDDLFDQLQGATV---FSKIDLRS------GYHQ 90
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I+D P A SFG F L +++ F+ +IV +DDIL
Sbjct: 91 LRIKDEDVPKTAFRSRYGHYQFIVMSFGLTNAPAVFMDLMNRVFREFLDTFVIVFIDDIL 150
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +++ + +HL +V KLYA F F Q+ +G+VV + VD IE
Sbjct: 151 IYSKTEAEHEEHLRMVLQTLRDNKLYAKFSKCEFWLKQVSFLGHVVSKAGVSVDPAKIEA 210
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V W PS V E R F+GLA YYR F + F
Sbjct: 211 VTGWTRPSTVSEVRSFLGLAGYYRRFVENF 240
Score = 41.6 bits (96), Expect = 0.19
Identities = 34/115 (29%), Positives = 52/115 (44%), Gaps = 4/115 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK++HF+ K Y A + +++YM P+ + W+ +G
Sbjct: 599 RLTKSAHFVPGKSTYTASKWAQLYMSEIVRLHGVPVSIVSDRDARFTSKFWKGLQTAMGT 658
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
F P ++G T R N V + C ++F G WD L L EF+YNN+Y
Sbjct: 659 RLDFSTAFHPQTDGQTERLNQ--VLEDMLRACALEFPGSWDSHLHLMEFAYNNSY 711
>UniRef100_Q6L3Q3 Hypothetical protein [Solanum demissum]
Length = 838
Score = 91.7 bits (226), Expect = 2e-16
Identities = 69/181 (38%), Positives = 81/181 (44%), Gaps = 59/181 (32%)
Query: 389 GVVSSASVRPSFYRTFYNCGDFRHISRYCPKPRILDLMQK*TRAVVPASNDKNSRGHPRG 448
G S RPSF RT YNC + RHI R CP R++D Q+ TRAVV A N N RG P+G
Sbjct: 223 GATPSTGNRPSFDRTCYNCREPRHIRRDCPHLRMMDFAQQQTRAVVLAGNCNNGRGRPQG 282
Query: 449 W*GGNHRGCEGRENGNVGRGALQPRKKVVCQDGRA*FYARLIWRDLMKESQVLFFSLPDG 508
GGN RG GR NGN G KV+ Y L+ RD M + VLF+
Sbjct: 283 VRGGNQRGRGGRGNGNAG--------KVIT-------YTILV-RDRM--ANVLFYP---- 320
Query: 509 **IFF*SYIFS*VTV*FALGFDEVCNIFYTPIHVSTLVGGSVIVTHVYQLYLVIFMGFQS 568
VG SVIVTHVY ++FMGFQ+
Sbjct: 321 -------------------------------------VGESVIVTHVYCACPILFMGFQT 343
Query: 569 W 569
W
Sbjct: 344 W 344
Score = 60.8 bits (146), Expect = 3e-07
Identities = 57/212 (26%), Positives = 90/212 (41%), Gaps = 68/212 (32%)
Query: 1 MVHTRINDIKLVAHINELVESSTIRGRSLFKGRERGRVEQARGGAPFEKSPREKAPYAPH 60
MV+TR N ++ VA +N E S + AP E +PR +AP A H
Sbjct: 1 MVNTRFNGVRPVAPVNTPAEESVAK-------------------APIENAPRNEAPSAHH 41
Query: 61 VDV*EIVEV*NEEEVELEEDAHVETICIPPLNLVLSQQIMFF*RDWTVLESFPQCKQLKL 120
++ E+ VE+EE+ E+ Q +++
Sbjct: 42 EEI--------EDNVEVEEE-----------------------------ENVGQEEEVHA 64
Query: 121 KSIILLL*RFLRLVE**ELMYFSSPNRPVMTGNEHKMLTKFLKLMSYMFFGSESEDTFKF 180
++I+LL + +P P + K+ + + + ++ ED ++F
Sbjct: 65 ETILLL-----------SVQATQAPANPPIAITVPKV-DETIGIDAFFRPYWVLEDAYEF 112
Query: 181 ILHYYERLNKLVIVKQYGVEIVTLQFRGEAKQ 212
ILH YERL+KL IV Q+GVE V Q +GEAKQ
Sbjct: 113 ILHCYERLHKLRIVHQHGVEFVAFQLQGEAKQ 144
Score = 42.7 bits (99), Expect = 0.087
Identities = 19/26 (73%), Positives = 24/26 (92%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMR 1289
RLTK++HFILVK YNAQQL+KVY++
Sbjct: 645 RLTKSTHFILVKIDYNAQQLAKVYVK 670
>UniRef100_Q60D63 Putative polyprotein [Oryza sativa]
Length = 1765
Score = 90.9 bits (224), Expect = 3e-16
Identities = 71/210 (33%), Positives = 106/210 (49%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A +SK+ L G +
Sbjct: 887 DKTKRMCVDYRALNEVTIKNKYPLPRIDDLFDQLKGAKV---FSKIDLRS------GYHQ 937
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + F+ ++V +DDIL
Sbjct: 938 LRIREEDIPKTAFTTRYGLYECTVMSFGLTNAPAFFMNLMNKVFMEFLDKFVVVFIDDIL 997
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ ES+ + HL +V + +LYA F F ++ +G+V+ + + VD N+E
Sbjct: 998 IYSESEEEHEQHLRLVLEKLKEHQLYAKFSKCDFWLTEVKFLGHVITAQGVAVDPSNVES 1057
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V W P V + R F+GLA YYR F + F
Sbjct: 1058 VTKWTPPKTVSQIRSFLGLAGYYRRFIENF 1087
Score = 47.8 bits (112), Expect = 0.003
Identities = 38/118 (32%), Positives = 55/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI V Y ++L+++Y+ R P + W+ E +G
Sbjct: 1431 RLTKVAHFIPVHTTYTGKRLAELYLSRIMCLHGVPKKIVSDRGSQFTSKFWQKLQEELGT 1490
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1491 RLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGAWDKSLPYAEFSYNNSYQAS 1546
>UniRef100_Q9XE85 Polyprotein [Sorghum bicolor]
Length = 1484
Score = 90.9 bits (224), Expect = 3e-16
Identities = 73/210 (34%), Positives = 107/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S MC DYR LN VT +NKY LP IDDLF+QL A +SK+ L G +
Sbjct: 607 DGSQRMCVDYRSLNEVTIKNKYPLPRIDDLFDQLRGACV---FSKIDLRS------GYHQ 657
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F + +++ + ++ ++V +DDIL
Sbjct: 658 LKIRNSDIPKTAFTTRYGLYEYTVMSFGLTNAPAYFMYMMNKVFMEYLDKFVVVFIDDIL 717
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
+F ++K + +HL +V KLYA F ++ +G+VV N I VD ++
Sbjct: 718 VFSKTKEEHAEHLRLVLQKLREHKLYAKRSKCEFWLEEVSFLGHVVSNGGIAVDPSKVKD 777
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V+NW P+ V E R F+GLA YYR F + F
Sbjct: 778 VLNWKPPTDVSEIRSFLGLAGYYRRFIEGF 807
>UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa]
Length = 1827
Score = 90.9 bits (224), Expect = 3e-16
Identities = 71/210 (33%), Positives = 107/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A +SK+ L G +
Sbjct: 937 DKTKRMCVDYRALNEVTIKNKYPLPRIDDLFDQLKGAKV---FSKIDLRS------GYHQ 987
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + F+ ++V +DDIL
Sbjct: 988 LRIREEDIPKTAFTTRYGLYECTVMSFGLTNAPAFFMNLMNKVFIEFLDKFVVVFIDDIL 1047
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +S+ + HL +V + +LYA F F ++ +G+V+ + + VD N+E
Sbjct: 1048 IYSKSEEEHEQHLRLVLEKLKEHQLYAKFSKCDFWLTEVKFLGHVITAQGVAVDPSNVES 1107
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V W P V + R F+GLASYYR F + F
Sbjct: 1108 VTKWTPPKTVSQIRSFLGLASYYRRFIENF 1137
Score = 47.8 bits (112), Expect = 0.003
Identities = 38/118 (32%), Positives = 55/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI V Y ++L+++Y+ R P + W+ E +G
Sbjct: 1493 RLTKVAHFIPVHTTYTGKRLAELYLARIMCLHGVPKKIVSDRGSQFTSKFWQKLQEEMGT 1552
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1553 RLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGAWDKSLPYAEFSYNNSYQAS 1608
>UniRef100_Q6L3S3 Putative gag-pol polyprotein [Solanum demissum]
Length = 1515
Score = 90.9 bits (224), Expect = 3e-16
Identities = 72/220 (32%), Positives = 106/220 (47%), Gaps = 40/220 (18%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S+ MC DYRQLN+VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 705 DGSLRMCIDYRQLNKVTIKNKYPLPRIDDLFDQLQGATC---FSKIDLRS------GYHQ 755
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGF 839
+++ P A + +G +EF + +T L
Sbjct: 756 LRVRERDIPKTAF----------RTRYGHYEFLVMSFGLTNAPAAFMDLMNRVFRPYLDM 805
Query: 840 ILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNER 898
+I+ +DDIL++ ++ D HL V ++LYA F F + +G++V +
Sbjct: 806 FVIIFIDDILIYSRNEEDHASHLRTVLQTLKDKELYAKFSKCEFWLKSVAFLGHIVSGDG 865
Query: 899 IMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
I VD + IE V NW P+ E R F+GLA YYR F + F
Sbjct: 866 IKVDTRKIEAVQNWPRPTSPTEIRSFLGLAGYYRRFVEGF 905
Score = 42.4 bits (98), Expect = 0.11
Identities = 32/118 (27%), Positives = 54/118 (45%), Gaps = 10/118 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWEN-----A* 1318
R+TK++HF+ V+ ++A+ +K+Y++ PI + W++
Sbjct: 1267 RMTKSAHFLPVRTTHSAEDYAKLYIQEIVRLHGVPISIISDRGAQFTAQFWKSFQKGLGS 1326
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
+V T F TI+ +R C IDF+ +WD L L EF+YNN+Y
Sbjct: 1327 KVSLSTAFHPQTDGQAERTIQTLEDMLRA-----CVIDFKSNWDDHLPLIEFAYNNSY 1379
>UniRef100_Q7XWU5 OSJNBa0065B15.6 protein [Oryza sativa]
Length = 1467
Score = 90.5 bits (223), Expect = 4e-16
Identities = 71/210 (33%), Positives = 106/210 (49%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 745 DHTQRMCVDYRALNDVTIKNKYPLPRIDDLFDQLKGATV---FSKIDLRS------GYHQ 795
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 796 LRIKEEDIPKTAFTTRYGLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDIL 855
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +K +HLC+ + +LYA F F ++ +G+V+ + VD N+E
Sbjct: 856 IYSRTKEVHEEHLCLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISAGGVAVDPSNVES 915
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V NW P V E R F+GLA YYR F + F
Sbjct: 916 VTNWKQPKTVSEIRNFLGLAGYYRRFIENF 945
Score = 45.1 bits (105), Expect = 0.017
Identities = 34/116 (29%), Positives = 46/116 (39%), Gaps = 43/116 (37%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*EVGHV 1323
RLTK +HFI VK Y+ +L+++YM R
Sbjct: 1253 RLTKVAHFIPVKTTYSGSRLAELYMAR--------------------------------- 1279
Query: 1324 THF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1280 --------ITDGQTERVNQ--ILEDMLRACALDFGGSWDKNLPYAEFSYNNSYQAS 1325
>UniRef100_Q7XLT1 OSJNBa0057M08.3 protein [Oryza sativa]
Length = 1664
Score = 90.1 bits (222), Expect = 5e-16
Identities = 70/210 (33%), Positives = 106/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 1061 DHTQRMCVDYRALNDVTIKNKYPLPRIDDLFDQLKGATV---FSKIDLRS------GYHQ 1111
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 1112 LRIKEEDIPKTAFTTRYRLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDIL 1171
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +K + +HLC+ + +LYA F F ++ +G+V+ + VD N+E
Sbjct: 1172 IYSRTKEEHEEHLCLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISAGGVAVDPSNVES 1231
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V NW P V E F+GLA YYR F + F
Sbjct: 1232 VTNWKQPKTVSEIHSFLGLAGYYRRFIENF 1261
Score = 38.9 bits (89), Expect = 1.3
Identities = 16/27 (59%), Positives = 18/27 (66%)
Query: 1353 CDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
C +DF G WDK L EFSYNN+Y S
Sbjct: 1573 CALDFGGSWDKNLPYAEFSYNNSYQAS 1599
>UniRef100_Q5WMS7 Putative polyprotein [Oryza sativa]
Length = 1131
Score = 89.7 bits (221), Expect = 6e-16
Identities = 71/213 (33%), Positives = 109/213 (50%), Gaps = 26/213 (12%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DY LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 564 DHTQRMCVDYHALNEVTIKNKYPLPRIDDLFDQLEGATV---FSKIDLRS------GYHQ 614
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 615 LRIREEDIPKTAFTTRYGLFECTVMSFGLTNSPAFFMNLMNKVFMEYLDKFVVVFIDDIL 674
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP----IGNVVMNERIMVDLQN 905
++ ++K + +HL + + +LYA F S C LP +G+V+ + VD N
Sbjct: 675 IYSKTKEEHEEHLRLALEKLREHQLYAKF---SKCEFWLPEVKFLGHVISAGGVAVDPSN 731
Query: 906 IEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
+E V+NW P ++E R F+GLA YYR F + F
Sbjct: 732 VESVLNWKQPKTILEIRSFLGLAGYYRRFIENF 764
>UniRef100_Q8W5E8 Putative polyprotein [Oryza sativa]
Length = 1416
Score = 89.7 bits (221), Expect = 6e-16
Identities = 81/253 (32%), Positives = 123/253 (48%), Gaps = 30/253 (11%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 550 DHTQRMCVDYRALNDVTIKNKYLLPRIDDLFDQLKGATV---FSKIDLRS------GYHQ 600
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 601 LRIKEEDIPKTAFTTRYGLFKCTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDIL 660
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +K + +HL + + +LYA F F ++ +G+V+ + VD N+E
Sbjct: 661 IYSRTKEEHEEHLRLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISAGGVAVDPSNVES 720
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAFDHNTYLGTTSVT*SLII--------NYY-- 958
V NW P V E R F+GLA YY+ E+ L ++ ++I Y
Sbjct: 721 VTNWKQPKTVSEIRSFLGLAGYYKWSEECEQSFQELKNRLISAPILILPDPKKGFQVYCD 780
Query: 959 ASQLILGVVLMQD 971
AS+L LG VLMQD
Sbjct: 781 ASKLGLGCVLMQD 793
Score = 51.2 bits (121), Expect = 2e-04
Identities = 40/118 (33%), Positives = 56/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI VK Y+ +L+++YM R P + W+ E +G
Sbjct: 1082 RLTKVAHFIPVKTTYSGSRLAELYMARIVCLHGVPKKIVSDRGSQFTSNFWKKLQEEMGS 1141
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1142 KLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGSWDKNLPYAEFSYNNSYQAS 1197
>UniRef100_Q6L3Y7 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 89.4 bits (220), Expect = 8e-16
Identities = 80/258 (31%), Positives = 118/258 (45%), Gaps = 50/258 (19%)
Query: 702 WLQQC*QSLKLKFKSFLIKVISILVILV*RPCNVC*E*DSSMIMCKDYRQLNRVTTRNKY 761
WL C SL + ++ + ++ VI E SS + KDYRQLN+VT +NKY
Sbjct: 459 WLHACYASLDCRTRAVKFQFLNEPVI----------EWSSSSTVPKDYRQLNKVTIKNKY 508
Query: 762 TLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTKKGIQD---PLWAL*VSSDIVWV 818
LP IDDLF+QL A+ +SK+ L G + +++ P A
Sbjct: 509 PLPRIDDLFDQLQGATC---FSKIDLRS------GYHQLRVRECDIPKTAF--------- 550
Query: 819 DKCSFGCHEFD*LCDQMT-----------------LGFILIVCVDDILLFLESK*DKVDH 861
+ +G +EF + +T L +IV +DDIL++ ++ D H
Sbjct: 551 -RTRYGHYEFLVMSFGLTNAPAAFMDLMNRVFKPYLDMFVIVFIDDILIYSRNEGDHASH 609
Query: 862 LCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEPVMNWV*PSFVIE 920
L V ++LYA F F + +G++V + I VD IE V NW P+ E
Sbjct: 610 LRTVLQTLKDKELYAKFSKCEFWLKSVAFLGHIVSGDGIKVDTWKIEAVQNWPRPTSPTE 669
Query: 921 SRGFMGLASYYRGFEDAF 938
R F+GLA YYR F + F
Sbjct: 670 IRSFLGLAGYYRRFVEGF 687
Score = 41.2 bits (95), Expect = 0.25
Identities = 31/113 (27%), Positives = 50/113 (43%), Gaps = 27/113 (23%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*EVGHV 1323
R+TK++HF+ VK + +K+Y++ G LS+
Sbjct: 1049 RMTKSAHFLPVKTTNTTEDYAKLYVQEIKGLGSKVNLST--------------------A 1088
Query: 1324 THF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
H P ++G E+ + + C IDF+G+WD L L EF+YNN+Y
Sbjct: 1089 FH-----PQTDGQA--EHTIQILEDMLRACVIDFKGNWDDHLPLIEFAYNNSY 1134
>UniRef100_Q6L3Y6 Putative gag-pol polyprotein [Solanum demissum]
Length = 1351
Score = 89.4 bits (220), Expect = 8e-16
Identities = 80/258 (31%), Positives = 118/258 (45%), Gaps = 50/258 (19%)
Query: 702 WLQQC*QSLKLKFKSFLIKVISILVILV*RPCNVC*E*DSSMIMCKDYRQLNRVTTRNKY 761
WL C SL + ++ + ++ VI E SS + KDYRQLN+VT +NKY
Sbjct: 459 WLHACYASLDCRTRAVKFQFLNEPVI----------EWSSSSTVPKDYRQLNKVTIKNKY 508
Query: 762 TLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTKKGIQD---PLWAL*VSSDIVWV 818
LP IDDLF+QL A+ +SK+ L G + +++ P A
Sbjct: 509 PLPRIDDLFDQLQGATC---FSKIDLRS------GYHQLRVRECDIPKTAF--------- 550
Query: 819 DKCSFGCHEFD*LCDQMT-----------------LGFILIVCVDDILLFLESK*DKVDH 861
+ +G +EF + +T L +IV +DDIL++ ++ D H
Sbjct: 551 -RTRYGHYEFLVMSFGLTNAPAAFMDLMNRVFKPYLDMFVIVFIDDILIYSRNEGDHASH 609
Query: 862 LCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEPVMNWV*PSFVIE 920
L V ++LYA F F + +G++V + I VD IE V NW P+ E
Sbjct: 610 LRTVLQTLKDKELYAKFSKCEFWLKSVAFLGHIVSGDGIKVDTWKIEAVQNWPRPTSPTE 669
Query: 921 SRGFMGLASYYRGFEDAF 938
R F+GLA YYR F + F
Sbjct: 670 IRSFLGLAGYYRRFVEGF 687
Score = 41.2 bits (95), Expect = 0.25
Identities = 31/113 (27%), Positives = 50/113 (43%), Gaps = 27/113 (23%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*EVGHV 1323
R+TK++HF+ VK + +K+Y++ G LS+
Sbjct: 1049 RMTKSAHFLPVKTTNTTEDYAKLYVQEIKGLGSKVNLST--------------------A 1088
Query: 1324 THF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
H P ++G E+ + + C IDF+G+WD L L EF+YNN+Y
Sbjct: 1089 FH-----PQTDGQA--EHTIQILEDMLRACVIDFKGNWDDHLPLIEFAYNNSY 1134
>UniRef100_Q65WS7 Putative polyprotein [Oryza sativa]
Length = 1717
Score = 89.4 bits (220), Expect = 8e-16
Identities = 69/210 (32%), Positives = 109/210 (51%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 851 DHTQRMCVDYRALNEVTIKNKYPLPRIDDLFDQLEGATV---FSKIDLRS------GYHQ 901
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 902 LRIREEDIPKTAFTTRYGLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKLVVVFIDDIL 961
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ ++K + +HL + + +LYA F F ++ +G+V+ + + VD N+E
Sbjct: 962 IYSKTKEEHEEHLRLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISSGGVAVDPSNVES 1021
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V++W P V E R F+GLA YYR F + F
Sbjct: 1022 VLSWKQPKTVSEIRSFLGLAGYYRRFIENF 1051
Score = 50.8 bits (120), Expect = 3e-04
Identities = 40/117 (34%), Positives = 54/117 (45%), Gaps = 26/117 (22%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*EVGHV 1323
RLTK +HFI VK Y +L+++YM R V + V E+G
Sbjct: 1407 RLTKVAHFIPVKTTYTGNKLAELYMAR-----------------VKLQV------EMGTR 1443
Query: 1324 THF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1444 LNFSTAYHPQTDGQTERVNQ--ILEDMLRACVLDFGGSWDKNLPYAEFSYNNSYQAS 1498
>UniRef100_Q6L3S2 Putative retrotransposon protein [Solanum demissum]
Length = 1602
Score = 89.4 bits (220), Expect = 8e-16
Identities = 71/220 (32%), Positives = 106/220 (47%), Gaps = 40/220 (18%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D S+ +C DYRQLN+VT +NKY LP IDDLF+QL A+ +SK+ L G +
Sbjct: 711 DGSLRICIDYRQLNKVTIKNKYPLPRIDDLFDQLQGATC---FSKIDLRS------GYHQ 761
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHEFD*LCDQMT-----------------LGF 839
+++ P A + +G +EF + +T L
Sbjct: 762 LRVRERDIPKTAF----------RTRYGHYEFLVMSFGLTNAPAAFMDLMNRVFRPYLDM 811
Query: 840 ILIVCVDDILLFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNER 898
+I+ +DDIL++ ++ D HL V ++LYA F F + +G++V +
Sbjct: 812 FVIIFIDDILIYSRNEEDHASHLRTVLQTLKDKELYAKFSKCEFWLKSVAFLGHIVSGDG 871
Query: 899 IMVDLQNIEPVMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
I VD + IE V NW P+ E R F+GLA YYR F + F
Sbjct: 872 IKVDTRKIEAVQNWPRPTSPTEIRSFLGLAGYYRRFVEGF 911
Score = 43.5 bits (101), Expect = 0.051
Identities = 33/118 (27%), Positives = 54/118 (44%), Gaps = 10/118 (8%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWEN-----A* 1318
R+TK++HF+ VK ++A+ +K+Y++ PI + W++
Sbjct: 1273 RMTKSAHFLPVKTTHSAEDYAKLYIQEIVRLHGVPISIISDRGAQFTAQFWKSFQKGLGS 1332
Query: 1319 EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNY 1376
+V T F TI+ +R C IDF+ +WD L L EF+YNN+Y
Sbjct: 1333 KVSLSTAFHPQTDGQAERTIQTLEDMLRA-----CVIDFKSNWDDHLPLIEFAYNNSY 1385
>UniRef100_Q7Y1P5 Putative polyprotein [Oryza sativa]
Length = 1436
Score = 89.0 bits (219), Expect = 1e-15
Identities = 70/210 (33%), Positives = 106/210 (50%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN VT +NKY LP IDDLF+QL A+ +SK+ + G +
Sbjct: 584 DHTQRMCVDYRALNDVTIKNKYPLPRIDDLFDQLKGATV---FSKI------DPRSGYHQ 634
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ +V +DDIL
Sbjct: 635 LRIKEEDIPKTAFTTRYGLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKFEVVFIDDIL 694
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +K + +HLC+ + +LYA F F ++ +G+V+ + VD N+E
Sbjct: 695 IYSRTKEEHEEHLCLALEKLREHQLYAKFSKCEFWLSEVKFLGHVISAGGVAVDPSNVES 754
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V NW P V E R F+GLA YYR F + F
Sbjct: 755 VTNWKQPKTVSEIRSFLGLAGYYRRFIENF 784
Score = 48.9 bits (115), Expect = 0.001
Identities = 39/118 (33%), Positives = 55/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI VK Y+ +L+++YM R P + W+ E +G
Sbjct: 1102 RLTKVAHFIPVKTTYSGSRLAELYMARIVCLHGVPKKIVSDRGSQFTSKFWKKLQEEMGS 1161
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L FSYNN+Y S
Sbjct: 1162 KLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGSWDKNLPYAGFSYNNSYQAS 1217
>UniRef100_Q7XWH7 OSJNBa0085C10.17 protein [Oryza sativa]
Length = 1437
Score = 89.0 bits (219), Expect = 1e-15
Identities = 69/210 (32%), Positives = 106/210 (49%), Gaps = 20/210 (9%)
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTK 799
D + MC DYR LN +T +N Y LP IDDLF+QL A+ +SK+ L G +
Sbjct: 623 DHTQRMCVDYRALNDLTIKNNYPLPRIDDLFDQLKGATV---FSKIDLRS------GYHQ 673
Query: 800 KGIQD---PLWAL*VSSDIVWVDKCSFGCHE----FD*LCDQMTLGFI---LIVCVDDIL 849
I++ P A + SFG F L +++ + ++ ++V +DDIL
Sbjct: 674 LRIKEEDIPKTAFTTRYGLFECTVMSFGLTNAPAFFMNLMNKVFMEYLDKFVVVFIDDIL 733
Query: 850 LFLESK*DKVDHLCIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEP 908
++ +K + +HL + + +LYA F FC ++ +G+V+ + VD N+E
Sbjct: 734 IYSRTKEEHEEHLRLALEKLREHQLYAKFSKCEFCLSEVKFLGHVISAGGVAVDPSNVES 793
Query: 909 VMNWV*PSFVIESRGFMGLASYYRGFEDAF 938
V NW P V E R F+GLA YYR F + F
Sbjct: 794 VTNWKQPKTVSEIRSFLGLAGYYRRFIENF 823
Score = 51.2 bits (121), Expect = 2e-04
Identities = 40/118 (33%), Positives = 56/118 (46%), Gaps = 4/118 (3%)
Query: 1264 RLTKTSHFILVKKHYNAQQLSKVYMRR**GCTECPILSS*TVVPVYIYVLWENA*E-VGH 1322
RLTK +HFI VK Y+ +L+++YM R P + W+ E +G
Sbjct: 1179 RLTKVAHFIPVKTTYSGSRLAELYMARIVCLHGVPKKIVSDRGSQFTSNFWKKLQEEMGS 1238
Query: 1323 VTHF*NNL-PSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSYNNNYTLS 1379
+F P ++G T R N + + C +DF G WDK L EFSYNN+Y S
Sbjct: 1239 KLNFSTAYHPQTDGQTERVNQ--ILEDMLRACALDFGGSWDKNLPYAEFSYNNSYQAS 1294
>UniRef100_Q688F1 Hypothetical protein OSJNBa0035I01.9 [Oryza sativa]
Length = 666
Score = 68.9 bits (167), Expect = 1e-09
Identities = 58/186 (31%), Positives = 94/186 (50%), Gaps = 25/186 (13%)
Query: 746 CKDYRQLNRVTTRNKYTLPLIDDLFNQLWRASFL*DYSKVCLPLVKN*A*GCTKKGIQDP 805
C +Y LN VT +NKY LP IDDLF+QL A +SK+ L GC + I++
Sbjct: 18 CVNYCALNDVTIKNKYPLPRIDDLFDQLKGAIV---FSKIDLRS------GCHQLRIKE- 67
Query: 806 LWAL*VSSDIVWVDKCSFGCHEFD*LCDQMTLGFI---LIVCVDDILLFLESK*DKVDHL 862
DI +F F L +++ L ++ ++V +DDIL++ ++K + +HL
Sbjct: 68 -------EDIPKTAFTAF----FMNLMNKVFLEYLDKFVVVFIDDILIYSQTKEEHEEHL 116
Query: 863 CIV*DVHGRQKLYA*FLSVSFC*LQLP-IGNVVMNERIMVDLQNIEPVMNWV*PSFVIES 921
+ + +LYA F F ++ +G+++ ++VD N+E V NW P V E
Sbjct: 117 RLALEKLREHQLYAKFSKCEFWLSEVKFLGHIISVGGVVVDPSNVESVTNWKQPKAVSEI 176
Query: 922 RGFMGL 927
R +GL
Sbjct: 177 RSILGL 182
Score = 39.7 bits (91), Expect = 0.73
Identities = 26/67 (38%), Positives = 34/67 (49%), Gaps = 6/67 (8%)
Query: 1313 LWENA*EVGHVTHF*NNLPSSNGWTIRENNPSVRGHIEGMCDIDFRGHWDKFLTLCEFSY 1372
+W + V HF +P+ NG T R N + + C +DF G WDK L EFSY
Sbjct: 334 IWVIIDRLTKVAHF---IPT-NGQTERVNQ--ILEDMLRACALDFGGSWDKNLPNAEFSY 387
Query: 1373 NNNYTLS 1379
NN+Y S
Sbjct: 388 NNSYQAS 394
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,200,191,904
Number of Sequences: 2790947
Number of extensions: 81523914
Number of successful extensions: 306150
Number of sequences better than 10.0: 782
Number of HSP's better than 10.0 without gapping: 686
Number of HSP's successfully gapped in prelim test: 97
Number of HSP's that attempted gapping in prelim test: 303079
Number of HSP's gapped (non-prelim): 2741
length of query: 1546
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1405
effective length of database: 454,526,306
effective search space: 638609459930
effective search space used: 638609459930
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 82 (36.2 bits)
Medicago: description of AC137826.3


