

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.12 - phase: 0 /pseudo
(346 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
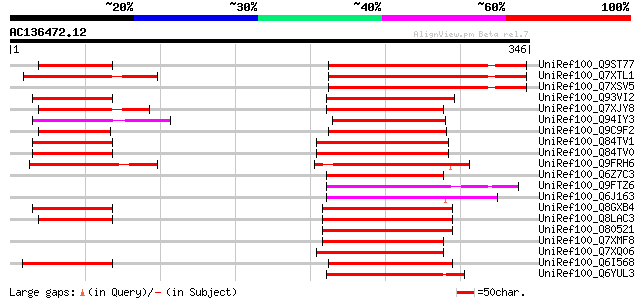
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ST77 CAA303720.1 protein [Oryza sativa] 110 4e-23
UniRef100_Q7XTL1 OSJNBa0070M12.19 protein [Oryza sativa] 110 4e-23
UniRef100_Q7XSV5 OSJNBa0039K24.1 protein [Oryza sativa] 110 4e-23
UniRef100_Q93VI2 Putative MtN21 [Oryza sativa] 108 2e-22
UniRef100_Q7XJY8 OSJNBb0088C09.3 protein [Oryza sativa] 105 1e-21
UniRef100_Q94IY3 Putative CAA303720.1 protein [Oryza sativa] 102 2e-20
UniRef100_Q9C9F2 MtN21-like protein; 91922-89607 [Arabidopsis th... 101 3e-20
UniRef100_Q84TV1 Nodulin-like protein [Gossypium hirsutum] 98 4e-19
UniRef100_Q84TV0 Putative nodulin protein [Gossypium hirsutum] 98 4e-19
UniRef100_Q9FRH6 MtN21 nodulin protein, putative [Arabidopsis th... 96 1e-18
UniRef100_Q6Z7C3 Putative nodulin MtN21 [Oryza sativa] 94 5e-18
UniRef100_Q9FTZ6 Putative nodulin-like protein 5NG4 [Oryza sativa] 92 2e-17
UniRef100_Q6J163 Nodulin-like protein 5NG4 [Pinus taeda] 90 1e-16
UniRef100_Q8GXB4 Putative nodulin protein, N21 [Arabidopsis thal... 89 2e-16
UniRef100_Q8LAC3 Putative nodulin protein, N21 [Arabidopsis thal... 89 2e-16
UniRef100_O80521 F14J9.4 protein [Arabidopsis thaliana] 89 2e-16
UniRef100_Q7XMF8 OSJNBa0028I23.24 protein [Oryza sativa] 88 3e-16
UniRef100_Q7XQ06 OSJNBa0036B21.8 protein [Oryza sativa] 87 5e-16
UniRef100_Q6I568 Hypothetical protein OSJNBb0014K18.13 [Oryza sa... 87 5e-16
UniRef100_Q6YUL3 Nodulin-like protein [Oryza sativa] 87 7e-16
>UniRef100_Q9ST77 CAA303720.1 protein [Oryza sativa]
Length = 376
Score = 110 bits (276), Expect = 4e-23
Identities = 58/132 (43%), Positives = 83/132 (61%), Gaps = 4/132 (3%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
SV Y+G++ SG M+ V+SWCV RGPLFASVF PLML++VA+ G +L+EK+H+G+++GA
Sbjct: 225 SVVYSGVLASGVMLVVLSWCVKRRGPLFASVFNPLMLVVVAVLGSLLLDEKMHVGTLLGA 284
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDN 332
LIV GLYAV+WGK +E + V N + V +VV + +Q + +
Sbjct: 285 ALIVVGLYAVLWGKGRETALEAAKVGDDNDNHHIHVVVVVPPEQAQP----QPHQQAEAD 340
Query: 333 EDSQTDEHEQQS 344
D+ T EQ S
Sbjct: 341 ADATTTACEQPS 352
Score = 66.2 bits (160), Expect = 1e-09
Identities = 30/50 (60%), Positives = 43/50 (86%), Gaps = 1/50 (2%)
Query: 20 MVMVQIAFAGVNVLYKLAVN-DGMSLRIVVAYRFIFATAFIAPIAFILER 68
MV+VQ+ FAGVN+ YKLAV DGM +R++VAYR++FA+A +AP+A+ +ER
Sbjct: 1 MVVVQVVFAGVNIFYKLAVVCDGMDMRVLVAYRYLFASAVLAPLAYFVER 50
>UniRef100_Q7XTL1 OSJNBa0070M12.19 protein [Oryza sativa]
Length = 412
Score = 110 bits (276), Expect = 4e-23
Identities = 58/132 (43%), Positives = 83/132 (61%), Gaps = 4/132 (3%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
SV Y+G++ SG M+ V+SWCV RGPLFASVF PLML++VA+ G +L+EK+H+G+++GA
Sbjct: 261 SVVYSGVLASGVMLVVLSWCVKRRGPLFASVFNPLMLVVVAVLGSLLLDEKMHVGTLLGA 320
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDN 332
LIV GLYAV+WGK +E + V N + V +VV + +Q + +
Sbjct: 321 ALIVVGLYAVLWGKGRETALEAAKVGDDNDNHHIHVVVVVPPEQAQP----QPHQQAEAD 376
Query: 333 EDSQTDEHEQQS 344
D+ T EQ S
Sbjct: 377 ADATTTACEQPS 388
Score = 81.6 bits (200), Expect = 3e-14
Identities = 43/90 (47%), Positives = 61/90 (67%), Gaps = 7/90 (7%)
Query: 10 VVQGLKPTLLMVMVQIAFAGVNVLYKLA-VNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
VVQGLKP MV+VQ+ FAGVN+ YKLA V DGM +R++VAYR++FA+A +AP+A+ +ER
Sbjct: 6 VVQGLKPVAAMVVVQVVFAGVNIFYKLAVVCDGMDMRVLVAYRYLFASAVLAPLAYFVER 65
Query: 69 LQEEEGKDDLDNTVSVISMRFIWGIISSKL 98
K+ T V+ + F+ G+ L
Sbjct: 66 ------KNRTKMTWRVLMLSFVCGLSGGSL 89
>UniRef100_Q7XSV5 OSJNBa0039K24.1 protein [Oryza sativa]
Length = 315
Score = 110 bits (276), Expect = 4e-23
Identities = 58/132 (43%), Positives = 83/132 (61%), Gaps = 4/132 (3%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
SV Y+G++ SG M+ V+SWCV RGPLFASVF PLML++VA+ G +L+EK+H+G+++GA
Sbjct: 164 SVVYSGVLASGVMLVVLSWCVKRRGPLFASVFNPLMLVVVAVLGSLLLDEKMHVGTLLGA 223
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKDN 332
LIV GLYAV+WGK +E + V N + V +VV + +Q + +
Sbjct: 224 ALIVVGLYAVLWGKGRETALEAAKVGDDNDNHHIHVVVVVPPEQAQP----QPHQQAEAD 279
Query: 333 EDSQTDEHEQQS 344
D+ T EQ S
Sbjct: 280 ADATTTACEQPS 291
>UniRef100_Q93VI2 Putative MtN21 [Oryza sativa]
Length = 380
Score = 108 bits (271), Expect = 2e-22
Identities = 49/85 (57%), Positives = 67/85 (78%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
YSV Y G+V SG A++SWC+ +RGPLF S+F+PLML++VA+ G IL+EK+H+GS IG
Sbjct: 254 YSVLYIGVVGSGIAFALMSWCIQVRGPLFVSMFSPLMLVVVAIVGWAILDEKIHVGSAIG 313
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQL 296
+VLIV GLY V+WGK++EM + L
Sbjct: 314 SVLIVAGLYMVLWGKAREMGSPSDL 338
Score = 63.5 bits (153), Expect = 8e-09
Identities = 30/53 (56%), Positives = 40/53 (74%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
PTL MVMVQ+ FAG+NV+ KLA++ GMS +++AYR I A F+AP A+ ER
Sbjct: 7 PTLAMVMVQLGFAGMNVVSKLALDTGMSPYVLIAYRNIIAAVFLAPFAYYFER 59
>UniRef100_Q7XJY8 OSJNBb0088C09.3 protein [Oryza sativa]
Length = 356
Score = 105 bits (263), Expect = 1e-21
Identities = 48/78 (61%), Positives = 62/78 (78%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
YS AYAG++ SG+ ++SWC+ +GPLF SVF+PLML+ VAL ILNE LHLGS++G
Sbjct: 253 YSSAYAGLIASGSAFPLLSWCLRKKGPLFISVFSPLMLIFVALMSSIILNEALHLGSVLG 312
Query: 272 AVLIVCGLYAVVWGKSKE 289
+VLIV GLY V+WGK+KE
Sbjct: 313 SVLIVGGLYMVLWGKAKE 330
Score = 61.6 bits (148), Expect = 3e-08
Identities = 32/74 (43%), Positives = 46/74 (61%), Gaps = 6/74 (8%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEEGKDDLD 79
MV +Q+ F+ + V KLA+NDGM R++VAYRF+FA F+ PIAF+ ER K
Sbjct: 12 MVSLQLLFSALQVFIKLALNDGMDARVLVAYRFMFAATFLCPIAFLRER------KKRPP 65
Query: 80 NTVSVISMRFIWGI 93
T+ V+ F+ G+
Sbjct: 66 LTMKVVLQLFLCGL 79
>UniRef100_Q94IY3 Putative CAA303720.1 protein [Oryza sativa]
Length = 358
Score = 102 bits (254), Expect = 2e-20
Identities = 44/75 (58%), Positives = 61/75 (80%)
Query: 216 YAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGAVLI 275
YAGIV SG + V+SWC+ RGP+F S+F+PLML++VA+ G IL EK+H+GS+IGAV+I
Sbjct: 238 YAGIVASGMVCTVMSWCIQERGPVFVSMFSPLMLIVVAVVGWGILGEKIHVGSVIGAVII 297
Query: 276 VCGLYAVVWGKSKEM 290
V GLY V+WGK +++
Sbjct: 298 VVGLYTVLWGKGRDL 312
Score = 60.5 bits (145), Expect = 7e-08
Identities = 34/92 (36%), Positives = 53/92 (56%), Gaps = 7/92 (7%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEEGK 75
P L MV+VQ+ AG+NV+ KL + GMS +++AYR A AF+APIAF++ER
Sbjct: 11 PLLAMVLVQLGLAGLNVMSKLTMASGMSPYVLLAYRNFIAAAFLAPIAFLVERA------ 64
Query: 76 DDLDNTVSVISMRFIWGIISSKLLLTSPDFNF 107
L+ + + +++ ++S L T P F
Sbjct: 65 -TLNQVLYFVGLKYSSPTVASALNNTLPAVTF 95
>UniRef100_Q9C9F2 MtN21-like protein; 91922-89607 [Arabidopsis thaliana]
Length = 329
Score = 101 bits (252), Expect = 3e-20
Identities = 47/79 (59%), Positives = 63/79 (79%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
++AYA I++SG +VAV +WC+ RGPLF SVF+P+ L++VAL G +L+E LHLGSIIG
Sbjct: 220 TIAYAAILISGMVVAVNAWCIESRGPLFVSVFSPVGLVIVALVGSFLLDETLHLGSIIGT 279
Query: 273 VLIVCGLYAVVWGKSKEMK 291
V+IV LY V+W K+KEMK
Sbjct: 280 VIIVGALYIVLWAKNKEMK 298
Score = 60.5 bits (145), Expect = 7e-08
Identities = 27/48 (56%), Positives = 38/48 (78%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILE 67
MV+VQIA AG+N+ +KLA+ DGM+ ++VAYR +FAT F+ PI FI +
Sbjct: 1 MVVVQIATAGLNIFFKLAMEDGMNPSVLVAYRLLFATLFMIPICFIFQ 48
>UniRef100_Q84TV1 Nodulin-like protein [Gossypium hirsutum]
Length = 376
Score = 97.8 bits (242), Expect = 4e-19
Identities = 42/88 (47%), Positives = 65/88 (73%)
Query: 205 LLCTLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKL 264
L ++ + YAGI+ + V+SWC+ RGPL+ SVF+PL+L++VA+ +L EKL
Sbjct: 244 LSSSMRLIAALYAGIICNAVAFCVMSWCIQKRGPLYVSVFSPLLLVIVAILSWALLREKL 303
Query: 265 HLGSIIGAVLIVCGLYAVVWGKSKEMKK 292
++G+++G++LIV GLYAV+WGK KEMK+
Sbjct: 304 YVGTVVGSLLIVGGLYAVLWGKDKEMKQ 331
Score = 57.8 bits (138), Expect = 4e-07
Identities = 29/53 (54%), Positives = 37/53 (69%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P L V++Q+ +AG+N+ KLA+ GM I+VAYR IFAT IAP AF LER
Sbjct: 7 PFLANVLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLER 59
>UniRef100_Q84TV0 Putative nodulin protein [Gossypium hirsutum]
Length = 374
Score = 97.8 bits (242), Expect = 4e-19
Identities = 42/88 (47%), Positives = 65/88 (73%)
Query: 205 LLCTLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKL 264
L ++ + YAGI+ + V+SWC+ RGPL+ SVF+PL+L++VA+ +L EKL
Sbjct: 244 LSSSMRLIAALYAGIICNAVAFCVMSWCIQKRGPLYVSVFSPLLLVIVAILSWALLREKL 303
Query: 265 HLGSIIGAVLIVCGLYAVVWGKSKEMKK 292
++G+++G++LIV GLYAV+WGK KEMK+
Sbjct: 304 YVGTVVGSLLIVGGLYAVLWGKDKEMKQ 331
Score = 57.8 bits (138), Expect = 4e-07
Identities = 29/53 (54%), Positives = 37/53 (69%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P L V++Q+ +AG+N+ KLA+ GM I+VAYR IFAT IAP AF LER
Sbjct: 7 PFLANVLLQVGYAGMNITSKLALESGMKPLILVAYRQIFATLAIAPFAFFLER 59
>UniRef100_Q9FRH6 MtN21 nodulin protein, putative [Arabidopsis thaliana]
Length = 345
Score = 96.3 bits (238), Expect = 1e-18
Identities = 51/107 (47%), Positives = 71/107 (65%), Gaps = 10/107 (9%)
Query: 204 HLLCTLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEK 263
+LL TL Y+GIVVSG +V +++WC+ +GPLF +VF+P+ L++VAL G L E
Sbjct: 233 NLLATL------YSGIVVSGMVVPLVAWCIATKGPLFVTVFSPIRLVIVALIGSFALEEP 286
Query: 264 LHLGSIIGAVLIVCGLYAVVWGKSKEMKK----KNQLVPSKNPHEFD 306
LHLGSIIGA+++V G+Y VVW K KE K + + +KN E D
Sbjct: 287 LHLGSIIGAMIMVGGVYLVVWCKMKEKKSASTTSDHIETNKNNKELD 333
Score = 60.8 bits (146), Expect = 5e-08
Identities = 30/85 (35%), Positives = 53/85 (62%), Gaps = 6/85 (7%)
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILERLQEEE 73
+K + MV VQ FAG+ +L+K+ V+DG +L+++VAYR FAT F+ P+A I +R + E
Sbjct: 1 MKSVVAMVAVQFIFAGMFILFKITVDDGTNLKVLVAYRLSFATIFMLPLALIFQRKKRPE 60
Query: 74 GKDDLDNTVSVISMRFIWGIISSKL 98
T ++ + F+ G++ + +
Sbjct: 61 ------FTWRLLLLAFVSGLLGAAI 79
>UniRef100_Q6Z7C3 Putative nodulin MtN21 [Oryza sativa]
Length = 386
Score = 94.0 bits (232), Expect = 5e-18
Identities = 41/78 (52%), Positives = 56/78 (71%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG V SG AV +WC+H GP+F +V+ P+ L+VA+ +L E+ HLG IIG
Sbjct: 262 FTILYAGFVASGVAFAVQTWCIHRGGPVFVAVYQPVQTLLVAVMASLLLGEQFHLGGIIG 321
Query: 272 AVLIVCGLYAVVWGKSKE 289
AVLIV GLY V+WGKS+E
Sbjct: 322 AVLIVAGLYLVLWGKSQE 339
Score = 39.3 bits (90), Expect = 0.16
Identities = 18/56 (32%), Positives = 35/56 (62%)
Query: 13 GLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
G + + M+ +Q+ +AG +V+ +LA++ G+S + YR + A +AP A+ LE+
Sbjct: 17 GARLHVAMLALQLGYAGFHVVSRLALDMGVSKLVFPVYRNLIALFLLAPFAYFLEK 72
>UniRef100_Q9FTZ6 Putative nodulin-like protein 5NG4 [Oryza sativa]
Length = 374
Score = 92.0 bits (227), Expect = 2e-17
Identities = 53/128 (41%), Positives = 73/128 (56%), Gaps = 8/128 (6%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG+V SG A+ WC+ GPLF +VF P+ + VA+ IL ++L+ G IIG
Sbjct: 253 FTILYAGLVASGVAFALQIWCIDRGGPLFTAVFQPVQTVAVAVMAAIILGDQLYSGGIIG 312
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPHEFDPVEIVVRSIEKDKSNLNNNNQMVKD 331
AVLIV GLY V+WGKS+E K KN N + PV+ I + L + KD
Sbjct: 313 AVLIVIGLYFVLWGKSEEKKSKN------NNLQDQPVQGGGDDIRRHL--LGQEDASRKD 364
Query: 332 NEDSQTDE 339
E + TDE
Sbjct: 365 EEAAVTDE 372
Score = 37.0 bits (84), Expect = 0.80
Identities = 15/49 (30%), Positives = 31/49 (62%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
++ +Q AG +++ + A+N G+S + + YR + + A +AP A+ LE+
Sbjct: 21 VLALQFLLAGFHIVSRAALNMGISKIVFIVYRNLISLALLAPFAYFLEK 69
>UniRef100_Q6J163 Nodulin-like protein 5NG4 [Pinus taeda]
Length = 410
Score = 89.7 bits (221), Expect = 1e-16
Identities = 43/120 (35%), Positives = 70/120 (57%), Gaps = 6/120 (5%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG V SG +V WC+ GP+F +V+ P+ + VA+ IL E+ +LG I G
Sbjct: 266 FTILYAGFVASGIAFSVQIWCIDRGGPVFVAVYQPVQTIAVAIMASIILGEQFYLGGIFG 325
Query: 272 AVLIVCGLYAVVWGKSKE-----MKKKNQLVPSKNPHEFD-PVEIVVRSIEKDKSNLNNN 325
A+LI+ GLY V+WGKS+E ++ K+ +VP P D +++ S K N +++
Sbjct: 326 AILIIIGLYLVLWGKSEEKRLGLLQAKSSMVPENQPDNMDQSATLIINSSNGIKPNTSSS 385
Score = 38.5 bits (88), Expect = 0.27
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 2/63 (3%)
Query: 8 CNVVQGLKPTL--LMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFI 65
CNV + L M+ +Q +AG +++ + A+N G+S + YR I A I P A+
Sbjct: 9 CNVFMSERVKLHAAMLALQFGYAGFHIVSRAALNMGVSKVVFPVYRNILALMLIGPCAYF 68
Query: 66 LER 68
LE+
Sbjct: 69 LEK 71
>UniRef100_Q8GXB4 Putative nodulin protein, N21 [Arabidopsis thaliana]
Length = 374
Score = 88.6 bits (218), Expect = 2e-16
Identities = 37/87 (42%), Positives = 60/87 (68%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L + S YAG+V S ++SW + +GPL+ SVF+PL+L++VA+ +L EKL+ G+
Sbjct: 249 LRFISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGT 308
Query: 269 IIGAVLIVCGLYAVVWGKSKEMKKKNQ 295
+G+ L+V GLY V+WGK +E+ +K +
Sbjct: 309 FMGSALVVIGLYGVLWGKDREVSEKEE 335
Score = 57.4 bits (137), Expect = 6e-07
Identities = 29/53 (54%), Positives = 37/53 (69%)
Query: 16 PTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
P L MV+VQI +AG+N+ K+A+ GM I+VAYR IFAT P+AF LER
Sbjct: 8 PFLAMVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLER 60
>UniRef100_Q8LAC3 Putative nodulin protein, N21 [Arabidopsis thaliana]
Length = 363
Score = 88.6 bits (218), Expect = 2e-16
Identities = 37/87 (42%), Positives = 60/87 (68%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L + S YAG+V S ++SW + +GPL+ SVF+PL+L++VA+ +L EKL+ G+
Sbjct: 238 LRFISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGT 297
Query: 269 IIGAVLIVCGLYAVVWGKSKEMKKKNQ 295
+G+ L+V GLY V+WGK +E+ +K +
Sbjct: 298 FMGSALVVIGLYGVLWGKDREVSEKEE 324
Score = 54.3 bits (129), Expect = 5e-06
Identities = 27/49 (55%), Positives = 35/49 (71%)
Query: 20 MVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
MV+VQI +AG+N+ K+A+ GM I+VAYR IFAT P+AF LER
Sbjct: 1 MVLVQIGYAGMNITSKMAMEAGMKPLILVAYRQIFATIATFPVAFFLER 49
>UniRef100_O80521 F14J9.4 protein [Arabidopsis thaliana]
Length = 385
Score = 88.6 bits (218), Expect = 2e-16
Identities = 37/87 (42%), Positives = 60/87 (68%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L + S YAG+V S ++SW + +GPL+ SVF+PL+L++VA+ +L EKL+ G+
Sbjct: 260 LRFISALYAGVVASALAFCLMSWAMQRKGPLYVSVFSPLLLVVVAIFSWALLEEKLYTGT 319
Query: 269 IIGAVLIVCGLYAVVWGKSKEMKKKNQ 295
+G+ L+V GLY V+WGK +E+ +K +
Sbjct: 320 FMGSALVVIGLYGVLWGKDREVSEKEE 346
>UniRef100_Q7XMF8 OSJNBa0028I23.24 protein [Oryza sativa]
Length = 325
Score = 88.2 bits (217), Expect = 3e-16
Identities = 39/81 (48%), Positives = 55/81 (67%)
Query: 209 LAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGS 268
L +V + GIV SG +SWCV RGP+F + FTPL+ ++ A +L E+LHLG+
Sbjct: 208 LQIITVLFVGIVGSGIGFLAMSWCVEQRGPVFTTAFTPLIQIIAAAINVIVLREQLHLGT 267
Query: 269 IIGAVLIVCGLYAVVWGKSKE 289
+IG+ L++ GLY V+WGKSKE
Sbjct: 268 VIGSALVIMGLYFVLWGKSKE 288
Score = 35.0 bits (79), Expect = 3.0
Identities = 14/33 (42%), Positives = 23/33 (69%)
Query: 36 LAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
+A+N+GM +++ R + AT F+APIA+ ER
Sbjct: 1 MALNEGMHATVLITLRQLIATLFLAPIAYFRER 33
>UniRef100_Q7XQ06 OSJNBa0036B21.8 protein [Oryza sativa]
Length = 255
Score = 87.4 bits (215), Expect = 5e-16
Identities = 39/85 (45%), Positives = 56/85 (65%)
Query: 205 LLCTLAYYSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKL 264
L L +V + GIV SG +SWCV RGP+F + FTPL+ ++ A +L+E+L
Sbjct: 136 LTTKLQIITVLFVGIVGSGIAFLAMSWCVEQRGPVFTTAFTPLIQIIAAAINVIVLHEQL 195
Query: 265 HLGSIIGAVLIVCGLYAVVWGKSKE 289
HLG +IG+ L++ GLY V+WGK+KE
Sbjct: 196 HLGIVIGSALVIIGLYFVLWGKNKE 220
Score = 33.9 bits (76), Expect = 6.8
Identities = 14/33 (42%), Positives = 22/33 (66%)
Query: 36 LAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
+A N+GM +++ R + AT F+APIA+ ER
Sbjct: 1 MAFNEGMRSTVLITLRQLIATLFLAPIAYFRER 33
>UniRef100_Q6I568 Hypothetical protein OSJNBb0014K18.13 [Oryza sativa]
Length = 420
Score = 87.4 bits (215), Expect = 5e-16
Identities = 43/83 (51%), Positives = 58/83 (69%)
Query: 213 SVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIGA 272
+ AYAGIV S V + RGP+FAS F+PLM+++VA+ G IL E ++LG IIG+
Sbjct: 270 AAAYAGIVTSSIAYYVQGLVMQSRGPVFASAFSPLMMIIVAIMGSFILAENIYLGGIIGS 329
Query: 273 VLIVCGLYAVVWGKSKEMKKKNQ 295
VLIV GLY+V+WGK KE +K +
Sbjct: 330 VLIVAGLYSVLWGKHKENAEKKE 352
Score = 59.3 bits (142), Expect = 2e-07
Identities = 28/60 (46%), Positives = 42/60 (69%)
Query: 9 NVVQGLKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
+++Q KP + MV +Q +AG+NV+ K+++N GMS ++V YR FAT IAP A +LER
Sbjct: 18 SLMQKCKPYVAMVSLQFGYAGMNVITKVSLNHGMSHYVLVVYRHAFATLSIAPFALVLER 77
>UniRef100_Q6YUL3 Nodulin-like protein [Oryza sativa]
Length = 356
Score = 87.0 bits (214), Expect = 7e-16
Identities = 41/92 (44%), Positives = 60/92 (64%), Gaps = 1/92 (1%)
Query: 212 YSVAYAGIVVSGAMVAVISWCVHMRGPLFASVFTPLMLLMVALAGCTILNEKLHLGSIIG 271
+++ YAG+V SG +++ WC+ G LF ++F P+ +MVA+ IL + L+ G IIG
Sbjct: 249 FTILYAGLVASGVALSLQIWCIDRGGALFTAIFQPVQTVMVAIMAAVILGDLLYTGGIIG 308
Query: 272 AVLIVCGLYAVVWGKSKEMKKKNQLVPSKNPH 303
AVLIV GLY V+WGK++E KK N P + H
Sbjct: 309 AVLIVIGLYLVLWGKNEE-KKSNSNQPDLSRH 339
Score = 35.8 bits (81), Expect = 1.8
Identities = 18/55 (32%), Positives = 31/55 (55%)
Query: 14 LKPTLLMVMVQIAFAGVNVLYKLAVNDGMSLRIVVAYRFIFATAFIAPIAFILER 68
LK L ++ +Q AG +++ + A+N G+S + YR A A + P A+ LE+
Sbjct: 8 LKLLLGVLALQCCLAGFHIVSRAALNMGISKIVFTVYRNCIALALLIPFAYFLEK 62
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.334 0.145 0.470
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 554,404,769
Number of Sequences: 2790947
Number of extensions: 23091548
Number of successful extensions: 123407
Number of sequences better than 10.0: 315
Number of HSP's better than 10.0 without gapping: 219
Number of HSP's successfully gapped in prelim test: 96
Number of HSP's that attempted gapping in prelim test: 122976
Number of HSP's gapped (non-prelim): 497
length of query: 346
length of database: 848,049,833
effective HSP length: 128
effective length of query: 218
effective length of database: 490,808,617
effective search space: 106996278506
effective search space used: 106996278506
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC136472.12


