

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.5 - phase: 0
(203 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
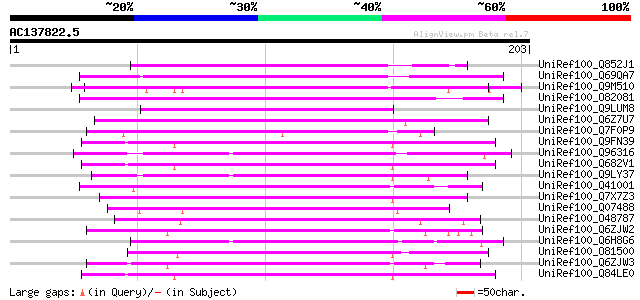
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q852J1 Putative blue copper-binding protein [Oryza sat... 105 9e-22
UniRef100_Q69QA7 Putative blue copper protein [Oryza sativa] 102 6e-21
UniRef100_Q9M510 Dicyanin [Lycopersicon esculentum] 99 9e-20
UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana] 96 4e-19
UniRef100_Q9LUM8 Blue copper-binding protein-like [Arabidopsis t... 94 2e-18
UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa] 91 2e-17
UniRef100_Q7F0P9 Putative blue copper protein [Oryza sativa] 91 2e-17
UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like pr... 90 3e-17
UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis th... 90 3e-17
UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis tha... 89 5e-17
UniRef100_Q9LY37 Stellacyanin (Uclacyanin 3)-like protein [Arabi... 88 2e-16
UniRef100_Q41001 Blue copper protein precursor [Pisum sativum] 87 2e-16
UniRef100_Q7X7Z3 OSJNBa0079A21.5 protein [Oryza sativa] 87 3e-16
UniRef100_Q07488 Blue copper protein precursor [Arabidopsis thal... 86 4e-16
UniRef100_O48787 Putative phytocyanin [Arabidopsis thaliana] 86 6e-16
UniRef100_Q6ZJW2 Putative blue copper binding protein [Oryza sat... 86 6e-16
UniRef100_Q6H8G6 Putative uclacyanin 3 [Oryza sativa] 86 6e-16
UniRef100_O81500 F9D12.16 protein [Arabidopsis thaliana] 84 2e-15
UniRef100_Q6ZJW3 Putative blue copper binding protein [Oryza sat... 84 2e-15
UniRef100_Q84LE0 Phytocyanin protein, PUP2 [Arabidopsis thaliana] 83 5e-15
>UniRef100_Q852J1 Putative blue copper-binding protein [Oryza sativa]
Length = 172
Score = 105 bits (261), Expect = 9e-22
Identities = 54/132 (40%), Positives = 72/132 (53%), Gaps = 11/132 (8%)
Query: 48 AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVP 107
A A + +VGD GW FDY W A K FR+GDTL F Y G VV V G+DFK+C+
Sbjct: 25 AAATEHMVGDGNGWILGFDYAAWAATKQFRVGDTLVFRYKGTNHTVVEVGGADFKACNKT 84
Query: 108 LTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSL 167
+A +SG+D++ + GRRW+ V DHC KL ITV + + +P+P
Sbjct: 85 ASANEWSSGEDRVALDKEGRRWFFCGVGDHCAKNMKLKITV---------IAAGAPAPGA 135
Query: 168 DLVTPEAPPSNA 179
P PPS+A
Sbjct: 136 SEAPP--PPSSA 145
>UniRef100_Q69QA7 Putative blue copper protein [Oryza sativa]
Length = 198
Score = 102 bits (254), Expect = 6e-21
Identities = 55/166 (33%), Positives = 79/166 (47%), Gaps = 9/166 (5%)
Query: 28 MALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 87
MA S AL ++ A K + VGD GW DY TW ++K F++GDTL FNY
Sbjct: 1 MASSLALVALLLVSCAVVAAAATK-YTVGDTSGWAMGADYTTWASDKKFKMGDTLVFNYA 59
Query: 88 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 147
GG +V V+ +D+ +C+ +SG + + T G+ ++I + HC NG KL +
Sbjct: 60 GGAHSVDEVSAADYAACTASNALQSDSSGTTTVTLKTAGKHYFICGIAGHCSNGMKLVVD 119
Query: 148 VQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 193
V SP+P+P TP P + PAS L P
Sbjct: 120 V--------AAASPAPAPKAPSTTPTTPSTTPATPASPGTSSGLTP 157
>UniRef100_Q9M510 Dicyanin [Lycopersicon esculentum]
Length = 332
Score = 98.6 bits (244), Expect = 9e-20
Identities = 57/168 (33%), Positives = 91/168 (53%), Gaps = 6/168 (3%)
Query: 25 TSTMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFD----YQTWTANKVFRLGD 80
T+T ++ A + AT S +VA+ ++VGD GW+ YQ W NK F++GD
Sbjct: 145 TATPVMAPAPSVSVAPATAPSPASVAQTYIVGDNLGWSVPTSGPNSYQRWANNKSFKVGD 204
Query: 81 TLTFNYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCEN 140
TL FN+V G NV V+ + + SC+ +++G +I +T G +Y+ + HC
Sbjct: 205 TLVFNFVNGTHNVAMVSKASYDSCNTTSPINTISNGPARIRLTNSGEHYYMCTFPRHCSL 264
Query: 141 GQKLFITVQPKQDGWSPVPSPSPSPSLDLV-TPEAPPSNAPWPASSVP 187
GQKL I V D +P PS + +PS V + ++P ++ P P++S P
Sbjct: 265 GQKLAINV-TGSDVTAPTPSTAATPSSPTVPSSDSPVNSPPAPSASAP 311
Score = 86.3 bits (212), Expect = 4e-16
Identities = 57/177 (32%), Positives = 83/177 (46%), Gaps = 14/177 (7%)
Query: 30 LSRALFLFAFIATIFSTMAVAKD-FVVGDEKGWTT----LFDYQTWTANKVFRLGDTLTF 84
LS + + + + +A+A+ VVGD GWT Y TW A K F +GD L F
Sbjct: 5 LSSLVVFGSILFALLQHVAMAQQTHVVGDTLGWTVPNGGAASYSTWAAGKSFVVGDILVF 64
Query: 85 NYVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKL 144
N+ G +V V+ F SC+ + T+G I +++ G +Y+ + HC GQKL
Sbjct: 65 NFRSGSHSVAEVSKGAFDSCNTSSPISISTNGPTNITLSSAGSHYYLCTFPSHCTLGQKL 124
Query: 145 FITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSV-PRRSLLPKKLFQIF 200
I V SP P P+P+ P A P AP P+ SV P + P + Q +
Sbjct: 125 AINVSGSA---SPAPQPAPA-----TPPTATPVMAPAPSVSVAPATAPSPASVAQTY 173
>UniRef100_O82081 Uclacyanin I [Arabidopsis thaliana]
Length = 261
Score = 96.3 bits (238), Expect = 4e-19
Identities = 54/166 (32%), Positives = 77/166 (45%), Gaps = 10/166 (6%)
Query: 28 MALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYV 87
MA L + + +AT + VA D +G GWT +TW A + F +GD L F+Y
Sbjct: 1 MASREMLIIISVLATTLIGLTVATDHTIGGPSGWTVGASLRTWAAGQTFAVGDNLVFSYP 60
Query: 88 GGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 147
+VV V +F SC +G + +TT G+R++I + HC G KL +
Sbjct: 61 AAFHDVVEVTKPEFDSCQAVKPLITFANGNSLVPLTTPGKRYFICGMPGHCSQGMKLEVN 120
Query: 148 VQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRSLLP 193
V P P P+ PSL NAP P+S +P + LLP
Sbjct: 121 VVPTATVAPTAPLPNTVPSL----------NAPSPSSVLPIQPLLP 156
>UniRef100_Q9LUM8 Blue copper-binding protein-like [Arabidopsis thaliana]
Length = 129
Score = 94.0 bits (232), Expect = 2e-18
Identities = 40/99 (40%), Positives = 56/99 (56%)
Query: 52 DFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVPLTAP 111
+ +VGD GW +Y WT + F +GD L FNY + NV++VN + + C +
Sbjct: 31 EHIVGDSNGWELFTNYTNWTQGREFHVGDVLVFNYKSDQHNVMQVNSTAYTDCGLDNYTT 90
Query: 112 VLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP 150
+ T G D II++ G+ W+I V DHC NGQKL I V P
Sbjct: 91 LFTKGNDSIILSEVGKLWFICGVDDHCVNGQKLSINVAP 129
>UniRef100_Q6Z7U7 Putative Blue copper protein [Oryza sativa]
Length = 246
Score = 90.9 bits (224), Expect = 2e-17
Identities = 45/164 (27%), Positives = 75/164 (45%), Gaps = 10/164 (6%)
Query: 34 LFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNV 93
L + A + ++ A A + VGD GWT DY +W +K F++GD+L F Y G V
Sbjct: 9 LTIAALLLAACASSAAATSYTVGDASGWTIGVDYTSWAGSKSFKVGDSLVFKYASGAHTV 68
Query: 94 VRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQD 153
V V+ + + +C+ +SG + + T G+ ++I ++ HC G K+ + V
Sbjct: 69 VEVSAAGYLACAAANALGSDSSGSTTVALKTPGKHYFICTIAGHCAGGMKMEVDVSGSSS 128
Query: 154 ----------GWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVP 187
G PS SP+ P P P+P++ +P
Sbjct: 129 SSSGGGGGGGGGGSTPSSPSSPTPTTPNPSTPTPTTPYPSTPMP 172
>UniRef100_Q7F0P9 Putative blue copper protein [Oryza sativa]
Length = 168
Score = 90.5 bits (223), Expect = 2e-17
Identities = 51/140 (36%), Positives = 71/140 (50%), Gaps = 7/140 (5%)
Query: 31 SRALFLFAFIATI--FSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVG 88
SR + L A ++ + M A D+ VGD GWT + W K F++GD L F Y
Sbjct: 3 SRQVLLLAIVSAVALLPAMVSATDYTVGDGHGWTLEYPSTNWADGKSFQIGDKLVFTYTK 62
Query: 89 GKDNVVRVNGSDFKSCS-VPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFIT 147
GK V V+G+ F +C+ T SG D + + G+RW+ +V +HCE G KL +
Sbjct: 63 GKHTVTEVDGAAFHACNRQGNTLMTWNSGNDTVALDKAGKRWFFCNVDNHCELGMKLVVD 122
Query: 148 VQPKQDGWSPVP-SPSPSPS 166
V D +P P SP P PS
Sbjct: 123 V---ADPNAPAPASPPPPPS 139
>UniRef100_Q9FN39 Similarity to phytocyanin/early nodulin-like protein [Arabidopsis
thaliana]
Length = 370
Score = 90.1 bits (222), Expect = 3e-17
Identities = 58/169 (34%), Positives = 83/169 (48%), Gaps = 8/169 (4%)
Query: 29 ALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNY 86
+L + + A AT FS +A A F VG W T +Y TW F++ D+L F Y
Sbjct: 7 SLCFSFLILASFATFFS-VADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSLYFKY 65
Query: 87 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 146
G D+V +V +DF C+V +G+ + + G ++IS DHC+ GQKL +
Sbjct: 66 AKGSDSVQQVMKADFDGCNVRNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIV 125
Query: 147 TV-----QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 190
V QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 126 VVLAVRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
>UniRef100_Q96316 Blue-copper binging protein III [Arabidopsis thaliana]
Length = 222
Score = 90.1 bits (222), Expect = 3e-17
Identities = 63/176 (35%), Positives = 81/176 (45%), Gaps = 16/176 (9%)
Query: 26 STMALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFN 85
ST+A + LFL A A +T F VGD GWT+ DY W K FR+GDTL F
Sbjct: 3 STVAAALLLFLAAVPAVFAAT------FKVGDISGWTSNLDYTVWLTGKTFRVGDTLEFV 56
Query: 86 YVGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLF 145
Y G +V V+ + + +C G KI +TT G ++ HC+NG KL
Sbjct: 57 Y-GLSHSVSVVDKAGYDNCDSSGATQNFADGDTKIDLTTVGTMHFLCPTFGHCKNGMKLA 115
Query: 146 ITVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPAS-----SVPRRSLLPKKL 196
+ V +P PS SP TP +PPS P+S S P SL P L
Sbjct: 116 VPVLAA----APSPSTPSSPPSTPSTPSSPPSTPSTPSSPPSPPSPPSPSLPPSSL 167
>UniRef100_Q682V1 Predicted GPI-anchored protein [Arabidopsis thaliana]
Length = 370
Score = 89.4 bits (220), Expect = 5e-17
Identities = 58/169 (34%), Positives = 83/169 (48%), Gaps = 8/169 (4%)
Query: 29 ALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNY 86
+L + + A AT FS +A A F VG W T +Y TW F++ D+L F Y
Sbjct: 7 SLCFSFLILASFATFFS-VADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSLYFKY 65
Query: 87 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 146
G D+V +V +DF C+V +G+ + + G ++IS DHC+ GQKL +
Sbjct: 66 EKGSDSVQQVMKADFDGCNVRNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIV 125
Query: 147 TV-----QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 190
V QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 126 VVLAVRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
>UniRef100_Q9LY37 Stellacyanin (Uclacyanin 3)-like protein [Arabidopsis thaliana]
Length = 187
Score = 87.8 bits (216), Expect = 2e-16
Identities = 54/154 (35%), Positives = 72/154 (46%), Gaps = 10/154 (6%)
Query: 33 ALFLFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDN 92
AL L + + + AV F VGD GWT +Y +W + K FR+GDTL F Y G +
Sbjct: 8 ALLLLLLLVAVPAVFAVT--FQVGDNDGWTIGVEYTSWVSEKTFRVGDTLEFKY-GPSHS 64
Query: 93 VVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV---- 148
V VN +D+ C + G KI +T G ++ HC G KL + V
Sbjct: 65 VAVVNKADYDGCETSRPTQSFSDGDTKIDLTKVGAIHFLCLTPGHCSLGMKLAVQVLAAV 124
Query: 149 --QPKQDGWSPVPSPS-PSPSLDLVTPEAPPSNA 179
+P +P PSPS PSPS +P P NA
Sbjct: 125 SLEPPPSPSAPSPSPSAPSPSPSAPSPSPSPGNA 158
>UniRef100_Q41001 Blue copper protein precursor [Pisum sativum]
Length = 189
Score = 87.4 bits (215), Expect = 2e-16
Identities = 56/159 (35%), Positives = 80/159 (50%), Gaps = 7/159 (4%)
Query: 28 MALSRALFLFAFIATIFSTM-AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNY 86
MA S AL L +A I + ++A + VGD GW DY TW ++K F +GD+L FNY
Sbjct: 1 MAFSNALVLCFLLAIINMALPSLATVYTVGDTSGWVIGGDYSTWASDKTFAVGDSLVFNY 60
Query: 87 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 146
G V V SD+KSC+ + ++G I + G+ ++I V H G KL I
Sbjct: 61 GAGAHTVDEVKESDYKSCTSGNSISTDSTGATTIPLKKAGKHYFICGVPGHSTGGMKLSI 120
Query: 147 TVQPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASS 185
V+ G S PS +PS S + PS+ PA++
Sbjct: 121 KVK-ASSGSSAAPSATPSSS-----GKGSPSSDDTPAAT 153
>UniRef100_Q7X7Z3 OSJNBa0079A21.5 protein [Oryza sativa]
Length = 181
Score = 87.0 bits (214), Expect = 3e-16
Identities = 51/147 (34%), Positives = 67/147 (44%), Gaps = 3/147 (2%)
Query: 36 LFAFIATIFSTMAVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVR 95
L A + + A AKD+ VGD GWTT DY W K F +GDTL F Y +VV
Sbjct: 9 LVAVLLAAAAAPASAKDYTVGDSSGWTTGVDYTAWARGKTFNIGDTLLFQYTSAGHSVVE 68
Query: 96 VNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV---QPKQ 152
V+ +D SCS G + +T G R++I T HC G KL +TV
Sbjct: 69 VSEADHTSCSAANPLRSYKDGTTIVTLTRSGTRYFICGSTGHCGAGMKLTVTVATLSGSA 128
Query: 153 DGWSPVPSPSPSPSLDLVTPEAPPSNA 179
G + + PS S + T S+A
Sbjct: 129 AGGTRLAKPSSSDADPTTTTTTRTSSA 155
>UniRef100_Q07488 Blue copper protein precursor [Arabidopsis thaliana]
Length = 196
Score = 86.3 bits (212), Expect = 4e-16
Identities = 50/144 (34%), Positives = 75/144 (51%), Gaps = 10/144 (6%)
Query: 39 FIATIFSTMAV-AKDFVVGDEKGWTTLFD---YQTWTANKVFRLGDTLTFNYVGGKDNVV 94
F+ +F+ + V A+D+ VGD+ WT D Y TW K FR+GD L F++ G+ +V
Sbjct: 10 FLVLVFAAVVVFAEDYDVGDDTEWTRPMDPEFYTTWATGKTFRVGDELEFDFAAGRHDVA 69
Query: 95 RVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQP---- 150
V+ + F++C +T KI++ T G +++I +V DHC GQKL ITV
Sbjct: 70 VVSEAAFENCEKEKPISHMTVPPVKIMLNTTGPQYFICTVGDHCRFGQKLSITVVAAGAT 129
Query: 151 --KQDGWSPVPSPSPSPSLDLVTP 172
G P+P +PS TP
Sbjct: 130 GGATPGAGATPAPGSTPSTGGTTP 153
>UniRef100_O48787 Putative phytocyanin [Arabidopsis thaliana]
Length = 206
Score = 85.9 bits (211), Expect = 6e-16
Identities = 56/157 (35%), Positives = 74/157 (46%), Gaps = 14/157 (8%)
Query: 42 TIFSTMAVAKDFVVGDEKGWTTLF-DYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSD 100
T+F VG+ KGWT + DY+ W +++VF++GDTL F Y +V V +D
Sbjct: 18 TLFGVAVGGTVHKVGNTKGWTMIGGDYEAWASSRVFQVGDTLVFAYNKDYHDVTEVTHND 77
Query: 101 FKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVP- 159
F+ C +G D I +T G + +I V HC+ GQKL I V P G VP
Sbjct: 78 FEMCESSKPLRRYKTGSDSISLTKPGLQHFICGVPGHCKKGQKLQIHVLPASLGHVAVPV 137
Query: 160 -----------SPSPSPSLDLVTPEAPP-SNAPWPAS 184
SPSPSP +D AP P PAS
Sbjct: 138 PGPVRSQSSSSSPSPSPLVDPPVNNAPQYQMGPTPAS 174
>UniRef100_Q6ZJW2 Putative blue copper binding protein [Oryza sativa]
Length = 188
Score = 85.9 bits (211), Expect = 6e-16
Identities = 57/161 (35%), Positives = 76/161 (46%), Gaps = 7/161 (4%)
Query: 31 SRALFLFAFIATIFSTMAVAKDFVVGDEKG-WTTLFDYQTWTANKVFRLGDTLTFNYVGG 89
+RAL + A A + T + VG G W T +Y W + FR+GD L F Y
Sbjct: 3 ARALLVVAMAAAVLGTAMGVTTYTVGAPAGSWDTRTNYAQWVSAITFRVGDQLVFKYSPA 62
Query: 90 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 149
+VV VN +D+ SCS SG D I + G R++I HC G K+ + V+
Sbjct: 63 AHDVVEVNKADYDSCSSSSPISTFNSGDDTIPLAAIGTRYFICGFPGHCTAGMKVAVKVE 122
Query: 150 PKQDGWSPVPSP-SPSPSLDLV-TPEA-PPSNA--PWPASS 185
G +P PSP +P P V P A PP+N P P SS
Sbjct: 123 -AATGSNPTPSPLAPLPRTPTVMAPNAMPPTNGGRPTPPSS 162
>UniRef100_Q6H8G6 Putative uclacyanin 3 [Oryza sativa]
Length = 202
Score = 85.9 bits (211), Expect = 6e-16
Identities = 50/147 (34%), Positives = 71/147 (48%), Gaps = 4/147 (2%)
Query: 48 AVAKDFVVGDEKGWTTLFDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSCSVP 107
A A D+ VGD GW++ DY TW +K F +GD+L F Y V V+ +D+ +CS
Sbjct: 23 AFAVDYTVGDTSGWSSGVDYDTWAKSKTFSVGDSLVFQY-SMMHTVAEVSSADYSACSAS 81
Query: 108 LTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQPKQDGWSPVPSPSPSPSL 167
+ + KI +T G R++I + HC G KL +TV +P P+ S SP
Sbjct: 82 NSIQSYSDQNTKIALTKPGTRYFICGTSGHCSGGMKLAVTVSAAA-ATTPTPTASSSPP- 139
Query: 168 DLVTPEAPPSNAPWPA-SSVPRRSLLP 193
TP P S+ SS P + P
Sbjct: 140 STATPATPSSDPGMDTPSSTPDATTTP 166
>UniRef100_O81500 F9D12.16 protein [Arabidopsis thaliana]
Length = 187
Score = 84.3 bits (207), Expect = 2e-15
Identities = 50/149 (33%), Positives = 72/149 (47%), Gaps = 11/149 (7%)
Query: 47 MAVAKDFVVGDEKGWTTL--FDYQTWTANKVFRLGDTLTFNYVGGKDNVVRVNGSDFKSC 104
++ A + VGD GWTT+ DY+ W + K F +GDT+ F Y NV+RV ++SC
Sbjct: 18 LSEAAVYKVGDSAGWTTIANVDYKLWASTKTFHIGDTVLFEYNPQFHNVMRVTHPMYRSC 77
Query: 105 SVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITV------QPKQDGWSPV 158
+ T+G D I +T +G ++ V HC GQKL + V P D P
Sbjct: 78 NTSKPISTFTTGNDSITLTNHGHHFFFCGVPGHCLAGQKLDLHVLLPASSTPLSD---PP 134
Query: 159 PSPSPSPSLDLVTPEAPPSNAPWPASSVP 187
S S SP + P +P A+S+P
Sbjct: 135 TSSSSSPPSTTIPAAGVPGPSPSLAASLP 163
>UniRef100_Q6ZJW3 Putative blue copper binding protein [Oryza sativa]
Length = 187
Score = 84.3 bits (207), Expect = 2e-15
Identities = 54/156 (34%), Positives = 74/156 (46%), Gaps = 8/156 (5%)
Query: 31 SRALFLFAFIATIFSTMAVAKDFVVGDEKG-WTTLFDYQTWTANKVFRLGDTLTFNYVGG 89
+RAL + A A + T A+ + VG G W +Y W +N FR GD + F Y
Sbjct: 3 ARALLVVAMAAAVLGT-ALGATYTVGAPSGSWDLRTNYDQWVSNINFRAGDQIVFKYSPA 61
Query: 90 KDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFITVQ 149
+VV VN +D+ SCS SG D I +T G R++I HC G K+ + V+
Sbjct: 62 AHDVVEVNKADYDSCSSSSPIATFNSGDDTIPLTATGTRYFICGFNGHCTGGMKVAVKVE 121
Query: 150 PKQDGWSPVPSP-SPSPSLDLVTPEAPPSNAPWPAS 184
G +P PSP +P P TP A NA P +
Sbjct: 122 -AATGSNPAPSPMTPRPR----TPTAMAPNAMPPTA 152
>UniRef100_Q84LE0 Phytocyanin protein, PUP2 [Arabidopsis thaliana]
Length = 370
Score = 82.8 bits (203), Expect = 5e-15
Identities = 55/169 (32%), Positives = 80/169 (46%), Gaps = 8/169 (4%)
Query: 29 ALSRALFLFAFIATIFSTMAVAKDFVVGDEKGWTT--LFDYQTWTANKVFRLGDTLTFNY 86
+L + + A AT FS +A A F VG W T +Y TW F++ D+ F Y
Sbjct: 7 SLCFSFLILASFATFFS-VADAWRFNVGGNGAWVTNPQENYNTWAERNRFQVNDSPYFKY 65
Query: 87 VGGKDNVVRVNGSDFKSCSVPLTAPVLTSGQDKIIITTYGRRWYISSVTDHCENGQKLFI 146
D+V +V +DF C+ +G+ + + G ++IS DHC+ GQKL +
Sbjct: 66 AKRSDSVQQVMKADFDGCNARNPIKNFENGESVVTLDRSGAFYFISGNQDHCQKGQKLIV 125
Query: 147 TV-----QPKQDGWSPVPSPSPSPSLDLVTPEAPPSNAPWPASSVPRRS 190
V QP SPVPS SP+ +P +P + A P+ S P RS
Sbjct: 126 VVLAVRNQPSAPAHSPVPSVSPTQPPKSHSPVSPVAPASAPSKSQPPRS 174
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 345,477,536
Number of Sequences: 2790947
Number of extensions: 15858620
Number of successful extensions: 87895
Number of sequences better than 10.0: 273
Number of HSP's better than 10.0 without gapping: 164
Number of HSP's successfully gapped in prelim test: 115
Number of HSP's that attempted gapping in prelim test: 86625
Number of HSP's gapped (non-prelim): 835
length of query: 203
length of database: 848,049,833
effective HSP length: 121
effective length of query: 82
effective length of database: 510,345,246
effective search space: 41848310172
effective search space used: 41848310172
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Medicago: description of AC137822.5


