

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137546.15 + phase: 0 /pseudo
(355 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
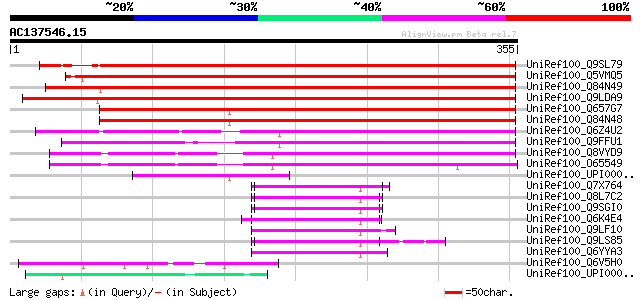
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SL79 Expressed protein [Arabidopsis thaliana] 489 e-137
UniRef100_Q5VMQ5 Putative CRS2-associated factor 1 [Oryza sativa] 475 e-133
UniRef100_Q84N49 CRS2-associated factor 1 [Zea mays] 473 e-132
UniRef100_Q9LDA9 F26F24.27 [Arabidopsis thaliana] 410 e-113
UniRef100_Q657G7 Putative CRS2-associated factor 2 [Oryza sativa] 346 5e-94
UniRef100_Q84N48 CRS2-associated factor 2 [Zea mays] 339 6e-92
UniRef100_Q6Z4U2 Putative CRS2-associated factor 1 [Oryza sativa] 263 7e-69
UniRef100_Q9FFU1 Emb|CAA18192.1 [Arabidopsis thaliana] 254 3e-66
UniRef100_Q8VYD9 Hypothetical protein At4g31010 [Arabidopsis tha... 240 4e-62
UniRef100_O65549 Hypothetical protein F6I18.80 [Arabidopsis thal... 234 2e-60
UniRef100_UPI000031FE0D UPI000031FE0D UniRef100 entry 84 8e-15
UniRef100_Q7X764 OSJNBa0060P14.12 protein [Oryza sativa] 65 4e-09
UniRef100_Q8L7C2 Hypothetical protein At3g01370 [Arabidopsis tha... 61 4e-08
UniRef100_Q9SGI0 T13O15.1 protein [Arabidopsis thaliana] 61 4e-08
UniRef100_Q6K4E4 Putative CRS1 [Oryza sativa] 59 2e-07
UniRef100_Q9LF10 Hypothetical protein T21H19_100 [Arabidopsis th... 53 1e-05
UniRef100_Q9LS85 Emb|CAB10230.1 [Arabidopsis thaliana] 52 2e-05
UniRef100_Q6YYA3 Putative CRS1 [Oryza sativa] 52 2e-05
UniRef100_Q6V5H0 Pollen coat oleosin-glycine rich protein [Arabi... 51 4e-05
UniRef100_UPI000042DD8F UPI000042DD8F UniRef100 entry 50 9e-05
>UniRef100_Q9SL79 Expressed protein [Arabidopsis thaliana]
Length = 701
Score = 489 bits (1258), Expect = e-137
Identities = 236/333 (70%), Positives = 271/333 (80%), Gaps = 15/333 (4%)
Query: 22 RPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFE 81
+P PN+ +KP+V P S P L + PE VK+SEDG++YVI GAPFE
Sbjct: 108 KPDSGPNRPKNKPRV-PDSPPQL-------------DAKPE-VKLSEDGLTYVINGAPFE 152
Query: 82 FKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKK 141
FKYSYTETPK KP+++REP + PFGP TM RPWTGR PLP S+K +EFDSF LPP KK
Sbjct: 153 FKYSYTETPKVKPLKLREPAYAPFGPTTMGRPWTGRAPLPQSQKTPREFDSFRLPPVGKK 212
Query: 142 GVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHN 201
G+KPVQ PGPF PG PRYV S+EE+LGEPLTKEE+ ELV S LK++RQLN+GRDG HN
Sbjct: 213 GLKPVQKPGPFRPGVGPRYVYSKEEILGEPLTKEEVRELVTSCLKTTRQLNMGRDGLTHN 272
Query: 202 MLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
ML+NIH WKRRRVCKIKC GVCTVDMDNVC+QLEEK GGKVIYRRGGV++LFRGRNYNH
Sbjct: 273 MLNNIHDLWKRRRVCKIKCKGVCTVDMDNVCEQLEEKIGGKVIYRRGGVLFLFRGRNYNH 332
Query: 262 KTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLV 321
+TRPRFPLMLWKPV PVYPRLIQQVPEGLT +EAT MR+KGR L PICKLGKNGVY +LV
Sbjct: 333 RTRPRFPLMLWKPVAPVYPRLIQQVPEGLTRQEATNMRRKGRELMPICKLGKNGVYCDLV 392
Query: 322 NNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
NV+EAFE CELVR++CQG+ SD+RKIGAKL+
Sbjct: 393 KNVKEAFEVCELVRIDCQGMKGSDFRKIGAKLK 425
>UniRef100_Q5VMQ5 Putative CRS2-associated factor 1 [Oryza sativa]
Length = 701
Score = 475 bits (1222), Expect = e-133
Identities = 225/321 (70%), Positives = 271/321 (84%), Gaps = 8/321 (2%)
Query: 40 SHPALKFSNI----PKQKLKPVNKTPE-NVKISEDGVSYVIEGAPFEFKYSYTETPKSKP 94
SHPA FS++ PK+ P N P V+++E G++Y ++GAPFEF+YSYTETP+++P
Sbjct: 49 SHPA--FSSVIRGRPKKVPIPENGEPAAGVRVTERGLAYHLDGAPFEFQYSYTETPRARP 106
Query: 95 VQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLP 154
V +RE PF+PFGP PRPWTGR PLP S+K+L EFDSF+LPPP KKGVKPVQSPGPFL
Sbjct: 107 VALREAPFLPFGPEVTPRPWTGRKPLPKSRKELPEFDSFMLPPPGKKGVKPVQSPGPFLA 166
Query: 155 GTSPRY-VMSREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRR 213
GT PRY SREEVLGEPLTKEE++ELV++TLK+ RQLN+GRDG HNML+NIH+HWKR+
Sbjct: 167 GTEPRYQAASREEVLGEPLTKEEVDELVKATLKTKRQLNIGRDGLTHNMLENIHSHWKRK 226
Query: 214 RVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLMLWK 273
RVCKIKC GVCTVDMDNVCQQLEEK GGKVI+ +GGVI+LFRGRNYN++TRP +PLMLWK
Sbjct: 227 RVCKIKCKGVCTVDMDNVCQQLEEKVGGKVIHHQGGVIFLFRGRNYNYRTRPIYPLMLWK 286
Query: 274 PVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECEL 333
P PVYPRL++++P+GLT +EA +MR++GR L PICKLGKNGVY NLV VREAFE C+L
Sbjct: 287 PAAPVYPRLVKKIPDGLTPDEAEDMRKRGRQLPPICKLGKNGVYLNLVKQVREAFEACDL 346
Query: 334 VRVNCQGLNKSDYRKIGAKLR 354
VRV+C GLNKSD RKIGAKL+
Sbjct: 347 VRVDCSGLNKSDCRKIGAKLK 367
>UniRef100_Q84N49 CRS2-associated factor 1 [Zea mays]
Length = 674
Score = 473 bits (1218), Expect = e-132
Identities = 223/333 (66%), Positives = 269/333 (79%), Gaps = 4/333 (1%)
Query: 26 KPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPE---NVKISEDGVSYVIEGAPFEF 82
+P P + + SHPA + + K P+ T E V++++ G+SY ++GAPFEF
Sbjct: 35 QPPTGPKRHRRAATSHPAFSAAARGRAKKIPIADTDEPAAGVRVTDRGISYRLDGAPFEF 94
Query: 83 KYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKG 142
+YSYTE P+++PV +RE PF+PFGP PRPWTGR PLP S+K+L EFDSFVLP P KKG
Sbjct: 95 QYSYTEAPRARPVALREAPFMPFGPEATPRPWTGRKPLPKSRKELPEFDSFVLPAPGKKG 154
Query: 143 VKPVQSPGPFLPGTSPRYV-MSREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHN 201
VKPVQSPGPFL G PRY +SRE++LGEPLTKEE++ELV+ +LKS RQLN+GRDG HN
Sbjct: 155 VKPVQSPGPFLAGMEPRYQSVSREDILGEPLTKEEVSELVKGSLKSKRQLNMGRDGLTHN 214
Query: 202 MLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
ML+NIH+HWKR+RVCKIKC GVCT+DMDN+C QLEEK GGKVI+R+GGVI+LFRGRNYN+
Sbjct: 215 MLENIHSHWKRKRVCKIKCKGVCTIDMDNICHQLEEKVGGKVIHRQGGVIFLFRGRNYNY 274
Query: 262 KTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLV 321
+TRP FPLMLWKPV PVYPRL+ +VP GLT +EATEMR +G L PICKLGKNGVY NLV
Sbjct: 275 RTRPCFPLMLWKPVAPVYPRLVTKVPGGLTPDEATEMRTRGHQLPPICKLGKNGVYANLV 334
Query: 322 NNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
N VREAFE C+LVRV+C GLNKSD RKIGAKL+
Sbjct: 335 NQVREAFEACDLVRVDCSGLNKSDCRKIGAKLK 367
>UniRef100_Q9LDA9 F26F24.27 [Arabidopsis thaliana]
Length = 564
Score = 410 bits (1054), Expect = e-113
Identities = 203/351 (57%), Positives = 252/351 (70%), Gaps = 6/351 (1%)
Query: 10 PISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQKLKPVNKT------PEN 63
PI S+R K N PQS+PALK + + KPV +
Sbjct: 35 PIPIPKYAPSNRNRNQKTNHQTDTNPKKPQSNPALKLPHHRTRYYKPVKEGVISSDGDRT 94
Query: 64 VKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
+ I + GVSY + GAPFEF++SY+ETPK KPV +REP F+PF P TMPRPWTG+ PL S
Sbjct: 95 ILIGDSGVSYQLPGAPFEFQFSYSETPKVKPVGIREPAFMPFAPPTMPRPWTGKAPLKKS 154
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRS 183
KKK+ FDSF PP K GVK V+ PGP G P+ M+REEVLGEPL + E L++
Sbjct: 155 KKKIPLFDSFNPPPAGKSGVKYVEMPGPLPFGRYPKEGMNREEVLGEPLKRWEKGMLIKP 214
Query: 184 TLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKV 243
+ +RQ+NLGRDGF HNML+ IH+HWKRRRVCK++C GV TVDM+NVC+ LEEKTGG++
Sbjct: 215 HMHDNRQVNLGRDGFTHNMLELIHSHWKRRRVCKVRCKGVPTVDMNNVCRVLEEKTGGEI 274
Query: 244 IYRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGR 303
I+R GGV+YLFRGRNYN++TRP++PLMLWKP PVYP+LIQ+VPEGLT EEA E R KG+
Sbjct: 275 IHRVGGVVYLFRGRNYNYRTRPQYPLMLWKPAAPVYPKLIQEVPEGLTKEEAHEFRVKGK 334
Query: 304 TLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
+L PICKL KNGVY +LV +VR+AFE LV+V+C GL SDY+KIGAKL+
Sbjct: 335 SLRPICKLSKNGVYVSLVKDVRDAFELSSLVKVDCPGLEPSDYKKIGAKLK 385
>UniRef100_Q657G7 Putative CRS2-associated factor 2 [Oryza sativa]
Length = 607
Score = 346 bits (888), Expect = 5e-94
Identities = 163/301 (54%), Positives = 218/301 (72%), Gaps = 10/301 (3%)
Query: 64 VKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
V + G+S+ + GAPF+F++SY+E P++ PV +REP F+PF P TMPRPWTG+ PL
Sbjct: 112 VVVGPSGLSFRLPGAPFDFRFSYSECPRAPPVAIREPAFLPFAPPTMPRPWTGKAPLLTK 171
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPF----------LPGTSPRYVMSREEVLGEPLT 173
++K + + P ++ + V + G L SP SREEVLGEPLT
Sbjct: 172 EEKARRRGVRLHTPLGEEAPRTVSAHGIMMEVRGRRKLDLARVSPGDGRSREEVLGEPLT 231
Query: 174 KEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQ 233
E+ +LV+ + +RQLN+GRDG HNML+ IH HW+R+ +CK++C GV TVDM N+C
Sbjct: 232 AAEVRDLVKPHISHNRQLNIGRDGLTHNMLEMIHCHWRRQEICKVRCRGVPTVDMKNLCY 291
Query: 234 QLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLE 293
LEEK+GGKVI+R GGV++L+RGRNYN +TRPR+PLMLWKP PVYP+LIQ+ PEGLT E
Sbjct: 292 HLEEKSGGKVIHRVGGVVFLYRGRNYNPRTRPRYPLMLWKPATPVYPKLIQEAPEGLTKE 351
Query: 294 EATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKL 353
EA EMR++G+ L PICKL KNG+Y LV +VR+AFE +LV+++C+GLN SDY+KIGAKL
Sbjct: 352 EADEMRRRGKDLLPICKLAKNGIYIYLVRDVRDAFEGSDLVKIDCEGLNPSDYKKIGAKL 411
Query: 354 R 354
R
Sbjct: 412 R 412
>UniRef100_Q84N48 CRS2-associated factor 2 [Zea mays]
Length = 611
Score = 339 bits (870), Expect = 6e-92
Identities = 160/301 (53%), Positives = 216/301 (71%), Gaps = 10/301 (3%)
Query: 64 VKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPS 123
V + G+S+ + GAPF+F++SY+E P++ P+ +REP F+PF P TMPRPWTG+ PL
Sbjct: 116 VVVGPSGLSFRLPGAPFDFQFSYSEAPRAPPLAIREPAFLPFAPPTMPRPWTGKAPLLTK 175
Query: 124 KKKLKEFDSFVLPPPHKKGVKPVQSPGPF----------LPGTSPRYVMSREEVLGEPLT 173
++K + + P ++ + V + G L SP SREEVLGEPLT
Sbjct: 176 EEKARRRGVRLHTPLGQETPQTVSAHGIMMEVRERRKMDLARVSPGDGRSREEVLGEPLT 235
Query: 174 KEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQ 233
E+ LV+ + +RQLN+GRDG HNML+ IH HW+R+ +CK++C GV TVDM N+C
Sbjct: 236 PSEVRALVKPHISHNRQLNIGRDGLTHNMLEMIHCHWRRQEICKVRCRGVPTVDMKNLCY 295
Query: 234 QLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLE 293
LEEK+GGKVI+R GGV++L+RGR+Y+ KTRPR+PLMLWKP PVYP+LI++ P+G T E
Sbjct: 296 HLEEKSGGKVIHRVGGVVFLYRGRHYDPKTRPRYPLMLWKPATPVYPKLIKEAPDGFTKE 355
Query: 294 EATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKL 353
EA EMR+KGR L PICKL KNG+Y LV +VR+AFE +LV+++C+GLN SDY+KIGAKL
Sbjct: 356 EADEMRRKGRDLLPICKLAKNGIYITLVKDVRDAFEGSDLVKIDCEGLNPSDYKKIGAKL 415
Query: 354 R 354
R
Sbjct: 416 R 416
>UniRef100_Q6Z4U2 Putative CRS2-associated factor 1 [Oryza sativa]
Length = 428
Score = 263 bits (671), Expect = 7e-69
Identities = 152/340 (44%), Positives = 199/340 (57%), Gaps = 20/340 (5%)
Query: 19 SSRRPTGKPNKNPSKPKVDPQSHPALKFSNIPKQK-LKPVNKTPENVKISEDGVSYVIEG 77
S RR +G ++ P H A K++ KP + P + + G
Sbjct: 18 SVRRLSGLLDRYGFVPPASLTPHSASDDGGAKKRRPKKPPYRPPSS--LDRGGRPAARSD 75
Query: 78 APFEFKYSYTET-PKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLP 136
PF+F++SYTE+ P KP+ +REP + PFGP + RPWTG P L++ +
Sbjct: 76 LPFDFRFSYTESSPGDKPIGLREPKYSPFGPGRLDRPWTGLCA-PAVDTTLRDAHADDPA 134
Query: 137 PPHKKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKS--SRQLNLG 194
P ++ ++ + RE VLGEPLT E LV KS +Q+NLG
Sbjct: 135 PAAERELEEARR-------------RERERVLGEPLTPAERAFLVSKCQKSRTKKQINLG 181
Query: 195 RDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLF 254
RDG HNML++IH HWK ++KC+GV TVDM NVC QLE+KTGG +I+R GG + L+
Sbjct: 182 RDGLTHNMLNDIHNHWKNDEAVRVKCLGVPTVDMQNVCHQLEDKTGGLIIHRHGGQLILY 241
Query: 255 RGRNYNHKTRPRFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKN 314
RGR+YN K RP PLMLWKP PVYPRLI+ EGLT+EE EMR+KG + + KL KN
Sbjct: 242 RGRHYNPKKRPVIPLMLWKPAEPVYPRLIKTTIEGLTVEETKEMRKKGLYVPVLTKLAKN 301
Query: 315 GVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKLR 354
G Y +LV VR+AF ELVR++ +GL KSDYRKIG KLR
Sbjct: 302 GYYASLVPMVRDAFLTDELVRIDSKGLPKSDYRKIGVKLR 341
>UniRef100_Q9FFU1 Emb|CAA18192.1 [Arabidopsis thaliana]
Length = 358
Score = 254 bits (648), Expect = 3e-66
Identities = 138/323 (42%), Positives = 190/323 (58%), Gaps = 35/323 (10%)
Query: 37 DPQSHPALKFSNIPKQKLKP-VNKTPENVKISEDGVSYVIEGAPFEFKYSYTET-PKSKP 94
DP P K + PK+K K K ++ ++ D VI PF+F+YSY+ET P+ +P
Sbjct: 35 DPPFSPLSKPTKPPKEKKKQKTKKQDQSSELVNDLKIPVISDLPFDFRYSYSETNPEIEP 94
Query: 95 VQMREPP-FVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFL 153
+ REP F PFGP + R WTG L E D
Sbjct: 95 IGFREPKRFSPFGPGRLDRKWTGTTALASP-----EIDQ--------------------- 128
Query: 154 PGTSPRYVMSREEVLGEPLTKEEINELVRSTLKS--SRQLNLGRDGFIHNMLDNIHAHWK 211
++V R VLGE LT++E+ EL+ S +RQ+NLG+ G HNM+D+IH HWK
Sbjct: 129 ----SQWVEERARVLGETLTEDEVTELIERYRHSDCTRQINLGKGGVTHNMIDDIHNHWK 184
Query: 212 RRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRPRFPLML 271
+ +IKC+GV T+DMDN+C LEEK+GGK++YR ++ L+RGRNY+ K+RP PLML
Sbjct: 185 KAEAVRIKCLGVPTLDMDNICFHLEEKSGGKIVYRNINILVLYRGRNYDPKSRPIIPLML 244
Query: 272 WKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEEC 331
WKP PP+YPRL++ V +GL EE EMR +G + KL +NGVY N+V VRE FE
Sbjct: 245 WKPHPPIYPRLVKNVADGLEFEETKEMRNRGLHSPALMKLTRNGVYVNVVGRVREEFETE 304
Query: 332 ELVRVNCQGLNKSDYRKIGAKLR 354
E+VR++C + SD ++IG KL+
Sbjct: 305 EIVRLDCTHVGMSDCKRIGVKLK 327
>UniRef100_Q8VYD9 Hypothetical protein At4g31010 [Arabidopsis thaliana]
Length = 405
Score = 240 bits (613), Expect = 4e-62
Identities = 134/329 (40%), Positives = 190/329 (57%), Gaps = 26/329 (7%)
Query: 29 KNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTE 88
++P +P N K+K KP + P ++ +GV V PF+F++SYTE
Sbjct: 38 QSPPPLSSSASENPDFNQKNNNKKKPKPQYRPPSSL----EGVKTVHSDLPFDFRFSYTE 93
Query: 89 TPKS-KPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQ 147
+ + +P+ +REP + PFGP + R WTG P K++ D P +K K
Sbjct: 94 SCSNVRPIGLREPKYSPFGPDRLDREWTG-VCAPAVNPKVESVDGVEDPKLEEKRRKV-- 150
Query: 148 SPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVR--STLKSSRQLNLGRDGFIHNMLDN 205
RE++ G LT+ E LV K+ RQ+NLGRDG HNML++
Sbjct: 151 ----------------REKIQGASLTEAERKFLVELCQRNKTKRQVNLGRDGLTHNMLND 194
Query: 206 IHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRP 265
++ HWK ++KC+GV T+DM NV LE+KT G+V+ + G + L+RGRNY+ K RP
Sbjct: 195 VYNHWKHAEAVRVKCLGVPTLDMKNVIFHLEDKTFGQVVSKHSGTLVLYRGRNYDPKKRP 254
Query: 266 RFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVR 325
+ PLMLWKP PVYPRLI+ +GL+++E MR+KG + + KL KNG Y +LV VR
Sbjct: 255 KIPLMLWKPHEPVYPRLIKTTIDGLSIDETKAMRKKGLAVPALTKLAKNGYYGSLVPMVR 314
Query: 326 EAFEECELVRVNCQGLNKSDYRKIGAKLR 354
+AF ELVR++C GL + DY+KIGAKLR
Sbjct: 315 DAFLVSELVRIDCLGLERKDYKKIGAKLR 343
>UniRef100_O65549 Hypothetical protein F6I18.80 [Arabidopsis thaliana]
Length = 392
Score = 234 bits (598), Expect = 2e-60
Identities = 135/343 (39%), Positives = 192/343 (55%), Gaps = 39/343 (11%)
Query: 29 KNPSKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPFEFKYSYTE 88
++P +P N K+K KP + P ++ +GV V PF+F++SYTE
Sbjct: 38 QSPPPLSSSASENPDFNQKNNNKKKPKPQYRPPSSL----EGVKTVHSDLPFDFRFSYTE 93
Query: 89 TPKS-KPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQ 147
+ + +P+ +REP + PFGP + R WTG P K++ D P +K K
Sbjct: 94 SCSNVRPIGLREPKYSPFGPDRLDREWTG-VCAPAVNPKVESVDGVEDPKLEEKRRKV-- 150
Query: 148 SPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVR--STLKSSRQLNLGRDGFIHNMLDN 205
RE++ G LT+ E LV K+ RQ+NLGRDG HNML++
Sbjct: 151 ----------------REKIQGASLTEAERKFLVELCQRNKTKRQVNLGRDGLTHNMLND 194
Query: 206 IHAHWKRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYNHKTRP 265
++ HWK ++KC+GV T+DM NV LE+KT G+V+ + G + L+RGRNY+ K RP
Sbjct: 195 VYNHWKHAEAVRVKCLGVPTLDMKNVIFHLEDKTFGQVVSKHSGTLVLYRGRNYDPKKRP 254
Query: 266 RFPLMLWKPVPPVYPRLIQQVPEGLTLEEATEMRQKGRTLTPICKLG------------- 312
+ PLMLWKP PVYPRLI+ +GL+++E MR+KG + + KLG
Sbjct: 255 KIPLMLWKPHEPVYPRLIKTTIDGLSIDETKAMRKKGLAVPALTKLGPYLFHAFLFLNSA 314
Query: 313 KNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRKIGAKLRI 355
KNG Y +LV VR+AF ELVR++C GL + DY+KIGAKLR+
Sbjct: 315 KNGYYGSLVPMVRDAFLVSELVRIDCLGLERKDYKKIGAKLRV 357
>UniRef100_UPI000031FE0D UPI000031FE0D UniRef100 entry
Length = 139
Score = 83.6 bits (205), Expect = 8e-15
Identities = 46/120 (38%), Positives = 67/120 (55%), Gaps = 10/120 (8%)
Query: 87 TETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKKLKEFDSFVLPPPHKKGVKPV 146
+E P++ P+ +REP F+PF P TMPRPWTG+ PL ++K + + P ++ + V
Sbjct: 1 SEAPRAPPLAIREPAFLPFAPPTMPRPWTGKAPLLTKEEKARRRGVRLHTPLGQETPQTV 60
Query: 147 QSPGPF----------LPGTSPRYVMSREEVLGEPLTKEEINELVRSTLKSSRQLNLGRD 196
+ G L SP SREEVLGEPLT E+ LV+ + +RQLN+G +
Sbjct: 61 SAHGIMMEVRERRKMDLARVSPGDGRSREEVLGEPLTPSEVRALVKPHISHNRQLNIGNN 120
>UniRef100_Q7X764 OSJNBa0060P14.12 protein [Oryza sativa]
Length = 1012
Score = 64.7 bits (156), Expect = 4e-09
Identities = 29/90 (32%), Positives = 51/90 (56%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L ++E+ L + ++ +L +G+ G +++ IH W+ + KI+C V ++M
Sbjct: 141 LPRDELRRLQGAGIRLRNRLKVGKAGVTEGIVNGIHERWRNAELVKIRCDDVSAMNMKRT 200
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
+ LE KTGG VI+R G I L+RG +Y +
Sbjct: 201 HEILERKTGGLVIWRSGSTIILYRGTDYKY 230
Score = 52.0 bits (123), Expect = 2e-05
Identities = 31/101 (30%), Positives = 51/101 (49%), Gaps = 4/101 (3%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +++EE L + LK L LGR G ++N+H HWK R + KI C D++
Sbjct: 578 EVISEEERYMLRKVGLKMKSFLLLGRRGVFDGTVENMHLHWKYRELVKIICKEHNIKDVE 637
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPR 266
+ LE ++GG ++ + I ++RG+NY + R
Sbjct: 638 YAARTLEAESGGILVAVERVSKAHAIIIYRGKNYQRPSTLR 678
Score = 33.9 bits (76), Expect = 7.0
Identities = 16/59 (27%), Positives = 30/59 (50%)
Query: 290 LTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNCQGLNKSDYRK 348
L +E ++ G L K+GK GV +VN + E + ELV++ C ++ + ++
Sbjct: 141 LPRDELRRLQGAGIRLRNRLKVGKAGVTEGIVNGIHERWRNAELVKIRCDDVSAMNMKR 199
>UniRef100_Q8L7C2 Hypothetical protein At3g01370 [Arabidopsis thaliana]
Length = 1011
Score = 61.2 bits (147), Expect = 4e-08
Identities = 30/90 (33%), Positives = 48/90 (53%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L E+ L ++ +++L +G+ G +++ IH W+ V KI C + ++M
Sbjct: 166 LPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHERWRTTEVVKIFCEDISRMNMKRT 225
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
LE KTGG VI+R G I L+RG NY +
Sbjct: 226 HDVLETKTGGLVIWRSGSKILLYRGVNYQY 255
Score = 51.6 bits (122), Expect = 3e-05
Identities = 31/94 (32%), Positives = 47/94 (49%), Gaps = 4/94 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T +E L + LK L LGR G ++N+H HWK R + KI C
Sbjct: 577 EGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIICNEYSIEAAH 636
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNY 259
V + LE ++GG ++ +G I ++RG+NY
Sbjct: 637 KVAEILEAESGGILVAVEMVSKGYAIIVYRGKNY 670
Score = 37.7 bits (86), Expect = 0.49
Identities = 22/82 (26%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Query: 271 LWKPVPPVYPRLIQQVPE--GLTLE--EATEMRQKGRTLTPICKLGKNGVYYNLVNNVRE 326
+WK + + ++VP LTL E +R G LT K+GK G+ +VN + E
Sbjct: 143 VWKKETEMERKKEEKVPSLAELTLPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHE 202
Query: 327 AFEECELVRVNCQGLNKSDYRK 348
+ E+V++ C+ +++ + ++
Sbjct: 203 RWRTTEVVKIFCEDISRMNMKR 224
Score = 35.4 bits (80), Expect = 2.4
Identities = 18/59 (30%), Positives = 32/59 (53%)
Query: 280 PRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNC 338
P+L EG+T +E +R+ G + P LG+ GV+ + N+ ++ ELV++ C
Sbjct: 569 PQLSDIDKEGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIIC 627
Score = 33.5 bits (75), Expect = 9.2
Identities = 29/119 (24%), Positives = 52/119 (43%), Gaps = 4/119 (3%)
Query: 145 PVQSPGPFLPGTSPRYVMSREEV---LGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHN 201
P+ G LP P Y + + LT +E+ + R LGR+ +
Sbjct: 348 PLPVDGDLLPAVVPDYRRPFRLLPYGVSPKLTDDEMTTIRRLGRPLPCHFALGRNRNLQG 407
Query: 202 MLDNIHAHWKRRRVCKIKCI-GVCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ I W++ + KI GV + + + ++L+ TGG +I R I L+RG+++
Sbjct: 408 LAVAIVKLWEKCELAKIAVKRGVQNTNSELMAEELKWLTGGTLISRDKDFIVLYRGKDF 466
>UniRef100_Q9SGI0 T13O15.1 protein [Arabidopsis thaliana]
Length = 1020
Score = 61.2 bits (147), Expect = 4e-08
Identities = 30/90 (33%), Positives = 48/90 (53%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L E+ L ++ +++L +G+ G +++ IH W+ V KI C + ++M
Sbjct: 166 LPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHERWRTTEVVKIFCEDISRMNMKRT 225
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNYNH 261
LE KTGG VI+R G I L+RG NY +
Sbjct: 226 HDVLETKTGGLVIWRSGSKILLYRGVNYQY 255
Score = 51.6 bits (122), Expect = 3e-05
Identities = 31/94 (32%), Positives = 47/94 (49%), Gaps = 4/94 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T +E L + LK L LGR G ++N+H HWK R + KI C
Sbjct: 586 EGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIICNEYSIEAAH 645
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNY 259
V + LE ++GG ++ +G I ++RG+NY
Sbjct: 646 KVAEILEAESGGILVAVEMVSKGYAIIVYRGKNY 679
Score = 37.7 bits (86), Expect = 0.49
Identities = 22/82 (26%), Positives = 43/82 (51%), Gaps = 4/82 (4%)
Query: 271 LWKPVPPVYPRLIQQVPE--GLTLE--EATEMRQKGRTLTPICKLGKNGVYYNLVNNVRE 326
+WK + + ++VP LTL E +R G LT K+GK G+ +VN + E
Sbjct: 143 VWKKETEMERKKEEKVPSLAELTLPPAELRRLRTVGIRLTKKLKIGKAGITEGIVNGIHE 202
Query: 327 AFEECELVRVNCQGLNKSDYRK 348
+ E+V++ C+ +++ + ++
Sbjct: 203 RWRTTEVVKIFCEDISRMNMKR 224
Score = 35.4 bits (80), Expect = 2.4
Identities = 18/59 (30%), Positives = 32/59 (53%)
Query: 280 PRLIQQVPEGLTLEEATEMRQKGRTLTPICKLGKNGVYYNLVNNVREAFEECELVRVNC 338
P+L EG+T +E +R+ G + P LG+ GV+ + N+ ++ ELV++ C
Sbjct: 578 PQLSDIDKEGITNDEKYMLRKIGLKMKPFLLLGRRGVFDGTIENMHLHWKYRELVKIIC 636
>UniRef100_Q6K4E4 Putative CRS1 [Oryza sativa]
Length = 947
Score = 59.3 bits (142), Expect = 2e-07
Identities = 30/98 (30%), Positives = 52/98 (52%)
Query: 163 SREEVLGEPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIG 222
S E+ + + E+ L L+ ++ +G G ++++IH W+ V K++ G
Sbjct: 332 SNTELAERTIPEHELRRLRDVALRMKERMRVGPGGVTQLIVESIHQKWRVEEVVKLRFEG 391
Query: 223 VCTVDMDNVCQQLEEKTGGKVIYRRGGVIYLFRGRNYN 260
+++M LEE+TGG VI+R G + L+RG NYN
Sbjct: 392 PPSLNMKRTHDILEERTGGIVIWRSGRSVVLYRGMNYN 429
Score = 50.8 bits (120), Expect = 6e-05
Identities = 32/94 (34%), Positives = 46/94 (48%), Gaps = 4/94 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T EE L R LK L LGR + N+H HWK R + K+ G +
Sbjct: 749 ETVTDEERFLLRRIGLKMKAFLMLGRREVFDGTVQNMHLHWKHRELVKVLVKGKSFPQVK 808
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNY 259
++ LE ++GG +I +G I L+RG+NY
Sbjct: 809 HIAISLEAESGGVLISVDKTTKGYAIILYRGKNY 842
Score = 38.1 bits (87), Expect = 0.37
Identities = 34/139 (24%), Positives = 60/139 (42%), Gaps = 8/139 (5%)
Query: 129 EFDSFV--LPPPHK--KGVKPVQSPGPFLPGTSPRYVMSRE---EVLGEPLTKEEINELV 181
+ DSF+ L P +K G P+ LPG P Y ++ L +E+ L
Sbjct: 488 DIDSFLDQLGPRYKDWSGRGPIPVDADLLPGVVPGYKTPFRLLPYMVKSTLRNKEMTALR 547
Query: 182 RSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCI-GVCTVDMDNVCQQLEEKTG 240
R +++ LGR+ + I W++ + KI GV D + +++ + TG
Sbjct: 548 RLARQTAPHFALGRNREHQGLATAIVKLWEKSSIAKIAIKRGVPNTCNDRMAEEIRKLTG 607
Query: 241 GKVIYRRGGVIYLFRGRNY 259
G ++ R I +RG ++
Sbjct: 608 GVLLSRNKEYIVFYRGNDF 626
>UniRef100_Q9LF10 Hypothetical protein T21H19_100 [Arabidopsis thaliana]
Length = 718
Score = 52.8 bits (125), Expect = 1e-05
Identities = 35/105 (33%), Positives = 51/105 (48%), Gaps = 7/105 (6%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT EE L R LK + L LGR G +++ +H HWK R V K+ + +
Sbjct: 568 EILTNEERECLRRIGLKMNSSLVLGRRGVFFGVMEGLHQHWKHREVAKVITMQKLFSRVV 627
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPRFPLM 270
+ LE ++ G +I + G I ++RG+NY RP LM
Sbjct: 628 YTAKALETESNGVLISIEKLKEGHAILIYRGKNYK---RPSSKLM 669
>UniRef100_Q9LS85 Emb|CAB10230.1 [Arabidopsis thaliana]
Length = 850
Score = 52.0 bits (123), Expect = 2e-05
Identities = 27/88 (30%), Positives = 47/88 (52%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMDNV 231
L + E+ L T +++ ++ + G +D I WK + ++K G ++M +
Sbjct: 190 LPESELRRLRNLTFRTASKMRIRGGGVTQVAVDAIKEKWKSAEIVRLKIEGASALNMRKM 249
Query: 232 CQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ LE+KTGG VI+R G I L+RG +Y
Sbjct: 250 HEILEKKTGGLVIWRSGTSISLYRGVSY 277
Score = 47.4 bits (111), Expect = 6e-04
Identities = 43/141 (30%), Positives = 65/141 (45%), Gaps = 8/141 (5%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E +T EE + LK L LGR G ++N+H HWK R + KI +
Sbjct: 605 ESITDEERFMFRKLGLKMKAFLLLGRRGVFDGTVENMHLHWKYRELVKIIVKAKTFDGVK 664
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTRPRFPLMLWKPVPPVYPRLIQ- 284
V LE ++GG ++ +G I ++RG++Y T R +L K R I+
Sbjct: 665 KVALALEAESGGILVSIDKVTKGYAIIVYRGQDYKRPTMLRPKNLLTK--RKALARSIEL 722
Query: 285 QVPEGLTLEEATEMRQKGRTL 305
Q EGL L+ + M+ K + L
Sbjct: 723 QRREGL-LKHISTMQAKAKQL 742
>UniRef100_Q6YYA3 Putative CRS1 [Oryza sativa]
Length = 725
Score = 52.0 bits (123), Expect = 2e-05
Identities = 31/99 (31%), Positives = 46/99 (46%), Gaps = 4/99 (4%)
Query: 170 EPLTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKCIGVCTVDMD 229
E LT+EE + LK + LGR G +++ IH HWK + V K+ +
Sbjct: 564 ELLTEEERRIFRKIGLKMDEHVLLGRRGVFEGVIEEIHQHWKHKEVVKVITKQNQASQIT 623
Query: 230 NVCQQLEEKTGGKVI----YRRGGVIYLFRGRNYNHKTR 264
LE +TGG +I + I L+RG+NY T+
Sbjct: 624 YTSMMLEVETGGTLIAIERFTTSHAIILYRGKNYRRPTK 662
Score = 45.8 bits (107), Expect = 0.002
Identities = 34/137 (24%), Positives = 59/137 (42%), Gaps = 14/137 (10%)
Query: 136 PPPH------KKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKE------EINELVRS 183
PPP KKG++ P + V+ RE+ P E E+ L R+
Sbjct: 108 PPPEREEEEEKKGIRRAV-PWAAARDEETKVVLRREKKTRVPTRAETELEAGELERLRRA 166
Query: 184 TLKSSRQLNLGRDGFIHNMLDNIHAHW-KRRRVCKIKCIGVCTVDMDNVCQQLEEKTGGK 242
R + G +++ + W K + + ++ + MD + LE KTGG
Sbjct: 167 ARGKERWARAKKAGITDEVVEEVRGQWAKGQELAGVRIVEPLRRCMDRAREILEIKTGGL 226
Query: 243 VIYRRGGVIYLFRGRNY 259
V++ RGG+ +++RG +Y
Sbjct: 227 VVWTRGGIHFVYRGSSY 243
Score = 37.0 bits (84), Expect = 0.83
Identities = 21/89 (23%), Positives = 42/89 (46%), Gaps = 1/89 (1%)
Query: 172 LTKEEINELVRSTLKSSRQLNLGRDGFIHNMLDNIHAHWKRRRVCKIKC-IGVCTVDMDN 230
L EE+ L + LGR+ + + I W++ + K+ +G+ + +
Sbjct: 354 LADEELTYLRKHARPLPTHFVLGRNTKLQGLAAAILKLWEKSLIAKVAVKVGIQNTNHEQ 413
Query: 231 VCQQLEEKTGGKVIYRRGGVIYLFRGRNY 259
+ + L+ TGG VI R I ++RG+++
Sbjct: 414 MARNLKRLTGGTVILRNKDYIIIYRGKDF 442
>UniRef100_Q6V5H0 Pollen coat oleosin-glycine rich protein [Arabidopsis arenosa]
Length = 1356
Score = 51.2 bits (121), Expect = 4e-05
Identities = 57/195 (29%), Positives = 79/195 (40%), Gaps = 23/195 (11%)
Query: 7 TTFPISASNADQSSRRPTGKPNKNPSKPKVDPQSHPALKFSNIP--KQKLKPVNKTPENV 64
TT P + S + S + T P K +KP P S P K S P K + KP++K+
Sbjct: 1039 TTKPTTKSTSKPSPKPTTKPPTKPITKPIAKPTSKPIAKPSTKPISKPETKPISKSTSKP 1098
Query: 65 KISEDG-VSYVIEGAP-----FEFKYSYTETPKSKPV--QMREPPFVPFG-PVTMPRPWT 115
KI S EG P + T P SKP +P P PVT P T
Sbjct: 1099 KIKTVAKPSSKEEGKPTTKPTTKSSSKPTAKPVSKPAAKSTPKPTSKPIAKPVTKP---T 1155
Query: 116 GRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSP--GPFLPGTSPRYVMSREEVLGEPLT 173
+P P+KK + PP K VKPV P P + V + + +P+
Sbjct: 1156 AKPTSNPTKKPAAK-------PPSKLTVKPVTKPTEKPTSKPIAKPAVKPTSKPIAKPVA 1208
Query: 174 KEEINELVRSTLKSS 188
K + +ST+K +
Sbjct: 1209 KPVSKPIAKSTVKQT 1223
Score = 37.7 bits (86), Expect = 0.49
Identities = 49/194 (25%), Positives = 73/194 (37%), Gaps = 30/194 (15%)
Query: 8 TFPISASNADQ-----SSRRPTGKPN----KNPSKPKVDPQSHPALKFSNIPKQKLKPVN 58
+FP ++ D+ +S +PT KP K+ SKP P + P K S K P
Sbjct: 919 SFPGGGASPDKLVPTKTSNKPTTKPGDKSAKSSSKPAAKPSTKPTSKPSTKSASKPSPKP 978
Query: 59 KTPENVKISEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPF-GPVTMP--RPWT 115
+ + K + FK + T KS +P P P+T P +P T
Sbjct: 979 TSKPSTKPT--------------FKPTAKPTSKSTAKPSTKPTTKPITKPITKPTSKPTT 1024
Query: 116 GRPPLPPSKKKLKEFDSFVLPPPHKKGVKPVQSPGPFLPGTSPRYVMSREEVL---GEPL 172
P SK+ K K KP P P P T P + + + +P+
Sbjct: 1025 KTVAKPSSKEAGKPTTKPTTKSTSKPSPKPTTKP-PTKPITKPIAKPTSKPIAKPSTKPI 1083
Query: 173 TKEEINELVRSTLK 186
+K E + +ST K
Sbjct: 1084 SKPETKPISKSTSK 1097
Score = 37.7 bits (86), Expect = 0.49
Identities = 43/168 (25%), Positives = 60/168 (35%), Gaps = 12/168 (7%)
Query: 22 RPTGKPNKNP-SKPKVDPQSHPALKFSNIPKQKLKPVNKTPENVKISEDGVSYVIEGAPF 80
+PT KP P +KP V P S P K + K KP+ K+ VK + V +
Sbjct: 1181 KPTEKPTSKPIAKPAVKPTSKPIAK--PVAKPVSKPIAKS--TVKQTSKPVGKPVTKPTA 1236
Query: 81 EFKYSYTETPKSKPVQMREPPFVPFG-PVTMPRPWTGRPPLPPSKKKL-KEFDSFVLPPP 138
+ P +KPV +P P P P +P P+ K + K P
Sbjct: 1237 KPAGKLASKPTAKPVA--KPTAKPVAKPAAKP---VAKPAAKPTSKLISKPVAKPASKPA 1291
Query: 139 HKKGVKPVQSPGPFLPGTSPRYVMSREEVLGEPLTKEEINELVRSTLK 186
K KP P P TS + +P K + + T K
Sbjct: 1292 SKPTTKPTSKPKPAAKSTSKPIAKPAVKPASKPAAKPTSKPITKPTSK 1339
>UniRef100_UPI000042DD8F UPI000042DD8F UniRef100 entry
Length = 844
Score = 50.1 bits (118), Expect = 9e-05
Identities = 49/175 (28%), Positives = 66/175 (37%), Gaps = 16/175 (9%)
Query: 12 SASNADQSSRRPTGKPN-KNPSKPK----VDPQSHPALKFSNIPKQKLKPVNKTPENVKI 66
+A ++DQ S + P K P KPK P++ P + + QK + K+ E
Sbjct: 541 TAPDSDQESTSKSSPPRVKPPLKPKPKFLAPPKNPPKVSHISPKTQKTSDLTKSKETSST 600
Query: 67 SEDGVSYVIEGAPFEFKYSYTETPKSKPVQMREPPFVPFGPVTMPRPWTGRPPLPPSKKK 126
S D + AP P S+ + PP P P PPLPPS
Sbjct: 601 SSDEEAPPAPPAPLPLPEELPPPPPSEELPPPPPPSEELPPPPPPSSGELPPPLPPS--- 657
Query: 127 LKEFDSFVLPPPHKKGVKPVQSPGPF-LPGTSPRYVMSREEVLGEPLTKEEINEL 180
F LPPP +P+ SP LP P S E L KEE + +
Sbjct: 658 ---FGELPLPPPPPPLEEPLSSPSDEPLPPPPPEVSSSS----NESLLKEEASSV 705
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.137 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 664,399,096
Number of Sequences: 2790947
Number of extensions: 32299044
Number of successful extensions: 122917
Number of sequences better than 10.0: 634
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 609
Number of HSP's that attempted gapping in prelim test: 120141
Number of HSP's gapped (non-prelim): 2090
length of query: 355
length of database: 848,049,833
effective HSP length: 128
effective length of query: 227
effective length of database: 490,808,617
effective search space: 111413556059
effective search space used: 111413556059
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC137546.15


