

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.1 - phase: 0 /pseudo
(381 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
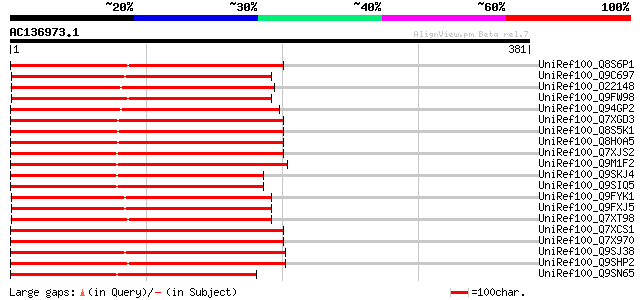
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa] 195 2e-48
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 188 2e-46
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 186 9e-46
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 181 3e-44
UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa] 179 1e-43
UniRef100_Q7XGD3 Putative retroelement [Oryza sativa] 175 2e-42
UniRef100_Q8S5K1 Putative retroelement [Oryza sativa] 175 2e-42
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 174 3e-42
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 173 8e-42
UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis tha... 171 4e-41
UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcrip... 168 3e-40
UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcrip... 168 3e-40
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 166 1e-39
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 166 1e-39
UniRef100_Q7XT98 OSJNBa0052O21.12 protein [Oryza sativa] 165 2e-39
UniRef100_Q7XCS1 Putative reverse transcriptase [Oryza sativa] 165 2e-39
UniRef100_Q7X970 BZIP-like protein [Oryza sativa] 165 2e-39
UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcrip... 165 2e-39
UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcrip... 164 5e-39
UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis tha... 161 3e-38
>UniRef100_Q8S6P1 Putative reverse transcriptase [Oryza sativa]
Length = 1509
Score = 195 bits (495), Expect = 2e-48
Identities = 91/202 (45%), Positives = 132/202 (65%), Gaps = 2/202 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP VGRS+ FS++K+R+WQ+I W K LS+AG+E++IK V Q IP++ M F
Sbjct: 950 KYLGLPVFVGRSRTKIFSYLKERIWQRIQGWKEKLLSRAGKEILIKAVAQVIPTFAMGCF 1009
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
L+ L D+I +M+ +WW + N+ ++WLSW KL++ KN GG+GF+D FN+AML
Sbjct: 1010 ELTKDLCDQISKMIAKYWWSNQEKDNK-MHWLSWNKLTLPKNMGGLGFRDIYIFNLAMLA 1068
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQGW+L PDSL S++ +A+Y+P + K N S+ WRSI V++ G W++G
Sbjct: 1069 KQGWRLIQDPDSLCSRVLRAKYFPLGDCFRPKQTSNVSYTWRSIQKGLRVLQNGMIWRVG 1128
Query: 180 AGFDIPIISEPWIGSGSSIPPV 201
G I I ++PWI G S P+
Sbjct: 1129 DGSKINIWADPWIPRGWSRKPM 1150
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 188 bits (477), Expect = 2e-46
Identities = 89/192 (46%), Positives = 131/192 (67%), Gaps = 2/192 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLGLP +G SK FSF++DR+ +I+ WS+K LSK G+EVMIK V +P YVMS FR
Sbjct: 534 YLGLPESLGGSKTKVFSFVRDRLQSRINGWSAKFLSKGGKEVMIKSVAATLPRYVMSCFR 593
Query: 62 LSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
L + ++ + FWW G S RG++W++W+KL K+DGG+GF++ FN A+L K
Sbjct: 594 LPKAITSKLTSAVAKFWWSSNGDS-RGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAK 652
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L T PDSL +K+FK RY+ SN LD+ ++PS+ WRS++SA+ +V +G ++G+
Sbjct: 653 QLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMISARSLVYKGLIKRVGS 712
Query: 181 GFDIPIISEPWI 192
G I + ++PWI
Sbjct: 713 GASISVWNDPWI 724
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 186 bits (472), Expect = 9e-46
Identities = 90/194 (46%), Positives = 132/194 (67%), Gaps = 2/194 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLGLP SK AT S++KDR+ +K+ W S LS G+E+++K V A+P+Y MS F+
Sbjct: 750 YLGLPESFQGSKVATLSYLKDRLGKKVLGWQSNFLSPGGKEILLKAVAMALPTYTMSCFK 809
Query: 62 LSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
+ T+ +IE +M FWW + RGL+W +W LS K GG+GFK+ AFN+A+LGK
Sbjct: 810 IPKTICQQIESVMAEFWWKNK-KEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGK 868
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W++ T+ DSL++K+FK+RY+ S+ L+A LG PSF W+SI A+V+++QG R IG
Sbjct: 869 QLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFAWKSIYEAQVLIKQGIRAVIGN 928
Query: 181 GFDIPIISEPWIGS 194
G I + ++PWIG+
Sbjct: 929 GETINVWTDPWIGA 942
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 181 bits (459), Expect = 3e-44
Identities = 83/192 (43%), Positives = 127/192 (65%), Gaps = 2/192 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P+ +G + F F+ +R+W++++ W+ + LS+AG E M+K V QAIP+YVMS FR
Sbjct: 765 YLGMPTEIGLATTNFFKFLPERIWKRVNGWTDRPLSRAGMETMLKAVAQAIPNYVMSCFR 824
Query: 62 LSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
+ ++ ++++ + WWG + ++W SW LS K GGMGF++F FN AMLG+
Sbjct: 825 IPVSICEKMKTCIADHWWGFEDGKKK-MHWKSWSWLSTPKFLGGMGFREFTTFNQAMLGR 883
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L T PDSL S++ K RY+PNS+F +A +PSF WRS+L + ++ +G RW +G
Sbjct: 884 QCWRLLTDPDSLCSRVLKGRYFPNSSFWEAAQPKSPSFTWRSLLFGRELLAKGVRWGVGD 943
Query: 181 GFDIPIISEPWI 192
G I I S+ WI
Sbjct: 944 GKTIKIFSDNWI 955
>UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa]
Length = 1833
Score = 179 bits (453), Expect = 1e-43
Identities = 80/199 (40%), Positives = 126/199 (63%), Gaps = 2/199 (1%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP+ GR K F I D++ +++S W+ K +S +EV+IK V QA+P+Y+M +F
Sbjct: 661 KYLGLPTPDGRMKAEQFQPINDKLSKRMSDWNEKYMSSGAKEVLIKSVAQALPTYIMGVF 720
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
++++ D+ M+ +FWWGH + R ++W++WEK+ K+ GG+GF+D FN A+L
Sbjct: 721 KMTDKFCDDYTRMVRNFWWGHD-NGERKVHWIAWEKILYPKSHGGLGFRDMKCFNQALLA 779
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
+Q W+L T PDSL +++FKA+YYPN N +D S VW+ I +++QG W++G
Sbjct: 780 RQAWRLLTSPDSLCARVFKAKYYPNGNLVDTAFPTVSSSVWKGIQHGLDLLKQGTIWRVG 839
Query: 180 AGFDIPIISEPWIGSGSSI 198
G +I I W+ G I
Sbjct: 840 NGHNIKIWRHKWVPHGEHI 858
>UniRef100_Q7XGD3 Putative retroelement [Oryza sativa]
Length = 1694
Score = 175 bits (443), Expect = 2e-42
Identities = 83/202 (41%), Positives = 125/202 (61%), Gaps = 2/202 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLG P+ GR K F +++R+W++I W K LS G+E++IK VLQA+P YVM +F
Sbjct: 1362 KYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGEKFLSTGGKEILIKAVLQAMPVYVMGLF 1421
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ +E+ +++ +FWWG R +W SW+ L K GGMGF+DF FN A+L
Sbjct: 1422 KLPESVCEELTKLVRNFWWG-AEKGMRRTHWKSWDCLIAQKLKGGMGFRDFRLFNQALLA 1480
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
+Q W+L +PDSL ++I KA YYPN + +D G N S WR+I ++++G W+IG
Sbjct: 1481 RQAWRLLDRPDSLCARILKAIYYPNGSLIDTSFGGNASPGWRAIEYGLELLKKGIIWRIG 1540
Query: 180 AGFDIPIISEPWIGSGSSIPPV 201
G + + +PWI S P+
Sbjct: 1541 NGKSVRLWRDPWIPRSYSRRPI 1562
>UniRef100_Q8S5K1 Putative retroelement [Oryza sativa]
Length = 1888
Score = 175 bits (443), Expect = 2e-42
Identities = 83/202 (41%), Positives = 125/202 (61%), Gaps = 2/202 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLG P+ GR K F +++R+W++I W K LS G+E++IK VLQA+P YVM +F
Sbjct: 1319 KYLGFPTPEGRMTKGKFQSLQERLWKRIMLWGEKFLSTGGKEILIKAVLQAMPVYVMGLF 1378
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ +E+ +++ +FWWG R +W SW+ L K GGMGF+DF FN A+L
Sbjct: 1379 KLPESVCEELTKLVRNFWWG-AEKGMRRTHWKSWDCLIAQKLKGGMGFRDFRLFNQALLA 1437
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
+Q W+L +PDSL ++I KA YYPN + +D G N S WR+I ++++G W+IG
Sbjct: 1438 RQAWRLLDRPDSLCARILKAIYYPNGSLIDTSFGGNASPGWRAIEYGLELLKKGIIWRIG 1497
Query: 180 AGFDIPIISEPWIGSGSSIPPV 201
G + + +PWI S P+
Sbjct: 1498 NGKSVRLWRDPWIPRSYSRRPI 1519
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 174 bits (442), Expect = 3e-42
Identities = 81/202 (40%), Positives = 124/202 (61%), Gaps = 2/202 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP+ GR K F +++R+W+KI W LS G+EV+IK V+QAIP YVM IF
Sbjct: 1118 KYLGLPTPGGRMHKGRFQSLRERIWKKILQWGENYLSSGGKEVLIKAVIQAIPVYVMGIF 1177
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ +++ + +FWWG R +W +W+ L+ K+ GG+GFKD FN A+L
Sbjct: 1178 KLPESVCEDLTSLTRNFWWG-AEKGKRKTHWKAWKSLTKSKSLGGLGFKDIRLFNQALLA 1236
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
+Q W+L PDSL +++ KA+YYPN + +D G N S W++I +V++G W+IG
Sbjct: 1237 RQAWRLIDNPDSLCARVLKAKYYPNGSIVDTSFGGNASPGWQAIEHGLELVKKGIIWRIG 1296
Query: 180 AGFDIPIISEPWIGSGSSIPPV 201
G + + +PW+ S P+
Sbjct: 1297 NGRSVRVWQDPWLPRDLSRRPI 1318
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 173 bits (438), Expect = 8e-42
Identities = 82/202 (40%), Positives = 129/202 (63%), Gaps = 2/202 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP + SK+ F FIK+++ +++ W +K LS+ G+EV++K + A+P YVMS F
Sbjct: 727 KYLGLPECLSGSKRDLFGFIKEKLQSRLTGWYAKTLSQGGKEVLLKSIALALPVYVMSCF 786
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L L ++ +M FWW + R ++WLSW++L++ K+ GG GFKD FN A+L
Sbjct: 787 KLPKNLCQKLTTVMMDFWW-NSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLA 845
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W++ + SL S++F++RY+ NS+FL A G PS+ WRSIL + ++ QG R IG
Sbjct: 846 KQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSILFGRELLMQGLRTVIG 905
Query: 180 AGFDIPIISEPWIGSGSSIPPV 201
G + ++ W+ GS+ P+
Sbjct: 906 NGQKTFVWTDKWLHDGSNRRPL 927
>UniRef100_Q9M1F2 Hypothetical protein F9K21.130 [Arabidopsis thaliana]
Length = 851
Score = 171 bits (432), Expect = 4e-41
Identities = 83/205 (40%), Positives = 132/205 (63%), Gaps = 2/205 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP GR KK F++I DRV ++ +SWS+K LS AG+E+++K V A+P Y MS F
Sbjct: 370 KYLGLPEQFGRKKKEMFNYIIDRVKERTASWSAKFLSPAGKEILLKSVALAMPVYAMSCF 429
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ EIE ++ +FWW S+ RG+ W++W++L K +GG+GF+D FN A+L
Sbjct: 430 KLPQGIVSEIESLLMNFWW-EKASNKRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLA 488
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W++ P+SL +++ KARY+ +++ +DAK S+ W S+LS ++R+G R+ IG
Sbjct: 489 KQAWRIIQYPNSLFARVMKARYFKDNSIIDAKTRSQQSYGWSSLLSGIALLRKGTRYVIG 548
Query: 180 AGFDIPIISEPWIGSGSSIPPVGDD 204
G I + + + S P + D+
Sbjct: 549 DGKTIRLGIDNVVDSHPPRPLLTDE 573
>UniRef100_Q9SKJ4 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1750
Score = 168 bits (425), Expect = 3e-40
Identities = 81/187 (43%), Positives = 123/187 (65%), Gaps = 2/187 (1%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP GR KK F +I DRV ++ S+WS++ LS AG+E+M+K V A+P Y MS F
Sbjct: 1131 KYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCF 1190
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ EIE ++ +FWW S+ RG+ W++W++L K +GG+GF+D FN A+L
Sbjct: 1191 KLPKGIVSEIESLLMNFWW-EKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLA 1249
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W+L P+SL +++ KARY+ + + LDAK+ S+ W S+L ++++G R IG
Sbjct: 1250 KQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIG 1309
Query: 180 AGFDIPI 186
G +I I
Sbjct: 1310 DGQNIRI 1316
>UniRef100_Q9SIQ5 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1524
Score = 168 bits (425), Expect = 3e-40
Identities = 81/187 (43%), Positives = 123/187 (65%), Gaps = 2/187 (1%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP GR KK F +I DRV ++ S+WS++ LS AG+E+M+K V A+P Y MS F
Sbjct: 905 KYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWSARFLSPAGKEIMLKSVALAMPVYAMSCF 964
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ EIE ++ +FWW S+ RG+ W++W++L K +GG+GF+D FN A+L
Sbjct: 965 KLPKGIVSEIESLLMNFWW-EKASNQRGIPWVAWKRLQYSKKEGGLGFRDLAKFNDALLA 1023
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W+L P+SL +++ KARY+ + + LDAK+ S+ W S+L ++++G R IG
Sbjct: 1024 KQAWRLIQYPNSLFARVMKARYFKDVSILDAKVRKQQSYGWASLLDGIALLKKGTRHLIG 1083
Query: 180 AGFDIPI 186
G +I I
Sbjct: 1084 DGQNIRI 1090
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 166 bits (419), Expect = 1e-39
Identities = 80/192 (41%), Positives = 123/192 (63%), Gaps = 2/192 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLGLP SK ++KDR+ +K+ W ++CLS+ G+EV++K V A+P + MS F+
Sbjct: 713 YLGLPECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFK 772
Query: 62 LSNTLLDEIEM-MNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
L T + +E M SFWW S R ++W SWE+L + K+ GG+GF+D +FN A+L K
Sbjct: 773 LPITTCENLESAMASFWWDSCDHS-RKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAK 831
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L PD L+S++ K+RY+ ++FLDA L PSF WRSIL + ++ +G + ++G
Sbjct: 832 QAWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGD 891
Query: 181 GFDIPIISEPWI 192
G + + +PWI
Sbjct: 892 GASLFVWIDPWI 903
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 166 bits (419), Expect = 1e-39
Identities = 80/192 (41%), Positives = 123/192 (63%), Gaps = 2/192 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLGLP SK ++KDR+ +K+ W ++CLS+ G+EV++K V A+P + MS F+
Sbjct: 716 YLGLPECFSGSKVDMLHYLKDRLKEKLDVWFTRCLSQGGKEVLLKSVALAMPVFAMSCFK 775
Query: 62 LSNTLLDEIEM-MNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
L T + +E M SFWW S R ++W SWE+L + K+ GG+GF+D +FN A+L K
Sbjct: 776 LPITTCENLESAMASFWWDSCDHS-RKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAK 834
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L PD L+S++ K+RY+ ++FLDA L PSF WRSIL + ++ +G + ++G
Sbjct: 835 QAWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPSFGWRSILFGRELLSKGLQKRVGD 894
Query: 181 GFDIPIISEPWI 192
G + + +PWI
Sbjct: 895 GASLFVWIDPWI 906
>UniRef100_Q7XT98 OSJNBa0052O21.12 protein [Oryza sativa]
Length = 427
Score = 165 bits (418), Expect = 2e-39
Identities = 76/192 (39%), Positives = 124/192 (64%), Gaps = 2/192 (1%)
Query: 2 YLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFR 61
YLG+P+ +GR+ +F F+ +RVW++++ W+ + +S+A +E M+K V QAIP+YVMS F+
Sbjct: 174 YLGMPTEIGRASTGSFHFLPERVWRRVNGWNDRPMSRARKETMLKAVAQAIPTYVMSCFK 233
Query: 62 LSNTLLDEIEM-MNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
L + ++++ + WWG + ++W SWE ++ K+ GGMGF+D FN AML +
Sbjct: 234 LPVSTCEKMKSCILDHWWGFEDGKKK-MHWRSWEWMTTPKSLGGMGFRDLGLFNQAMLAR 292
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
QGW++ T SL +++ K RY+PNS+ +A SF W SIL + ++R+G RW IG
Sbjct: 293 QGWRIVTDLVSLCARVLKGRYFPNSDLWNAPKPTATSFTWPSILFGRDLLRKGVRWGIGN 352
Query: 181 GFDIPIISEPWI 192
G + I+ + WI
Sbjct: 353 GSSVKILKDHWI 364
>UniRef100_Q7XCS1 Putative reverse transcriptase [Oryza sativa]
Length = 791
Score = 165 bits (418), Expect = 2e-39
Identities = 73/201 (36%), Positives = 122/201 (60%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
+YL P+ GR K F ++ ++W+++ W LS G+EV+IK V+QAIP YVM IF
Sbjct: 159 RYLEFPTPEGRMHKGRFQSLQAKIWKRVIQWGENHLSTGGKEVLIKAVIQAIPVYVMGIF 218
Query: 61 RLSNTLLDEIEMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
+L +++D++ + +W + R +W +W+ L+ K+ GG+GF+D+ FN A+L +
Sbjct: 219 KLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWDSLTKPKSLGGLGFRDYWLFNQALLAR 278
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L T PDSL +++ KA+Y+P+ + +D G N S WRSI ++++G W++G
Sbjct: 279 QAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSIEYGLDLLKKGIIWRVGN 338
Query: 181 GFDIPIISEPWIGSGSSIPPV 201
G I I +PW+ S P+
Sbjct: 339 GNSIRIWRDPWLPRDHSRRPI 359
>UniRef100_Q7X970 BZIP-like protein [Oryza sativa]
Length = 2367
Score = 165 bits (418), Expect = 2e-39
Identities = 73/201 (36%), Positives = 122/201 (60%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
+YLG P+ GR K F ++ ++W+++ W LS G+EV+IK V+QAIP YVM IF
Sbjct: 1483 RYLGFPTPEGRMHKGRFQSLQAKIWKRVIQWGENHLSTGGKEVLIKAVIQAIPVYVMGIF 1542
Query: 61 RLSNTLLDEIEMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGK 120
+L +++D++ + +W + R +W +W+ L+ K+ GG+GF+D+ FN A+L +
Sbjct: 1543 KLPESVIDDLTKLTKNFWWDSMNGQRKTHWKAWDSLTKPKSLGGLGFRDYRLFNQALLAR 1602
Query: 121 QGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGA 180
Q W+L T PDSL +++ KA+Y+P+ + +D G N S WRSI ++++G W++G
Sbjct: 1603 QAWRLITYPDSLCARVLKAKYFPHGSLIDTSFGSNSSPAWRSIEYGLDLLKKGIIWRVGN 1662
Query: 181 GFDIPIISEPWIGSGSSIPPV 201
G I I + W+ S P+
Sbjct: 1663 GNSIRIWRDSWLPRDHSRRPI 1683
>UniRef100_Q9SJ38 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1229
Score = 165 bits (417), Expect = 2e-39
Identities = 80/203 (39%), Positives = 127/203 (62%), Gaps = 2/203 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP GR K+ F I D++ QK SW+S+ LS+AG++VM+K VL ++P Y MS F
Sbjct: 644 KYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWASRFLSQAGKQVMLKAVLASMPLYSMSCF 703
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L + L +I+ ++ FWW R +W++W KL+ KN GG+GF+D N ++L
Sbjct: 704 KLPSALCRKIQSLLTRFWWDTKPDV-RKTSWVAWSKLTNPKNAGGLGFRDIERCNDSLLA 762
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
K GW+L P+SL+S+I +Y +S+F++ KL PS WRSI++ + ++++G W I
Sbjct: 763 KLGWRLLNSPESLLSRILLGKYCHSSSFMECKLPSQPSHGWRSIIAGREILKEGLGWLIT 822
Query: 180 AGFDIPIISEPWIGSGSSIPPVG 202
G + I ++PW+ + P+G
Sbjct: 823 NGEKVSIWNDPWLSISKPLVPIG 845
>UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 773
Score = 164 bits (414), Expect = 5e-39
Identities = 80/203 (39%), Positives = 124/203 (60%), Gaps = 2/203 (0%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
+YLGLP VGR+K FSFI + QK+ +W +K LS AG+EV+IK ++ AIP+Y MS F
Sbjct: 204 RYLGLPEFVGRNKTNAFSFIAQTMDQKMDNWYNKLLSPAGKEVLIKSIVTAIPTYSMSCF 263
Query: 61 RLSNTLLDEI-EMMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
L L+ +I M FWW + ++ + W++W KL+ K GG+ +D FN+A+L
Sbjct: 264 LLPMRLIHQITSAMRWFWWSNTKVKHK-IPWVAWSKLNDPKKMGGLAIRDLKDFNIALLA 322
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W++ +P SL++++FKA+Y+P LDAK S+ W+SIL ++ +G ++ G
Sbjct: 323 KQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQSSYAWKSILHGTKLISRGLKYIAG 382
Query: 180 AGFDIPIISEPWIGSGSSIPPVG 202
G +I + + W+ PPVG
Sbjct: 383 NGNNIQLWKDNWLPLNPPRPPVG 405
>UniRef100_Q9SN65 Hypothetical protein F25I24.40 [Arabidopsis thaliana]
Length = 1294
Score = 161 bits (407), Expect = 3e-38
Identities = 76/182 (41%), Positives = 120/182 (65%), Gaps = 2/182 (1%)
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
KYLGLP GR K+ F++I +RV ++ SSWS+K LS AG+E+M+K V ++P Y MS F
Sbjct: 1111 KYLGLPEQFGRKKRDMFNYIIERVKKRTSSWSAKYLSPAGKEIMLKSVAMSMPVYAMSCF 1170
Query: 61 RLSNTLLDEIE-MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLG 119
+L ++ EIE ++ +FWW + R + W++W++L K +GG+GF+D FN A+L
Sbjct: 1171 KLPLNIVSEIEALLMNFWW-EKNAKKREIPWIAWKRLQYSKKEGGLGFRDLAKFNDALLA 1229
Query: 120 KQGWQLQTKPDSLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIG 179
KQ W++ P+SL ++I KARY+ + LDAK S+ W S+L+ V+++G+R+ +G
Sbjct: 1230 KQVWRMINNPNSLFARIMKARYFREDSILDAKRQRYQSYGWTSMLAGLDVIKKGSRFIVG 1289
Query: 180 AG 181
G
Sbjct: 1290 DG 1291
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.339 0.148 0.518
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 622,102,004
Number of Sequences: 2790947
Number of extensions: 24765667
Number of successful extensions: 66670
Number of sequences better than 10.0: 194
Number of HSP's better than 10.0 without gapping: 168
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 66378
Number of HSP's gapped (non-prelim): 223
length of query: 381
length of database: 848,049,833
effective HSP length: 129
effective length of query: 252
effective length of database: 488,017,670
effective search space: 122980452840
effective search space used: 122980452840
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 76 (33.9 bits)
Medicago: description of AC136973.1


