

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.10 - phase: 0 /pseudo
(820 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
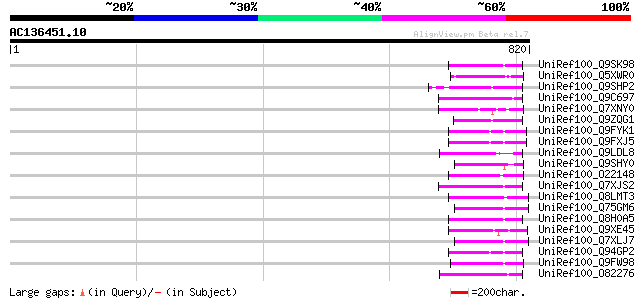
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SK98 Very similar to retrotransposon reverse transcr... 83 4e-14
UniRef100_Q5XWR0 Hypothetical protein [Solanum tuberosum] 79 4e-13
UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcrip... 78 1e-12
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 76 4e-12
UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa] 74 2e-11
UniRef100_Q9ZQG1 Putative non-LTR retroelement reverse transcrip... 74 2e-11
UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana] 73 4e-11
UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana] 73 4e-11
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 72 5e-11
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 72 5e-11
UniRef100_O22148 Putative non-LTR retroelement reverse transcrip... 72 5e-11
UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcrip... 72 7e-11
UniRef100_Q8LMT3 Putative retroelement [Oryza sativa] 72 9e-11
UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcrip... 70 2e-10
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 70 3e-10
UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcri... 69 4e-10
UniRef100_Q7XLJ7 OSJNBa0009K15.13 protein [Oryza sativa] 69 7e-10
UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa] 68 1e-09
UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcrip... 67 2e-09
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 67 3e-09
>UniRef100_Q9SK98 Very similar to retrotransposon reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 82.8 bits (203), Expect = 4e-14
Identities = 47/118 (39%), Positives = 64/118 (53%), Gaps = 3/118 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ ND +KI W+ W +CL KE GGLG R I FN ALL K +W +L + L+ R+
Sbjct: 673 FWWSANDHQRKIHWLSWEKLCLPKEHGGLGFRDIGLFNQALLAKQAWRLLQFPDCLFARL 732
Query: 754 LVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
+ +RY G L D G S WR+I LH + + VGNG++I W D W+
Sbjct: 733 IKSRYFPVGEFLDSDVGSRPSFGWRSI--LHGRDLLCRGLVKRVGNGKSIRVWIDYWL 788
>UniRef100_Q5XWR0 Hypothetical protein [Solanum tuberosum]
Length = 327
Score = 79.3 bits (194), Expect = 4e-13
Identities = 47/117 (40%), Positives = 62/117 (52%), Gaps = 5/117 (4%)
Query: 697 GGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVA 756
G N DHK I V W V L K+ GGLG++ + N LL KW W E LW V+ A
Sbjct: 114 GNNKDHK-IHLVKWAKVTLPKQYGGLGIKDLALHNKCLLLKWHWRYNQEAAGLWKEVIQA 172
Query: 757 RYGEEGGRLLDGGRTA--SAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
+YG + + T+ + WR I L E +FG + VGNGE+I FW D+W+G
Sbjct: 173 KYGSDSHWCSNKVSTSYGTGVWRDIRKLW-EIFFG-DASLMVGNGEHIQFWKDIWLG 227
>UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 773
Score = 77.8 bits (190), Expect = 1e-12
Identities = 52/149 (34%), Positives = 75/149 (49%), Gaps = 13/149 (8%)
Query: 663 SMSCHLCLFTLFPSSRLPQVSSPLLNLF*IIFFLGGNDDHK-KITWVDWNYVCLSKEVGG 721
SMSC L P + Q++S + +F N K KI WV W+ + K++GG
Sbjct: 259 SMSCFL-----LPMRLIHQITSAMR------WFWWSNTKVKHKIPWVAWSKLNDPKKMGG 307
Query: 722 LGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIAT 781
L +R +++FN ALL K SW +L + SL RV A+Y + RLLD T+ + + +
Sbjct: 308 LAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPK-ERLLDAKATSQSSYAWKSI 366
Query: 782 LHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
LH +K GNG NI W D W+
Sbjct: 367 LHGTKLISRGLKYIAGNGNNIQLWKDNWL 395
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 76.3 bits (186), Expect = 4e-12
Identities = 45/134 (33%), Positives = 71/134 (52%), Gaps = 3/134 (2%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
RLP+ + L F+ N D + + W+ W+ +C SK GGLG R + +FNSALL K
Sbjct: 593 RLPKAITSKLTSAVAKFWWSSNGDSRGMHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAK 652
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRLLD-GGRTASAWWRAIATLHSEAWFGTNVKRSV 796
W ++ +SL+ +V RY + L + S WR++ + S + G +KR V
Sbjct: 653 QLWRLITAPDSLFAKVFKGRYFRKSNPLDSIKSYSPSYGWRSMISARSLVYKGL-IKR-V 710
Query: 797 GNGENIYFWSDVWV 810
G+G +I W+D W+
Sbjct: 711 GSGASISVWNDPWI 724
>UniRef100_Q7XNY0 OSJNBb0015N08.11 protein [Oryza sativa]
Length = 1026
Score = 73.6 bits (179), Expect = 2e-11
Identities = 45/139 (32%), Positives = 69/139 (49%), Gaps = 10/139 (7%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
RLP+ ++ F F + +K V+W VC ++ GGLGV + N A+LGK
Sbjct: 615 RLPKGVQERIDYFRKRFLWQEDQGIRKYHLVNWPLVCSPRDQGGLGVLDLEAMNKAMLGK 674
Query: 738 WSWMVLVEKESLWYRVLVARYGEE----GGRLLDGGRTASAWWRAIATLHSEAWFGTNVK 793
W W L +E W ++ A+Y + G RL G +S +W+ + + + F +
Sbjct: 675 WIWR-LENEEGWWQEIIYAKYCSDKPLSGLRLKAG---SSHFWQGVMEVKDD--FFSFCT 728
Query: 794 RSVGNGENIYFWSDVWVGG 812
+ VGNGE FW D W+GG
Sbjct: 729 KIVGNGEKTLFWEDSWLGG 747
>UniRef100_Q9ZQG1 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1138
Score = 73.6 bits (179), Expect = 2e-11
Identities = 38/111 (34%), Positives = 57/111 (51%), Gaps = 2/111 (1%)
Query: 701 DHKKIT-WVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYG 759
+HK+ T WV W +CL+KE GGLG R I FN ALL K SW +L L+ R +RY
Sbjct: 607 EHKRKTHWVSWEKLCLAKESGGLGFRDIESFNQALLAKQSWRILQYPSFLFARFFKSRYF 666
Query: 760 EEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
++ L+ + + L +++ VGNG+++ W D W+
Sbjct: 667 DD-EEFLEADLGVRPSYACCSILFGRELLAKGLRKEVGNGKSLNVWMDPWI 716
>UniRef100_Q9FYK1 F21J9.30 [Arabidopsis thaliana]
Length = 1270
Score = 72.8 bits (177), Expect = 4e-11
Identities = 42/125 (33%), Positives = 64/125 (50%), Gaps = 5/125 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ D +KI W W +CL K+ GGLG R I+ FN ALL K +W +L + L R+
Sbjct: 788 FWWDSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRL 847
Query: 754 LVARYGEEGGRLLDG--GRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
L +RY + LD + S WR+I L +++ VG+G +++ W D W+
Sbjct: 848 LKSRY-FDATDFLDAALSQRPSFGWRSI--LFGRELLSKGLQKRVGDGASLFVWIDPWID 904
Query: 812 GVSFK 816
F+
Sbjct: 905 DNGFR 909
>UniRef100_Q9FXJ5 F5A9.24 [Arabidopsis thaliana]
Length = 1254
Score = 72.8 bits (177), Expect = 4e-11
Identities = 42/125 (33%), Positives = 64/125 (50%), Gaps = 5/125 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ D +KI W W +CL K+ GGLG R I+ FN ALL K +W +L + L R+
Sbjct: 791 FWWDSCDHSRKIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRL 850
Query: 754 LVARYGEEGGRLLDG--GRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
L +RY + LD + S WR+I L +++ VG+G +++ W D W+
Sbjct: 851 LKSRY-FDATDFLDAALSQRPSFGWRSI--LFGRELLSKGLQKRVGDGASLFVWIDPWID 907
Query: 812 GVSFK 816
F+
Sbjct: 908 DNGFR 912
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 72.4 bits (176), Expect = 5e-11
Identities = 44/132 (33%), Positives = 65/132 (48%), Gaps = 22/132 (16%)
Query: 679 LPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKW 738
LPQ + L+ +F LG + + KK+ V W+ VCL K GGLG+R + N AL+ K
Sbjct: 487 LPQSTLEGLDKLARVFLLGSSAEKKKLHLVAWDRVCLPKSEGGLGIRTSKCMNKALVSKV 546
Query: 739 SWMVLVEKESLWYRVLVARYGEEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGN 798
W ++ ++ SLW R+L ++Y ++ G S W VGN
Sbjct: 547 GWRLINDRYSLWARILRSKYRVGLREVVSRG---SRW-------------------VVGN 584
Query: 799 GENIYFWSDVWV 810
G +I FWSD W+
Sbjct: 585 GRDILFWSDNWL 596
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 72.4 bits (176), Expect = 5e-11
Identities = 47/119 (39%), Positives = 59/119 (49%), Gaps = 16/119 (13%)
Query: 703 KKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARY--GE 760
KK V W+ VC K+ GGLGVR + N AL+ K W +L EK SLW VL +Y GE
Sbjct: 191 KKQHLVKWSKVCSPKKEGGLGVRAAKSMNRALISKVGWRLLQEKNSLWTLVLQKKYHVGE 250
Query: 761 -EGGRLLDGGRTASAWWRAIA------TLHSEAWFGTNVKRSVGNGENIYFWSDVWVGG 812
R L + S+ WR+IA H W G+G+ I FW+D WV G
Sbjct: 251 IRDSRWLIPKGSWSSTWRSIAIGLRDVVSHGVGWI-------PGDGQQIRFWTDRWVSG 302
>UniRef100_O22148 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1374
Score = 72.4 bits (176), Expect = 5e-11
Identities = 39/120 (32%), Positives = 61/120 (50%), Gaps = 5/120 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ + + + W W ++ K VGGLG + I FN ALLGK W ++ EK+SL +V
Sbjct: 825 FWWKNKKEGRGLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKV 884
Query: 754 LVARYGEEGGRLLD--GGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVG 811
+RY + L G R + AW + ++ ++ +GNGE I W+D W+G
Sbjct: 885 FKSRYFSKSDPLNAPLGSRPSFAW---KSIYEAQVLIKQGIRAVIGNGETINVWTDPWIG 941
>UniRef100_Q7XJS2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1344
Score = 72.0 bits (175), Expect = 7e-11
Identities = 43/134 (32%), Positives = 65/134 (48%), Gaps = 3/134 (2%)
Query: 678 RLPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGK 737
+LP+ L + F+ +KI W+ W + L K+ GG G + ++ FN ALL K
Sbjct: 787 KLPKNLCQKLTTVMMDFWWNSMQQKRKIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAK 846
Query: 738 WSWMVLVEKESLWYRVLVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSV 796
+W VL EK SL+ RV +RY L G S WR+I L ++ +
Sbjct: 847 QAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPSYAWRSI--LFGRELLMQGLRTVI 904
Query: 797 GNGENIYFWSDVWV 810
GNG+ + W+D W+
Sbjct: 905 GNGQKTFVWTDKWL 918
>UniRef100_Q8LMT3 Putative retroelement [Oryza sativa]
Length = 764
Score = 71.6 bits (174), Expect = 9e-11
Identities = 40/127 (31%), Positives = 61/127 (47%), Gaps = 3/127 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
FF G + +K + W +C KE GGLG R+ N ALL KW + + KE +
Sbjct: 362 FFWEGLEKKRKYHMIKWEALCRPKEFGGLGFIDTRKMNIALLCKWIYRLESGKEDPCCVL 421
Query: 754 LVARYGEEGGRLLDG-GRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVGG 812
L +Y ++GG +S +W+ + + + W VGNG+ FWSDVW+G
Sbjct: 422 LRNKYMKDGGGFFQSKAEESSQFWKGLHEV--KKWMDLGSSYKVGNGKATNFWSDVWIGE 479
Query: 813 VSFKERF 819
K ++
Sbjct: 480 TPLKTQY 486
>UniRef100_Q75GM6 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1614
Score = 70.5 bits (171), Expect = 2e-10
Identities = 39/116 (33%), Positives = 55/116 (46%), Gaps = 2/116 (1%)
Query: 704 KITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGG 763
K V W + K+ GGLG R N+ALL KW + E +SL +L +Y ++GG
Sbjct: 1460 KYHMVKWEDLAFPKDYGGLGFTETRRMNTALLAKWIMKIESEDDSLCIELLRRKYLQDGG 1519
Query: 764 RLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVGGVSFKERF 819
R AS +W+ + L+ W VG+G +I FW DVW G + F
Sbjct: 1520 FFQCKERYASQFWKGL--LNIRRWLSLGSVWQVGDGSHISFWRDVWWGQCPLRTLF 1573
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 69.7 bits (169), Expect = 3e-10
Identities = 40/118 (33%), Positives = 58/118 (48%), Gaps = 3/118 (2%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ G +K W W + SK +GGLG + IR FN ALL + +W ++ +SL RV
Sbjct: 1194 FWWGAEKGKRKTHWKAWKSLTKSKSLGGLGFKDIRLFNQALLARQAWRLIDNPDSLCARV 1253
Query: 754 LVARYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
L A+Y G + G AS W+AI H + +GNG ++ W D W+
Sbjct: 1254 LKAKYYPNGSIVDTSFGGNASPGWQAIE--HGLELVKKGIIWRIGNGRSVRVWQDPWL 1309
>UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 732
Score = 69.3 bits (168), Expect = 4e-10
Identities = 39/129 (30%), Positives = 61/129 (47%), Gaps = 8/129 (6%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F G + +K + W VC + GGLG+R+ ++ N ALL K W ++ + SLW R+
Sbjct: 350 FLWGSSVTQRKQHLISWKRVCKPRSEGGLGIRKAQDMNKALLSKVGWRLIQDYHSLWARI 409
Query: 754 LVARYGEEGGRLLDGGRT-----ASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDV 808
+ Y + R DG T S+ WR++A E + +G+G I FW D
Sbjct: 410 MRCNYRVQDVR--DGAWTKVRSVCSSTWRSVALGMREVVI-PGLSWVIGDGREILFWMDK 466
Query: 809 WVGGVSFKE 817
W+ + E
Sbjct: 467 WLTNIPLAE 475
>UniRef100_Q7XLJ7 OSJNBa0009K15.13 protein [Oryza sativa]
Length = 779
Score = 68.6 bits (166), Expect = 7e-10
Identities = 39/118 (33%), Positives = 58/118 (49%), Gaps = 3/118 (2%)
Query: 703 KKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARY-GEE 761
KK + W +C KE GGLG + N ALL KW + + +S +L +Y G+
Sbjct: 481 KKYHLIKWEALCRPKEDGGLGFIDTKIMNEALLCKWIYKLESGNDSPCCNLLRKKYLGQG 540
Query: 762 GGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVGGVSFKERF 819
GG S +W+++ L + W ++GNGE + FW DVW+G V K +F
Sbjct: 541 GGFFQSSPEGGSQFWKSL--LEVKNWMKLGSAYNIGNGEAVRFWDDVWLGEVPLKIQF 596
>UniRef100_Q94GP2 Putative reverse transcriptase [Oryza sativa]
Length = 1833
Score = 67.8 bits (164), Expect = 1e-09
Identities = 40/119 (33%), Positives = 58/119 (48%), Gaps = 5/119 (4%)
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F+ G ++ +K+ W+ W + K GGLG R ++ FN ALL + +W +L +SL RV
Sbjct: 737 FWWGHDNGERKVHWIAWEKILYPKSHGGLGFRDMKCFNQALLARQAWRLLTSPDSLCARV 796
Query: 754 LVARYGEEGGRLLDGG--RTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWV 810
A+Y G L+D +S+ W+ I H VGNG NI W WV
Sbjct: 797 FKAKY-YPNGNLVDTAFPTVSSSVWKGIQ--HGLDLLKQGTIWRVGNGHNIKIWRHKWV 852
>UniRef100_Q9FW98 Putative non-LTR retroelement reverse transcriptase [Oryza sativa]
Length = 1382
Score = 67.0 bits (162), Expect = 2e-09
Identities = 38/117 (32%), Positives = 58/117 (49%), Gaps = 3/117 (2%)
Query: 697 GGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVA 756
G D KK+ W W+++ K +GG+G R FN A+LG+ W +L + +SL RVL
Sbjct: 843 GFEDGKKKMHWKSWSWLSTPKFLGGMGFREFTTFNQAMLGRQCWRLLTDPDSLCSRVLKG 902
Query: 757 RYGEEGGRL-LDGGRTASAWWRAIATLHSEAWFGTNVKRSVGNGENIYFWSDVWVGG 812
RY ++ S WR++ L V+ VG+G+ I +SD W+ G
Sbjct: 903 RYFPNSSFWEAAQPKSPSFTWRSL--LFGRELLAKGVRWGVGDGKTIKIFSDNWIPG 957
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 66.6 bits (161), Expect = 3e-09
Identities = 44/135 (32%), Positives = 64/135 (46%), Gaps = 4/135 (2%)
Query: 679 LPQVSSPLLNLF*IIFFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKW 738
LP + L+ + F G + KK + W +C K GG+G+R R+ N AL+ K
Sbjct: 673 LPVSTLDTLDRYSRTFLWGSTMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKV 732
Query: 739 SWMVLVEKESLWYRVLVARY---GEEGGRLLDGGRTASAWWRAIATLHSEAWFGTNVKRS 795
W +L +KESLW RV+ +Y G + L S+ WR++A E V
Sbjct: 733 GWRLLQDKESLWARVVRKKYKVGGVQDTSWLKPQPRWSSTWRSVAVGLREV-VVKGVGWV 791
Query: 796 VGNGENIYFWSDVWV 810
G+G I FW D W+
Sbjct: 792 PGDGCTIRFWLDRWL 806
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.361 0.164 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,230,395,492
Number of Sequences: 2790947
Number of extensions: 47934557
Number of successful extensions: 152045
Number of sequences better than 10.0: 141
Number of HSP's better than 10.0 without gapping: 118
Number of HSP's successfully gapped in prelim test: 25
Number of HSP's that attempted gapping in prelim test: 151783
Number of HSP's gapped (non-prelim): 186
length of query: 820
length of database: 848,049,833
effective HSP length: 136
effective length of query: 684
effective length of database: 468,481,041
effective search space: 320441032044
effective search space used: 320441032044
T: 11
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 79 (35.0 bits)
Medicago: description of AC136451.10


