

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
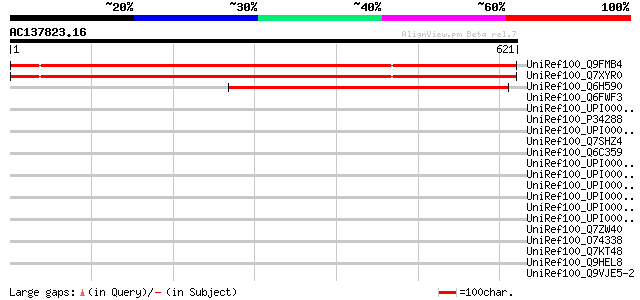
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FMB4 Arabidopsis thaliana genomic DNA, chromosome 5,... 874 0.0
UniRef100_Q7XYR0 At5g38880 [Arabidopsis thaliana] 874 0.0
UniRef100_Q6H590 Hypothetical protein OSJNBa0048A13.17 [Oryza sa... 454 e-126
UniRef100_Q6FWF3 Similar to tr|Q07457 Saccharomyces cerevisiae Y... 45 0.006
UniRef100_UPI000021B39A UPI000021B39A UniRef100 entry 45 0.008
UniRef100_P34288 GTPase-activating protein GAP [Caenorhabditis e... 44 0.014
UniRef100_UPI000042FE9D UPI000042FE9D UniRef100 entry 43 0.024
UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa] 42 0.041
UniRef100_Q6C359 Similar to DEHA0C09658g Debaryomyces hansenii I... 42 0.054
UniRef100_UPI00003C22C3 UPI00003C22C3 UniRef100 entry 41 0.091
UniRef100_UPI0000433454 UPI0000433454 UniRef100 entry 40 0.16
UniRef100_UPI0000433453 UPI0000433453 UniRef100 entry 40 0.16
UniRef100_UPI000042F508 UPI000042F508 UniRef100 entry 40 0.16
UniRef100_UPI000027C715 Transcription factor 12 [Homo sapiens] 40 0.16
UniRef100_UPI000004DF67 UPI000004DF67 UniRef100 entry 40 0.16
UniRef100_Q7ZW40 Similar to vesicle docking protein p115 [Brachy... 40 0.16
UniRef100_O74338 SPBC1A4.05 protein [Schizosaccharomyces pombe] 40 0.16
UniRef100_Q7KT48 CG5020-PD, isoform D [Drosophila melanogaster] 40 0.20
UniRef100_Q9HEL8 Hypothetical protein 12F11.080 [Neurospora crassa] 40 0.20
UniRef100_Q9VJE5-2 Splice isoform B of Q9VJE5 [Drosophila melano... 40 0.20
>UniRef100_Q9FMB4 Arabidopsis thaliana genomic DNA, chromosome 5, TAC clone:K15E6
[Arabidopsis thaliana]
Length = 796
Score = 874 bits (2258), Expect = 0.0
Identities = 441/620 (71%), Positives = 524/620 (84%), Gaps = 3/620 (0%)
Query: 1 MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST 60
+LEAYD+QCDEA+RIFAEYHKRL Y+NQA DAQRS V+SS E+++S SA +E+EAVYST
Sbjct: 178 LLEAYDQQCDEATRIFAEYHKRLQVYVNQANDAQRS-VNSSNEVLSSLSANSEREAVYST 236
Query: 61 VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG 120
VKG+KS+DDVI++ETTRE+NIR C+ L + M+++I++SFPAYEG+G+ S P+ E AKLG
Sbjct: 237 VKGTKSADDVILMETTRERNIRIVCDLLASRMIERIRNSFPAYEGNGICSLPELETAKLG 296
Query: 121 FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY 180
F++DG+I DE++TVIVN L+ PPLLLQAI AYT +K+ ISRE+EKIDVRADAE+LRYK+
Sbjct: 297 FEYDGEITDEMKTVIVNSLRGPPLLLQAIAAYTLRIKTLISREMEKIDVRADAEMLRYKF 356
Query: 181 ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN 240
ENN V D SSSD SSPL Y GNGK+G D GS NQLLERQKAHVQQFLATEDALN
Sbjct: 357 ENNRVTDNSSSDVSSPLSYQFNGNGKIGTDTHFQGSNNQLLERQKAHVQQFLATEDALNK 416
Query: 241 AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN 300
AAEARDLC K + RLHG D + S +G T+Q+ +LRQ +LDVW KERE +GL+ASLN
Sbjct: 417 AAEARDLCHKFINRLHGSADTATHSF-VGGTTQSGSNLRQFELDVWGKEREAAGLRASLN 475
Query: 301 TLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQ 360
TL+SEIQRLNKLCAERKEAEDSLKKKWKKIEEFD RRSELE+IYT LLKAN DA +FW+Q
Sbjct: 476 TLLSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDARRSELETIYTTLLKANMDAVAFWNQ 535
Query: 361 QPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAI 420
QP AREYA +T+IPA VV+ SNSAKD IEKEVSAF++SPDNSLYMLP++PQ LLE++
Sbjct: 536 QPLAAREYASATVIPASEVVVDISNSAKDFIEKEVSAFFQSPDNSLYMLPATPQGLLESM 595
Query: 421 GSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASI 480
G++GS+G EAVA AE +AA+LTARAGARDPSAIPSICR+SAALQY A GLEGSDA LAS+
Sbjct: 596 GANGSTGPEAVAYAEKNAALLTARAGARDPSAIPSICRISAALQYPA-GLEGSDASLASV 654
Query: 481 LESLEFCLKRRGSEASVLEDLLKAINLVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSL 540
LESLEFCL+ RGSEA VLEDL KAI+LVHIR+DLV+SGH+LL+HA+ QQ YERTTN+ L
Sbjct: 655 LESLEFCLRVRGSEACVLEDLAKAIDLVHIRQDLVESGHSLLDHAFRAQQKYERTTNYCL 714
Query: 541 NLAAEQERAVMEKWLPELKTGVLNAQQSLEACKYVWGLLDEWWEQPASTVVDWATVDGSN 600
+LA+EQE + ++WLPEL+T V NAQ S E CKYV GLLDEWWEQPASTVVDW TVDG +
Sbjct: 715 DLASEQENTISDQWLPELRTAVQNAQASSEHCKYVRGLLDEWWEQPASTVVDWVTVDGQS 774
Query: 601 VAFWHNHVKKLLTCYDQELL 620
VA W NHVK+LL YD+E L
Sbjct: 775 VAAWQNHVKQLLAFYDKESL 794
>UniRef100_Q7XYR0 At5g38880 [Arabidopsis thaliana]
Length = 775
Score = 874 bits (2258), Expect = 0.0
Identities = 441/620 (71%), Positives = 524/620 (84%), Gaps = 3/620 (0%)
Query: 1 MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST 60
+LEAYD+QCDEA+RIFAEYHKRL Y+NQA DAQRS V+SS E+++S SA +E+EAVYST
Sbjct: 157 LLEAYDQQCDEATRIFAEYHKRLQVYVNQANDAQRS-VNSSNEVLSSLSANSEREAVYST 215
Query: 61 VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG 120
VKG+KS+DDVI++ETTRE+NIR C+ L + M+++I++SFPAYEG+G+ S P+ E AKLG
Sbjct: 216 VKGTKSADDVILMETTRERNIRIVCDLLASRMIERIRNSFPAYEGNGICSLPELETAKLG 275
Query: 121 FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY 180
F++DG+I DE++TVIVN L+ PPLLLQAI AYT +K+ ISRE+EKIDVRADAE+LRYK+
Sbjct: 276 FEYDGEITDEMKTVIVNSLRGPPLLLQAIAAYTLRIKTLISREMEKIDVRADAEMLRYKF 335
Query: 181 ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN 240
ENN V D SSSD SSPL Y GNGK+G D GS NQLLERQKAHVQQFLATEDALN
Sbjct: 336 ENNRVTDNSSSDVSSPLSYQFNGNGKIGTDTHFQGSNNQLLERQKAHVQQFLATEDALNK 395
Query: 241 AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN 300
AAEARDLC K + RLHG D + S +G T+Q+ +LRQ +LDVW KERE +GL+ASLN
Sbjct: 396 AAEARDLCHKFINRLHGSADTATHSF-VGGTTQSGSNLRQFELDVWGKEREAAGLRASLN 454
Query: 301 TLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQ 360
TL+SEIQRLNKLCAERKEAEDSLKKKWKKIEEFD RRSELE+IYT LLKAN DA +FW+Q
Sbjct: 455 TLLSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDARRSELETIYTTLLKANMDAVAFWNQ 514
Query: 361 QPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAI 420
QP AREYA +T+IPA VV+ SNSAKD IEKEVSAF++SPDNSLYMLP++PQ LLE++
Sbjct: 515 QPLAAREYASATVIPASEVVVDISNSAKDFIEKEVSAFFQSPDNSLYMLPATPQGLLESM 574
Query: 421 GSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASI 480
G++GS+G EAVA AE +AA+LTARAGARDPSAIPSICR+SAALQY A GLEGSDA LAS+
Sbjct: 575 GANGSTGPEAVAYAEKNAALLTARAGARDPSAIPSICRISAALQYPA-GLEGSDASLASV 633
Query: 481 LESLEFCLKRRGSEASVLEDLLKAINLVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSL 540
LESLEFCL+ RGSEA VLEDL KAI+LVHIR+DLV+SGH+LL+HA+ QQ YERTTN+ L
Sbjct: 634 LESLEFCLRVRGSEACVLEDLAKAIDLVHIRQDLVESGHSLLDHAFRAQQKYERTTNYCL 693
Query: 541 NLAAEQERAVMEKWLPELKTGVLNAQQSLEACKYVWGLLDEWWEQPASTVVDWATVDGSN 600
+LA+EQE + ++WLPEL+T V NAQ S E CKYV GLLDEWWEQPASTVVDW TVDG +
Sbjct: 694 DLASEQENTISDQWLPELRTAVQNAQASSEHCKYVRGLLDEWWEQPASTVVDWVTVDGQS 753
Query: 601 VAFWHNHVKKLLTCYDQELL 620
VA W NHVK+LL YD+E L
Sbjct: 754 VAAWQNHVKQLLAFYDKESL 773
>UniRef100_Q6H590 Hypothetical protein OSJNBa0048A13.17 [Oryza sativa]
Length = 365
Score = 454 bits (1169), Expect = e-126
Identities = 231/345 (66%), Positives = 280/345 (80%), Gaps = 2/345 (0%)
Query: 269 GATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWK 328
G TSQN+ + R L+LDVWAKEREV+GLKASLNTL SE+QRL KLCAE KEAEDSLKKKWK
Sbjct: 4 GNTSQNMTNSRHLELDVWAKEREVAGLKASLNTLTSEVQRLYKLCAEWKEAEDSLKKKWK 63
Query: 329 KIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAK 388
KIEEFD RRSELE IY ALL+AN +A++FW QQP +AR YA TIIPAC+AVV+ S +++
Sbjct: 64 KIEEFDARRSELECIYNALLRANMEASTFWEQQPLSARGYASRTIIPACNAVVDMSTNSR 123
Query: 389 DLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGAR 448
DLIE+E+SAF +S DNSL LP++PQALLEA+GS+G++G EAVA AE AA+LTARAGAR
Sbjct: 124 DLIERELSAFGQSLDNSLCRLPATPQALLEALGSNGATGSEAVAAAEKHAALLTARAGAR 183
Query: 449 DPSAIPSICRVSAALQY--AAGGLEGSDAGLASILESLEFCLKRRGSEASVLEDLLKAIN 506
DPSA+PSICR+S ALQY + G EG+D+GLAS+L SLEFCLK GSEAS+LEDL KAIN
Sbjct: 184 DPSAVPSICRISTALQYNSVSPGTEGTDSGLASVLNSLEFCLKPCGSEASILEDLSKAIN 243
Query: 507 LVHIRRDLVQSGHALLNHAYFVQQDYERTTNFSLNLAAEQERAVMEKWLPELKTGVLNAQ 566
LVH RR+LV++ LLN A+ QQ+YER N+ L LA EQE+ V E+WLPEL+ V AQ
Sbjct: 244 LVHTRRNLVENDRVLLNRAHRAQQEYERVANYCLKLAGEQEKMVSERWLPELRNAVQEAQ 303
Query: 567 QSLEACKYVWGLLDEWWEQPASTVVDWATVDGSNVAFWHNHVKKL 611
+ E C+ V GL+DEW+EQPA+TVVDW T+DG +V W N VK+L
Sbjct: 304 RCFEDCRRVRGLVDEWYEQPAATVVDWVTIDGQSVGAWINLVKQL 348
>UniRef100_Q6FWF3 Similar to tr|Q07457 Saccharomyces cerevisiae YDL074c BRE1 [Candida
glabrata]
Length = 693
Score = 45.1 bits (105), Expect = 0.006
Identities = 80/389 (20%), Positives = 150/389 (37%), Gaps = 64/389 (16%)
Query: 18 EYHKRLC-YYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDDVIVIETT 76
E K LC IN + D + ++ ++ N S S D + +E
Sbjct: 88 ESEKTLCDEMINAGESVDKERTDEFLNLLRKYTNSNGS---------SDSKIDSLGLELQ 138
Query: 77 REKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLGFDFDGQIPDEVRTVIV 136
R + + D+I S Y G V S + ++ + F+ Q D+ I
Sbjct: 139 RANKTKSELRLQNKKLTDEIDSLKAYYHGL-VRSYDREDSMTVKRVFNKQKTDDNTDNIS 197
Query: 137 NCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADA-----------EILRYKYENNIV 185
N + P + +T +H+K+++ E E ++ D+ + L +YEN I
Sbjct: 198 NTEQRPTSVSPVLTNGATHVKNEVKTEQEALNNHTDSSQSSGITEEEKKKLFLQYENKIT 257
Query: 186 MDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLAT--EDALNNAAE 243
D+ S N L + + QL E++ A + ++ + +N E
Sbjct: 258 -DLESH------------NSSLNRIIEELENYKQLNEKELAQTRLEISNLLSEKHSNEEE 304
Query: 244 ARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVG----------------SLRQLQLDVWA 287
DL ++ K TD+T + + Q + +L L+ A
Sbjct: 305 REDLLHQIEKLKASNTDLTLTNESFLSKFQELAKEKDTFQEKISSDFEKTLESLKAQNLA 364
Query: 288 KEREVSGLKASLNTLMSEIQRLNK------LCAERKEAEDSLKKKWKKIEEFDGRRSELE 341
E+++ ++ + + L+S++ L L ++ K+A D L+ +W+KIE R+
Sbjct: 365 LEKDLVRVRTTRDELISKVAILEAETSKSVLISDLKQALDILRDQWEKIE----FRNNQS 420
Query: 342 SIYTALLKANTDAASFWSQQPS-TAREYA 369
ALLK D S + + S T ++Y+
Sbjct: 421 PSSDALLKEIQDLESAFKELSSLTHKKYS 449
>UniRef100_UPI000021B39A UPI000021B39A UniRef100 entry
Length = 1173
Score = 44.7 bits (104), Expect = 0.008
Identities = 54/236 (22%), Positives = 104/236 (43%), Gaps = 17/236 (7%)
Query: 221 LERQKAHVQQFLATEDALNNA-AEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLR 279
LE+ K Q +++A ++ A+ R+ E L H S + AT ++ L
Sbjct: 336 LEKHKLETQSLATSKEATDSELAQLRETLEGLQAE-HAAKLNESEAALTKATEEHAAKLE 394
Query: 280 QLQLDVWAKERE-VSGLKAS----LNTLMSEIQRLNKLCAERKEA-EDSLKKKWKKIEEF 333
LQ + ++ ++ L+A L+ ++ K AE K + S+ + KKI++
Sbjct: 395 ALQQSLEEQQHAAIAALEAKHREELSKSSADAGDYEKAVAEMKSIHQSSISELQKKIDDL 454
Query: 334 DGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEK 393
+ ELE+ + A L + T++ STAR AL I A +ET+ A + +
Sbjct: 455 TASQQELEAAHEAKLLSETES--------STARLSALEAEIADLKAKLETAEQAAEAAKA 506
Query: 394 EVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARD 449
++ A + +L S +A L+A + + +EA+ ++ + A + +D
Sbjct: 507 DLEA-KTTLLPTLESRVSELEAELDAAKQAATKAEEALKESQAQLETVLAESKEKD 561
>UniRef100_P34288 GTPase-activating protein GAP [Caenorhabditis elegans]
Length = 1317
Score = 43.9 bits (102), Expect = 0.014
Identities = 50/167 (29%), Positives = 71/167 (41%), Gaps = 16/167 (9%)
Query: 296 KASLNTLMSEIQRLNK--------LCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTAL 347
KA+++ L + Q L+ L E +E ++KK + + R EL Y+
Sbjct: 1081 KAAIDALANHSQHLHLASSPAFEVLSEETREKIRRMQKKQSWHDTKELRSGELLKTYSPT 1140
Query: 348 LKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLY 407
K TDA S S ST LST P A + NS+ D + S R+P S
Sbjct: 1141 -KDLTDALSCTSDY-STTSSAPLSTNPPLAVACADQPNSSSDYASSDPSPCARNPSTSPA 1198
Query: 408 MLPSS---PQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPS 451
PS+ A L A SSG S Q + +I L + G+RDP+
Sbjct: 1199 SRPSNLAISPAQLHATSSSGQSHQPMSRSQKIR---LRTKLGSRDPA 1242
>UniRef100_UPI000042FE9D UPI000042FE9D UniRef100 entry
Length = 1103
Score = 43.1 bits (100), Expect = 0.024
Identities = 99/492 (20%), Positives = 184/492 (37%), Gaps = 63/492 (12%)
Query: 10 DEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDD 69
+E +R+ E +L Q A+ S + + ++ S +E E + D+
Sbjct: 485 EERNRLMMEMVSKL-----QKMGAEYSSIQTQLQSARSELTSSENELCSLKSSLQTAQDE 539
Query: 70 VIVIETTR--------EKNIRKACESLVAYMVDKIQSSFPAY--EGSGVLSNPQAEAAKL 119
+ + TT E KA ++ I++S EG G A+ A L
Sbjct: 540 LSRLRTTASDRDTAFAELESSKASLQTRLDELETIRNSLETKLAEGEGEADKSLADVAGL 599
Query: 120 GFDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYK 179
G D ++ D++ + N L S + + +K+ + +EIE + A+
Sbjct: 600 G-DKVKELEDQI-VELENKLGSEKGVWKQEEVKWQEIKAALEKEIEDLKNAAEG------ 651
Query: 180 YENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALN 239
+ G A S+ +L + AH Q +
Sbjct: 652 ----------------------HAEGLSRASSSAEASKEELGALRIAHEQLTTTYAGLVE 689
Query: 240 NAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGS------LRQLQLDVWAKEREVS 293
++ + E L ++L TD + ++S N S + +L+ + KE+EV
Sbjct: 690 ASSRHPEDVENLQRQLKEATDKHGALLLEASSSTNDQSAELKAQIEELKKEKEEKEKEVE 749
Query: 294 GLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIE-EFDGRRSELESIYTALLKANT 352
LK +L + E++ L E +E E L K +E E ++E E YTA +
Sbjct: 750 ELKGTLEEVEQEVELLKNARREGEERERQLNKIIAGLEAEIVELKAEFEDRYTAGYEDAK 809
Query: 353 DAA--SFWSQQPSTAREYAL--STIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYM 408
AA + S E ++ S + A +A +E+ +++ ++A + S ++L +
Sbjct: 810 RAAGEEHHKELASIRSEISVTHSRLQDAHTAELESLKASQSTTLATLNADHSSQTSALGL 869
Query: 409 -------LPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVSA 461
QA LE++ + E V + +AR DP + +V A
Sbjct: 870 SLQAANAQVEQDQAKLESVSEERDALAEQVERLKAELEGASARGDEVDPEVEAELKKVKA 929
Query: 462 ALQYAAGGLEGS 473
LQ+ + L G+
Sbjct: 930 ELQHVSDELAGA 941
>UniRef100_Q7SHZ4 Hypothetical protein [Neurospora crassa]
Length = 4007
Score = 42.4 bits (98), Expect = 0.041
Identities = 67/292 (22%), Positives = 119/292 (39%), Gaps = 29/292 (9%)
Query: 221 LERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSI-GIGATSQNVGSLR 279
LE A +++ A L AE +KL K+ D + I + +T + +G
Sbjct: 2310 LEETMAEIEKLSADNKQLT--AEISSYKDKL-KQSQTEADALNNDIKDMKSTKEKLGQDA 2366
Query: 280 QLQLDVWA-KEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEE-FDGRR 337
+ + V A K +E+ GLK S+N L +I N +++E D LK K D R
Sbjct: 2367 KAKETVLAEKMKEIQGLKDSINRLNQDISTKNATLDDKREIIDQLKDDIKTANSTIDTLR 2426
Query: 338 SELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSA 397
+++ + DA + AR+ L+ + A + + N+A +E A
Sbjct: 2427 KDVK---------DKDAILAHKTKDVVARDAELAKL----KAEIASKNAALAKKTEEAKA 2473
Query: 398 FYRSPDNSLYMLPSSPQALLEAIGSSGSS-GQEAVANAEISAAILTARAGARDPSAIPSI 456
F + ++ L + L + + + + Q+ ++++ I + +
Sbjct: 2474 F----EKNVQTLTDQAKGLNQDVATKTTQLAQDRATISKLNKDIFDLKTDV--TKLKQEL 2527
Query: 457 CRVSAALQYAAGGLEGSDAGLASILESL---EFCLKRRGSEASVLEDLLKAI 505
A L AG + DAGLA + E L E L ++ EAS LE +K +
Sbjct: 2528 STKDANLTQKAGEIGSRDAGLAKLREELRAKEAALAKKTEEASSLEKNVKKL 2579
Score = 40.0 bits (92), Expect = 0.20
Identities = 57/296 (19%), Positives = 116/296 (38%), Gaps = 19/296 (6%)
Query: 148 AITAYTSHLKSQISREIEKIDVRADAEILRYKYENNIVMDVSSSDGSSPLQYPLYGNGKL 207
A+ HL+ ++ R +D+RAD + E+ +D+ + G
Sbjct: 3271 ALREKVEHLEGEVGRRRRSLDLRADKILELTNSESAARLDLDKQRARNK-SLEETNTGLR 3329
Query: 208 GADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIG 267
GGS +L + +Q L E N+ A RD C+ L +H ++T
Sbjct: 3330 NRAAKEGGSLGRLGDEVTKLSRQLLEKE---NDLASLRDECDGLQTDIH---NLTRALAA 3383
Query: 268 IGA-TSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEI----QRLNKLCAERKEAEDS 322
GA SQ + L+ ++ ++ + +A++NTL S++ Q + L + +++
Sbjct: 3384 EGANVSQRDEQIALLKQELTTRQAALDAKQAAINTLESQLTEAQQAYDILASSNTTSQEE 3443
Query: 323 LKKKWKKIE-EFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVV 381
L + + +E+ S+ + + N D + +Q E + T++
Sbjct: 3444 LARSAAATQARLLACEAEIASLRSEITNLNEDITAKKTQ--IADNEKRIDTLLREAG--- 3498
Query: 382 ETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEIS 437
TS + ++ ++ +N +L L +A SS SS + ++ S
Sbjct: 3499 -TSEAQLARMKMTIAELQEEQENQQRLLDEYQSRLAQAATSSSSSSSSSDTASDTS 3553
>UniRef100_Q6C359 Similar to DEHA0C09658g Debaryomyces hansenii IPF 1836.1 [Yarrowia
lipolytica]
Length = 1906
Score = 42.0 bits (97), Expect = 0.054
Identities = 59/247 (23%), Positives = 101/247 (40%), Gaps = 40/247 (16%)
Query: 217 QNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVG 276
QN E QK+H Q +D + A+E +L KL K T+ +SR+ + +++
Sbjct: 961 QNSYDELQKSHEQLSSVGKDNESLASELAELKTKLSKI---ETESSSRADKVSELEKSLS 1017
Query: 277 SLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKW--------K 328
+ V A++ +VSG + T I+RL + +ER D LK
Sbjct: 1018 AAEAQSKSVAAEKEKVSG---QIATHEETIKRLKEELSERTAELDKLKSDLASSEKDLAS 1074
Query: 329 KIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTI------IPACSAVVE 382
K ++ + +E+E + + L AN+ A STA+E + T AC + +
Sbjct: 1075 KTKDVSAKDTEIEKLKSELETANSKLA-------STAKEVEILTSELKAAKSDACDSETK 1127
Query: 383 TSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILT 442
+L+E++ + + A L A SS SG +A LT
Sbjct: 1128 IKAVESELVEQKSKVEHLN-------------AELAAKSSSVESGAAELAEKVALVESLT 1174
Query: 443 ARAGARD 449
A+ ++D
Sbjct: 1175 AKLESKD 1181
>UniRef100_UPI00003C22C3 UPI00003C22C3 UniRef100 entry
Length = 1515
Score = 41.2 bits (95), Expect = 0.091
Identities = 36/145 (24%), Positives = 62/145 (41%), Gaps = 6/145 (4%)
Query: 346 ALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNS 405
A +K + S S P+ + + +T P SN++KD+ EK + S +
Sbjct: 617 AAVKVTSQVGSAASNAPNASIAASSTTCAPG-----PASNASKDISEKIAAILGSSSTSQ 671
Query: 406 LYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVSAALQY 465
ML L A +SGS AA+L++ AG S +P+ A+ +
Sbjct: 672 TTMLGPQGLPLAPAATASGSGAGSGAVPLGSIAAMLSSFAGQTSASNVPT-GSADASART 730
Query: 466 AAGGLEGSDAGLASILESLEFCLKR 490
+ +G + L+S+L+ LE K+
Sbjct: 731 STPAAQGQGSALSSLLQQLEEAKKK 755
>UniRef100_UPI0000433454 UPI0000433454 UniRef100 entry
Length = 1457
Score = 40.4 bits (93), Expect = 0.16
Identities = 48/224 (21%), Positives = 94/224 (41%), Gaps = 28/224 (12%)
Query: 280 QLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERK-------------------EAE 320
QLQ + + R + GLKAS L + ++R + AE++ E
Sbjct: 498 QLQGTLSEERRLIEGLKASQKELSTNLERAERSVAEKEKQVESLDAELRNSTIRMEDELR 557
Query: 321 DSLKKKWKKIEEFDGRRSEL-ESIYTALLKANTDAA---SFWSQQPSTAREYALSTIIPA 376
+SL K ++++E + R + E++ AL++ + + S+ SQ E ST + A
Sbjct: 558 ESLSAKDEQLKELNARLDSVKETLEEALVERDRQHSLLRSYESQINDLEMELRNSTDLDA 617
Query: 377 CSAV---VETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVAN 433
+ V + +DL++ +V+ RS + + + +E +G +E +A+
Sbjct: 618 LQDLEKRVLSLTKERDLLQLQVNDLTRSMEELKDSINRGRNSEMEKVGRERDEARETIAS 677
Query: 434 AEISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGL 477
+S A+ AR D + + +L+ A +E L
Sbjct: 678 --LSRALEEARERQSDKATVTDDTSTKESLERRASKIEERSVSL 719
>UniRef100_UPI0000433453 UPI0000433453 UniRef100 entry
Length = 1682
Score = 40.4 bits (93), Expect = 0.16
Identities = 48/224 (21%), Positives = 94/224 (41%), Gaps = 28/224 (12%)
Query: 280 QLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERK-------------------EAE 320
QLQ + + R + GLKAS L + ++R + AE++ E
Sbjct: 742 QLQGTLSEERRLIEGLKASQKELSTNLERAERSVAEKEKQVESLDAELRNSTIRMEDELR 801
Query: 321 DSLKKKWKKIEEFDGRRSEL-ESIYTALLKANTDAA---SFWSQQPSTAREYALSTIIPA 376
+SL K ++++E + R + E++ AL++ + + S+ SQ E ST + A
Sbjct: 802 ESLSAKDEQLKELNARLDSVKETLEEALVERDRQHSLLRSYESQINDLEMELRNSTDLDA 861
Query: 377 CSAV---VETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVAN 433
+ V + +DL++ +V+ RS + + + +E +G +E +A+
Sbjct: 862 LQDLEKRVLSLTKERDLLQLQVNDLTRSMEELKDSINRGRNSEMEKVGRERDEARETIAS 921
Query: 434 AEISAAILTARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGL 477
+S A+ AR D + + +L+ A +E L
Sbjct: 922 --LSRALEEARERQSDKATVTDDTSTKESLERRASKIEERSVSL 963
>UniRef100_UPI000042F508 UPI000042F508 UniRef100 entry
Length = 764
Score = 40.4 bits (93), Expect = 0.16
Identities = 33/134 (24%), Positives = 63/134 (46%), Gaps = 17/134 (12%)
Query: 217 QNQLLERQ----KAHVQQFLATEDALNNAAEARDLCE------KLMKRLHGGTDVTSRSI 266
+N+ L ++ KAH +F +E +NN R L E +L+++L +S
Sbjct: 229 ENEKLRKEIKSLKAHESEFAESE--VNNRETHRKLKEVERANKELIRKLEN-----LQSE 281
Query: 267 GIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKK 326
Q+ L +L+ +KE++++ KA L L + + +L A+ + L+
Sbjct: 282 HAADKEQHYKELAELERVSLSKEKDLARFKAELENLQNSVSEKTELRADNVHLSNQLEAS 341
Query: 327 WKKIEEFDGRRSEL 340
WK+I + + + SEL
Sbjct: 342 WKEIADLENQLSEL 355
>UniRef100_UPI000027C715 Transcription factor 12 [Homo sapiens]
Length = 1905
Score = 40.4 bits (93), Expect = 0.16
Identities = 70/343 (20%), Positives = 131/343 (37%), Gaps = 34/343 (9%)
Query: 29 QARDAQRSGVDSSVEMVNSFSAKNEKEAVYSTVKGSKSSDDVIVIETTREKNIR----KA 84
Q +D Q + +M +KEA TV G K + T K +R K
Sbjct: 1188 QVKDLQEQNAEMEKKMSELKIQIQKKEADRKTVTGLKELQQELERSTQECKRLREKLAKT 1247
Query: 85 CESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLGFDFDG---QIPDEVRTVIVNCLKS 141
L + + Q + + + Q + +++ + D + D RT ++ S
Sbjct: 1248 ETELQTTVEELFQVKIEREQYQTEIRDLQDQLSEMHDELDTAKKSVADGERTAMMAVTPS 1307
Query: 142 PPLLLQAITAYTSHLKSQISR------EIEKIDVRADAEILRYKYENNIVMDVSSSDGSS 195
P L ++ A HLK ++ E E + R + E+ K + +V++ D
Sbjct: 1308 P-LFDFSVWADMMHLKLELQEILLAKEEQEDVLKRRERELTALK--GALKEEVAAHDQEV 1364
Query: 196 PLQYPLYGN--GKLGADVPPGGSQNQLLERQKAHVQQF------------LATEDALNNA 241
Y G+L + + + R+KA V+ L TE A
Sbjct: 1365 DKLKEQYEKEIGRLQLSLEEAKQSSAAVVREKAEVEAAKGAVEGQAGRLTLETERLRRRA 1424
Query: 242 AEARDLCEKLMKRLHGGTDVTSRSIG--IGATSQNVGSLRQLQLDVWAKEREVSGLKASL 299
E + KL R+ + +G +G + L + ++ +E E+S +L
Sbjct: 1425 QELENEVAKL-NRIIDEAKLQESRLGDRVGRLEKEKKQLEESLAEIREQEEEMSRANRAL 1483
Query: 300 NTLMSEIQR-LNKLCAERKEAEDSLKKKWKKIEEFDGRRSELE 341
+ ++QR LN+L KE E+ L+++ + E+F ++ +E
Sbjct: 1484 TVRLEDVQRNLNRLSVTHKELEEMLQEERSQKEQFKNMKNNIE 1526
>UniRef100_UPI000004DF67 UPI000004DF67 UniRef100 entry
Length = 1038
Score = 40.4 bits (93), Expect = 0.16
Identities = 33/134 (24%), Positives = 63/134 (46%), Gaps = 17/134 (12%)
Query: 217 QNQLLERQ----KAHVQQFLATEDALNNAAEARDLCE------KLMKRLHGGTDVTSRSI 266
+N+ L ++ KAH +F +E +NN R L E +L+++L +S
Sbjct: 503 ENEKLRKEIKSLKAHESEFAESE--VNNRETHRKLKEVERANKELIRKLEN-----LQSE 555
Query: 267 GIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKK 326
Q+ L +L+ +KE++++ KA L L + + +L A+ + L+
Sbjct: 556 HAADKEQHYKELAELERVSLSKEKDLARFKAELENLQNSVSEKTELRADNVHLSNQLEAS 615
Query: 327 WKKIEEFDGRRSEL 340
WK+I + + + SEL
Sbjct: 616 WKEIADLENQLSEL 629
>UniRef100_Q7ZW40 Similar to vesicle docking protein p115 [Brachydanio rerio]
Length = 956
Score = 40.4 bits (93), Expect = 0.16
Identities = 66/283 (23%), Positives = 122/283 (42%), Gaps = 62/283 (21%)
Query: 148 AITAYTSHLKSQISREIEKIDVRADAEILRYKYENNIVMDVSSSDGSSPLQYPLYGNGKL 207
A+T+ T +L+S I++++ +I ++K + NI+ KL
Sbjct: 684 ALTSQTENLESTITQQLSQIQ--------QHKDQYNIL------------------KLKL 717
Query: 208 GADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSR-SI 266
G D G+Q + AHV E L+ E + ++ + L T +T + S+
Sbjct: 718 GKDTQQAGNQ-----AEAAHVNGLQPEE--LSQLRERLEEVQRQNQLLQ--TQLTEKDSL 768
Query: 267 GIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKE-------- 318
++ V S+ +V +EV GLKA L + +EI +L ER+E
Sbjct: 769 ITSLKAEGVPSVEDSTSEVTELRKEVEGLKAQLQSQTAEITQLQN---ERQELLRGSEPT 825
Query: 319 ----AEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPSTAREYALSTII 374
AE+S+ K +KI E + R L + T K + + + S ++ A +T
Sbjct: 826 VAVPAEESVYK--EKIAELESR---LSAQTTENQKLKGEVKTLLESKESMEKDLASAT-- 878
Query: 375 PACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALL 417
+ SA+++ + K+ +++EV + D+ L +L Q +L
Sbjct: 879 -STSAILQ---AEKNKLQQEVQESKKEQDDLLMLLADQDQKIL 917
>UniRef100_O74338 SPBC1A4.05 protein [Schizosaccharomyces pombe]
Length = 700
Score = 40.4 bits (93), Expect = 0.16
Identities = 53/204 (25%), Positives = 86/204 (41%), Gaps = 26/204 (12%)
Query: 279 RQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRS 338
RQLQ++V E + + L + +N L + +LN D +K K ++E + R
Sbjct: 504 RQLQMNVEELEGKKADLTSKINRLDKDFVKLN---TTYSLLTDQVKTKQLALQEIESRVI 560
Query: 339 ELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAF 398
LE L K + A S++ E+ P+ SAVV +IEK+
Sbjct: 561 RLEERLNMLQKLSMQPAI------SSSSEFVPIESHPSSSAVVPIEEPKNSIIEKK-RVP 613
Query: 399 YRSPDNSLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICR 458
+ + NS+ + + P + SGSS + V + TA R PS R
Sbjct: 614 HNAARNSVNLEVTLPNSKKRFSSFSGSSSKLPVRPS-------TALTDKRKPSWSR---R 663
Query: 459 VSAALQYAAGGLE------GSDAG 476
++AA+ +++G E SDAG
Sbjct: 664 LAAAIGFSSGSPEKKHVITSSDAG 687
>UniRef100_Q7KT48 CG5020-PD, isoform D [Drosophila melanogaster]
Length = 1677
Score = 40.0 bits (92), Expect = 0.20
Identities = 63/297 (21%), Positives = 116/297 (38%), Gaps = 29/297 (9%)
Query: 206 KLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRS 265
KL ++ SQ + + + Q L + AA L E+ K H +T
Sbjct: 845 KLHDEISQLKSQAEETQSELKSTQSNLEAKSKQLEAANG-SLEEEAKKSGHLLEQITKLK 903
Query: 266 IGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKK 325
+G T +L DV +K +++ A+L +++NK AE + L+
Sbjct: 904 SEVGETQ---AALSSCHTDVESKTKQLEAANAAL-------EKVNKEYAESRAEASDLQD 953
Query: 326 KWKKIE-----EFDGRRSELESIYTALLKAN----------TDAASFWSQQPSTAREYAL 370
K K+I E RS +++T L K + T A WSQ+ +E L
Sbjct: 954 KVKEITDTLHAELQAERSSSSALHTKLSKFSDEIATGHKELTSKADAWSQE-MLQKEKEL 1012
Query: 371 STIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEA 430
+ ++ K E++ +F S N + + LE + ++ ++
Sbjct: 1013 QELRQQLQDSQDSQTKLKAEGERKEKSFEESIKNLQEEVTKAKTENLELSTGTQTTIKDL 1072
Query: 431 VANAEISAAILT--ARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASILESLE 485
EI+ A L + + D I + + A+Q A + ++A L+++LE L+
Sbjct: 1073 QERLEITNAELQHKEKMASEDAQKIADLKTLVEAIQVANANISATNAELSTVLEVLQ 1129
>UniRef100_Q9HEL8 Hypothetical protein 12F11.080 [Neurospora crassa]
Length = 602
Score = 40.0 bits (92), Expect = 0.20
Identities = 55/249 (22%), Positives = 99/249 (39%), Gaps = 37/249 (14%)
Query: 157 KSQISR--EIEKIDVR-----ADAEILRYKYENNIVMDVSSSDGSSPLQYPLYGNGKLGA 209
+ +SR E+EK + R ADAE K E + + DG G+
Sbjct: 156 RKHVSRIEELEKENKRLAKEAADAEKRWQKAEEELADFREAEDGKQ--------GGEGAE 207
Query: 210 DVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRSIGIG 269
+V ++ L+RQ +Q LA N A GG S SI +G
Sbjct: 208 NVEKLKNELAALQRQNTQLQSQLARAGG-NGAG--------------GGAHGRSPSISVG 252
Query: 270 ATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKKKWKK 329
+ QL+ + +K + ++ ++ L +++++L+ + E +L++K +
Sbjct: 253 GGGSPSAATAQLEAQLQSKSATIETMELEMSRLRAQVEKLSASASGPSEQIAALEEKLAR 312
Query: 330 IEEFDG-RRSELESIYTALLKANTDAASFWSQQPSTAREYALSTIIPACSAVVETSNSAK 388
E+ G + EL + L + + +A SQ+ S T + VE +N K
Sbjct: 313 AEKAAGLAQQELADLKQNLDRVSKEAVKEGSQRASA------ETKLRTLEKEVEEANKTK 366
Query: 389 DLIEKEVSA 397
D + K+ A
Sbjct: 367 DELAKKAEA 375
>UniRef100_Q9VJE5-2 Splice isoform B of Q9VJE5 [Drosophila melanogaster]
Length = 1689
Score = 40.0 bits (92), Expect = 0.20
Identities = 63/297 (21%), Positives = 116/297 (38%), Gaps = 29/297 (9%)
Query: 206 KLGADVPPGGSQNQLLERQKAHVQQFLATEDALNNAAEARDLCEKLMKRLHGGTDVTSRS 265
KL ++ SQ + + + Q L + AA L E+ K H +T
Sbjct: 857 KLHDEISQLKSQAEETQSELKSTQSNLEAKSKQLEAANG-SLEEEAKKSGHLLEQITKLK 915
Query: 266 IGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQRLNKLCAERKEAEDSLKK 325
+G T +L DV +K +++ A+L +++NK AE + L+
Sbjct: 916 SEVGETQ---AALSSCHTDVESKTKQLEAANAAL-------EKVNKEYAESRAEASDLQD 965
Query: 326 KWKKIE-----EFDGRRSELESIYTALLKAN----------TDAASFWSQQPSTAREYAL 370
K K+I E RS +++T L K + T A WSQ+ +E L
Sbjct: 966 KVKEITDTLHAELQAERSSSSALHTKLSKFSDEIATGHKELTSKADAWSQE-MLQKEKEL 1024
Query: 371 STIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSSGSSGQEA 430
+ ++ K E++ +F S N + + LE + ++ ++
Sbjct: 1025 QELRQQLQDSQDSQTKLKAEGERKEKSFEESIKNLQEEVTKAKTENLELSTGTQTTIKDL 1084
Query: 431 VANAEISAAILT--ARAGARDPSAIPSICRVSAALQYAAGGLEGSDAGLASILESLE 485
EI+ A L + + D I + + A+Q A + ++A L+++LE L+
Sbjct: 1085 QERLEITNAELQHKEKMASEDAQKIADLKTLVEAIQVANANISATNAELSTVLEVLQ 1141
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 951,511,817
Number of Sequences: 2790947
Number of extensions: 37916970
Number of successful extensions: 107635
Number of sequences better than 10.0: 169
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 164
Number of HSP's that attempted gapping in prelim test: 107406
Number of HSP's gapped (non-prelim): 410
length of query: 621
length of database: 848,049,833
effective HSP length: 134
effective length of query: 487
effective length of database: 474,062,935
effective search space: 230868649345
effective search space used: 230868649345
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 78 (34.7 bits)
Medicago: description of AC137823.16


