

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137667.13 - phase: 0 /pseudo
(267 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
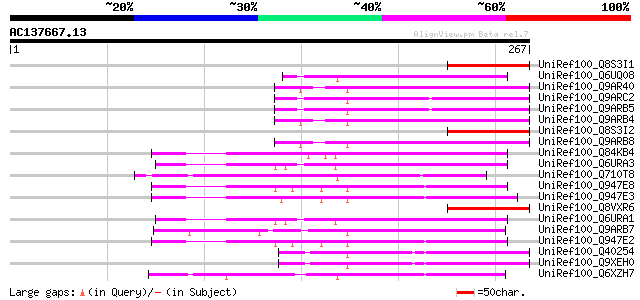
UniRef100_Q8S3I1 NBS-LRR-Toll resistance gene analog protein [Me... 70 7e-11
UniRef100_Q6UQ08 Putative TIR-NBS type R protein 11 [Malus domes... 69 1e-10
UniRef100_Q9AR40 Nbi-B protein [Linum usitatissimum] 68 2e-10
UniRef100_Q9ARC2 Ngc-A protein [Linum usitatissimum] 68 2e-10
UniRef100_Q9ARB5 N2-A protein [Linum usitatissimum] 68 2e-10
UniRef100_Q9ARB4 N2-B protein [Linum usitatissimum] 68 2e-10
UniRef100_Q8S3I2 NBS-LRR-Toll resistance gene analog protein [Me... 68 2e-10
UniRef100_Q9ARB8 N1-B protein [Linum usitatissimum] 68 3e-10
UniRef100_Q84KB4 MRGH5 [Cucumis melo] 67 4e-10
UniRef100_Q6URA3 Putative TIR-NBS type R protein 4 [Malus baccata] 67 5e-10
UniRef100_Q710T8 TIR/NBS/LRR protein [Populus deltoides] 67 6e-10
UniRef100_Q947E8 Resistance gene analog PU3 [Helianthus annuus] 65 1e-09
UniRef100_Q947E3 Resistance gene analog NBS5 [Helianthus annuus] 65 1e-09
UniRef100_Q8VXR6 RGA-G protein [Cicer arietinum] 65 2e-09
UniRef100_Q6URA1 Putative TIR-NBS type R protein 4 [Malus baccata] 65 2e-09
UniRef100_Q9ARB7 N1-C protein [Linum usitatissimum] 65 2e-09
UniRef100_Q947E2 Resistance gene analog NBS6 [Helianthus annuus] 65 2e-09
UniRef100_Q40254 L6tr [Linum usitatissimum] 64 3e-09
UniRef100_Q9XEH0 Flax rust resistance protein [Linum usitatissimum] 64 3e-09
UniRef100_Q6XZH7 Nematode resistance-like protein [Solanum tuber... 64 3e-09
>UniRef100_Q8S3I1 NBS-LRR-Toll resistance gene analog protein [Medicago sativa]
Length = 170
Score = 69.7 bits (169), Expect = 7e-11
Identities = 30/42 (71%), Positives = 35/42 (82%)
Query: 226 GGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDGPISAQ 267
GG+GK +AT LYNKI HQFPV +IDDL K+YRHDGP+SAQ
Sbjct: 1 GGVGKTTLATVLYNKISHQFPVCSIIDDLSKMYRHDGPVSAQ 42
>UniRef100_Q6UQ08 Putative TIR-NBS type R protein 11 [Malus domestica]
Length = 634
Score = 69.3 bits (168), Expect = 1e-10
Identities = 40/121 (33%), Positives = 60/121 (49%), Gaps = 8/121 (6%)
Query: 141 DSNKTLRWCKDGGKLKHKSPISGWDVR-----HNDAQIEKIVEEIMNILGYKSTSLPNYL 195
D K RW L S +SGWD+R H I I ++ L K + Y
Sbjct: 224 DEKKVERW---RAALTEASNLSGWDLRNTLDGHEAKFIRMITNDVTTKLNNKYFDVAPYQ 280
Query: 196 AGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLR 255
G+D+ ++ +L + DDVRV+G+ GMGGIGK IA A+YN + +F ++ +R
Sbjct: 281 VGIDTRVLDISNYLGIGDSDDVRVIGISGMGGIGKTTIAQAIYNIFYERFEGKSFLEKVR 340
Query: 256 K 256
+
Sbjct: 341 E 341
>UniRef100_Q9AR40 Nbi-B protein [Linum usitatissimum]
Length = 1108
Score = 68.2 bits (165), Expect = 2e-10
Identities = 44/138 (31%), Positives = 71/138 (50%), Gaps = 13/138 (9%)
Query: 137 NMNKDSNKTLRW---CKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTS 190
N+ D L W +D GK+K GW + Q ++KI I L T
Sbjct: 153 NLKHDPETILEWKEALQDVGKMK------GWHINELTGQGAVVDKIFTTIEFHLRANYTL 206
Query: 191 LPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFL 250
+ L G+DS EE+ + + LD R++G+ GMGG+GK +A A++NK+ QF
Sbjct: 207 ATDELVGIDSSVEEVMELMNLDHSTSERIIGIYGMGGLGKTTLAKAVFNKVSMQFERCCF 266
Query: 251 IDDLRK-IYRHDGPISAQ 267
+D++R+ + R+DG ++ Q
Sbjct: 267 LDNIRETLLRNDGVVALQ 284
>UniRef100_Q9ARC2 Ngc-A protein [Linum usitatissimum]
Length = 1075
Score = 68.2 bits (165), Expect = 2e-10
Identities = 42/135 (31%), Positives = 71/135 (52%), Gaps = 8/135 (5%)
Query: 137 NMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTSLPN 193
N+ D L W G L+ + GW + Q ++KI E+ L T +
Sbjct: 153 NLKHDPETILEW---KGALQEVGKMKGWHISELTGQGAVVDKIFTEVELHLRANYTLATD 209
Query: 194 YLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDD 253
L G+D +E+ K L LDS + +++G+ GMG +GK +ATA+YNK+ QF +D+
Sbjct: 210 ELVGIDFSVDEMVKLLNLDSTSE-KIIGIYGMGRLGKTTLATAVYNKVSMQFERCCFLDN 268
Query: 254 LRK-IYRHDGPISAQ 267
+R+ + ++DG ++ Q
Sbjct: 269 IRETLLKNDGVVALQ 283
>UniRef100_Q9ARB5 N2-A protein [Linum usitatissimum]
Length = 1075
Score = 68.2 bits (165), Expect = 2e-10
Identities = 42/135 (31%), Positives = 71/135 (52%), Gaps = 8/135 (5%)
Query: 137 NMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTSLPN 193
N+ D L W G L+ + GW + Q ++KI E+ L T +
Sbjct: 153 NLKHDPETILEW---KGALQEVGKMKGWHISELTGQGAVVDKIFTEVELHLRANYTLATD 209
Query: 194 YLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDD 253
L G+D +E+ K L LDS + +++G+ GMG +GK +ATA+YNK+ QF +D+
Sbjct: 210 ELVGIDFSVDEMVKLLNLDSTSE-KIIGIYGMGRLGKTTLATAVYNKVSMQFERCCFLDN 268
Query: 254 LRK-IYRHDGPISAQ 267
+R+ + ++DG ++ Q
Sbjct: 269 IRETLLKNDGVVALQ 283
>UniRef100_Q9ARB4 N2-B protein [Linum usitatissimum]
Length = 1108
Score = 68.2 bits (165), Expect = 2e-10
Identities = 44/138 (31%), Positives = 71/138 (50%), Gaps = 13/138 (9%)
Query: 137 NMNKDSNKTLRW---CKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTS 190
N+ D L W +D GK+K GW + Q ++KI I L T
Sbjct: 153 NLKHDPETILEWKEALQDVGKMK------GWHINELTGQGAVVDKIFTTIEFHLRANYTL 206
Query: 191 LPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFL 250
+ L G+DS EE+ + + LD R++G+ GMGG+GK +A A++NK+ QF
Sbjct: 207 ATDELVGIDSSVEEVMELMNLDHSTSERIIGIYGMGGLGKTTLAKAVFNKVSMQFERCCF 266
Query: 251 IDDLRK-IYRHDGPISAQ 267
+D++R+ + R+DG ++ Q
Sbjct: 267 LDNIRETLLRNDGVVALQ 284
>UniRef100_Q8S3I2 NBS-LRR-Toll resistance gene analog protein [Medicago sativa]
Length = 170
Score = 68.2 bits (165), Expect = 2e-10
Identities = 31/42 (73%), Positives = 35/42 (82%)
Query: 226 GGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDGPISAQ 267
GG+GK +ATALYNKI HQF V LIDD+ KIYRHDGP+SAQ
Sbjct: 1 GGVGKTTLATALYNKISHQFSVYCLIDDVSKIYRHDGPMSAQ 42
>UniRef100_Q9ARB8 N1-B protein [Linum usitatissimum]
Length = 1108
Score = 67.8 bits (164), Expect = 3e-10
Identities = 44/138 (31%), Positives = 71/138 (50%), Gaps = 13/138 (9%)
Query: 137 NMNKDSNKTLRW---CKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTS 190
NM D L W +D GK+K GW + Q ++KI I L T
Sbjct: 153 NMKHDPETILEWKEALQDVGKMK------GWHINELTGQGAVVDKIFTTIEFHLRANYTL 206
Query: 191 LPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFL 250
+ L G+DS EE+ + + LD R++G+ GMGG+GK +A A++N++ QF
Sbjct: 207 ATDELVGIDSSVEEVMELMNLDHSTSERIIGIYGMGGLGKTTLAKAVFNQVSMQFERCCF 266
Query: 251 IDDLRK-IYRHDGPISAQ 267
+D++R+ + R+DG ++ Q
Sbjct: 267 LDNIRETLLRNDGVVALQ 284
>UniRef100_Q84KB4 MRGH5 [Cucumis melo]
Length = 1092
Score = 67.4 bits (163), Expect = 4e-10
Identities = 60/203 (29%), Positives = 97/203 (47%), Gaps = 40/203 (19%)
Query: 74 RDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLK 133
++SLI IV+FS+NYASST +CL ++ I+ K + +L
Sbjct: 72 QNSLISIVIFSENYASST--------------------WCLDELVEIMECKKSKGQKVLP 111
Query: 134 PFSNMN-KDSNKTLRWCKDG-----GKLKHKSPI-----------SGWDV--RHNDAQIE 174
F ++ D K W ++G K PI SGW + R I+
Sbjct: 112 IFYKVDPSDVRKQNGWFREGLAKHEANFMEKIPIWRDALTTAANLSGWHLGARKEAHLIQ 171
Query: 175 KIVEEIMNILGY-KSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAI 233
IV+E+++IL + K + +L G+DS E L + + + V ++G+ G+GGIGK +
Sbjct: 172 DIVKEVLSILNHTKPLNANEHLVGIDSKIEFLYRKEEMYKSECVNMLGIYGIGGIGKTTL 231
Query: 234 ATALYNKIFHQFPVLFLIDDLRK 256
A ALY+K+ QF + D+R+
Sbjct: 232 AKALYDKMASQFEGCCYLRDVRE 254
>UniRef100_Q6URA3 Putative TIR-NBS type R protein 4 [Malus baccata]
Length = 726
Score = 67.0 bits (162), Expect = 5e-10
Identities = 55/204 (26%), Positives = 88/204 (42%), Gaps = 46/204 (22%)
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPF 135
S I I+VFS+ YA S+ +CL ++ I+ +++L F
Sbjct: 179 SRISIIVFSRRYADSS--------------------WCLEELVKIMECRRTLGQLVLPIF 218
Query: 136 -----SNMNK-------------DSNKTLRWCKDGGKLKHKSPISGWDV-----RHNDAQ 172
SN+ K D K RW L S +SGWD+ RH
Sbjct: 219 YDVDPSNVRKLTGSFAQSFLKHTDEKKVERW---RAALTEASNLSGWDLKNTLDRHEAKF 275
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKA 232
I I ++ L + ++ Y G+D+ + +L + DDVRV+G+ GMGGIGK
Sbjct: 276 IRMITNQVTVKLNNRYFNVAPYQVGIDTRVLNISNYLGIGDSDDVRVIGISGMGGIGKTT 335
Query: 233 IATALYNKIFHQFPVLFLIDDLRK 256
I A+YN+ + +F ++ +R+
Sbjct: 336 IVKAIYNEFYERFEGKSFLEKVRE 359
>UniRef100_Q710T8 TIR/NBS/LRR protein [Populus deltoides]
Length = 1147
Score = 66.6 bits (161), Expect = 6e-10
Identities = 56/187 (29%), Positives = 92/187 (48%), Gaps = 12/187 (6%)
Query: 65 HWIRAPWSNRDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR* 124
H++RA ++S I I VFSK YASS C N K K + + +
Sbjct: 85 HFLRAI---QESKISIAVFSKGYASSRW--CLNELVEILKCKKRKTGQIVLPIFYDIDPS 139
Query: 125 DIKKEIMLKPFSNMNKDSNKTLRWCKDGGK-LKHKSPISGWDVR-----HNDAQIEKIVE 178
D++K+ + + + + K+ K L+ +SGW++ H I++I++
Sbjct: 140 DVRKQNGSFAEAFVKHEERFEEKLVKEWRKALEEAGNLSGWNLNDMANGHEAKFIKEIIK 199
Query: 179 EIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALY 238
++N L K +P +L GMD L + L + DDVR+VG+ GM GIGK IA A++
Sbjct: 200 VVLNKLEPKYLYVPEHLVGMDQLARNIFDFLSA-ATDDVRIVGIHGMPGIGKTTIAQAVF 258
Query: 239 NKIFHQF 245
N++ + F
Sbjct: 259 NQLCYGF 265
>UniRef100_Q947E8 Resistance gene analog PU3 [Helianthus annuus]
Length = 770
Score = 65.5 bits (158), Expect = 1e-09
Identities = 57/203 (28%), Positives = 100/203 (49%), Gaps = 41/203 (20%)
Query: 74 RDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLK 133
++S I +VVFS+NYA S+ +CL + I+ D + +I++
Sbjct: 135 QESRIAVVVFSQNYADSS--------------------WCLDELAHIMECMDTRGQIVIP 174
Query: 134 PF-----SNMNKDSNK---TLRWCKDGGKLKHKS---------PISGWDVRHNDAQ---I 173
F S++ K K R K K K +S +SGW + N + I
Sbjct: 175 IFYFVDPSDVRKQKGKYGKAFRKHKRENKQKVESWRKALEKAGNLSGWVINENSHEAKCI 234
Query: 174 EKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAI 233
++IV I + L ST++ L G+++ ++L+ L ++S DVR++G+ G+GG GK +
Sbjct: 235 KEIVATISSRLPTLSTNVNKDLIGIETRLQDLKSKLKMES-GDVRIIGIWGVGGGGKTTL 293
Query: 234 ATALYNKIFHQFPVLFLIDDLRK 256
A+A Y +I H+F L+ ++R+
Sbjct: 294 ASAAYAEISHRFEAHCLLQNIRE 316
>UniRef100_Q947E3 Resistance gene analog NBS5 [Helianthus annuus]
Length = 285
Score = 65.5 bits (158), Expect = 1e-09
Identities = 56/208 (26%), Positives = 100/208 (47%), Gaps = 41/208 (19%)
Query: 74 RDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLK 133
++S I ++VFS+NYA S+ +CL + I+ D +I++
Sbjct: 99 QESRIAVIVFSENYADSS--------------------WCLDELQQIIECMDTNGQIVIP 138
Query: 134 PFSNM--------NKDSNKTLRWCKDGGKLKHKS---------PISGWDVRHNDAQ---I 173
F ++ N K R K K K +S +SGW + N + I
Sbjct: 139 IFYHVDPSDVRKQNGKYGKAFRKHKKENKQKVESWRKALEKAGNLSGWVINENSHEAKCI 198
Query: 174 EKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAI 233
++IV I + L ST++ L GM++ ++L+ L ++S VR++G+ G+GG GK +
Sbjct: 199 KEIVGTISSRLPTLSTNVNKDLIGMETRLQDLKSKLKMES-GGVRIIGIWGVGGGGKTTL 257
Query: 234 ATALYNKIFHQFPVLFLIDDLRKIYRHD 261
A+A Y +I H+F L++++R+ R +
Sbjct: 258 ASAAYAEISHRFEAHCLLENIREESRKE 285
>UniRef100_Q8VXR6 RGA-G protein [Cicer arietinum]
Length = 169
Score = 65.1 bits (157), Expect = 2e-09
Identities = 30/42 (71%), Positives = 32/42 (75%)
Query: 226 GGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDGPISAQ 267
GG+GK +AT LYNKI HQFPV I DL KIYRHDGPI AQ
Sbjct: 1 GGVGKTTLATVLYNKISHQFPVYCRIADLSKIYRHDGPIGAQ 42
>UniRef100_Q6URA1 Putative TIR-NBS type R protein 4 [Malus baccata]
Length = 726
Score = 64.7 bits (156), Expect = 2e-09
Identities = 54/204 (26%), Positives = 87/204 (42%), Gaps = 46/204 (22%)
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPF 135
S I I+VFS+ YA S+ +CL ++ I+ +++L F
Sbjct: 179 SRISIIVFSRRYADSS--------------------WCLEELVKIMECRRTLGQLVLPIF 218
Query: 136 -----SNMNK-------------DSNKTLRWCKDGGKLKHKSPISGWDV-----RHNDAQ 172
SN+ K D K RW L S +SGWD+ RH
Sbjct: 219 YDVDPSNVRKLTGSFAQSFLKHTDEKKVERW---RAALTEASNLSGWDLKNTLDRHEAKF 275
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKA 232
I I ++ L + ++ Y G+D+ + +L + DDVRV+G+ G GGIGK
Sbjct: 276 IRMITNQVTVKLNNRYFNVAPYQVGIDTRVLNISNYLGIGDSDDVRVIGISGSGGIGKTT 335
Query: 233 IATALYNKIFHQFPVLFLIDDLRK 256
I A+YN+ + +F ++ +R+
Sbjct: 336 IVKAIYNEFYERFEGKSFLEKVRE 359
>UniRef100_Q9ARB7 N1-C protein [Linum usitatissimum]
Length = 1120
Score = 64.7 bits (156), Expect = 2e-09
Identities = 51/193 (26%), Positives = 92/193 (47%), Gaps = 17/193 (8%)
Query: 75 DSLIFIVVFSKNYASST--IGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIK----- 127
+S I+I + + NYASS + + + KG + + L + D++
Sbjct: 84 ESKIYIPILTPNYASSKWCLQELAKMVECWKSGGGAKGQHIILPVFLFVDPRDVRHTESG 143
Query: 128 --KEIMLKPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMN 182
KE + D L W + L+ + G+ V +D I+KI+ E+
Sbjct: 144 SYKEAFEQ--HRQKHDPETVLEWKE---ALQEVGKMKGYHVTESDGHGSIIDKILTEVEL 198
Query: 183 ILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIF 242
LG T + + L G+DSL +E+ L LDS +++G+ GMGG+GK +A A+Y+K+
Sbjct: 199 HLGANYTLVTDELVGIDSLVDEVVGLLNLDSSTSEKIIGIHGMGGLGKTTLAKAVYDKVS 258
Query: 243 HQFPVLFLIDDLR 255
+F + ++++R
Sbjct: 259 TKFERCYFLENIR 271
>UniRef100_Q947E2 Resistance gene analog NBS6 [Helianthus annuus]
Length = 303
Score = 64.7 bits (156), Expect = 2e-09
Identities = 57/203 (28%), Positives = 100/203 (49%), Gaps = 41/203 (20%)
Query: 74 RDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLK 133
++S I +VVFS+NYA S+ +CL + I+ D + +I++
Sbjct: 97 QESRIALVVFSENYADSS--------------------WCLDELAHIMECMDTRGQIVIP 136
Query: 134 PF-----SNMNKDSNK---TLRWCKDGGKLKHKS---------PISGWDVRHNDAQ---I 173
F S++ K K R K K K +S +SGW + N + I
Sbjct: 137 IFYFVDPSDVRKQKGKYGKAFRKHKRENKQKVESWRKALEKAGNLSGWVINENSHEAKRI 196
Query: 174 EKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAI 233
++IV I + L ST++ L G+++ ++L+ L ++S DVR++G+ G+GG GK +
Sbjct: 197 KEIVATISSRLPTLSTNVNKDLIGIETRLQDLKSTLKMES-GDVRIIGIWGVGGGGKTTL 255
Query: 234 ATALYNKIFHQFPVLFLIDDLRK 256
A+A Y +I H+F L+ ++R+
Sbjct: 256 ASAAYAEISHRFEAHCLLQNIRE 278
>UniRef100_Q40254 L6tr [Linum usitatissimum]
Length = 705
Score = 64.3 bits (155), Expect = 3e-09
Identities = 49/134 (36%), Positives = 73/134 (53%), Gaps = 9/134 (6%)
Query: 139 NKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTSLP-NY 194
NK +T++ KD LK + GW + ND Q +K+ +I + + ++ L +
Sbjct: 179 NKFDGQTIQNWKDA--LKKVGDLKGWHIGKNDKQGAIADKVSADIWSHISKENLILETDE 236
Query: 195 LAGMDS-LTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDD 253
L G+D +T LEK L LDS ++V +VG+ GMGGIGK A A+YNKI F ID+
Sbjct: 237 LVGIDDHITAVLEK-LSLDS-ENVTMVGLYGMGGIGKTTTAKAVYNKISSCFDCCCFIDN 294
Query: 254 LRKIYRHDGPISAQ 267
+R+ DG + Q
Sbjct: 295 IRETQEKDGVVVLQ 308
>UniRef100_Q9XEH0 Flax rust resistance protein [Linum usitatissimum]
Length = 1294
Score = 64.3 bits (155), Expect = 3e-09
Identities = 49/134 (36%), Positives = 73/134 (53%), Gaps = 9/134 (6%)
Query: 139 NKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTSLP-NY 194
NK +T++ KD LK + GW + ND Q +K+ +I + + ++ L +
Sbjct: 179 NKFDGQTIQNWKDA--LKKVGDLKGWHIGKNDKQGAIADKVSADIWSHISKENLILETDE 236
Query: 195 LAGMDS-LTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDD 253
L G+D +T LEK L LDS ++V +VG+ GMGGIGK A A+YNKI F ID+
Sbjct: 237 LVGIDDHITAVLEK-LSLDS-ENVTMVGLYGMGGIGKTTTAKAVYNKISSCFDCCCFIDN 294
Query: 254 LRKIYRHDGPISAQ 267
+R+ DG + Q
Sbjct: 295 IRETQEKDGVVVLQ 308
>UniRef100_Q6XZH7 Nematode resistance-like protein [Solanum tuberosum]
Length = 1121
Score = 64.3 bits (155), Expect = 3e-09
Identities = 59/195 (30%), Positives = 91/195 (46%), Gaps = 19/195 (9%)
Query: 72 SNRDSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKG-----MFCLFSMMLILMR*DI 126
S +S I +++FSKNYA+ST C + K+ KG +F + + +I
Sbjct: 68 SIEESRIALIIFSKNYANSTW--CLDELTKIIECKNVKGQIVVPVFYDVDPSTVRRQKNI 125
Query: 127 KKEIMLKPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVR-----HNDAQIEKIVEEIM 181
E K + +D K R L+ + ISGWD+ H IEKI E+IM
Sbjct: 126 FGEAFSKHEARFEEDKVKKWR-----AALEEAANISGWDLPNTSNGHEARVIEKITEDIM 180
Query: 182 NILG-YKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNK 240
LG + S + GM+S ++ K L + S VR +G+ GM G+GK +A +Y+
Sbjct: 181 VRLGSQRHASNARNVVGMESHMHQVYKMLGIGS-GGVRFLGILGMSGVGKTTLARVIYDN 239
Query: 241 IFHQFPVLFLIDDLR 255
I QF + ++R
Sbjct: 240 IQSQFEGACFLHEVR 254
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.339 0.149 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 412,278,775
Number of Sequences: 2790947
Number of extensions: 16117944
Number of successful extensions: 70095
Number of sequences better than 10.0: 514
Number of HSP's better than 10.0 without gapping: 391
Number of HSP's successfully gapped in prelim test: 123
Number of HSP's that attempted gapping in prelim test: 69395
Number of HSP's gapped (non-prelim): 697
length of query: 267
length of database: 848,049,833
effective HSP length: 125
effective length of query: 142
effective length of database: 499,181,458
effective search space: 70883767036
effective search space used: 70883767036
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC137667.13


