

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.6 - phase: 0
(456 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
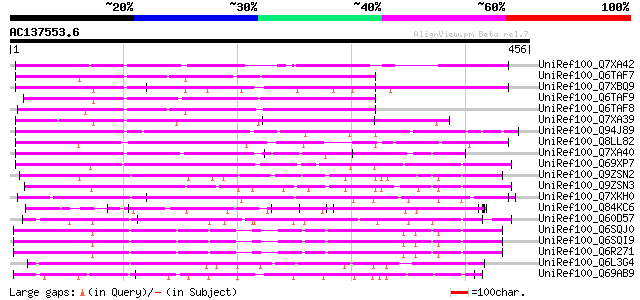
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XA42 Putative disease resistance protein RGA1 [Solan... 172 2e-41
UniRef100_Q6TAF7 Blight resistance protein T118 [Solanum tarijense] 159 2e-37
UniRef100_Q7XBQ9 Disease resistance protein RGA2 [Solanum bulboc... 157 8e-37
UniRef100_Q6TAF9 Blight resistance protein SH10 [Solanum tuberosum] 156 1e-36
UniRef100_Q6TAF8 Blight resistance protein SH20 [Solanum tuberosum] 152 2e-35
UniRef100_Q7XA39 Putative disease resistance protein RGA4 [Solan... 151 3e-35
UniRef100_Q94J89 Putative NBS-LRR type resistance protein [Oryza... 141 3e-32
UniRef100_Q8LL82 NBS-LRR-like protein [Oryza sativa] 138 3e-31
UniRef100_Q7XA40 Putative disease resistance protein RGA3 [Solan... 136 1e-30
UniRef100_Q69XP7 Putative disease resistance protein [Oryza sativa] 130 1e-28
UniRef100_Q9ZSN2 NBS-LRR-like protein cD8 [Phaseolus vulgaris] 127 9e-28
UniRef100_Q9ZSN3 NBS-LRR-like protein cD7 [Phaseolus vulgaris] 125 2e-27
UniRef100_Q7XKH0 OSJNBa0083D01.14 protein [Oryza sativa] 124 6e-27
UniRef100_Q84KC6 NBS-LRR disease resistance protein homologue [H... 124 6e-27
UniRef100_Q60D57 Putative disease resistance protein I2C-5 [Sola... 122 2e-26
UniRef100_Q6SQJ0 NBS-LRR type disease resistance protein Hom-F [... 121 5e-26
UniRef100_Q6SQI9 NBS-LRR type disease resistance protein Hom-B [... 120 1e-25
UniRef100_Q6R271 Disease resistance protein [Glycine max] 120 1e-25
UniRef100_Q6L3G4 Putative plant disease resistant protein [Solan... 115 2e-24
UniRef100_Q69AB9 CC-NBS-LRR [Helianthus annuus] 115 2e-24
>UniRef100_Q7XA42 Putative disease resistance protein RGA1 [Solanum bulbocastanum]
Length = 1025
Score = 172 bits (435), Expect = 2e-41
Identities = 128/435 (29%), Positives = 201/435 (45%), Gaps = 86/435 (19%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F + G L +L NL ++ I L ++ D +E NLS+ L+ L L+WD +
Sbjct: 673 FVIGKRKGHQLGELKNLNLYGSISITKLDRVKKDTDAKEANLSAKANLHSLCLSWDLDGK 732
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+E VL AL+PH +L ++G+ G+ +P+WM S+L +V +++ C NCS LPP
Sbjct: 733 HRYDSE-VLEALKPHSNLKYLEINGFGGIRLPDWMNQ-SVLKNVVSIRIRGCENCSCLPP 790
Query: 125 LGKLPFLNTLYL-SQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L + +V+Y++D+ + FPSL ++ ++D NL+ +L++EG +
Sbjct: 791 FGELPCLESLELHTGSADVEYVEDNVHP----GRFPSLRKLVIWDFSNLKGLLKMEGEKQ 846
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L +++ P F +P+L SVK LK ++ DA
Sbjct: 847 FPVLEEMTFYWCPMFVIPTLSSVK---------------------------TLKVIVTDA 879
Query: 244 FHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSV 303
TVL +S+LR+L L I D + S+P +F L +L+ L +L LP S+
Sbjct: 880 ----TVL-RSISNLRALTSLDISDNVEATSLPEEMFKSLANLKYLKISFFRNLKELPTSL 934
Query: 304 TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRD 363
+L +L+ L +C + L SL E + G
Sbjct: 935 ASLNALKSLKFEFC---------DALESLPEEGVKG------------------------ 961
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
SL EL +S L LP+ L L L I +CP + R R
Sbjct: 962 -------------LTSLTELSVSNCMMLKCLPEGLQHLTALTTLTITQCPIVFKRCERGI 1008
Query: 424 GEDWYKIAHVPILSL 438
GEDW+KIAH+P L+L
Sbjct: 1009 GEDWHKIAHIPYLTL 1023
Score = 42.0 bits (97), Expect = 0.036
Identities = 40/153 (26%), Positives = 70/153 (45%), Gaps = 7/153 (4%)
Query: 247 LTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTL 306
L LP+ + L L L + ++ ++P + L +L+ L C SL+ LP+ + L
Sbjct: 583 LNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCK-LQNLQTLDLHYCDSLSCLPKQTSKL 641
Query: 307 TSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPS 366
SL+ L++ C P + +L L+ +S +R+G + L+NL+L S
Sbjct: 642 GSLRNLLLDGCSLTSTPPRIGLLTCLKSLSCFVIGKRKG-----HQLGELKNLNLYGSIS 696
Query: 367 LRSLPDWLGDTLSLQELEISKFPKLTSLPDNFD 399
+ L DT +E +S L SL ++D
Sbjct: 697 ITKLDRVKKDT-DAKEANLSAKANLHSLCLSWD 728
Score = 35.4 bits (80), Expect = 3.4
Identities = 22/66 (33%), Positives = 38/66 (57%), Gaps = 1/66 (1%)
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
L+ L+ L+LR+ +L LP +GD + L+ L++S ++ +LP +L+NLQ L +
Sbjct: 567 LQKFVSLRVLNLRN-SNLNQLPSSIGDLVHLRYLDLSGNFRIRNLPKRLCKLQNLQTLDL 625
Query: 410 DRCPRL 415
C L
Sbjct: 626 HYCDSL 631
>UniRef100_Q6TAF7 Blight resistance protein T118 [Solanum tarijense]
Length = 948
Score = 159 bits (401), Expect = 2e-37
Identities = 108/325 (33%), Positives = 171/325 (52%), Gaps = 15/325 (4%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWD---R 61
F V+ G+ L +L +L ++ I L+ ++N + +E NLS+ L+ L + WD R
Sbjct: 627 FVVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDER 686
Query: 62 NTNSTNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQ 121
+ EVL AL+PH +LT +SG+RG+ +P+WM S+L +V +++ C NCS
Sbjct: 687 PHRYESEEVEVLEALKPHSNLTCLTISGFRGIRLPDWMNH-SVLKNIVLIEISGCKNCSC 745
Query: 122 LPPLGKLPFLNTLYLSQMTNVKYIDDSPYEIS-----TENAFPSLTEMTLFDLPNLERVL 176
LPP G LP L +L L + + +Y+++ ++ T FPSL ++ + NL+ ++
Sbjct: 746 LPPFGDLPCLESLQLYR-GSAEYVEEVDIDVEDSGFPTRIRFPSLRKLCICKFDNLKGLV 804
Query: 177 RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNL 236
+ EG E L ++ I+ P +P+L S + ++ + SF ++ + NL
Sbjct: 805 KKEGGEQFPVLEEMEIRYCP---IPTLSSNLKALTSLNISDNKE-ATSFPEEMFKSLANL 860
Query: 237 KELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSL 296
K L I F L LP L+SL +L+ L I C LESIP GL SL L C L
Sbjct: 861 KYLNISHFKNLKELPTSLASLNALKSLKIQWCCALESIPEEGVKGLTSLTELIVKFCKML 920
Query: 297 NSLPQSVTTLTSLQRLIIHYCPELI 321
LP+ + LT+L R+ I CP+LI
Sbjct: 921 KCLPEGLQHLTALTRVKIWGCPQLI 945
Score = 37.7 bits (86), Expect = 0.69
Identities = 29/88 (32%), Positives = 44/88 (49%), Gaps = 6/88 (6%)
Query: 256 SLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIH 315
SLR L Y +K E +P+++ L+ LR + + SLP+ + L +LQ L +
Sbjct: 525 SLRVLNLSY----SKFEELPSSIG-DLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQ 579
Query: 316 YCPEL-ILPANMNMLNSLREVSIMGGDR 342
YC L LP + L SLR + + G R
Sbjct: 580 YCTRLCCLPKQTSKLGSLRNLLLHGCHR 607
Score = 36.6 bits (83), Expect = 1.5
Identities = 21/58 (36%), Positives = 32/58 (54%), Gaps = 3/58 (5%)
Query: 358 NLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
NLS F LP +GD + L+ +++S ++ SLP +L+NLQ L + C RL
Sbjct: 530 NLSYSKF---EELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRL 584
Score = 34.7 bits (78), Expect = 5.8
Identities = 47/193 (24%), Positives = 83/193 (42%), Gaps = 21/193 (10%)
Query: 232 KMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRIL-SF 290
K+ NL+ L + +L LP + S L SL L + C++L P + L L+ L F
Sbjct: 569 KLQNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRI-GSLTCLKTLGQF 627
Query: 291 VICHSLNSLPQSVTTLTSLQRLIIHYCPEL-----ILPANMNMLNSLREVSIMGGDRRRG 345
V+ + +L + I + + AN++ +L +S+ D R
Sbjct: 628 VVKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDERP 687
Query: 346 IYNGLEDIPLLQN---------LSLRDFPSLRSLPDWLGDTL--SLQELEISKFPKLTSL 394
E++ +L+ L++ F +R LPDW+ ++ ++ +EIS + L
Sbjct: 688 HRYESEEVEVLEALKPHSNLTCLTISGFRGIR-LPDWMNHSVLKNIVLIEISGCKNCSCL 746
Query: 395 PDNFDQ--LENLQ 405
P D LE+LQ
Sbjct: 747 PPFGDLPCLESLQ 759
Score = 34.3 bits (77), Expect = 7.6
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Query: 231 GKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF 290
G + +L+ + + ++ LP +L L++L+ L + C +L +P L SLR L
Sbjct: 544 GDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRLCCLPKQT-SKLGSLRNLLL 602
Query: 291 VICHSLNSLPQSVTTLTSLQRL 312
CH L P + +LT L+ L
Sbjct: 603 HGCHRLTRTPPRIGSLTCLKTL 624
>UniRef100_Q7XBQ9 Disease resistance protein RGA2 [Solanum bulbocastanum]
Length = 970
Score = 157 bits (396), Expect = 8e-37
Identities = 111/328 (33%), Positives = 170/328 (50%), Gaps = 21/328 (6%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D +E NLS+ L+ L ++W+
Sbjct: 628 FVVGRKKGYQLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGP 687
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+EEV L AL+PH +LT ++ G+RG+++P WM S+L +V + + N NCS L
Sbjct: 688 HIYESEEVKVLEALKPHSNLTSLKIYGFRGIHLPEWMNH-SVLKNIVSILISNFRNCSCL 746
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYIDDSPYEIS----TENAFPSLTEMTLFDLPNLERVLR 177
PP G LP L +L L +V+Y+++ ++ T FPSL ++ ++D +L+ +L+
Sbjct: 747 PPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRKLDIWDFGSLKGLLK 806
Query: 178 IEGVEMLSQLSKLSIQSIPQFELPS----LPSVKEVYVGGETEEDIDHEASFLRDIAGKM 233
EG E L ++ I P L S L S++ Y T SF ++ +
Sbjct: 807 KEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVAT--------SFPEEMFKNL 858
Query: 234 PNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
NLK L I + L LP L+SL +L+ L I C LES+P GL SL L C
Sbjct: 859 ANLKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLPEEGLEGLSSLTELFVEHC 918
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPELI 321
+ L LP+ + LT+L L I CP+LI
Sbjct: 919 NMLKCLPEGLQHLTTLTSLKIRGCPQLI 946
Score = 72.0 bits (175), Expect = 3e-11
Identities = 90/356 (25%), Positives = 144/356 (40%), Gaps = 62/356 (17%)
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPY-EISTENAFPSLTEMTLFDLPNL---ERVL 176
QL LG L ++ +S + VK D+ +S + SL+ P++ E V
Sbjct: 637 QLGELGNLNLYGSIKISHLERVKNDKDAKEANLSAKGNLHSLSMSWNNFGPHIYESEEVK 696
Query: 177 RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNL 236
+E ++ S L+ L I LP E ++H + N+
Sbjct: 697 VLEALKPHSNLTSLKIYGFRGIHLP---------------EWMNHSV---------LKNI 732
Query: 237 KELMIDAFHQLTVLP--NELSSLRSLEELY-IIDCNKLESIPNNVFYGLI------SLRI 287
++I F + LP +L L SLE + D +E + +V G SLR
Sbjct: 733 VSILISNFRNCSCLPPFGDLPCLESLELHWGSADVEYVEEVDIDVHSGFPTRIRFPSLRK 792
Query: 288 LSFVICHSLNSL--PQSVTTLTSLQRLIIHYCPELILPANMNMLNSLR------------ 333
L SL L + L+ +IIH CP L L +N+ L SLR
Sbjct: 793 LDIWDFGSLKGLLKKEGEEQFPVLEEMIIHECPFLTLSSNLRALTSLRICYNKVATSFPE 852
Query: 334 ----------EVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPD-WLGDTLSLQE 382
++I + + + L + L++L ++ +L SLP+ L SL E
Sbjct: 853 EMFKNLANLKYLTISRCNNLKELPTSLASLNALKSLKIQLCCALESLPEEGLEGLSSLTE 912
Query: 383 LEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
L + L LP+ L L L I CP+L+ R + GEDW+KI+H+P +++
Sbjct: 913 LFVEHCNMLKCLPEGLQHLTTLTSLKIRGCPQLIKRCEKGIGEDWHKISHIPNVNI 968
>UniRef100_Q6TAF9 Blight resistance protein SH10 [Solanum tuberosum]
Length = 948
Score = 156 bits (395), Expect = 1e-36
Identities = 105/313 (33%), Positives = 158/313 (49%), Gaps = 9/313 (2%)
Query: 13 GFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTNSTNSAEE 71
G+ L L ++ ++ I L+ ++N D +E NLS+ L+ L + W R +EE
Sbjct: 638 GYQLGKLRDVNLYGSIEITHLERVKNVMDAKEANLSAKGNLHSLIMNWSRKGPHIYESEE 697
Query: 72 V--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLGKLP 129
V + AL+PH +LT +SG+RG P WM S+L +V +++ C NCS LPP G+LP
Sbjct: 698 VRVIEALKPHPNLTCLTISGFRGFRFPEWMNH-SVLKNVVSIEISGCKNCSCLPPFGELP 756
Query: 130 FLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQLS 188
L L L + V+Y+D T FPSL ++ + + PNL+ +L+ EG E L
Sbjct: 757 CLKRLELQKGSAEVEYVDSG---FPTRRRFPSLRKLFIGEFPNLKGLLKKEGEEKFPVLE 813
Query: 189 KLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLT 248
+++I F +L S + + S +I NLK L I F+ L
Sbjct: 814 RMTIFYCHMFVYTTLSSNFRALTSLHISHN-NEATSLPEEIFKSFANLKYLKISLFYNLK 872
Query: 249 VLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTS 308
LP+ L+ L +L+ L I C+ LES+P GL SL L C L LP+ + LT+
Sbjct: 873 ELPSSLACLNALKTLEIHSCSALESLPEEGVKGLTSLTELFVYDCEMLKFLPEGLQHLTA 932
Query: 309 LQRLIIHYCPELI 321
L L + CP+LI
Sbjct: 933 LTSLKLRRCPQLI 945
Score = 40.0 bits (92), Expect = 0.14
Identities = 46/193 (23%), Positives = 86/193 (43%), Gaps = 20/193 (10%)
Query: 232 KMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFV 291
K+ NL+ L + + L+ LP E S L SL L+ C++L S+P + L L+ L ++
Sbjct: 572 KLQNLQTLDLHNCYSLSCLPKEPSKLGSLRNLFFHGCDELNSMPPRI-GSLTFLKTLKWI 630
Query: 292 IC------HSLNSLPQ-------SVTTLTSLQRLIIHYCPELILPANMN--MLNSLREVS 336
C + L L +T L ++ ++ L N++ ++N R+
Sbjct: 631 CCGIQKKGYQLGKLRDVNLYGSIEITHLERVKNVMDAKEANLSAKGNLHSLIMNWSRKGP 690
Query: 337 IMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTL--SLQELEISKFPKLTSL 394
+ + L+ P L L++ F R P+W+ ++ ++ +EIS + L
Sbjct: 691 HIYESEEVRVIEALKPHPNLTCLTISGFRGFR-FPEWMNHSVLKNVVSIEISGCKNCSCL 749
Query: 395 PDNFDQLENLQKL 407
P F +L L++L
Sbjct: 750 PP-FGELPCLKRL 761
>UniRef100_Q6TAF8 Blight resistance protein SH20 [Solanum tuberosum]
Length = 947
Score = 152 bits (384), Expect = 2e-35
Identities = 106/323 (32%), Positives = 170/323 (51%), Gaps = 16/323 (4%)
Query: 8 VRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNTNS- 65
V+ G+ L +L +L ++ I L+ ++N + +E NLS+ L+ L + WD + +
Sbjct: 629 VKRKKGYQLGELGSLNLYGSIKISHLERVKNDKEAKEANLSAKENLHSLSMKWDDDEHPH 688
Query: 66 --TNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLP 123
+ EVL AL+PH +LT ++SG+RG+ +P+WM S+L +V +++ C NCS LP
Sbjct: 689 RYESEEVEVLEALKPHSNLTCLKISGFRGIRLPDWMNH-SVLKNIVLIEISGCKNCSCLP 747
Query: 124 PLGKLPFLNTLYLSQMTNVKYIDDSPYEIS----TENAFPSLTEMTLFDLPNLERVLRIE 179
P G LP L +L L + + +Y+++ ++ T PSL ++ + NL+ +L+ E
Sbjct: 748 PFGDLPCLESLELYR-GSAEYVEEVDIDVDSGFPTRIRLPSLRKLCICKFDNLKGLLKKE 806
Query: 180 GVEMLSQLSKLSIQSIPQFEL-PSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKE 238
G E L ++ I+ P L P+L ++ + + E SF ++ + NLK
Sbjct: 807 GGEQFPVLEEMEIRYCPIPTLSPNLKALTSLNISDNKEA-----TSFPEEMFKSLANLKY 861
Query: 239 LMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNS 298
L I F L LP L+SL +L+ L I C LE+IP GL SL L L
Sbjct: 862 LNISHFKNLKELPTSLASLNALKSLKIQWCCALENIPKEGVKGLTSLTELIVKFSKVLKC 921
Query: 299 LPQSVTTLTSLQRLIIHYCPELI 321
LP+ + LT+L RL I CP+LI
Sbjct: 922 LPEGLHHLTALTRLKIWGCPQLI 944
Score = 38.1 bits (87), Expect = 0.52
Identities = 31/113 (27%), Positives = 51/113 (44%), Gaps = 11/113 (9%)
Query: 313 IIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLR---- 368
+IH +L AN + N +RE+++ E + L+ F SLR
Sbjct: 473 LIHDLAXSLLSANTSSSN-IREINVESYTHMMMSIGFSEVVSSYSPSLLQKFVSLRVLNL 531
Query: 369 ------SLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
LP +GD + L+ +++S ++ SLP +L+NLQ L + C RL
Sbjct: 532 SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCKLQNLQTLDLQYCTRL 584
Score = 35.8 bits (81), Expect = 2.6
Identities = 30/128 (23%), Positives = 60/128 (46%), Gaps = 3/128 (2%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ + LP+ + L L + + + ++ S+P + L
Sbjct: 513 SSYSPSLLQKFVSLRVLNL-SYSKFEELPSSIGDLVHLRYMDLSNNIEIRSLPKQLCK-L 570
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELI-LPANMNMLNSLREVSIMGGD 341
+L+ L C L LP+ + L SL+ L++H C L P + L L+ +
Sbjct: 571 QNLQTLDLQYCTRLCCLPKQTSKLGSLRNLLLHGCHRLTRTPPRIGSLTCLKTLGQSVVK 630
Query: 342 RRRGIYNG 349
R++G G
Sbjct: 631 RKKGYQLG 638
>UniRef100_Q7XA39 Putative disease resistance protein RGA4 [Solanum bulbocastanum]
Length = 988
Score = 151 bits (382), Expect = 3e-35
Identities = 107/339 (31%), Positives = 169/339 (49%), Gaps = 26/339 (7%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL ++ I L+ ++N D E NLS+ L L ++WD +
Sbjct: 628 FIVGSKKGYQLGELKNLNLCGSISITHLERVKNDTDA-EANLSAKANLQSLSMSWDNDGP 686
Query: 65 STNSAEEV--LGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
+ ++EV L AL+PH +L + + G P+W+ S+L +++ V++ +C NC L
Sbjct: 687 NRYESKEVKVLEALKPHPNLKYLEIIAFGGFRFPSWINH-SVLEKVISVRIKSCKNCLCL 745
Query: 123 PPLGKLPFLNTLYLSQ-MTNVKYI--DDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
PP G+LP L L L V+Y+ DD ST +FPSL ++ ++ +L+ +++ E
Sbjct: 746 PPFGELPCLENLELQNGSAEVEYVEEDDVHSRFSTRRSFPSLKKLRIWFFRSLKGLMKEE 805
Query: 180 GVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEE-----------------DIDHE 222
G E L +++I P F P+L SVK++ V G T ++
Sbjct: 806 GEEKFPMLEEMAILYCPLFVFPTLSSVKKLEVHGNTNTRGLSSISNLSTLTSLRIGANYR 865
Query: 223 ASFL-RDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYG 281
A+ L ++ + NL+ L F L LP L+SL +L+ L I C+ LES P G
Sbjct: 866 ATSLPEEMFTSLTNLEFLSFFDFKNLKDLPTSLTSLNALKRLQIESCDSLESFPEQGLEG 925
Query: 282 LISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL 320
L SL L C L LP+ + LT+L L + CPE+
Sbjct: 926 LTSLTQLFVKYCKMLKCLPEGLQHLTALTNLGVSGCPEV 964
Score = 52.4 bits (124), Expect = 3e-05
Identities = 43/168 (25%), Positives = 83/168 (48%), Gaps = 7/168 (4%)
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
+S+ + K +L+ L + ++ +L LP+ + L L L + CN S+P + L
Sbjct: 516 SSYSPSLLKKFVSLRVLNL-SYSKLEQLPSSIGDLLHLRYLDL-SCNNFRSLPERLCK-L 572
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDR 342
+L+ L C+SLN LP+ + L+SL+ L++ CP P + +L L+ +
Sbjct: 573 QNLQTLDVHNCYSLNCLPKQTSKLSSLRHLVVDGCPLTSTPPRIGLLTCLKTLGFFIVGS 632
Query: 343 RRGIYNG-LEDIPLLQNLSLRDFPSLRSLPDW---LGDTLSLQELEIS 386
++G G L+++ L ++S+ +++ D L +LQ L +S
Sbjct: 633 KKGYQLGELKNLNLCGSISITHLERVKNDTDAEANLSAKANLQSLSMS 680
Score = 34.7 bits (78), Expect = 5.8
Identities = 22/62 (35%), Positives = 34/62 (54%), Gaps = 2/62 (3%)
Query: 364 FPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRLARRT 423
+ L LP +GD L L+ L++S SLP+ +L+NLQ L + C L N L ++T
Sbjct: 536 YSKLEQLPSSIGDLLHLRYLDLS-CNNFRSLPERLCKLQNLQTLDVHNCYSL-NCLPKQT 593
Query: 424 GE 425
+
Sbjct: 594 SK 595
>UniRef100_Q94J89 Putative NBS-LRR type resistance protein [Oryza sativa]
Length = 1110
Score = 141 bits (356), Expect = 3e-32
Identities = 135/454 (29%), Positives = 213/454 (46%), Gaps = 25/454 (5%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNT 63
F V+ +G ++ +L N+ L I+GL N+ NG D L + L LHL WD +
Sbjct: 667 FVVQKRSGHNVTELNNMDELQGQLSIRGLNNVPNGQDAVCAKLRNKEHLRTLHLIWDEDC 726
Query: 64 NSTNSAE-EVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQL 122
S S + EVL L+PH DL + G+ G+ P+W+ S L +L + + NC ++L
Sbjct: 727 ESNPSEQQEVLEGLQPHLDLKELVIKGFPGVRFPSWLAS-SFLPKLQTIHICNC-RSTRL 784
Query: 123 PPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVE 182
P LG+LPFL L ++ +T V + FP+L ++ L D+PNL + +
Sbjct: 785 PALGQLPFLKYLVIAGVTEVTQLSSEFTGFGQPKGFPALEDLLLEDMPNLSEWIFDVADQ 844
Query: 183 MLSQLSKLSIQSIPQF-ELPSLPS-VKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELM 240
+ QL++L + PQ +LP +PS ++ +++ +E ++ + P L
Sbjct: 845 LFPQLTELGLIKCPQLKKLPPIPSTLRTLWI---SESGLESLPELQNNSCPSSPT--SLY 899
Query: 241 IDAFHQLTVLPNELSSLR--SLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLN- 297
I+ LT L L + R +L+ L I C L S+P F LISLR L C L
Sbjct: 900 INDCPNLTSLRVGLLAYRPTALKSLTIAHCEGLVSLPEECFRPLISLRSLHIYECPCLVP 959
Query: 298 -SLPQSVTTLTSLQRLIIHYCPEL--ILPANMNMLNSLREVSIMG-GDRRRGIYNGLEDI 353
+ + TS++ + ++ C L +L ++ L LR I D GL
Sbjct: 960 WTALEGGLLPTSIEDIRLNSCTPLASVLLNGLSYLPHLRHFEIADCPDINNFPAEGLPH- 1018
Query: 354 PLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCP 413
LQ L + L+ LP L + SL+ L IS P + SLP + L +L I CP
Sbjct: 1019 -TLQFLEISCCDDLQCLPPGLHNISSLETLRISNCPGVESLPKEGLPM-GLNELYIKGCP 1076
Query: 414 RLVNRLARRTGEDWYKIAHVPILSLRLESDVVHP 447
+ + + + GE KIAH I + ++ DV+ P
Sbjct: 1077 Q-IKQQCQEGGEYHAKIAH--IRDIEIDGDVIVP 1107
Score = 37.4 bits (85), Expect = 0.89
Identities = 43/158 (27%), Positives = 71/158 (44%), Gaps = 7/158 (4%)
Query: 252 NELSSLRSLEELYIIDCNK--LESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSL 309
N L R L L II K + +P+ +F L LR+L + L LP+S+ L L
Sbjct: 534 NPLYGFRKLRTLTIIHGYKSRMSQLPHGLFMKLEYLRVLD-MHGQGLKELPESIGNLKQL 592
Query: 310 QRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRS 369
+ L + LPA++ L +L+ + + + R + G+ + L++L L S
Sbjct: 593 RFLDLSSTEIETLPASLVKLYNLQILKLSDCNFLREVPQGITRLINLRHLEAS--TRLLS 650
Query: 370 LPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKL 407
+G + LQELE +F N +L N+ +L
Sbjct: 651 RIHGIGSLVCLQELE--EFVVQKRSGHNVTELNNMDEL 686
>UniRef100_Q8LL82 NBS-LRR-like protein [Oryza sativa]
Length = 1108
Score = 138 bits (348), Expect = 3e-31
Identities = 134/475 (28%), Positives = 201/475 (42%), Gaps = 85/475 (17%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAW---- 59
F V + G+ +++L + G ++ IK L+++ + + E LS ++ L L W
Sbjct: 661 FVVHKDKGYKVSELKAMNKIGGHICIKNLESVSSAEEADEALLSEKAHISILDLIWSSSR 720
Query: 60 DRNTNSTNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINC 119
D + N E L +L PH +L + + G P+W IL L + L +C NC
Sbjct: 721 DFTSEEANQDIETLTSLEPHDELKELTVKAFAGFEFPHW-----ILSHLQTIHLSDCTNC 775
Query: 120 SQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIE 179
S LP LG+LP L + + + I D S FPSL E+ D PNLER +
Sbjct: 776 SILPALGQLPLLKVIIIGGFPTIIKIGDEFSGSSEVKGFPSLKELVFEDTPNLERWTSTQ 835
Query: 180 GVEMLSQLSKLSIQSIPQF-ELPSLPSV---KEVYVGGETEEDIDHEASFLRDIAGKMPN 235
E L L +L + P+ ELP LPS ++ G + H FL P+
Sbjct: 836 DGEFLPFLRELQVLDCPKVTELPLLPSTLVELKISEAGFSVLPEVHAPRFL-------PS 888
Query: 236 LKELMIDAFHQLTVLPNELSS--LRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
L L I LT L L S L +L++L I +C +L P
Sbjct: 889 LTRLQIHKCPNLTSLQQGLLSQQLSALQQLTITNCPELIHPPT----------------- 931
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPEL-------ILPANMNMLNSLREVSIMGGDRRRGI 346
+ + TLT+LQ L I+ CP L +LP M+ LR S + +
Sbjct: 932 -------EGLRTLTALQSLHIYDCPRLATAEHRGLLP---RMIEDLRITSC--SNIINPL 979
Query: 347 YNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLE---- 402
+ L ++ L+NL + D SL + P+ L T L++LEI L SLP +
Sbjct: 980 LDELNELFALKNLVIADCVSLNTFPEKLPAT--LKKLEIFNCSNLASLPACLQEASCLKT 1037
Query: 403 -------------------NLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSL 438
+L++L I CP L R +GEDW KI+H+ I+ +
Sbjct: 1038 MTILNCVSIKCLPAHGLPLSLEELYIKECPFLAERCQENSGEDWPKISHIAIIEI 1092
>UniRef100_Q7XA40 Putative disease resistance protein RGA3 [Solanum bulbocastanum]
Length = 947
Score = 136 bits (343), Expect = 1e-30
Identities = 94/300 (31%), Positives = 154/300 (51%), Gaps = 16/300 (5%)
Query: 6 FTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTN 64
F V G+ L +L NL + I L+ ++N + +E NLS+ L+ L ++WDR
Sbjct: 636 FVVGERKGYQLGELRNLNLRGAISITHLERVKNDMEAKEANLSAKANLHSLSMSWDRPNR 695
Query: 65 STNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
+ +VL AL+PH +L + + G +P+WM S+L +V + + C NCS LPP
Sbjct: 696 YESEEVKVLEALKPHPNLKYLEIIDFCGFCLPDWMNH-SVLKNVVSILISGCENCSCLPP 754
Query: 125 LGKLPFLNTLYLSQ-MTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEM 183
G+LP L +L L V+Y++DS + T FPSL ++ + NL+ + R++G E
Sbjct: 755 FGELPCLESLELQDGSVEVEYVEDSGF--LTRRRFPSLRKLHIGGFCNLKGLQRMKGAEQ 812
Query: 184 LSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDA 243
L ++ I P F P+L SVK++ + GE +A L I+ + L L I +
Sbjct: 813 FPVLEEMKISDCPMFVFPTLSSVKKLEIWGEA------DAGGLSSIS-NLSTLTSLKIFS 865
Query: 244 FHQLTVLPNELSSLRSLEELYIIDCNKLESIPN--NVFYGLISLRILSFVICHSLNSLPQ 301
H +T L E+ ++LE L + + LE++ L +L+ L C++L SLP+
Sbjct: 866 NHTVTSLLEEM--FKNLENLIYLSVSFLENLKELPTSLASLNNLKCLDIRYCYALESLPE 923
Score = 52.8 bits (125), Expect = 2e-05
Identities = 48/176 (27%), Positives = 85/176 (48%), Gaps = 14/176 (7%)
Query: 225 FLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLIS 284
F R ++ ++ NL F QL +L LR L+ + NK+ S+P + L +
Sbjct: 531 FKRFVSLRVLNLSN---SEFEQLPSSVGDLVHLRYLD----LSGNKICSLPKRLCK-LQN 582
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
L+ L C SL+ LP+ + L SL+ L++ +CP +P + +L L+ + R+
Sbjct: 583 LQTLDLYNCQSLSCLPKQTSKLCSLRNLVLDHCPLTSMPPRIGLLTCLKTLGYFVVGERK 642
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQ 400
G G L+NL+LR S+ L + + + + +E +S L SL ++D+
Sbjct: 643 GYQLG-----ELRNLNLRGAISITHL-ERVKNDMEAKEANLSAKANLHSLSMSWDR 692
>UniRef100_Q69XP7 Putative disease resistance protein [Oryza sativa]
Length = 1171
Score = 130 bits (326), Expect = 1e-28
Identities = 135/478 (28%), Positives = 206/478 (42%), Gaps = 52/478 (10%)
Query: 6 FTVRPNAGFSLADLPNLQP-GDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNT 63
F V+ +G L +L + L ++ L+N+ + + +L + L L W
Sbjct: 691 FHVKKESGHRLGELKEMNNIRGRLSVRFLENVEHQQQAVDAHLDCKEHVKHLQLEWSDLP 750
Query: 64 NSTNSA--EEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQ 121
S +VL ALRPH DL ++GY+G+ P W + + + L V L NC+ Q
Sbjct: 751 RPITSELDSDVLEALRPHPDLDRLNITGYKGLRSPTWF-ETNWMKALTSVILENCMGWVQ 809
Query: 122 LPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGV 181
LPPLG+LP L L L M V I + Y FP L E+ +PN E+ IE
Sbjct: 810 LPPLGQLPLLEDLVLRNMHAVGQIGEEFYGNGEMKGFPKLEEIVFDGMPNWEKWSGIEDG 869
Query: 182 EMLSQLSKLSIQSIPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELM 240
+L L++L I P+ E P L + +V V ++ +S L D + L+
Sbjct: 870 SLLPCLTRLYIAKCPKLQEAPPLNARPKVEVAITSD---SLPSSCLFDSLMASASYLILL 926
Query: 241 IDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNS-- 298
++ L+ L + L +EEL + C + +P F GL SL++L C +L S
Sbjct: 927 VNCCSFLSSLNTD--QLSHVEELNVKSCT--DPMPACGFIGLSSLKVLRISNCSALLSSV 982
Query: 299 -------------------------------LPQSVTTLTSLQRLIIHYCPE---LILPA 324
LP+ + LT+L L+I+ C L L
Sbjct: 983 CVEAGEELDTCFFPQSLSELEIVDSNIQSSLLPRYLQGLTNLSVLVINSCDSMDLLSLAY 1042
Query: 325 NMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELE 384
+ L SL + I + +G E++ L+ L + D + LP L +SL+ L
Sbjct: 1043 GTHHLTSLEAIIIKDCIFLSSL-DGFENLIALRKLVVADCKNFCFLPADLNALISLKTLA 1101
Query: 385 ISKFPKLTSLPDNFDQLENLQKLCIDRC-PRLVNRLARRTGEDWYKIAHVPILSLRLE 441
I PK+ LP N +LQ + + P L +L RR G +W KIAHVP L +E
Sbjct: 1102 IYGCPKMKFLPQN-GVPASLQLILLSLLHPELDRQLQRREGTEWDKIAHVPEKKLEVE 1158
>UniRef100_Q9ZSN2 NBS-LRR-like protein cD8 [Phaseolus vulgaris]
Length = 900
Score = 127 bits (318), Expect = 9e-28
Identities = 144/514 (28%), Positives = 211/514 (41%), Gaps = 101/514 (19%)
Query: 9 RPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWD--RNTNS 65
+ ++ F++ L L L IK L+N+ N D +L + L L L W+ RN
Sbjct: 387 KSSSEFNIQQLGQLDLHGELSIKNLENIVNPCDALAADLKNKTHLVMLDLKWNLKRNNED 446
Query: 66 TNSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPL 125
EVL L+P + L ++GY G P W++D +L +V + C C LP L
Sbjct: 447 PIKEREVLENLQPSKHLEHLSINGYSGTQFPRWLSDTFVLN-VVSLSFYKCKYCQWLPSL 505
Query: 126 GKLPFLNTLYLSQMTNVKYIDDSPYEISTE-----------------------NAFPSLT 162
G L L L + + + ID Y S+ AFP L
Sbjct: 506 GLLTSLKHLKVRSLDEIVRIDADFYGNSSSAFASLETLIFYDMKEWEEWQCMTGAFPCLQ 565
Query: 163 EMTLFDLPNLERVL-------------------------RIEGVEMLSQ--------LSK 189
+++L D P L+ L IEGVEM + L
Sbjct: 566 DLSLHDCPKLKGHLPDLPHLKDRFITCCRQLVASTPSGVEIEGVEMETSSFDMIGHHLQS 625
Query: 190 LSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTV 249
L I S P +P + V E + D +F D+ P L EL++ L +
Sbjct: 626 LRIISCPGMNIP-INYCYHFLVNLEISKCCDSLTNFPLDL---FPKLHELILSNCRNLQI 681
Query: 250 LPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC--HSLNSLPQSVTTLT 307
+ E L+ L I C++ ES PN GL++ +I IC L S+P+ ++ L
Sbjct: 682 ISQEHPH-HHLKSLSIYHCSEFESFPNE---GLLAPQIQEIYICAMEKLKSMPKRMSDLL 737
Query: 308 -SLQRLIIHYCPEL-----ILPAN---MNMLN-----------------SLREVSIMGGD 341
SL L I+ CPEL LP+N M +LN S++ +SI D
Sbjct: 738 PSLDYLFIYDCPELELSEGCLPSNIKEMCLLNCSKLVASLKKGGWGTNPSIQVLSINEVD 797
Query: 342 RRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLPDNFD 399
G + + Q L ++D P L+ L D+ G SLQ+L I P L LP+
Sbjct: 798 GECFPDEGFLPLSITQ-LEIKDCPKLKKL-DYRGLCHLSSLQKLGIENCPILQCLPEE-G 854
Query: 400 QLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHV 433
E++ +L I+ CP L R + GEDW KIAH+
Sbjct: 855 LPESISELRIESCPLLNQRCKKEEGEDWKKIAHI 888
>UniRef100_Q9ZSN3 NBS-LRR-like protein cD7 [Phaseolus vulgaris]
Length = 813
Score = 125 bits (315), Expect = 2e-27
Identities = 146/520 (28%), Positives = 218/520 (41%), Gaps = 106/520 (20%)
Query: 14 FSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRNTNSTNSAE-- 70
FS+ L L L I+ L+N+ N D +L + L L+L W+ NS +S +
Sbjct: 304 FSIQQLGQLDLHGQLSIENLENIVNPCDALAADLKNKTHLVGLNLKWNLKRNSEDSIKHR 363
Query: 71 EVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLGKLPF 130
EVL L+P R L ++GY G P W++D +L +V + L C C LP LG L
Sbjct: 364 EVLENLQPSRHLEFLLINGYFGTQFPRWLSDTFVLNVVVSLCLYKCKYCQWLPSLGLLTS 423
Query: 131 LNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQLSKL 190
L L + + + ID Y S+ +AF SL + +D+ E + G L L
Sbjct: 424 LKHLTIEGLDEILRIDAEFYGNSS-SAFASLETLIFYDMKEWEEWQCMTGA--FPSLQYL 480
Query: 191 SIQSIPQFE--LPSLPSVKEVYVG-----------GETEEDIDHEASFLRDIAGKMPNLK 237
S+Q+ P+ + LP LP +K +++ G E ++ E S I + +LK
Sbjct: 481 SLQNCPKLKGHLPDLPHLKHLFIKRCRXLVASIPRGVEIEGVEMETSSFDMIGNHLQSLK 540
Query: 238 EL-------------------------------MIDAFHQLTVLPNELSSLRSLE----- 261
L +D F +L L +L+ R+L+
Sbjct: 541 ILDCPGMNIPINHWYHFLLNLVISESCDSLTNFPLDLFPKLHEL--DLTYCRNLQIISQE 598
Query: 262 -------ELYIIDCNKLESIPNNVFYGLISLRILSFVIC--HSLNSLPQSVTTLT-SLQR 311
L I DC++ ES PN GL+ +I I L S+P+ ++ L SL
Sbjct: 599 HPHHHLKSLSICDCSEFESFPNE---GLLVPQIQKIYITAMEKLKSMPKRMSDLLPSLDY 655
Query: 312 LIIHYCPEL-----ILPAN---MNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQ------ 357
L I CPEL LP+N M +LN + V+ + ++G + I LL
Sbjct: 656 LSIRDCPELELSEGCLPSNIKEMRLLNCSKLVASL----KKGGWGTNPSIQLLSINEVDG 711
Query: 358 --------------NLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLPDNFDQL 401
L ++D P L+ L D+ G SL EL I P L LP+
Sbjct: 712 ECFPDEGFLPLSITQLEIKDCPKLKKL-DYRGLCHLSSLHELVIENCPILQCLPEE-GLP 769
Query: 402 ENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLRLE 441
E++ L I+ CP L + GEDW KIAH+ + L E
Sbjct: 770 ESISYLRIESCPLLKQWCKKEEGEDWIKIAHIKSILLDCE 809
Score = 38.9 bits (89), Expect = 0.31
Identities = 51/178 (28%), Positives = 80/178 (44%), Gaps = 29/178 (16%)
Query: 253 ELSSLRSLEELYIIDCNK-LESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQR 311
EL S L + CN ++ +P+ + LI LR L S+ LP S+ +L +LQ
Sbjct: 186 ELISNFKFLRLLSLSCNPYIKEMPDTII-DLIHLRSLDLSNT-SIERLPDSMCSLCNLQV 243
Query: 312 LIIHYCPEL-ILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSL 370
L + YCP L LP+ ++ L+ LR + + G R+ P+L L++L
Sbjct: 244 LKLKYCPFLKELPSTLHELSKLRCLELKGTTLRKA--------PML-------LGKLKNL 288
Query: 371 PDWLGD---TLSLQELEISKFPKLTSLPDNFDQL--ENLQKLCIDRCPRLVNRLARRT 423
W+G S E I + +L D QL ENL+ + ++ C L L +T
Sbjct: 289 QVWMGGFEVGKSTSEFSIQQLGQL----DLHGQLSIENLENI-VNPCDALAADLKNKT 341
>UniRef100_Q7XKH0 OSJNBa0083D01.14 protein [Oryza sativa]
Length = 1079
Score = 124 bits (311), Expect = 6e-27
Identities = 124/442 (28%), Positives = 194/442 (43%), Gaps = 18/442 (4%)
Query: 8 VRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNTNST 66
V P GF ++L +L+ L I+ L+N+ N + L L L L W + +
Sbjct: 525 VPPKCGFIASELMDLKDLRYLCIRCLENV-NADEATLAKLGEKENLIMLSLTWKNSQQES 583
Query: 67 NSAEEVLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLG 126
++ E VL L+PH +LT ++ GY G P W+ + +I+ L + + NC LPPLG
Sbjct: 584 DTEERVLNNLQPHMNLTKLKIKGYNGSRSPCWLGNTTIIN-LTYLYISNCSYWQHLPPLG 642
Query: 127 KLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQ 186
+LP L LYL + +VK ID S Y FPSL + + LP LE + +EG + +
Sbjct: 643 ELPSLKYLYLICLNSVKRIDSSFYGCERPFGFPSLEYLFIEHLPALEEWVEMEGEHLFPR 702
Query: 187 LSKLSIQSIPQF-ELPSLPSVKEVYVGGETEEDIDHEASFLRDIA-GKMPNLKELMIDAF 244
L L ++ + +P+LPS HE + A + P+L L I
Sbjct: 703 LKALVVRHCKELRNVPTLPSTVNYLEMDSVGLTTLHEPYVPNENAEPQKPSLSRLKICHC 762
Query: 245 HQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVT 304
L L +L+ SLEEL+I C L +P + L L+ ++ + C L P ++
Sbjct: 763 PYLETL-EQLNQFLSLEELHIEHCENLVQLPMDHLQMLSFLKHMTVLGCPKLMVPPATIR 821
Query: 305 TLTSLQRLIIHYCP--ELILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLR 362
++L + C E L ++ L SL + + G D +E L LS
Sbjct: 822 LPLPTKKLHVGSCGTYETCLVNSLCGLTSLTTLMLYGCD--IAALPPVEVCKSLIALSCL 879
Query: 363 DFPSLRSLPDWLG--DTLSLQELEISKFPKLTSLP----DNFDQLENLQKLCIDRCPRLV 416
+ S L D G + SL EL++ KL LP F E+ Q + C +
Sbjct: 880 EIVSCHELADLNGMEELTSLTELKVIGCNKLEELPVVSSQRFQASEHNQ--VVTACTSYL 937
Query: 417 NRLARRTGEDWYKIAHVPILSL 438
+L R D + + P+ S+
Sbjct: 938 RKLKRLQISDPFVLQWAPLRSV 959
Score = 72.8 bits (177), Expect = 2e-11
Identities = 91/346 (26%), Positives = 149/346 (42%), Gaps = 48/346 (13%)
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEG 180
Q P L +L + YL + + N F SL E+ + NL + L ++
Sbjct: 750 QKPSLSRLKICHCPYLETLEQL-------------NQFLSLEELHIEHCENLVQ-LPMDH 795
Query: 181 VEMLSQLSKLSIQSIPQFELPS------LPSVKEVYVGGETEEDIDHEASFLRDIAGKMP 234
++MLS L +++ P+ +P LP+ K+++VG +E + + G +
Sbjct: 796 LQMLSFLKHMTVLGCPKLMVPPATIRLPLPT-KKLHVGSCGT----YETCLVNSLCG-LT 849
Query: 235 NLKELMIDAFHQLTVLPNEL-SSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
+L LM+ + P E+ SL +L L I+ C++L + N L SL L + C
Sbjct: 850 SLTTLMLYGCDIAALPPVEVCKSLIALSCLEIVSCHELADL--NGMEELTSLTELKVIGC 907
Query: 294 HSLNSLP-------------QSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGG 340
+ L LP Q VT TS R + L S+ V+ M
Sbjct: 908 NKLEELPVVSSQRFQASEHNQVVTACTSYLRKLKRLQISDPFVLQWAPLRSVTSVTNMTI 967
Query: 341 DRRRGIYNG--LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNF 398
+ R + +++ LQ + +RD L LP + SL+ LE ++ + SLP+
Sbjct: 968 NSCRCLPEEWLMQNCNNLQRIGVRDASHLEFLPSIMASLTSLESLEFTRVMLIQSLPELP 1027
Query: 399 DQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLRLESDV 444
L LQ L + P L+ R + G DW+KIAH+P LR+ D+
Sbjct: 1028 SSLRRLQILGCN--PVLMRRCRKSRGRDWHKIAHIP--DLRIVEDI 1069
>UniRef100_Q84KC6 NBS-LRR disease resistance protein homologue [Hordeum vulgare]
Length = 1262
Score = 124 bits (311), Expect = 6e-27
Identities = 118/410 (28%), Positives = 191/410 (45%), Gaps = 40/410 (9%)
Query: 15 SLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMRLNRLHLAWDRNTNSTNSAEEVLG 74
SL L NLQ D K L++L E NL + L+ N S E LG
Sbjct: 869 SLGSLKNLQTLDLSGCKKLESLPESLGSLE-NLQILNLS--------NCFKLESLPESLG 919
Query: 75 ALRPHRDLTGFRLSGYRGMNIPNWMTDISILGR-------LVDVKLMNCINCSQLPP-LG 126
RL + +NI +W T++ L + L + L C+ LP LG
Sbjct: 920 -----------RLKNLQTLNI-SWCTELVFLPKNLGNLKNLPRLDLSGCMKLESLPDSLG 967
Query: 127 KLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEGVEMLSQ 186
L L TL LS+ ++ + +S + L L LP E + ++ ++ L
Sbjct: 968 SLENLETLNLSKCFKLESLPESLGGLQNLQTLDLLVCHKLESLP--ESLGGLKNLQTLQL 1025
Query: 187 LSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQ 246
++S+P+ SL +K + + + L + G + NL L + ++
Sbjct: 1026 SFCHKLESLPE----SLGGLKNLQT---LTLSVCDKLESLPESLGSLKNLHTLKLQVCYK 1078
Query: 247 LTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTL 306
L LP L S+++L L + C+ LESIP +V L +L+IL+ C L S+P+S+ +L
Sbjct: 1079 LKSLPESLGSIKNLHTLNLSVCHNLESIPESVG-SLENLQILNLSNCFKLESIPKSLGSL 1137
Query: 307 TSLQRLIIHYCPELI-LPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFP 365
+LQ LI+ +C L+ LP N+ L +L+ + + G + + + L + LQ L+L +
Sbjct: 1138 KNLQTLILSWCTRLVSLPKNLGNLKNLQTLDLSGCKKLESLPDSLGSLENLQTLNLSNCF 1197
Query: 366 SLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
L SLP+ LG LQ L + + KL SLP++ L++LQ L + CP+L
Sbjct: 1198 KLESLPEILGSLKKLQTLNLFRCGKLESLPESLGSLKHLQTLVLIDCPKL 1247
Score = 114 bits (285), Expect = 6e-24
Identities = 104/334 (31%), Positives = 164/334 (48%), Gaps = 18/334 (5%)
Query: 87 LSGYRGMN-IPNWMTDISILGRLVDVKLMNCINCSQLPP-LGKLPFLNTLYLSQMTNVKY 144
LSG RG++ IP+ + L LV + L C N +P LG L L TL LS ++
Sbjct: 617 LSGSRGISEIPS---SVGKLVSLVHLDLSYCTNVKVIPKALGILRNLQTLDLSWCEKLES 673
Query: 145 IDDSPYEISTENAFPSLTEMTLFDLPNLERVL-RIEGVEMLSQLSKLSIQSIPQFELPSL 203
+ +S + + L F+L L L ++ V+ L S ++S+P+ SL
Sbjct: 674 LPES---LGSVQNLQRLNLSNCFELEALPESLGSLKDVQTLDLSSCYKLESLPE----SL 726
Query: 204 PSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEEL 263
S+K V + ++ L G++ NL+ + + +L P SL +L+ L
Sbjct: 727 GSLKNVQT---LDLSRCYKLVSLPKNLGRLKNLRTIDLSGCKKLETFPESFGSLENLQIL 783
Query: 264 YIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL-IL 322
+ +C +LES+P + F L +L+ L+ V C L SLP+S+ L +LQ L C +L +
Sbjct: 784 NLSNCFELESLPES-FGSLKNLQTLNLVECKKLESLPESLGGLKNLQTLDFSVCHKLESV 842
Query: 323 PANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQE 382
P ++ LN+L+ + + D + L + LQ L L L SLP+ LG +LQ
Sbjct: 843 PESLGGLNNLQTLKLSVCDNLVSLLKSLGSLKNLQTLDLSGCKKLESLPESLGSLENLQI 902
Query: 383 LEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLV 416
L +S KL SLP++ +L+NLQ L I C LV
Sbjct: 903 LNLSNCFKLESLPESLGRLKNLQTLNISWCTELV 936
Score = 112 bits (279), Expect = 3e-23
Identities = 95/316 (30%), Positives = 156/316 (49%), Gaps = 14/316 (4%)
Query: 105 LGRLVDVKLMNCINC-SQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTE 163
L L +KL C N S L LG L L TL LS K ++ P + + L
Sbjct: 849 LNNLQTLKLSVCDNLVSLLKSLGSLKNLQTLDLS---GCKKLESLPESLGSLENLQILNL 905
Query: 164 MTLFDLPNLERVL-RIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHE 222
F L +L L R++ ++ L+ + +P+ L +L ++ + + G + +
Sbjct: 906 SNCFKLESLPESLGRLKNLQTLNISWCTELVFLPK-NLGNLKNLPRLDLSGCMKLES--- 961
Query: 223 ASFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGL 282
L D G + NL+ L + +L LP L L++L+ L ++ C+KLES+P ++ GL
Sbjct: 962 ---LPDSLGSLENLETLNLSKCFKLESLPESLGGLQNLQTLDLLVCHKLESLPESLG-GL 1017
Query: 283 ISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPEL-ILPANMNMLNSLREVSIMGGD 341
+L+ L CH L SLP+S+ L +LQ L + C +L LP ++ L +L + +
Sbjct: 1018 KNLQTLQLSFCHKLESLPESLGGLKNLQTLTLSVCDKLESLPESLGSLKNLHTLKLQVCY 1077
Query: 342 RRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQL 401
+ + + L I L L+L +L S+P+ +G +LQ L +S KL S+P + L
Sbjct: 1078 KLKSLPESLGSIKNLHTLNLSVCHNLESIPESVGSLENLQILNLSNCFKLESIPKSLGSL 1137
Query: 402 ENLQKLCIDRCPRLVN 417
+NLQ L + C RLV+
Sbjct: 1138 KNLQTLILSWCTRLVS 1153
Score = 88.6 bits (218), Expect = 3e-16
Identities = 64/214 (29%), Positives = 109/214 (50%), Gaps = 26/214 (12%)
Query: 231 GKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSF 290
G + NL+ L + +L LP L S+++L+ L + +C +LE++P ++ L ++ L
Sbjct: 655 GILRNLQTLDLSWCEKLESLPESLGSVQNLQRLNLSNCFELEALPESLG-SLKDVQTLDL 713
Query: 291 VICHSLNSLPQSVTTLTSLQRLIIHYCPELI-LPANMNMLNSLREVSIMGGDRRRGI--- 346
C+ L SLP+S+ +L ++Q L + C +L+ LP N+ L +LR + + G +
Sbjct: 714 SSCYKLESLPESLGSLKNVQTLDLSRCYKLVSLPKNLGRLKNLRTIDLSGCKKLETFPES 773
Query: 347 YNGLEDIPLL---------------------QNLSLRDFPSLRSLPDWLGDTLSLQELEI 385
+ LE++ +L Q L+L + L SLP+ LG +LQ L+
Sbjct: 774 FGSLENLQILNLSNCFELESLPESFGSLKNLQTLNLVECKKLESLPESLGGLKNLQTLDF 833
Query: 386 SKFPKLTSLPDNFDQLENLQKLCIDRCPRLVNRL 419
S KL S+P++ L NLQ L + C LV+ L
Sbjct: 834 SVCHKLESVPESLGGLNNLQTLKLSVCDNLVSLL 867
Score = 73.6 bits (179), Expect = 1e-11
Identities = 48/166 (28%), Positives = 88/166 (52%), Gaps = 4/166 (2%)
Query: 255 SSLRSLEELYIIDCNKLES--IPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRL 312
S+L L++L ++ KL+ P ++ L L L+ ++ +P SV L SL L
Sbjct: 581 SALGQLKQLEVLIAQKLQDRQFPESITR-LSKLHYLNLSGSRGISEIPSSVGKLVSLVHL 639
Query: 313 IIHYCPEL-ILPANMNMLNSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLP 371
+ YC + ++P + +L +L+ + + ++ + L + LQ L+L + L +LP
Sbjct: 640 DLSYCTNVKVIPKALGILRNLQTLDLSWCEKLESLPESLGSVQNLQRLNLSNCFELEALP 699
Query: 372 DWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRLVN 417
+ LG +Q L++S KL SLP++ L+N+Q L + RC +LV+
Sbjct: 700 ESLGSLKDVQTLDLSSCYKLESLPESLGSLKNVQTLDLSRCYKLVS 745
Score = 50.4 bits (119), Expect = 1e-04
Identities = 72/281 (25%), Positives = 118/281 (41%), Gaps = 32/281 (11%)
Query: 15 SLADLPNLQPGDNLHIKGLQNLRNG-GDVREPNLSSMRLNRLHLAWDRNTNSTNSAEEVL 73
SL L NLQ D L L++L G ++ NL +++L+ H S E L
Sbjct: 989 SLGGLQNLQTLDLLVCHKLESLPESLGGLK--NLQTLQLSFCH--------KLESLPESL 1038
Query: 74 GALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP-LGKLPFLN 132
G L+ + LT + + + L L +KL C LP LG + L+
Sbjct: 1039 GGLKNLQTLTLSVCD-----KLESLPESLGSLKNLHTLKLQVCYKLKSLPESLGSIKNLH 1093
Query: 133 TLYLSQMTNVKYIDDSPYEIST------ENAFP-SLTEMTLFDLPNLERVLRIEGVEMLS 185
TL LS N++ I +S + N F +L L NL+ ++ ++S
Sbjct: 1094 TLNLSVCHNLESIPESVGSLENLQILNLSNCFKLESIPKSLGSLKNLQTLILSWCTRLVS 1153
Query: 186 QLSKL----SIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASF----LRDIAGKMPNLK 237
L ++Q++ L S+ + E + ++ F L +I G + L+
Sbjct: 1154 LPKNLGNLKNLQTLDLSGCKKLESLPDSLGSLENLQTLNLSNCFKLESLPEILGSLKKLQ 1213
Query: 238 ELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNV 278
L + +L LP L SL+ L+ L +IDC KLE +P ++
Sbjct: 1214 TLNLFRCGKLESLPESLGSLKHLQTLVLIDCPKLEYLPKSL 1254
Score = 48.9 bits (115), Expect = 3e-04
Identities = 34/128 (26%), Positives = 63/128 (48%), Gaps = 1/128 (0%)
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
LR+L C S+ ++ L L+ LI + P ++ L+ L +++ G
Sbjct: 566 LRVLDLSGC-SIKDFASALGQLKQLEVLIAQKLQDRQFPESITRLSKLHYLNLSGSRGIS 624
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENL 404
I + + + L +L L +++ +P LG +LQ L++S KL SLP++ ++NL
Sbjct: 625 EIPSSVGKLVSLVHLDLSYCTNVKVIPKALGILRNLQTLDLSWCEKLESLPESLGSVQNL 684
Query: 405 QKLCIDRC 412
Q+L + C
Sbjct: 685 QRLNLSNC 692
>UniRef100_Q60D57 Putative disease resistance protein I2C-5 [Solanum demissum]
Length = 1399
Score = 122 bits (307), Expect = 2e-26
Identities = 125/472 (26%), Positives = 211/472 (44%), Gaps = 76/472 (16%)
Query: 12 AGFSLADLPNLQPGDNLHIK-GLQNLRNGGDVREPNLSSMR----LNRLHLAWDRNTNST 66
+G+ + DL Q NL+ + L+N D RE + MR +++L L W ++++
Sbjct: 621 SGWRMEDLGEAQ---NLYGSLSVVELQNVVDRREAVKAKMREKNHVDKLSLEWSESSSAD 677
Query: 67 NSAEE--VLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPP 124
NS E +L L PH+++ +++GYRG PNW+ D L +LV + ++NC NCS LP
Sbjct: 678 NSQTERDILDELSPHKNIKEVKITGYRGTKFPNWLADPLFL-KLVQLSVVNCKNCSSLPS 736
Query: 125 LGKLPFLNTLYLSQMTNVKYIDDSPY-EISTENAFPSLTEMTLFDLPNLER--------- 174
LG+LP L L +S M + + + Y +S++ F SL ++ D+P ++
Sbjct: 737 LGQLPCLKFLSISGMHGITELSEEFYGSLSSKKPFNSLVDLRFEDMPEWKQWHVLGSGEF 796
Query: 175 ----VLRIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGG----ETEEDIDHEASFL 226
L+I+ LS + + + + +FE+ P K + + G + E+ E +
Sbjct: 797 AILEKLKIKNCPELSLETPIQLSCLKRFEVIGCP--KRIRISGCKKLKFEDLTLDECDCI 854
Query: 227 RDIAGK-MPNLKELMIDAFHQLT--VLPNELSSL-----------------RSLEELYII 266
DI+ + +P + L + H LT ++P SL + L II
Sbjct: 855 DDISPELLPTARTLTVSNCHNLTRFLIPTATESLDIWNCDNIDKLSVSCGGTQMTSLKII 914
Query: 267 DCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELI---LP 323
C KL+ +P + L SL+ L C + S P+ +LQ L I+ C +L+
Sbjct: 915 YCKKLKWLPERMQELLPSLKDLILEKCPEIESFPEGGLPF-NLQLLFINNCKKLVNRRKE 973
Query: 324 ANMNMLNSLREVSIMGGDRRRGIYNG-------------LEDIPLLQNLSLRDFPSLRSL 370
+ L L+E++I I G + ++ L + L+ SL+ L
Sbjct: 974 WRLQRLPYLKELTISHDGSDEEIVGGENWELPSSIQTLRINNVKTLSSQHLKSLTSLQYL 1033
Query: 371 -------PDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
L SLQ L+I + P L SLP++ +L +L I CP L
Sbjct: 1034 EILGKLPQGQLSHLTSLQSLQIIRCPNLQSLPES-ALPSSLSQLAIYGCPNL 1084
Score = 57.4 bits (137), Expect = 8e-07
Identities = 85/336 (25%), Positives = 138/336 (40%), Gaps = 76/336 (22%)
Query: 113 LMNCINCSQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNL 172
+ NC N +L + +L + +K++ + E+ PSL ++ L P +
Sbjct: 890 IWNCDNIDKLSVSCGGTQMTSLKIIYCKKLKWLPERMQEL-----LPSLKDLILEKCPEI 944
Query: 173 ERVLRIEGVEMLSQL-----SKLSIQSIPQFELPSLPSVKEVYVG--GETEEDIDHEASF 225
E G+ QL K + ++ L LP +KE+ + G EE + E
Sbjct: 945 ESFPE-GGLPFNLQLLFINNCKKLVNRRKEWRLQRLPYLKELTISHDGSDEEIVGGENWE 1003
Query: 226 LRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISL 285
L +++ L I+ + T+ L SL SL+ L I L +P L SL
Sbjct: 1004 LPS------SIQTLRIN--NVKTLSSQHLKSLTSLQYLEI-----LGKLPQGQLSHLTSL 1050
Query: 286 RILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRRG 345
+ L + C +L SLP+S +SL +L I+ CP L + + +SL +++I+G
Sbjct: 1051 QSLQIIRCPNLQSLPESALP-SSLSQLAIYGCPNLQSLSESALPSSLSKLTIIG------ 1103
Query: 346 IYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQ 405
P LQ+L ++ PS SL EL IS+ P LT+L + FD+
Sbjct: 1104 -------CPNLQSLPVKGMPS------------SLSELHISECPLLTALLE-FDK----- 1138
Query: 406 KLCIDRCPRLVNRLARRTGEDWYKIAHVPILSLRLE 441
GE W IA P +++ E
Sbjct: 1139 ------------------GEYWSNIAQFPTININRE 1156
Score = 43.1 bits (100), Expect = 0.016
Identities = 59/247 (23%), Positives = 110/247 (43%), Gaps = 29/247 (11%)
Query: 169 LPNLERVLRIEGVEMLSQLSKLSI-QSIPQFELPSLPSVKEVYVGGETEEDIDHEASF-- 225
+P + +++ G + +L S+ + +P PS+ +++E+++ + D+ AS
Sbjct: 458 IPKDDGMIQDSGNQYFLELRSRSLFEKVPN---PSVGNIEELFLMHDLVNDLAQIASSKL 514
Query: 226 -LRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLIS 284
+R ++ N+KEL D F +L +L R L+ I K++ +P++V GL +
Sbjct: 515 CIRVLSLSHYNIKELPNDLFIKLKLL-------RFLD----ISQTKIKRLPDSVC-GLYN 562
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREV----SIMGG 340
L+ L C L LP + L +L L I L +P +++ L SLR + ++ G
Sbjct: 563 LKTLLLSSCDYLEELPLQMEKLINLCHLDISNTSRLKMPLHLSKLKSLRVLVGAKFLLSG 622
Query: 341 DR------RRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSL 394
R + +Y L + L + R+ + D LSL+ E S +
Sbjct: 623 WRMEDLGEAQNLYGSLSVVELQNVVDRREAVKAKMREKNHVDKLSLEWSESSSADNSQTE 682
Query: 395 PDNFDQL 401
D D+L
Sbjct: 683 RDILDEL 689
>UniRef100_Q6SQJ0 NBS-LRR type disease resistance protein Hom-F [Glycine max]
Length = 1124
Score = 121 bits (303), Expect = 5e-26
Identities = 128/462 (27%), Positives = 202/462 (43%), Gaps = 68/462 (14%)
Query: 4 TTFTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRN 62
++F V + FS+ L L +L I+ LQN+ N D +L + L L L WD +
Sbjct: 692 SSFNVGKSREFSIQQLGELNLHGSLSIRQLQNVENPSDALAVDLKNKTHLVELELEWDSD 751
Query: 63 TNSTNSAEE--VLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCS 120
N +S +E V+ L+P + L +S Y G P W+ + S+L R+V + L NC
Sbjct: 752 WNPDDSTKERDVIENLQPSKHLEKLTMSNYGGKQFPRWLFNNSLL-RVVSLTLKNCKGFL 810
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEG 180
LPPLG+LP L L + + + I+ + + S+ +F SL + D+ E +G
Sbjct: 811 CLPPLGRLPSLKELSIEGLDGIVSIN-ADFLGSSSCSFTSLESLEFSDMKEWEE-WECKG 868
Query: 181 VE-MLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKEL 239
V +L +LSI+ P+ + G E + H L L
Sbjct: 869 VTGAFPRLRRLSIERCPKLK-------------GHLPEQLCH--------------LNSL 901
Query: 240 MIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSL 299
I + LT +P ++ + L+EL I +C L+ I L L LS C L SL
Sbjct: 902 KISGWDSLTTIPLDIFPI--LKELQIWECPNLQRISQG--QALNHLETLSMRECPQLESL 957
Query: 300 PQSV-TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR--------GIYNGL 350
P+ + L SL L I CP++ + + ++L+ + + GG + G + L
Sbjct: 958 PEGMHVLLPSLDSLWIKDCPKVEMFPEGGLPSNLKSMGLYGGSYKLISLLKSALGGNHSL 1017
Query: 351 EDIPL-----------------LQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKL 391
E + + L NL +R+ L+ L D+ G SL+ L + P+L
Sbjct: 1018 ERLVIGGVDVECLPDEGVLPHSLVNLWIRECGDLKRL-DYRGLCHLSSLKTLTLWDCPRL 1076
Query: 392 TSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHV 433
LP+ +++ L I CP L R GEDW KIAH+
Sbjct: 1077 ECLPEE-GLPKSISTLGILNCPLLKQRCREPEGEDWPKIAHI 1117
>UniRef100_Q6SQI9 NBS-LRR type disease resistance protein Hom-B [Glycine max]
Length = 1124
Score = 120 bits (300), Expect = 1e-25
Identities = 128/462 (27%), Positives = 201/462 (42%), Gaps = 68/462 (14%)
Query: 4 TTFTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRN 62
++F V + FS+ L L +L I+ LQN+ N D +L + L L L WD +
Sbjct: 692 SSFNVGKSREFSIQQLGELNLHGSLSIRQLQNVENPSDALAVDLKNKTHLVELELEWDSD 751
Query: 63 TNSTNSAEE--VLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCS 120
N +S +E V+ L+P + L +S Y G P W+ + S+L R+V + L NC
Sbjct: 752 WNPDDSTKERDVIENLQPSKHLEKLTMSNYGGKQFPRWLFNNSLL-RVVSLTLKNCKGFL 810
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEG 180
LPPLG+LP L L + + + I+ + + S+ +F SL + D+ E +G
Sbjct: 811 CLPPLGRLPSLKELSIEGLDGIVSIN-ADFFGSSSCSFTSLESLEFSDMKEWEE-WECKG 868
Query: 181 VE-MLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKEL 239
V +L +LSI P+ + G E + H L L
Sbjct: 869 VTGAFPRLQRLSIMRCPKLK-------------GHLPEQLCH--------------LNYL 901
Query: 240 MIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSL 299
I + LT +P ++ + L+EL I +C L+ I L L LS C L SL
Sbjct: 902 KISGWDSLTTIPLDIFPI--LKELQIWECPNLQRISQG--QALNHLETLSMRECPQLESL 957
Query: 300 PQSV-TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR--------GIYNGL 350
P+ + L SL L I CP++ + + ++L+ + + GG + G + L
Sbjct: 958 PEGMHVLLPSLDSLWIDDCPKVEMFPEGGLPSNLKSMGLYGGSYKLISLLKSALGGNHSL 1017
Query: 351 EDIPL-----------------LQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKL 391
E + + L NL +R+ L+ L D+ G SL+ L + P+L
Sbjct: 1018 ERLVIGGVDVECLPDEGVLPHSLVNLWIRECGDLKRL-DYKGLCHLSSLKTLTLWDCPRL 1076
Query: 392 TSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHV 433
LP+ +++ L I CP L R GEDW KIAH+
Sbjct: 1077 QCLPEE-GLPKSISTLGILNCPLLKQRCREPEGEDWPKIAHI 1117
Score = 34.7 bits (78), Expect = 5.8
Identities = 28/99 (28%), Positives = 48/99 (48%), Gaps = 1/99 (1%)
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
LR+LS ++L +P SV L L L + + + LP ++ L +L+ + + G + +
Sbjct: 593 LRVLSLSGYYNLTKVPNSVGNLKYLSSLDLSHTEIVKLPESICSLYNLQILKLNGCEHLK 652
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQEL 383
+ + L + L L L D +R +P LG LQ L
Sbjct: 653 ELPSNLHKLTDLHRLELID-TEVRKVPAHLGKLKYLQVL 690
>UniRef100_Q6R271 Disease resistance protein [Glycine max]
Length = 1129
Score = 120 bits (300), Expect = 1e-25
Identities = 128/462 (27%), Positives = 201/462 (42%), Gaps = 68/462 (14%)
Query: 4 TTFTVRPNAGFSLADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSM-RLNRLHLAWDRN 62
++F V + FS+ L L +L I+ LQN+ N D +L + L L L WD +
Sbjct: 692 SSFNVGKSREFSIQQLGELNLHGSLSIRQLQNVENPSDALAVDLKNKTHLVELELEWDSD 751
Query: 63 TNSTNSAEE--VLGALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCS 120
N +S +E V+ L+P + L +S Y G P W+ + S+L R+V + L NC
Sbjct: 752 WNPDDSTKERDVIENLQPSKHLEKLTMSNYGGKQFPRWLFNNSLL-RVVSLTLKNCKGFL 810
Query: 121 QLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTENAFPSLTEMTLFDLPNLERVLRIEG 180
LPPLG+LP L L + + + I+ + + S+ +F SL + D+ E +G
Sbjct: 811 CLPPLGRLPSLKELSIEGLDGIVSIN-ADFFGSSSCSFTSLESLEFSDMKEWEE-WECKG 868
Query: 181 VE-MLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEASFLRDIAGKMPNLKEL 239
V +L +LSI P+ + G E + H L L
Sbjct: 869 VTGAFPRLQRLSIMRCPKLK-------------GHLPEQLCH--------------LNYL 901
Query: 240 MIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSL 299
I + LT +P ++ + L+EL I +C L+ I L L LS C L SL
Sbjct: 902 KISGWDSLTTIPLDIFPI--LKELQIWECPNLQRISQG--QALNHLETLSMRECPQLESL 957
Query: 300 PQSV-TTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR--------GIYNGL 350
P+ + L SL L I CP++ + + ++L+ + + GG + G + L
Sbjct: 958 PEGMHVLLPSLDSLWIDDCPKVEMFPEGGLPSNLKSMGLYGGSYKLISLLKSALGGNHSL 1017
Query: 351 EDIPL-----------------LQNLSLRDFPSLRSLPDWLG--DTLSLQELEISKFPKL 391
E + + L NL +R+ L+ L D+ G SL+ L + P+L
Sbjct: 1018 ERLVIGGVDVECLPDEGVLPHSLVNLWIRECGDLKRL-DYKGLCHLSSLKTLTLWDCPRL 1076
Query: 392 TSLPDNFDQLENLQKLCIDRCPRLVNRLARRTGEDWYKIAHV 433
LP+ +++ L I CP L R GEDW KIAH+
Sbjct: 1077 QCLPEE-GLPKSISTLGILNCPLLKQRCREPEGEDWPKIAHI 1117
Score = 34.7 bits (78), Expect = 5.8
Identities = 28/99 (28%), Positives = 48/99 (48%), Gaps = 1/99 (1%)
Query: 285 LRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILPANMNMLNSLREVSIMGGDRRR 344
LR+LS ++L +P SV L L L + + + LP ++ L +L+ + + G + +
Sbjct: 593 LRVLSLSGYYNLTKVPNSVGNLKYLSSLDLSHTEIVKLPESICSLYNLQILKLNGCEHLK 652
Query: 345 GIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQEL 383
+ + L + L L L D +R +P LG LQ L
Sbjct: 653 ELPSNLHKLTDLHRLELID-TEVRKVPAHLGKLKYLQVL 690
>UniRef100_Q6L3G4 Putative plant disease resistant protein [Solanum demissum]
Length = 1406
Score = 115 bits (289), Expect = 2e-24
Identities = 123/443 (27%), Positives = 204/443 (45%), Gaps = 61/443 (13%)
Query: 16 LADLPNLQPGDNLHIKGLQNLRNGGDVREPNLSSMR-LNRLHLAWDRNTNSTNSAE-EVL 73
L +L NL ++ + LQN+ + + N+ + L L W + ++ E ++L
Sbjct: 793 LGELHNLH--GSISVLELQNVVDRREALNANMMKKEHVEMLSLEWSESIADSSQTEGDIL 850
Query: 74 GALRPHRDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLMNCINCSQLPPLGKLPFLNT 133
L+P+ ++ ++GYRG PNWM D S L +LV V L NC NC+ LP LG+LP L
Sbjct: 851 DKLQPNTNIKELEIAGYRGTKFPNWMADHSFL-KLVGVSLSNCNNCASLPALGQLPSLKF 909
Query: 134 LYLSQMTNVKYIDDSPY-EISTENAFPSLTEMTLFDLPN-------------------LE 173
L + M + + + Y +S++ F SL ++ ++P +E
Sbjct: 910 LTVRGMHRITEVSEEFYGTLSSKKPFNSLEKLEFAEMPEWKQWHVLGKGEFPALHDFLIE 969
Query: 174 RVLRIEG--VEMLSQLSKLSIQSIPQF--ELP-SLPSVKEVYVGGETEEDI--DHEASFL 226
++ G E L L L I P+ E P L ++KE V + + D F
Sbjct: 970 DCPKLIGKLPEKLCSLRGLRISKCPELSPETPIQLSNLKEFKVVASPKVGVLFDDAQLFT 1029
Query: 227 RDIAGKMPNLKELMIDAFHQLTVLPNELSSLRS-LEELYIIDCNKLESIPNNVFYGLISL 285
+ G M + EL I H LT LP +S L S L+++ I C KL+ + + G ++
Sbjct: 1030 SQLQG-MKQIVELCIHDCHSLTFLP--ISILPSTLKKIEIYHCRKLKLEASMISRGDCNM 1086
Query: 286 RILSFVI--CHSLNSLPQSVTTLTSLQRLIIHYCP---ELILPANMNML-----NSLREV 335
+ + VI C S++ + S + L ++ CP L++P L +L +
Sbjct: 1087 FLENLVIYGCDSIDDI--SPELVPRSHYLSVNSCPNLTRLLIPTETEKLYIWHCKNLEIL 1144
Query: 336 SIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWLGDTL-SLQELEISKFPKLTSL 394
S+ G + +L+NLS+RD L+ LP+ + + + SL+ELE+ ++ S
Sbjct: 1145 SVASGTQ-----------TMLRNLSIRDCEKLKWLPECMQELIPSLKELELWFCTEIVSF 1193
Query: 395 PDNFDQLENLQKLCIDRCPRLVN 417
P+ NLQ L I C +LVN
Sbjct: 1194 PEGGLPF-NLQVLRIHYCKKLVN 1215
Score = 44.3 bits (103), Expect = 0.007
Identities = 32/87 (36%), Positives = 45/87 (50%), Gaps = 4/87 (4%)
Query: 235 NLKELMIDAFHQLTVLPNE-LSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVIC 293
+L L + H+L LP E L L SL +L+I C++L+S+P + SL L+ C
Sbjct: 1295 SLSRLTLFGNHELHSLPIEGLRQLTSLRDLFISSCDQLQSVPESALPS--SLSELTIQNC 1352
Query: 294 HSLNSLPQSVTTLTSLQRLIIHYCPEL 320
H L LP TS+ L I+ CP L
Sbjct: 1353 HKLQYLPVKGMP-TSISSLSIYDCPLL 1378
Score = 40.8 bits (94), Expect = 0.081
Identities = 79/314 (25%), Positives = 129/314 (40%), Gaps = 54/314 (17%)
Query: 176 LRIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEE-DIDH--EASFLRDIAGK 232
L I G + + +S + + S P++ + + ETE+ I H L +G
Sbjct: 1091 LVIYGCDSIDDISPELVPRSHYLSVNSCPNLTRLLIPTETEKLYIWHCKNLEILSVASGT 1150
Query: 233 MPNLKELMIDAFHQLTVLPNELSSL-RSLEELYIIDCNKLESIPNNVFYGL-ISLRILSF 290
L+ L I +L LP + L SL+EL + C ++ S P GL +L++L
Sbjct: 1151 QTMLRNLSIRDCEKLKWLPECMQELIPSLKELELWFCTEIVSFPEG---GLPFNLQVLRI 1207
Query: 291 VICHSLNSLPQS--VTTLTSLQRL-IIHYCPELI-----LPANMNMLN------------ 330
C L + + + L L+ L I+H +L LP ++ L
Sbjct: 1208 HYCKKLVNARKEWHLQRLPCLRELTILHDGSDLAGENWELPCSIRRLTVSNLKTLSSQLF 1267
Query: 331 -SLREVSIMGGDRRRGIYNGLED-IPL-LQNLSLRDFPSLRSLP-DWLGDTLSLQELEIS 386
SL + + I + LE+ +P+ L L+L L SLP + L SL++L IS
Sbjct: 1268 KSLTSLEYLSTGNSLQIQSLLEEGLPISLSRLTLFGNHELHSLPIEGLRQLTSLRDLFIS 1327
Query: 387 KFPKLTSLPD-------------NFDQLE---------NLQKLCIDRCPRLVNRLARRTG 424
+L S+P+ N +L+ ++ L I CP L L G
Sbjct: 1328 SCDQLQSVPESALPSSLSELTIQNCHKLQYLPVKGMPTSISSLSIYDCPLLKPLLEFDKG 1387
Query: 425 EDWYKIAHVPILSL 438
E W KIAH+ +++
Sbjct: 1388 EYWPKIAHISTINI 1401
Score = 37.4 bits (85), Expect = 0.89
Identities = 30/92 (32%), Positives = 51/92 (54%), Gaps = 7/92 (7%)
Query: 249 VLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFVICHSLNSLPQSVTTLTS 308
+LP L+SLR+L + +++ +PN++F L LRIL + ++ LP S+ L +
Sbjct: 671 ILPR-LTSLRALS----LSHYRIKELPNDLFITLKLLRILD-LSQTAIRKLPDSICALYN 724
Query: 309 LQRLIIHYCPEL-ILPANMNMLNSLREVSIMG 339
L+ L++ C L LP +M L +LR + G
Sbjct: 725 LEILLLSSCIYLEELPPHMEKLINLRHLDTTG 756
>UniRef100_Q69AB9 CC-NBS-LRR [Helianthus annuus]
Length = 1302
Score = 115 bits (289), Expect = 2e-24
Identities = 131/461 (28%), Positives = 205/461 (44%), Gaps = 68/461 (14%)
Query: 4 TTFTVRPNAGFSLADLPNLQPGDNLH----IKGLQNLRNGGDVREPNLSSMRLNRLHLAW 59
T + + GF++ +L L NLH ++GL +++ RE NLS ++ L L W
Sbjct: 680 TRIIIEGDDGFAINELKGLT---NLHGKVSLEGLHKVQSAKHAREANLSLKKITGLKLQW 736
Query: 60 ----DRNTNSTNSAEEVLGALRPH-RDLTGFRLSGYRGMNIPNWMTDISILGRLVDVKLM 114
D + T+ EEVL L+P+ L + Y G I NW+ D S LV+V +
Sbjct: 737 VDVFDGSRMDTHE-EEVLNELKPNSHTLKTLSVVSYGGTQISNWVGDCSF-HELVNVSIR 794
Query: 115 NCINCSQLPPLGKLPFLNTLYLSQMTNVKYIDDSPYEISTE--NAFPSLTEMTLFDLPNL 172
C C+ LPP G LP L L + M VK I E++ NAF SL + D+
Sbjct: 795 GCKRCTSLPPFGLLPSLKRLQIQGMDEVKIIG---LELTGNDVNAFRSLEVLIFQDMSVW 851
Query: 173 E--RVLRIEGVEMLSQLSKLSIQSIPQF---ELPSLPSVKEVYVGGETEEDIDHEASFLR 227
E + + + L +LSI S P+ L +LPS+K + + + LR
Sbjct: 852 EGWSTINEGSAAVFTCLKELSIISCPKLINVSLQALPSLKVLKIDRCGD-------GVLR 904
Query: 228 DIAGKMPNLKELMIDAFHQLT--VLPNELSSLRSLEELYIIDCNKLESI---PNNVFYGL 282
+ ++ +L I + LT V + L+ +EEL I CN+++ + L
Sbjct: 905 GLVQVASSVTKLRISSILGLTYKVWRGVIRYLKEVEELSIRGCNEIKYLWESETEASKLL 964
Query: 283 ISLRILSFVICHSLNSLPQ-------SVTTLTSLQRLIIHYC---PELILPANMNML--- 329
+ L+ LS C L SL + +TL SL+ L + YC L P ++ L
Sbjct: 965 VRLKELSLWGCSGLVSLEEKEEDGNFGSSTLLSLRSLDVSYCSSIKRLCCPNSIESLYIG 1024
Query: 330 ---------------NSLREVSIMGGDRRRGIYNGLEDIPLLQNLSLRDFPSLRSLPDWL 374
N L+ +SI D G N + +P+L+ L + + +LRS+ + L
Sbjct: 1025 DCSVITDVYLPKEGGNKLKSLSIRNCDNFEGKIN-TQSMPMLEPLHIWAWENLRSISE-L 1082
Query: 375 GDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCIDRCPRL 415
++ L L I +P + SLP+ QL NL +L I +C L
Sbjct: 1083 SNSTHLTSLYIESYPHIVSLPEL--QLSNLTRLEIGKCDNL 1121
Score = 50.4 bits (119), Expect = 1e-04
Identities = 78/331 (23%), Positives = 128/331 (38%), Gaps = 74/331 (22%)
Query: 111 VKLMNCINCSQLPPLGKLPFLNTLYLSQ-------MTNVKYIDDSPYEISTENAFPSLTE 163
+K + C N + +G + +YL + +++ D+ +I+T++ P L
Sbjct: 1009 IKRLCCPNSIESLYIGDCSVITDVYLPKEGGNKLKSLSIRNCDNFEGKINTQS-MPMLEP 1067
Query: 164 MTLFDLPNLERVLRIEGVEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDHEA 223
+ ++ NL + + + L+ L I+S P + SLP ++
Sbjct: 1068 LHIWAWENLRSISELSNS---THLTSLYIESYPH--IVSLPELQ---------------- 1106
Query: 224 SFLRDIAGKMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLI 283
+ NL L I L LP ELS+L SL I C LES+ L
Sbjct: 1107 ---------LSNLTRLEIGKCDNLESLP-ELSNLTSLS---IWTCESLESLSE-----LS 1148
Query: 284 SLRILSFVICHSLNSLPQSVTTLTSLQRLIIHYCPELILP-------------------- 323
+L LS C L SLP+ + L L+ L+I CP + +
Sbjct: 1149 NLTFLSISDCKRLVSLPE-LKNLALLKDLVIKECPCIDVSIHCVHWPPKLCSLELEGLKK 1207
Query: 324 -----ANMNMLNSLREVSIMGGDRRRGIYNGLEDIPL-LQNLSLRDFPSLRSLPDWLGDT 377
++N SL ++++ G R P L +L + F +L SL L
Sbjct: 1208 PISEWGDLNFPTSLVDLTLYGEPHVRNFSQLSHLFPSSLTSLDITGFDNLESLSTGLQHL 1267
Query: 378 LSLQELEISKFPKLTSLPDNFDQLENLQKLC 408
SLQ L I PK+ LP+ ++ Q+ C
Sbjct: 1268 TSLQHLAIFSCPKVNDLPETLPKVTIYQRRC 1298
Score = 35.0 bits (79), Expect = 4.4
Identities = 22/66 (33%), Positives = 37/66 (55%), Gaps = 2/66 (3%)
Query: 350 LEDIPLLQNLSLRDFPSLRSLPDWLGDTLSLQELEISKFPKLTSLPDNFDQLENLQKLCI 409
L + LL+ LSL F + +P+++G L+ L +S+ ++ +LP+N L NLQ L +
Sbjct: 576 LPSLTLLRVLSLSRF-RITEVPEFIGGLKHLRYLNLSR-TRIKALPENIGNLYNLQTLIV 633
Query: 410 DRCPRL 415
C L
Sbjct: 634 FGCKSL 639
Score = 34.7 bits (78), Expect = 5.8
Identities = 63/257 (24%), Positives = 111/257 (42%), Gaps = 41/257 (15%)
Query: 181 VEMLSQLSKLSIQSIPQFELPSLPSVKEVYVGGETEEDIDH---------EASFLRDIAG 231
V++L L+ L + S+ +F + +P ++GG + H L + G
Sbjct: 573 VDLLPSLTLLRVLSLSRFRITEVPE----FIGG-----LKHLRYLNLSRTRIKALPENIG 623
Query: 232 KMPNLKELMIDAFHQLTVLPNELSSLRSLEELYIIDCNKLESIPNNVFYGLISLRILSFV 291
+ NL+ L++ LT LP S L+ L D LE +P + L SL+ L+ +
Sbjct: 624 NLYNLQTLIVFGCKSLTKLPESFSKLKKLLHFDTRDTPLLEKLPLGI-GELGSLQTLTRI 682
Query: 292 ICH-----SLNSLPQSVTTL---TSLQRLIIHYCPELILPANMNMLN----SLREVSIMG 339
I ++N L + +T L SL+ L + AN+++ L+ V +
Sbjct: 683 IIEGDDGFAINEL-KGLTNLHGKVSLEGLHKVQSAKHAREANLSLKKITGLKLQWVDVFD 741
Query: 340 GDR----RRGIYNGLE-DIPLLQNLSLRDFPSLRSLPDWLGDTL--SLQELEISKFPKLT 392
G R + N L+ + L+ LS+ + + + +W+GD L + I + T
Sbjct: 742 GSRMDTHEEEVLNELKPNSHTLKTLSVVSYGGTQ-ISNWVGDCSFHELVNVSIRGCKRCT 800
Query: 393 SLPDNFDQLENLQKLCI 409
SLP F L +L++L I
Sbjct: 801 SLPP-FGLLPSLKRLQI 816
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.139 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 747,526,493
Number of Sequences: 2790947
Number of extensions: 30971767
Number of successful extensions: 88283
Number of sequences better than 10.0: 1776
Number of HSP's better than 10.0 without gapping: 312
Number of HSP's successfully gapped in prelim test: 1501
Number of HSP's that attempted gapping in prelim test: 79691
Number of HSP's gapped (non-prelim): 5530
length of query: 456
length of database: 848,049,833
effective HSP length: 131
effective length of query: 325
effective length of database: 482,435,776
effective search space: 156791627200
effective search space used: 156791627200
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC137553.6


