

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.10 + phase: 0
(330 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
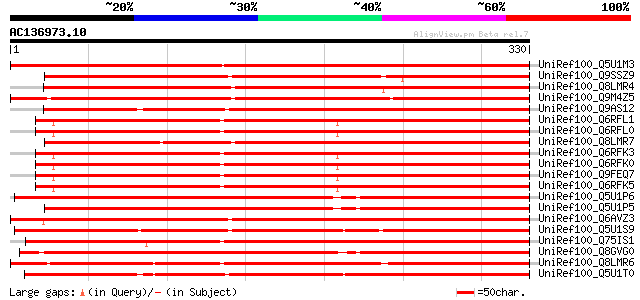
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q5U1M3 Class III peroxidase 70 precursor [Oryza sativa] 358 8e-98
UniRef100_Q9SSZ9 Peroxidase 1 [Scutellaria baicalensis] 320 3e-86
UniRef100_Q8LMR4 Putative peroxidase [Oryza sativa] 318 1e-85
UniRef100_Q9M4Z5 Peroxidase prx12 precursor [Spinacia oleracea] 312 9e-84
UniRef100_Q9AS12 Putative peroxidase [Oryza sativa] 311 2e-83
UniRef100_Q6RFL1 Peroxidase [Zea mays] 308 1e-82
UniRef100_Q6RFL0 Peroxidase [Zea mays] 308 1e-82
UniRef100_Q8LMR7 Putative peroxidase [Oryza sativa] 308 2e-82
UniRef100_Q6RFK3 Peroxidase [Zea mays] 307 2e-82
UniRef100_Q6RFK0 Peroxidase [Zea mays] 307 3e-82
UniRef100_Q9FEQ7 Peroxidase [Zea mays] 307 3e-82
UniRef100_Q6RFK5 Peroxidase [Zea mays] 306 5e-82
UniRef100_Q5U1P6 Class III peroxidase 47 precursor [Oryza sativa] 304 2e-81
UniRef100_Q5U1P5 Class III peroxidase 48 precursor [Oryza sativa] 304 2e-81
UniRef100_Q6AVZ3 Putative peroxidase [Oryza sativa] 302 9e-81
UniRef100_Q5U1S9 Class III peroxidase 14 precursor [Oryza sativa] 302 9e-81
UniRef100_Q75IS1 Peroxidase [Oryza sativa] 300 3e-80
UniRef100_Q8GVG0 Putative peroxidase [Oryza sativa] 300 5e-80
UniRef100_Q8LMR6 Putative peroxidase [Oryza sativa] 297 2e-79
UniRef100_Q5U1T0 Class III peroxidase 13 precursor [Oryza sativa] 295 9e-79
>UniRef100_Q5U1M3 Class III peroxidase 70 precursor [Oryza sativa]
Length = 335
Score = 358 bits (920), Expect = 8e-98
Identities = 185/333 (55%), Positives = 241/333 (71%), Gaps = 4/333 (1%)
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
M S + +++V+ T LK GFY+ +C E IVR AV +AV+ +PG+AAG
Sbjct: 1 MRSPWMVFAWAAAMVAVAAASPVPTKLKVGFYEHSCPQAEEIVRNAVRRAVARDPGLAAG 60
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
LIRMHFHDCFVRGCDGS+L++S PG +E+D ANNPS+RGFEV+++AKA +EA CP+TV
Sbjct: 61 LIRMHFHDCFVRGCDGSILINSTPGHVAEKDSVANNPSMRGFEVVDDAKAIVEAHCPRTV 120
Query: 121 SCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEV-TQNLPPPTFSAEQLIDNF 179
SCADILAFAARDSA ++G +DY VPSGRRDGRVS+ DEV N+P PTFS QL+ +F
Sbjct: 121 SCADILAFAARDSAH-LAGATVDYPVPSGRRDGRVSVSDEVLADNVPAPTFSLAQLVASF 179
Query: 180 DRKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQ-DPSMDPNFARLLKSKCPPP 238
+RKGL+ D+MVTLSGAH+IG SHCSSF+ RLY+F+ + DP++DP +A LK +CPP
Sbjct: 180 ERKGLTADDMVTLSGAHTIGRSHCSSFTARLYNFSGEAGRTDPAIDPAYAAELKRRCPPA 239
Query: 239 QSQSINPTVV-LDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAI 297
++PT V LD TP DN YYK + +R +L SDQ LL+S T +V ++ +
Sbjct: 240 TDDQMDPTTVPLDPVTPASFDNQYYKNVLKHRVVLNSDQALLDSPWTAGVVKLHSAVEKV 299
Query: 298 WNVKFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
+ VKFA AMV MG++DVLTG EGEIRE+C +VN
Sbjct: 300 FQVKFAAAMVKMGNIDVLTGDEGEIREKCFMVN 332
>UniRef100_Q9SSZ9 Peroxidase 1 [Scutellaria baicalensis]
Length = 322
Score = 320 bits (820), Expect = 3e-86
Identities = 167/310 (53%), Positives = 212/310 (67%), Gaps = 7/310 (2%)
Query: 23 SSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDS 82
S L+ GFY+ +C E IV++ V A + GIAAGLIR+HFHDCFVRGCDGSVL+DS
Sbjct: 17 SEAQLQKGFYQLSCGFAETIVKQEVRNAFFRDSGIAAGLIRLHFHDCFVRGCDGSVLIDS 76
Query: 83 IPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRI 142
+E+D P NNPSLRGFEV++ K ++E +CP VSCADILA+AARDS G +
Sbjct: 77 TGSNTAEKDSPPNNPSLRGFEVVDAIKRRLEVSCPGVVSCADILAYAARDSVEITRG--L 134
Query: 143 DYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSH 202
Y V +GRRDGRVS+ E NLPPP+F+ +QL F KGLS DEMVTLSGAH++G SH
Sbjct: 135 GYDVLAGRRDGRVSLASEALSNLPPPSFNVDQLTRAFANKGLSQDEMVTLSGAHTLGRSH 194
Query: 203 CSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVV--LDGSTPNDLDNM 260
C+SF+ RLY+F+ + QDP++D +A LK +CP S NP +V +D TP D
Sbjct: 195 CTSFNNRLYNFSTSSMQDPTLDLAYASQLKQQCP---QGSANPNLVVPMDPPTPAVSDVS 251
Query: 261 YYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEG 320
YY+ + NRGL TSDQTLL S TR VL+NA++ +W KFA AMV MG++ V+TG G
Sbjct: 252 YYRGVLANRGLFTSDQTLLTSPQTRAQVLQNAQNQFLWWRKFAGAMVSMGNIGVITGGAG 311
Query: 321 EIRERCSVVN 330
EIR C V+N
Sbjct: 312 EIRRDCRVIN 321
>UniRef100_Q8LMR4 Putative peroxidase [Oryza sativa]
Length = 319
Score = 318 bits (815), Expect = 1e-85
Identities = 166/312 (53%), Positives = 204/312 (65%), Gaps = 5/312 (1%)
Query: 22 ASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLD 81
A+ SL FY TC E IVR+ V +A+ N G AAGL+RMHFHDCFVRGCDGSVLL+
Sbjct: 10 ANDGSLHPNFYAATCPQAETIVRQEVTRALYTNIGFAAGLVRMHFHDCFVRGCDGSVLLE 69
Query: 82 SIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGR 141
S +ERD P NNPSLRGFEVI+ AKA++EAACP VSCAD+LA+AARD G R
Sbjct: 70 STSDNVAERDSPINNPSLRGFEVIDAAKARLEAACPGVVSCADVLAYAARDGVALTGGPR 129
Query: 142 IDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVS 201
Y VP GRRDG S+ EV N+P PTF+ +QL +F KGL+ +EMVTLSGAH++G +
Sbjct: 130 --YDVPGGRRDGTASLEPEVADNIPAPTFTLDQLTQSFAAKGLTQEEMVTLSGAHTVGRA 187
Query: 202 HCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCP---PPQSQSINPTVVLDGSTPNDLD 258
HC+SFS RLY+F+ T DPS+DP L+ CP P + V ++ TPN D
Sbjct: 188 HCTSFSDRLYNFSATGAADPSVDPALLPQLRRACPAAGPDGAVDAGLVVPMEPRTPNGFD 247
Query: 259 NMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGS 318
+YY + NR L TSDQ LL+S T V + A W +KFA AMV MG ++VLTG
Sbjct: 248 ALYYWAVLRNRALFTSDQALLSSPPTAAQVRQTAYGGYPWKLKFAAAMVKMGQIEVLTGG 307
Query: 319 EGEIRERCSVVN 330
GEIR +CS VN
Sbjct: 308 SGEIRTKCSAVN 319
>UniRef100_Q9M4Z5 Peroxidase prx12 precursor [Spinacia oleracea]
Length = 331
Score = 312 bits (799), Expect = 9e-84
Identities = 168/330 (50%), Positives = 216/330 (64%), Gaps = 5/330 (1%)
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
+HS + L V+ +S L + L+ GFY +C S E IVR V K + G+A G
Sbjct: 7 IHSSVRFLVLFSVLSCLSVQLEAQ--LQVGFYCESCPSAERIVREEVMKGFMNDKGVAPG 64
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
L+RMHFHDCFVRGCDGSVL+DS +E+D PANNPSLRGFEVI+ AK ++EA C V
Sbjct: 65 LVRMHFHDCFVRGCDGSVLIDSTSSNTAEKDSPANNPSLRGFEVIDSAKTRLEAECKGVV 124
Query: 121 SCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFD 180
SCADILAFAARDS G R Y VPSGR+DGRVS+ E QN+P TF+ +L +F
Sbjct: 125 SCADILAFAARDSVAMTRGQR--YDVPSGRKDGRVSLVSEGFQNIPGFTFNVTRLTQSFA 182
Query: 181 RKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQS 240
K L+ +EMVTLSGAH+IG SHC+S S RLY+F+ T DP++D +A L+ +CP +
Sbjct: 183 NKNLTQEEMVTLSGAHTIGRSHCTSVSNRLYNFSGTNGADPTLDSKYAGQLQQQCPQGST 242
Query: 241 QSINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNV 300
S N V++D +P D YY+ + N+GL SDQTLL T V +N R+ +W
Sbjct: 243 NS-NQVVLMDPVSPFITDVNYYQDVLANKGLFRSDQTLLTDSNTANEVNQNGRNQFLWMR 301
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
KFA AMV+MG ++VLTG+ GEIR CSV+N
Sbjct: 302 KFAAAMVNMGQIEVLTGTNGEIRTNCSVIN 331
>UniRef100_Q9AS12 Putative peroxidase [Oryza sativa]
Length = 351
Score = 311 bits (796), Expect = 2e-83
Identities = 165/311 (53%), Positives = 203/311 (65%), Gaps = 7/311 (2%)
Query: 22 ASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLD 81
A LK GFY TC S E +V++AV A N G+A GLIR+HFHDCFVRGCD SVL+D
Sbjct: 21 AVGAGLKVGFYNKTCPSAERLVQQAVAAAFKNNSGVAPGLIRLHFHDCFVRGCDASVLID 80
Query: 82 SIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGR 141
G +E+ P NNPSLRGFEVI+ AKA +EAACP+ VSCADILAFAARDS G
Sbjct: 81 ---GNDTEKTAPPNNPSLRGFEVIDAAKAAVEAACPRVVSCADILAFAARDSVALT--GN 135
Query: 142 IDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVS 201
+ Y VP+GRRDG VSI + NLPPPTF+A +L+ F K L+ ++MV LSGAH+IGVS
Sbjct: 136 VTYKVPAGRRDGNVSIAQDALDNLPPPTFNATELVGRFANKSLTAEDMVVLSGAHTIGVS 195
Query: 202 HCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-NPTVVLDGSTPNDLDNM 260
HC SF+ RLY+F DP++ +A LL++ CP SQ N TV +D TP LDN
Sbjct: 196 HCDSFTSRLYNFTGVGDADPAISAAYAFLLRAVCPSNSSQFFPNTTVDMDVITPAALDNK 255
Query: 261 YYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTG-SE 319
YY + NN GL TSD LL + R V + + W KF KAMV MG ++V TG ++
Sbjct: 256 YYVGVANNLGLFTSDHALLTNATLRASVDEFVKSETRWKSKFVKAMVKMGGIEVKTGTTQ 315
Query: 320 GEIRERCSVVN 330
GE+R C VVN
Sbjct: 316 GEVRLNCRVVN 326
>UniRef100_Q6RFL1 Peroxidase [Zea mays]
Length = 357
Score = 308 bits (790), Expect = 1e-82
Identities = 164/326 (50%), Positives = 212/326 (64%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP+GRRDGR+S E NLPPP F+A +L D F K LS++++V LSG
Sbjct: 132 --VLSGGLGYQVPAGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLSIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
TV +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q6RFL0 Peroxidase [Zea mays]
Length = 357
Score = 308 bits (789), Expect = 1e-82
Identities = 164/326 (50%), Positives = 212/326 (64%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKAALEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP+GRRDGR+S E NLPPP F+A +L D F K LS++++V LSG
Sbjct: 132 --VLSGGLGYQVPAGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLSIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
TV +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q8LMR7 Putative peroxidase [Oryza sativa]
Length = 311
Score = 308 bits (788), Expect = 2e-82
Identities = 164/308 (53%), Positives = 199/308 (64%), Gaps = 3/308 (0%)
Query: 23 SSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDS 82
S L+ G+Y T C + E IV+ V+KAVS NPG+AAGL+R+HFHDCFVRGCD SVLLDS
Sbjct: 7 SQAQLQVGYYDTLCPAAEIIVQEEVSKAVSGNPGMAAGLVRLHFHDCFVRGCDASVLLDS 66
Query: 83 IPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRI 142
G ++E+D P N SLRGFEVI+ AK+++E AC VSCAD+LAFAARD+ V G
Sbjct: 67 TQGNRAEKDAPPNT-SLRGFEVIDSAKSRLETACFGVVSCADVLAFAARDALALVGGNA- 124
Query: 143 DYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSH 202
Y VP GRRDG VS+ E NLPPP+ + QL F KGL+ EMV LSGAH+IGVSH
Sbjct: 125 -YQVPGGRRDGNVSVAQETNGNLPPPSANVAQLNQMFGAKGLTQAEMVALSGAHTIGVSH 183
Query: 203 CSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVLDGSTPNDLDNMYY 262
CSSFS RLYS QDPSMDP++ L ++CP Q Q V +D TPN D YY
Sbjct: 184 CSSFSNRLYSSGPNAGQDPSMDPSYVAALTTQCPQQQGQPAAGMVPMDAVTPNAFDTNYY 243
Query: 263 KRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGEI 322
+ NRGLL+SDQ LL T V+ + + FA AMV MGS+ VLTG+ G I
Sbjct: 244 AAIVANRGLLSSDQALLADQTTAAQVVGYTNNPDSFQTDFAAAMVKMGSIGVLTGNAGTI 303
Query: 323 RERCSVVN 330
R C V +
Sbjct: 304 RTNCRVAS 311
>UniRef100_Q6RFK3 Peroxidase [Zea mays]
Length = 357
Score = 307 bits (787), Expect = 2e-82
Identities = 163/326 (50%), Positives = 212/326 (65%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP+GRRDGR+S E NLPPP F+A +L D F K L+++++V LSG
Sbjct: 132 --VLSGGLGYQVPAGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
TV +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q6RFK0 Peroxidase [Zea mays]
Length = 357
Score = 307 bits (786), Expect = 3e-82
Identities = 163/326 (50%), Positives = 212/326 (65%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKAALEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP+GRRDGR+S E NLPPP F+A +L D F K L+++++V LSG
Sbjct: 132 --VLSGGLGYQVPAGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
TV +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q9FEQ7 Peroxidase [Zea mays]
Length = 357
Score = 307 bits (786), Expect = 3e-82
Identities = 163/326 (50%), Positives = 211/326 (64%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP GRRDGR+S E NLPPP F+A +L D F K L+++++V LSG
Sbjct: 132 --VLSGGLGYQVPGGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
TV +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTVFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q6RFK5 Peroxidase [Zea mays]
Length = 357
Score = 306 bits (784), Expect = 5e-82
Identities = 162/326 (49%), Positives = 212/326 (64%), Gaps = 14/326 (4%)
Query: 17 VSTTLASSTS--LKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGC 74
++TTL S+T+ L GFY TC + E IV++ V A N G+A LIRMHFHDCFVRGC
Sbjct: 12 LATTLLSATAACLDVGFYDRTCPTAETIVQQTVAAAFRNNSGVAPALIRMHFHDCFVRGC 71
Query: 75 DGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSA 134
DGSVL+D++ + +E+D P NNPSLR F+V++ AKA +EA CP VSCAD+LAFAARDS
Sbjct: 72 DGSVLIDTVGNLTAEKDAPPNNPSLRFFDVVDRAKASLEAQCPGVVSCADVLAFAARDSV 131
Query: 135 RKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSG 194
V G + Y VP+GRRDGR+S E NLPPP F+A +L D F K L+++++V LSG
Sbjct: 132 --VLSGGLGYQVPAGRRDGRISNDTEALNNLPPPFFNATELADRFASKNLTIEDLVVLSG 189
Query: 195 AHSIGVSHCSSFS---------KRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI-N 244
AH+IGVSHCS F+ RLY+F+ DP++ +A LLKS CP SQ N
Sbjct: 190 AHTIGVSHCSGFAGPTDLNGPVDRLYNFSSPDGIDPTLSKAYAFLLKSICPANTSQFFPN 249
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
T+ +D TP DN YY L NN GL SD LL + + +V R A + KFA+
Sbjct: 250 TTLFMDLITPERFDNKYYVGLTNNLGLFKSDVALLTNATMKALVDSFVRSEATFRTKFAR 309
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ MG ++VLTG++GEIR C V+N
Sbjct: 310 SMIKMGQIEVLTGTQGEIRRNCRVIN 335
>UniRef100_Q5U1P6 Class III peroxidase 47 precursor [Oryza sativa]
Length = 332
Score = 304 bits (779), Expect = 2e-81
Identities = 165/329 (50%), Positives = 207/329 (62%), Gaps = 8/329 (2%)
Query: 4 ILSIATLVIVILSVSTTLASST-SLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLI 62
++S A L+ T + ST LK G+Y C EAIVR AV A+ +PG+ AGLI
Sbjct: 9 LVSFAMLMAAAAGFYTPPSPSTCGLKVGYYHDKCPHAEAIVRGAVGAAILRDPGVGAGLI 68
Query: 63 RMHFHDCFVRGCDGSVLLDSIPGI-QSERDHPANNPSLRGFEVINEAKAQIEAACPKTVS 121
RM FHDCFV GCD SVLLD P Q E+ P NNPSLRGFEVI+ AK +EAACP VS
Sbjct: 69 RMLFHDCFVEGCDASVLLDPTPANPQPEKLAPPNNPSLRGFEVIDAAKTAVEAACPGVVS 128
Query: 122 CADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDR 181
CADI+AFAARD++ +S R+ + +PSGR DGR S LPPP F+ QL+ NF
Sbjct: 129 CADIVAFAARDASFFLSNSRVSFDMPSGRLDGRYSNASRTLDFLPPPKFNLGQLVANFAA 188
Query: 182 KGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQ 241
KGLSV++MV L+G+H++G SHCSSF L P D +DP+FA L+ +CP S
Sbjct: 189 KGLSVEDMVVLAGSHTVGRSHCSSF----VPDRLAVPSD--IDPSFAATLRGQCPASPSS 242
Query: 242 SINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVK 301
+PTVV D TPN LDN YYK + ++GL TSD +LL S T +MVL NA W +
Sbjct: 243 GNDPTVVQDVETPNKLDNQYYKNVLAHKGLFTSDASLLTSPATMKMVLDNANIPGWWEDR 302
Query: 302 FAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F KAMV + +++V TG GE+R C VN
Sbjct: 303 FQKAMVKLAAVEVKTGGNGEVRRNCRAVN 331
>UniRef100_Q5U1P5 Class III peroxidase 48 precursor [Oryza sativa]
Length = 339
Score = 304 bits (778), Expect = 2e-81
Identities = 164/309 (53%), Positives = 202/309 (65%), Gaps = 7/309 (2%)
Query: 23 SSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDS 82
S+ LK G+Y C EAIV+ V A+ +PG+ AGLIRM FHDCFV GCD SVLLD
Sbjct: 37 STCGLKIGYYHDKCPHAEAIVKGVVAAALHRDPGVGAGLIRMLFHDCFVEGCDASVLLDP 96
Query: 83 IPGI-QSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGR 141
P Q E+ P NNPSLRGFEVI+ AK +EAACP VSCADI+AFAARD++ +S R
Sbjct: 97 TPANPQPEKLAPPNNPSLRGFEVIDAAKDAVEAACPGVVSCADIVAFAARDASFFLSDSR 156
Query: 142 IDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVS 201
+ + +PSGR DGR S LPPPTF+ QL+ NF KGLSV++MV LSGAH+IG+S
Sbjct: 157 VSFDIPSGRLDGRYSNASRALDFLPPPTFNLGQLVANFAAKGLSVEDMVVLSGAHTIGLS 216
Query: 202 HCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVLDGSTPNDLDNMY 261
HCSSF S L D +DP+FA +L+++CP S S +PTVV D TPN LDN Y
Sbjct: 217 HCSSF----VSDRLAVASD--IDPSFAAVLRAQCPASPSSSNDPTVVQDVVTPNKLDNQY 270
Query: 262 YKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGE 321
YK + +R L TSD +LL S T +MV+ NA W +F AMV M +++V TGS GE
Sbjct: 271 YKNVLAHRALFTSDASLLASPATAKMVVDNANIPGWWEDRFKTAMVKMAAVEVKTGSNGE 330
Query: 322 IRERCSVVN 330
IR C VN
Sbjct: 331 IRRHCRAVN 339
>UniRef100_Q6AVZ3 Putative peroxidase [Oryza sativa]
Length = 344
Score = 302 bits (773), Expect = 9e-81
Identities = 164/336 (48%), Positives = 215/336 (63%), Gaps = 8/336 (2%)
Query: 1 MHSILSIATLVIVILSVSTT----LASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPG 56
+H ++ A L+ +++ +S+ + ++ L GFY +C EAIVR V KA PG
Sbjct: 11 LHVPMAAAALLGMLMMMSSAPEMKVEAAGGLSVGFYAESCPKAEAIVRDTVTKAFEKAPG 70
Query: 57 IAAGLIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAAC 116
A LIR+ FHDCFVRGCD SVLL+S PG ++ERD+ ANNPSL GF+V+++AK +E C
Sbjct: 71 TPADLIRLFFHDCFVRGCDASVLLESTPGNKAERDNKANNPSLDGFDVVDDAKDLLEKEC 130
Query: 117 PKTVSCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLI 176
P TVSCADIL+ ARDSA G +D+ +P+GRRDG VS DEV N+P P F A+ L+
Sbjct: 131 PHTVSCADILSLVARDSAYLAGG--LDFEIPTGRRDGFVSKEDEVLSNVPHPEFGAKDLL 188
Query: 177 DNFDRKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCP 236
NF KG + +EMVTLSGAHSIG SHCSSF+ RLY + T+ DPSM +A +KSKCP
Sbjct: 189 KNFTAKGFTAEEMVTLSGAHSIGTSHCSSFTNRLYKYYGTYGTDPSMPAAYAADMKSKCP 248
Query: 237 PPQSQSINPTVV-LDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMV-LKNARH 294
P + + T+V LD TP +DN YY+ + SD LL++ T +V L A
Sbjct: 249 PETAAQQDATMVQLDDVTPFKMDNQYYRNVLAGNVTFASDVALLDTPETAALVRLYAAGD 308
Query: 295 AAIWNVKFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
A W +FA A+V + LDVLTG EGEIR CS +N
Sbjct: 309 PAAWLARFAAALVKVSKLDVLTGGEGEIRLNCSRIN 344
>UniRef100_Q5U1S9 Class III peroxidase 14 precursor [Oryza sativa]
Length = 356
Score = 302 bits (773), Expect = 9e-81
Identities = 162/327 (49%), Positives = 210/327 (63%), Gaps = 6/327 (1%)
Query: 4 ILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIR 63
+++ + + + + AS L+ GFY T+C + E +VR+AV A + + GIAAGLIR
Sbjct: 6 VVAAVAVALGVCLLQLPAASRGQLQVGFYNTSCPNAETLVRQAVTNAFANDSGIAAGLIR 65
Query: 64 MHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCA 123
+HFHDCFVRGCD SVLL S P +ERD NNPSLRGF+VI+ AKA +E +C +TVSCA
Sbjct: 66 LHFHDCFVRGCDASVLLTS-PNNTAERDAAPNNPSLRGFQVIDAAKAAVEQSCARTVSCA 124
Query: 124 DILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKG 183
DI+AFAARDS G + Y VPSGRRDG VS+ + NLP PTF+A QL+ +F K
Sbjct: 125 DIVAFAARDSVNLTGG--VSYQVPSGRRDGNVSVAQDAIDNLPQPTFTAAQLVASFANKS 182
Query: 184 LSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI 243
L+ +EMV LSGAH++G S CSSF R+++ N T D + P +A LL++ C P + S
Sbjct: 183 LTAEEMVVLSGAHTVGRSFCSSFLARIWN-NTTPIVDTGLSPGYAALLRALC--PSNASA 239
Query: 244 NPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFA 303
T +D STP LDN YYK L N GL SD L + V A + +W KF
Sbjct: 240 TATTAIDVSTPATLDNNYYKLLPLNLGLFFSDNQLRVNATLGASVSSFAANETLWKEKFV 299
Query: 304 KAMVHMGSLDVLTGSEGEIRERCSVVN 330
AMV MGS++VLTGS+GE+R CSVVN
Sbjct: 300 AAMVKMGSIEVLTGSQGEVRLNCSVVN 326
>UniRef100_Q75IS1 Peroxidase [Oryza sativa]
Length = 359
Score = 300 bits (769), Expect = 3e-80
Identities = 156/324 (48%), Positives = 207/324 (63%), Gaps = 6/324 (1%)
Query: 11 VIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCF 70
V+ L + + + L GFY TTC + E ++++ V A + G+A +IRMHFHDCF
Sbjct: 10 VVAALISAAAVGARACLDVGFYDTTCPTAETLIQQVVAAAFRNDSGVAPAMIRMHFHDCF 69
Query: 71 VRGCDGSVLLDSIPG--IQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAF 128
VRGCDGSVL+D++PG ++E+D NNPSLR F+VI+ AK+ +EAACP VSCAD++AF
Sbjct: 70 VRGCDGSVLIDTVPGSTTRAEKDAAPNNPSLRFFDVIDRAKSAVEAACPGVVSCADVVAF 129
Query: 129 AARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDE 188
ARD V G + Y VP+GRRDGR S+ D+ LPPPT +A L+ NF K L+ ++
Sbjct: 130 MARDGV--VLSGGLGYQVPAGRRDGRTSLEDDALNFLPPPTSTAADLVANFTAKNLTAED 187
Query: 189 MVTLSGAHSIGVSHCSSFSKRLYSF-NLTFPQDPSMDPNFARLLKSKCPPPQSQSI-NPT 246
MV LSGAH+IGVSHC SF+ R+Y+F N T DPS+ +A LLK CPP +Q+ T
Sbjct: 188 MVVLSGAHTIGVSHCDSFTNRIYNFPNTTDGIDPSLSKAYAFLLKGICPPNSNQTFPTTT 247
Query: 247 VVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAM 306
+D TP DN YY L NN GL SD LL + V R A + +KFA+AM
Sbjct: 248 TFMDILTPTKFDNRYYVGLTNNLGLFQSDAALLTDAALKATVNSFVRSEATFRLKFARAM 307
Query: 307 VHMGSLDVLTGSEGEIRERCSVVN 330
+ MG + VL+G++GEIR C VVN
Sbjct: 308 IKMGQIGVLSGTQGEIRLNCRVVN 331
>UniRef100_Q8GVG0 Putative peroxidase [Oryza sativa]
Length = 322
Score = 300 bits (767), Expect = 5e-80
Identities = 161/325 (49%), Positives = 209/325 (63%), Gaps = 10/325 (3%)
Query: 7 IATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHF 66
+A V +L+++ A L+ G+YK C+ E +VR V AV NPG+ AG++RM F
Sbjct: 6 VAMWVACVLAMAA--ACQGRLRVGYYKRKCAPAEYVVRAVVGNAVRQNPGVGAGIVRMFF 63
Query: 67 HDCFVRGCDGSVLLD-SIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADI 125
HDCFV+GCD SVLLD + Q E+ P N PSLRGFEVI+ AKA +E ACP VSCADI
Sbjct: 64 HDCFVQGCDASVLLDPTAANPQPEKLGPPNFPSLRGFEVIDAAKAAVEKACPGVVSCADI 123
Query: 126 LAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLS 185
+AFAARD++ +SGG I Y +P+GR DGRVS+ +E LPPP F+ QL+ +F KGL
Sbjct: 124 IAFAARDASFFLSGGGISYRIPAGRLDGRVSLANETLAFLPPPVFNLTQLVASFQAKGLD 183
Query: 186 VDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINP 245
D+MVTLSGAH+IG SHCSSF+ R L+ P D MDP A L+SKCP + + +P
Sbjct: 184 ADDMVTLSGAHTIGRSHCSSFADR-----LSPPSD--MDPGLAAALRSKCPASPNFTDDP 236
Query: 246 TVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKA 305
TV D TP+ +D YY+ + + + L SD LL S T MV +NA W +FA+A
Sbjct: 237 TVAQDAVTPDRMDRQYYRNVLDRKVLFDSDAALLASRPTAAMVARNAAARGRWERRFARA 296
Query: 306 MVHMGSLDVLTGSEGEIRERCSVVN 330
MV MG ++V T + GEIR C VVN
Sbjct: 297 MVKMGGIEVKTAANGEIRRMCRVVN 321
>UniRef100_Q8LMR6 Putative peroxidase [Oryza sativa]
Length = 322
Score = 297 bits (761), Expect = 2e-79
Identities = 162/330 (49%), Positives = 206/330 (62%), Gaps = 8/330 (2%)
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
M ++++A V+V LS+ L+ GFY +C E IVR V KAVS N G+AAG
Sbjct: 1 MELVVAVAGAVVVALSLCIGGVQG-QLQVGFYDQSCPQAEVIVRDEVGKAVSANVGLAAG 59
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
L+RMHFHDCFV+GCD SVLLDS +E+D N SLRGFEV++ AK ++E+AC V
Sbjct: 60 LVRMHFHDCFVKGCDASVLLDSTANSTAEKD-AIPNKSLRGFEVVDSAKRRLESACKGVV 118
Query: 121 SCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFD 180
SCADILAFAARDS V G Y VP+GRRDG S+ + NLP PT QL +F
Sbjct: 119 SCADILAFAARDSV--VLAGGTPYRVPAGRRDGNTSVASDAMANLPRPTSDVAQLTQSFA 176
Query: 181 RKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQS 240
GLS D+MV LSGAH+IGV+HCSSFS RLY +N + QDP+++ A L CP
Sbjct: 177 THGLSQDDMVILSGAHTIGVAHCSSFSSRLYGYNSSTGQDPALNAAMASRLSRSCP---- 232
Query: 241 QSINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNV 300
Q TV +D + N D YY+ L RG+L SDQTL T +V +NA + ++
Sbjct: 233 QGSANTVAMDDGSENTFDTSYYQNLLAGRGVLASDQTLTADNATAALVAQNAYNMYLFAT 292
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
KF +AMV MG++ VLTGS+G+IR C V N
Sbjct: 293 KFGQAMVKMGAIQVLTGSDGQIRTNCRVAN 322
>UniRef100_Q5U1T0 Class III peroxidase 13 precursor [Oryza sativa]
Length = 347
Score = 295 bits (756), Expect = 9e-79
Identities = 161/321 (50%), Positives = 207/321 (64%), Gaps = 7/321 (2%)
Query: 10 LVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDC 69
LV +++ A S LK GFY TC S EA+V++AV A N G+AAGLIR+HFHDC
Sbjct: 7 LVFFLVAFFPGAAVSAGLKVGFYNETCPSAEALVQQAVAAAFKNNSGVAAGLIRLHFHDC 66
Query: 70 FVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFA 129
FVRGCD SVL++ G +ER N SLRGFEVI+ AKA +EAACP TVSCADILAFA
Sbjct: 67 FVRGCDASVLIN---GSTTERS-AGPNASLRGFEVIDAAKAAVEAACPSTVSCADILAFA 122
Query: 130 ARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEM 189
ARD + G +DY VP+GRRDG VSI + NLPPPT +A++L D F K L++++M
Sbjct: 123 ARDGIKLT--GNVDYQVPAGRRDGNVSIAQDALDNLPPPTATAKELTDKFANKSLTLEDM 180
Query: 190 VTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVL 249
V LSGAH++G S CSSF R+++ N T D + P +A LL++ CP + S T +
Sbjct: 181 VVLSGAHTVGRSFCSSFLDRIWN-NTTAIVDTGLSPGYAALLRALCPSNANASTPITTAI 239
Query: 250 DGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHM 309
D STP LDN YYK L + GL SD L + +V + A + W +FA AMV M
Sbjct: 240 DVSTPATLDNNYYKLLPLDLGLFFSDNQLRVNATMNALVTRFAANETEWKQRFADAMVKM 299
Query: 310 GSLDVLTGSEGEIRERCSVVN 330
G+++VLTG G+IR C+VVN
Sbjct: 300 GNIEVLTGGAGQIRLNCNVVN 320
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.133 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 533,298,604
Number of Sequences: 2790947
Number of extensions: 21595899
Number of successful extensions: 55952
Number of sequences better than 10.0: 604
Number of HSP's better than 10.0 without gapping: 511
Number of HSP's successfully gapped in prelim test: 93
Number of HSP's that attempted gapping in prelim test: 53815
Number of HSP's gapped (non-prelim): 633
length of query: 330
length of database: 848,049,833
effective HSP length: 128
effective length of query: 202
effective length of database: 490,808,617
effective search space: 99143340634
effective search space used: 99143340634
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 75 (33.5 bits)
Medicago: description of AC136973.10


