

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
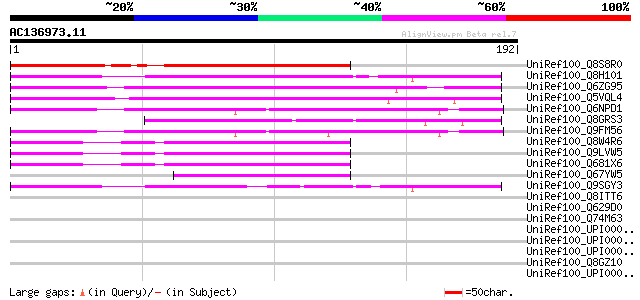
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S8R0 Predicted protein [Arabidopsis thaliana] 113 2e-24
UniRef100_Q8H101 Hypothetical protein At1g10660 [Arabidopsis tha... 100 2e-20
UniRef100_Q6ZG95 Hypothetical protein OJ1007_D04.9 [Oryza sativa] 99 8e-20
UniRef100_Q5VQL4 Hypothetical protein P0583G08.26 [Oryza sativa] 90 4e-17
UniRef100_Q6NPD1 At5g62960 [Arabidopsis thaliana] 85 1e-15
UniRef100_Q8GRS3 Hypothetical protein OJ1477_F01.109 [Oryza sativa] 82 6e-15
UniRef100_Q9FM56 Gb|AAF17656.1 [Arabidopsis thaliana] 77 2e-13
UniRef100_Q8W4R6 AT3g27760/MGF10_16 [Arabidopsis thaliana] 66 6e-10
UniRef100_Q9LVW5 Gb|AAF17656.1 [Arabidopsis thaliana] 66 6e-10
UniRef100_Q681X6 MRNA, complete cds, clone: RAFL21-92-C23 [Arabi... 66 6e-10
UniRef100_Q67YW5 MRNA, complete cds, clone: RAFL24-05-D16 [Arabi... 57 2e-07
UniRef100_Q9SGY3 F20B24.10 [Arabidopsis thaliana] 52 1e-05
UniRef100_Q8ITT6 High-affinity serotonin transporter [Manduca se... 37 0.21
UniRef100_Q629D0 Hypothetical protein CBG00076 [Caenorhabditis b... 36 0.47
UniRef100_Q74M63 NEQ510 [Nanoarchaeum equitans] 36 0.61
UniRef100_UPI000021F438 UPI000021F438 UniRef100 entry 35 1.0
UniRef100_UPI000021ECCF UPI000021ECCF UniRef100 entry 35 1.0
UniRef100_UPI00002BFDC0 UPI00002BFDC0 UniRef100 entry 35 1.0
UniRef100_Q8GZ10 Hypothetical protein [Arabidopsis thaliana] 35 1.4
UniRef100_UPI000042D911 UPI000042D911 UniRef100 entry 34 2.3
>UniRef100_Q8S8R0 Predicted protein [Arabidopsis thaliana]
Length = 289
Score = 113 bits (283), Expect = 2e-24
Identities = 61/129 (47%), Positives = 81/129 (62%), Gaps = 10/129 (7%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
WYDF+CFAIV +I+ +LW L + D++ +SLL S+NR +
Sbjct: 7 WYDFICFAIVAAAIVTSLWFLSRRDRGCVVIDDTSH--DSLLPLYR--SNNR------GS 56
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
++LW SCW +HP LL TR SF+S+A LL D+ ++DASIF YYTEWTF LV+IYFA+
Sbjct: 57 ARLWASCWTRLHPGWLLFTRSTSFLSMAALLAWDVIKWDASIFVYYTEWTFMLVIIYFAM 116
Query: 121 GTTVSAYGC 129
G S YGC
Sbjct: 117 GIVASVYGC 125
>UniRef100_Q8H101 Hypothetical protein At1g10660 [Arabidopsis thaliana]
Length = 320
Score = 100 bits (250), Expect = 2e-20
Identities = 63/198 (31%), Positives = 95/198 (47%), Gaps = 32/198 (16%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C I+ I+ A ++W EG Q +S R G +
Sbjct: 14 WRVLLCALILLAPIVLAAVLIWKYEGKRRRQRESQ----------------RELPGTLFQ 57
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ WT+C++ +HPL LL R+FSFV++ LL ++ A IFY+YT+WTFTLV +YF
Sbjct: 58 DEAWTTCFKRIHPLWLLAFRVFSFVAMLTLLISNVVRDGAGIFYFYTQWTFTLVTLYFGY 117
Query: 121 GTTVSAYGCWKVLNKPPPLQNGEMTEFLRRD------------LETKGSIFTFQSRYAEE 168
+ +S YGC + NK +G M + L+ +G+ +R +E
Sbjct: 118 ASVLSVYGCC-IYNKE---ASGNMESYTSIGDTEQGTYRPPIALDGEGNTSKASNRPSEA 173
Query: 169 EFEQTAGFWGYLMQITFQ 186
+TAGFW Y+ QI FQ
Sbjct: 174 PARKTAGFWVYIFQILFQ 191
>UniRef100_Q6ZG95 Hypothetical protein OJ1007_D04.9 [Oryza sativa]
Length = 362
Score = 98.6 bits (244), Expect = 8e-20
Identities = 61/188 (32%), Positives = 88/188 (46%), Gaps = 14/188 (7%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W VC V + A ++W +EG + S R G +
Sbjct: 47 WRAAVCALSVLACMAVAACLVWRHEGPGAERRPGGAS------GGGGGSKERRRPGVLYD 100
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ W C R +HP LL RL SF L LL + + +IFYYYT+WTF LV IYF L
Sbjct: 101 DEAWRPCLRDIHPAWLLGYRLISFFVLLSLLIVIVISDGGTIFYYYTQWTFILVTIYFGL 160
Query: 121 GTTVSAYGCWKVLNKPPPLQNGEMT--EFLRRDLETKGSIFTFQSRYAEEEFEQTAGFWG 178
GT +S YGC K+ ++ + +M ++ TK ++ E++ + AGFWG
Sbjct: 161 GTALSIYGCSKLADENVVTERTDMELGSYVAHGAGTKPNL------NGEDDTGEIAGFWG 214
Query: 179 YLMQITFQ 186
YL+QI +Q
Sbjct: 215 YLLQIIYQ 222
>UniRef100_Q5VQL4 Hypothetical protein P0583G08.26 [Oryza sativa]
Length = 333
Score = 89.7 bits (221), Expect = 4e-17
Identities = 61/189 (32%), Positives = 91/189 (47%), Gaps = 8/189 (4%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C V ++ A +++W EG S+ + + +S + R A G V
Sbjct: 14 WRFMLCAVWVYSCMVLACFLIWKYEGPSSQDGNGDGGEDS-----EDARPPRAASGVVYL 68
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
W C +HP LL R+ SF LA LL +D+ S+F YYT+WTF LV +YF L
Sbjct: 69 EDCWKPCLEQIHPGWLLAFRVVSFFILASLLAVDVVVDGWSVFLYYTQWTFLLVTLYFGL 128
Query: 121 GTTVSAYGCWKVLNKPPPLQNG--EMTEFLRRDLETKGSIFTFQSRYAE-EEFEQTAGFW 177
G+ +S YGC++ K ++G T + E+ Y++ ++ AGFW
Sbjct: 129 GSVLSIYGCYQYSYKNGDNRSGADHGTYIIAPAGESVYDQSIKNPCYSKMHGGKEIAGFW 188
Query: 178 GYLMQITFQ 186
GYL QI FQ
Sbjct: 189 GYLFQIMFQ 197
>UniRef100_Q6NPD1 At5g62960 [Arabidopsis thaliana]
Length = 347
Score = 84.7 bits (208), Expect = 1e-15
Identities = 53/189 (28%), Positives = 92/189 (48%), Gaps = 17/189 (8%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C + ++ + ++++ EG +SD + G+V
Sbjct: 38 WRVMICCIWMAIATVITAFLIFKYEGFRRKRSD----------VGEVDGGEKEWSGNVYE 87
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSF-VSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFA 119
+ W C R +HP LL R+ +F V L ML+ + + + +IF+YYT+WTF L+ +YF
Sbjct: 88 DETWRPCLRNIHPAWLLAFRVVAFFVLLVMLIVIGLVD-GPTIFFYYTQWTFGLITLYFG 146
Query: 120 LGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFTF-QSRYAEEEFEQTAGFWG 178
LG+ +S +GC++ + + + +KG+ T QS+Y+ AGFWG
Sbjct: 147 LGSLLSLHGCYQYNKRAAGDRVDSIEAIDSERARSKGADNTIQQSQYS----SNPAGFWG 202
Query: 179 YLMQITFQV 187
Y+ QI FQ+
Sbjct: 203 YVFQIIFQM 211
>UniRef100_Q8GRS3 Hypothetical protein OJ1477_F01.109 [Oryza sativa]
Length = 390
Score = 82.4 bits (202), Expect = 6e-15
Identities = 52/145 (35%), Positives = 74/145 (50%), Gaps = 12/145 (8%)
Query: 52 RVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTF 111
R A + + LW CW +HP LL R F+ + LL + + +F++YT WTF
Sbjct: 113 RRAAAPLRGTDLWVPCWARLHPGWLLGYRAFALAAAVALLVRLLVGHGIDVFFFYT-WTF 171
Query: 112 TLVMIYFALGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKG-SIFTFQSRYAEEEF 170
LV IYFA T +SA+GCW V +K + E EFL D+E + S + + + EE+
Sbjct: 172 LLVTIYFAFATAISAHGCW-VYSKKNLKKADESHEFLSDDVENREFSTSSGEMKRDEEKI 230
Query: 171 ---------EQTAGFWGYLMQITFQ 186
E+ AG WG MQI +Q
Sbjct: 231 TNYHEQIANEKRAGLWGRCMQIIYQ 255
>UniRef100_Q9FM56 Gb|AAF17656.1 [Arabidopsis thaliana]
Length = 317
Score = 77.4 bits (189), Expect = 2e-13
Identities = 53/197 (26%), Positives = 92/197 (45%), Gaps = 25/197 (12%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C + ++ + ++++ EG +SD + G+V
Sbjct: 38 WRVMICCIWMAIATVITAFLIFKYEGFRRKRSD----------VGEVDGGEKEWSGNVYE 87
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSF-VSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFA 119
+ W C R +HP LL R+ +F V L ML+ + + + +IF+YYT+WTF L+ +YF
Sbjct: 88 DETWRPCLRNIHPAWLLAFRVVAFFVLLVMLIVIGLVD-GPTIFFYYTQWTFGLITLYFG 146
Query: 120 --------LGTTVSAYGCWKVLNKPPPLQNGEMTEFLRRDLETKGSIFTF-QSRYAEEEF 170
LG+ +S +GC++ + + + +KG+ T QS+Y+
Sbjct: 147 VMFCHKNLLGSLLSLHGCYQYNKRAAGDRVDSIEAIDSERARSKGADNTIQQSQYS---- 202
Query: 171 EQTAGFWGYLMQITFQV 187
AGFWGY+ QI FQ+
Sbjct: 203 SNPAGFWGYVFQIIFQM 219
>UniRef100_Q8W4R6 AT3g27760/MGF10_16 [Arabidopsis thaliana]
Length = 315
Score = 65.9 bits (159), Expect = 6e-10
Identities = 41/129 (31%), Positives = 57/129 (43%), Gaps = 17/129 (13%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C V V ++ +L VLW E SS V PS + +
Sbjct: 18 WRVLLCAIWVIVPMIVSLLVLWKYEDSS--------------VQTQPSLNGNDVL---CI 60
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+W C+ +HP LL R+ F L I+YYYT+WTFTL+ IYF +
Sbjct: 61 DDVWRPCFERIHPGWLLGFRVLGFCFLLANNIARFANRGWRIYYYYTQWTFTLIAIYFGM 120
Query: 121 GTTVSAYGC 129
G+ +S YGC
Sbjct: 121 GSLLSIYGC 129
>UniRef100_Q9LVW5 Gb|AAF17656.1 [Arabidopsis thaliana]
Length = 343
Score = 65.9 bits (159), Expect = 6e-10
Identities = 41/129 (31%), Positives = 57/129 (43%), Gaps = 17/129 (13%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C V V ++ +L VLW E SS V PS + +
Sbjct: 18 WRVLLCAIWVIVPMIVSLLVLWKYEDSS--------------VQTQPSLNGNDVL---CI 60
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+W C+ +HP LL R+ F L I+YYYT+WTFTL+ IYF +
Sbjct: 61 DDVWRPCFERIHPGWLLGFRVLGFCFLLANNIARFANRGWRIYYYYTQWTFTLIAIYFGM 120
Query: 121 GTTVSAYGC 129
G+ +S YGC
Sbjct: 121 GSLLSIYGC 129
>UniRef100_Q681X6 MRNA, complete cds, clone: RAFL21-92-C23 [Arabidopsis thaliana]
Length = 315
Score = 65.9 bits (159), Expect = 6e-10
Identities = 41/129 (31%), Positives = 57/129 (43%), Gaps = 17/129 (13%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C V V ++ +L VLW E SS V PS + +
Sbjct: 18 WRVLLCAIWVIVPMIVSLLVLWKYEDSS--------------VQTQPSLNGNDVL---CI 60
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+W C+ +HP LL R+ F L I+YYYT+WTFTL+ IYF +
Sbjct: 61 DDVWRPCFERIHPGWLLGFRVLGFCFLLANNIARFANRGWRIYYYYTQWTFTLIAIYFGM 120
Query: 121 GTTVSAYGC 129
G+ +S YGC
Sbjct: 121 GSLLSIYGC 129
>UniRef100_Q67YW5 MRNA, complete cds, clone: RAFL24-05-D16 [Arabidopsis thaliana]
Length = 272
Score = 57.4 bits (137), Expect = 2e-07
Identities = 27/67 (40%), Positives = 37/67 (54%)
Query: 63 LWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALGT 122
L+ C+ +HP LL R+ F L I+YYYT+WTFTL+ IYF +G+
Sbjct: 20 LYGPCFERIHPGWLLGFRVLGFCFLLANNIARFANRGWRIYYYYTQWTFTLIAIYFGMGS 79
Query: 123 TVSAYGC 129
+S YGC
Sbjct: 80 LLSIYGC 86
>UniRef100_Q9SGY3 F20B24.10 [Arabidopsis thaliana]
Length = 409
Score = 51.6 bits (122), Expect = 1e-05
Identities = 50/198 (25%), Positives = 81/198 (40%), Gaps = 40/198 (20%)
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W +C I+ I+ A ++W EG Q +S R G +
Sbjct: 48 WRVLLCALILLAPIVLAAVLIWKYEGKRRRQRESQ----------------RELPGTLFQ 91
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ WT+C++ +HPL LL R+ + +L + S+ E T TLV+ +
Sbjct: 92 DEAWTTCFKRIHPLWLLAFRVDIYSCHTLL-------WGTSLGNSVVEVT-TLVLDTKSY 143
Query: 121 GTTVSAYGCWKVLNKPPPLQNGEMTEFLRRD------------LETKGSIFTFQSRYAEE 168
+ +S YGC + NK +G M + L+ +G+ +R +E
Sbjct: 144 ASVLSVYGCC-IYNKEA---SGNMESYTSIGDTEQGTYRPPIALDGEGNTSKASNRPSEA 199
Query: 169 EFEQTAGFWGYLMQITFQ 186
+TAGFW Y+ QI FQ
Sbjct: 200 PARKTAGFWVYIFQILFQ 217
>UniRef100_Q8ITT6 High-affinity serotonin transporter [Manduca sexta]
Length = 587
Score = 37.4 bits (85), Expect = 0.21
Identities = 28/92 (30%), Positives = 41/92 (44%), Gaps = 15/92 (16%)
Query: 48 SSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYT 107
S D R +GH + W +CW + P+ LL +FS ++ HE Y Y
Sbjct: 480 SEDVRTMLGH-TPGWFWRTCWSYISPVFLLVLFVFSVLA---------HEEMLGGEYTYP 529
Query: 108 EWTFTLVMIYFALGTTVSA---YGCWKVLNKP 136
W+ T+ + GTTVS Y +K+L P
Sbjct: 530 SWSITVGWV--MTGTTVSCIPLYIIYKLLITP 559
>UniRef100_Q629D0 Hypothetical protein CBG00076 [Caenorhabditis briggsae]
Length = 394
Score = 36.2 bits (82), Expect = 0.47
Identities = 25/85 (29%), Positives = 45/85 (52%), Gaps = 7/85 (8%)
Query: 30 SQSDSNIFVESLLV------ANSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFS 83
+++DS +FV+ + + A + NR+ I H+S S+L W V PL L+ + ++
Sbjct: 285 NKTDSTVFVDHITLFSLCFGAVGAKATNRLVIAHMSKSELRLWDWIYVGPLALMLNQYYN 344
Query: 84 FV-SLAMLLYLDIHEYDASIFYYYT 107
+V +LL++ AS+F Y T
Sbjct: 345 YVFDEYILLWIATIYCYASLFIYCT 369
>UniRef100_Q74M63 NEQ510 [Nanoarchaeum equitans]
Length = 146
Score = 35.8 bits (81), Expect = 0.61
Identities = 21/76 (27%), Positives = 38/76 (49%), Gaps = 7/76 (9%)
Query: 72 HPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFALG-------TTV 124
+PL+ L +F ++ L L +Y+A+I +Y +F +V++ F++ V
Sbjct: 68 YPLLYLLVPIFLAFTIESLKDLINRDYEATIVFYLLRVSFIIVLLLFSMYYIILSPIALV 127
Query: 125 SAYGCWKVLNKPPPLQ 140
+Y WK+L K P Q
Sbjct: 128 FSYLSWKILEKSPNRQ 143
>UniRef100_UPI000021F438 UPI000021F438 UniRef100 entry
Length = 634
Score = 35.0 bits (79), Expect = 1.0
Identities = 26/93 (27%), Positives = 42/93 (44%), Gaps = 8/93 (8%)
Query: 48 SSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLA--MLLYL----DIHEYDAS 101
+ D IGH + W WR V PL++L LF FV L+Y D E+ S
Sbjct: 515 NKDIEFMIGH-KPNIFWQVTWRVVSPLIMLVIFLFFFVIEVNKQLMYSVWDPDYEEFPKS 573
Query: 102 IFYYYTEWTFTLVMIYFALG-TTVSAYGCWKVL 133
Y +W + +V+I + T+ + +K++
Sbjct: 574 QKVPYPDWVYAVVVIVAGVPCLTIPCFAIYKLI 606
>UniRef100_UPI000021ECCF UPI000021ECCF UniRef100 entry
Length = 501
Score = 35.0 bits (79), Expect = 1.0
Identities = 26/93 (27%), Positives = 42/93 (44%), Gaps = 8/93 (8%)
Query: 48 SSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLA--MLLYL----DIHEYDAS 101
+ D IGH + W WR V PL++L LF FV L+Y D E+ S
Sbjct: 382 NKDIEFMIGH-KPNIFWQVTWRVVSPLIMLVIFLFFFVIEVNKQLMYSVWDPDYEEFPKS 440
Query: 102 IFYYYTEWTFTLVMIYFALG-TTVSAYGCWKVL 133
Y +W + +V+I + T+ + +K++
Sbjct: 441 QKVPYPDWVYAVVVIVAGVPCLTIPCFAIYKLI 473
>UniRef100_UPI00002BFDC0 UPI00002BFDC0 UniRef100 entry
Length = 278
Score = 35.0 bits (79), Expect = 1.0
Identities = 35/131 (26%), Positives = 64/131 (48%), Gaps = 10/131 (7%)
Query: 1 WYDFVCFAIVGVSILGALW-VLWTNEGSSTSQSDSNIFVESLLVANSP----SSDNRVA- 54
+ F+ + V +S +G L +L T + ST+ + +I + +V N+ + NR+
Sbjct: 147 YQSFIILSSVAISFVGVLLGLLITGKSFSTTMTGLSIVTLAGIVVNNNIVLIDTFNRLKE 206
Query: 55 -IGHVSTSQ-LWTSCWRGVHPLVLLT-TRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTF 111
H+ S+ + +C + + P++L + T +F + LAM + LDI D + +W
Sbjct: 207 EAPHLEASKHIIEACKQRLRPIILTSLTTIFGLLPLAMGISLDIISRDILVGSRIVDWWS 266
Query: 112 TL-VMIYFALG 121
L V I F LG
Sbjct: 267 NLAVSIVFGLG 277
>UniRef100_Q8GZ10 Hypothetical protein [Arabidopsis thaliana]
Length = 209
Score = 34.7 bits (78), Expect = 1.4
Identities = 13/36 (36%), Positives = 24/36 (66%)
Query: 60 TSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
T +LW+S WRG+ + +L T FSF+ ++ ++L +
Sbjct: 11 TRKLWSSLWRGIKTIFVLFTMFFSFLLVSAPIFLAV 46
>UniRef100_UPI000042D911 UPI000042D911 UniRef100 entry
Length = 474
Score = 33.9 bits (76), Expect = 2.3
Identities = 18/57 (31%), Positives = 32/57 (55%), Gaps = 5/57 (8%)
Query: 76 LLTTRLFSFVSLAMLLYLDIHEY---DASIFY--YYTEWTFTLVMIYFALGTTVSAY 127
L+ RL F+ L ++L LD+ Y ++++ Y T WT+ + +IY+ TT + Y
Sbjct: 56 LIYLRLSVFLYLLIMLGLDVSNYYKLGYAMYWGVYVTNWTYAITIIYYFFATTSTIY 112
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 312,076,837
Number of Sequences: 2790947
Number of extensions: 11504790
Number of successful extensions: 38715
Number of sequences better than 10.0: 37
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 22
Number of HSP's that attempted gapping in prelim test: 38682
Number of HSP's gapped (non-prelim): 39
length of query: 192
length of database: 848,049,833
effective HSP length: 120
effective length of query: 72
effective length of database: 513,136,193
effective search space: 36945805896
effective search space used: 36945805896
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 71 (32.0 bits)
Medicago: description of AC136973.11


