

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136286.2 - phase: 0
(251 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
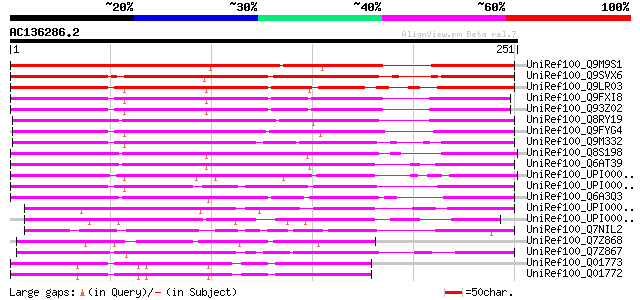
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M9S1 F14L17.20 protein [Arabidopsis thaliana] 238 8e-62
UniRef100_Q9SVX6 Hypothetical protein F15B8.190 [Arabidopsis tha... 231 1e-59
UniRef100_Q9LR03 F10A5.18 [Arabidopsis thaliana] 190 3e-47
UniRef100_Q9FXI8 F6F9.4 protein [Arabidopsis thaliana] 186 4e-46
UniRef100_Q93Z02 At1g19900/F6F9_4 [Arabidopsis thaliana] 186 4e-46
UniRef100_Q8RY19 AT5g19580/T20D1_100 [Arabidopsis thaliana] 184 1e-45
UniRef100_Q9FYG4 F1N21.11 [Arabidopsis thaliana] 184 2e-45
UniRef100_Q9M332 Hypothetical protein F5K20_250 [Arabidopsis tha... 181 1e-44
UniRef100_Q8S198 Glyoxal oxidase-like [Oryza sativa] 178 1e-43
UniRef100_Q6AT39 Putative glyoxal oxidase [Oryza sativa] 163 3e-39
UniRef100_UPI000021A206 UPI000021A206 UniRef100 entry 117 3e-25
UniRef100_UPI000023EF34 UPI000023EF34 UniRef100 entry 110 3e-23
UniRef100_Q6A3Q3 Glyoxal oxidase [Botrytis cinerea] 104 2e-21
UniRef100_UPI000042F76B UPI000042F76B UniRef100 entry 97 3e-19
UniRef100_UPI000042C13A UPI000042C13A UniRef100 entry 88 2e-16
UniRef100_Q7NIL2 Glr2171 protein [Gloeobacter violaceus] 86 1e-15
UniRef100_Q7Z868 Glyoxaloxidase 1 [Ustilago maydis] 84 3e-15
UniRef100_Q7Z867 Glyoxaloxidase 2 [Ustilago maydis] 84 3e-15
UniRef100_Q01773 Glyoxal oxidase precursor [Phanerochaete chryso... 80 5e-14
UniRef100_Q01772 Glyoxal oxidase precursor [Phanerochaete chryso... 80 5e-14
>UniRef100_Q9M9S1 F14L17.20 protein [Arabidopsis thaliana]
Length = 564
Score = 238 bits (608), Expect = 8e-62
Identities = 129/252 (51%), Positives = 164/252 (64%), Gaps = 26/252 (10%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP PRVM DMLLLP G+V+I+NGAANGTAGWE+A N VL P+LY P + +FE+
Sbjct: 337 MEQMPSPRVMSDMLLLPNGDVLIINGAANGTAGWEDATNAVLNPILYLPEEPDQTRRFEI 396
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAK-YPTELSLDAYYPDYLRPELDT 119
L P PRMYHS+++LL DGR+LVGGSNPHR Y+F A+ YPTELSL+AY P YL P+
Sbjct: 397 LTPTRIPRMYHSASLLLSDGRVLVGGSNPHRNYNFTARPYPTELSLEAYLPRYLDPQYAR 456
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVN-RIRVSMVAPSFTTHSFAMNQRLLFL 178
+RP I+ VE+ + L Y F+V+F + + + V +VAPSF+THS AMNQRLL L
Sbjct: 457 VRPTIITVELAGNML-YGQAFAVTFAIPAFGMFDGGVSVRLVAPSFSTHSTAMNQRLLVL 515
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
V + + ++ YKA V GP + VAPPGYYM+FV+H
Sbjct: 516 RVRRVSQ-----------------------LSVFAYKADVDGPTNSYVAPPGYYMMFVVH 552
Query: 239 VGIPSVATWVHV 250
GIPSVA WV +
Sbjct: 553 RGIPSVAVWVKI 564
>UniRef100_Q9SVX6 Hypothetical protein F15B8.190 [Arabidopsis thaliana]
Length = 547
Score = 231 bits (590), Expect = 1e-59
Identities = 124/251 (49%), Positives = 168/251 (66%), Gaps = 29/251 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP+PRVM DMLLLPTG+VII+NGA GTAGWE A +P++ PV+Y+P D+ F +
Sbjct: 324 METMPLPRVMGDMLLLPTGDVIIVNGAGAGTAGWEKARDPIIQPVIYQP-FDH---LFTV 379
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELDT 119
++ S PRMYHSSA+LLPDGR+LVGGSNPH Y+F +YPT+LSL+AY P YL D
Sbjct: 380 MSTPSRPRMYHSSAILLPDGRVLVGGSNPHVYYNFTNVEYPTDLSLEAYSPPYLFFTSDP 439
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLE 179
+RP I+ + LSY+ LF+V F + + V+ + V +VAPSFTTHSFAMNQR++ L+
Sbjct: 440 IRPKILLTS--DKVLSYKRLFNVDFSIAQFLTVDLLSVRIVAPSFTTHSFAMNQRMVILK 497
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
+ ++ +DQ + NS Y+ + GP + +APPGYYM+F++H
Sbjct: 498 LLSV------TRDQ---------------LTNS-YRVSALGPSTAEIAPPGYYMIFLVHA 535
Query: 240 GIPSVATWVHV 250
GIPS A WV +
Sbjct: 536 GIPSSAAWVQI 546
>UniRef100_Q9LR03 F10A5.18 [Arabidopsis thaliana]
Length = 547
Score = 190 bits (483), Expect = 3e-47
Identities = 114/253 (45%), Positives = 156/253 (61%), Gaps = 30/253 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVM DM+LLP GNV+++NG ++GTA WE PVL+P LY P D P +FE
Sbjct: 321 VEKMPRARVMGDMMLLPDGNVLLINGGSSGTAAWELGREPVLHPDLYHP--DKPVGSRFE 378
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELD 118
+ P++ PRMYHS A LL DGRILVGGSNPH Y+F +PTEL L+A+ P YL +
Sbjct: 379 VQNPSTIPRMYHSIATLLRDGRILVGGSNPHAFYNFTGVLFPTELRLEAFSPSYLDTKYS 438
Query: 119 TLRPVIVAVEVVNSTLSYESLFSVSFLLR-EVKDVNRIRVSMVAPSFTTHSFAMNQRLLF 177
+LRP IV +T++Y + + F++ VK + ++V+M+ PSFTTHSF+M+QRLL
Sbjct: 439 SLRPSIVDPR-PQTTVNYGRVLRLRFIVSGRVK--SPVKVTMLFPSFTTHSFSMHQRLL- 494
Query: 178 LEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVI 237
L+ V++ F G S +Y+ VR P S +APPGYYM+FV+
Sbjct: 495 ----VLDHVIS---------FKLGIS--------KIYEVRVRTPSSAILAPPGYYMVFVV 533
Query: 238 HVGIPSVATWVHV 250
+ IPS WV +
Sbjct: 534 NQDIPSEGLWVRL 546
>UniRef100_Q9FXI8 F6F9.4 protein [Arabidopsis thaliana]
Length = 504
Score = 186 bits (473), Expect = 4e-46
Identities = 109/250 (43%), Positives = 147/250 (58%), Gaps = 28/250 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVM DM+ LP G+V+++NG + GTA WE PVL P LY P +NP +FE
Sbjct: 278 VEKMPHARVMGDMIPLPNGDVLLINGGSFGTAAWELGRTPVLAPDLYHP--ENPVGSRFE 335
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELD 118
L P + PRMYHS+A+LL DGR+LVGGSNPH Y++ +PTELSL+A+ P YL+ E
Sbjct: 336 SLRPTTIPRMYHSAAILLRDGRVLVGGSNPHAFYNYTGVLFPTELSLEAFSPVYLQREFS 395
Query: 119 TLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
LRP I++ E S + Y + + F + + +V+MV P+FTTHSFAMNQR+L L
Sbjct: 396 NLRPKIISPE-PQSMIKYGTNLKLKFSVTG-EVTTPAKVTMVFPTFTTHSFAMNQRVLVL 453
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
+ K + +Y+ VR P S N+A PGYYM+FV++
Sbjct: 454 DNVKFTR----------------------KGKSPMYEVQVRTPRSANIAWPGYYMIFVVN 491
Query: 239 VGIPSVATWV 248
IPS WV
Sbjct: 492 QDIPSEGVWV 501
>UniRef100_Q93Z02 At1g19900/F6F9_4 [Arabidopsis thaliana]
Length = 548
Score = 186 bits (473), Expect = 4e-46
Identities = 109/250 (43%), Positives = 147/250 (58%), Gaps = 28/250 (11%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVM DM+ LP G+V+++NG + GTA WE PVL P LY P +NP +FE
Sbjct: 322 VEKMPHARVMGDMIPLPNGDVLLINGGSFGTAAWELGRTPVLAPDLYHP--ENPVGSRFE 379
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELD 118
L P + PRMYHS+A+LL DGR+LVGGSNPH Y++ +PTELSL+A+ P YL+ E
Sbjct: 380 SLRPTTIPRMYHSAAILLRDGRVLVGGSNPHAFYNYTGVLFPTELSLEAFSPVYLQREFS 439
Query: 119 TLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
LRP I++ E S + Y + + F + + +V+MV P+FTTHSFAMNQR+L L
Sbjct: 440 NLRPKIISPE-PQSMIKYGTNLKLKFSVTG-EVTTPAKVTMVFPTFTTHSFAMNQRVLVL 497
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
+ K + +Y+ VR P S N+A PGYYM+FV++
Sbjct: 498 DNVKFTR----------------------KGKSPMYEVQVRTPRSANIAWPGYYMIFVVN 535
Query: 239 VGIPSVATWV 248
IPS WV
Sbjct: 536 QDIPSEGVWV 545
>UniRef100_Q8RY19 AT5g19580/T20D1_100 [Arabidopsis thaliana]
Length = 594
Score = 184 bits (468), Expect = 1e-45
Identities = 96/250 (38%), Positives = 141/250 (56%), Gaps = 27/250 (10%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELL 61
E+MP PR+M D ++LP G+++++NGA G +GW +P P+LYKP +F L
Sbjct: 370 EMMPTPRIMSDTVILPNGDILLVNGAKRGCSGWGYGKDPAFAPLLYKPHAARG-KRFRQL 428
Query: 62 APASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTLR 121
P + PRMYHSSA++LPDG++LVGGSN + Y + ++PTEL ++ + P YL P L +R
Sbjct: 429 KPTTIPRMYHSSAIILPDGKVLVGGSNTNDGYKYNVEFPTELRVEKFSPPYLDPALANIR 488
Query: 122 PVIVAVEVVNSTLSYESLFSVSFLLREV-KDVNRIRVSMVAPSFTTHSFAMNQRLLFLEV 180
P IV + Y F+V L+E ++V+M+AP+FTTHS +MN R+L L V
Sbjct: 489 PKIVTTGTPKQ-VKYGQFFNVKVDLKEKGATKGNLKVTMLAPAFTTHSISMNMRMLILGV 547
Query: 181 TALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVG 240
+ K A + Y PP+ N+APPGYY++F I+ G
Sbjct: 548 NNV------------------------KPAGAGYDIQAVAPPNGNIAPPGYYLIFAIYKG 583
Query: 241 IPSVATWVHV 250
+PS W+ V
Sbjct: 584 VPSTGEWIQV 593
>UniRef100_Q9FYG4 F1N21.11 [Arabidopsis thaliana]
Length = 615
Score = 184 bits (467), Expect = 2e-45
Identities = 98/251 (39%), Positives = 141/251 (56%), Gaps = 28/251 (11%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFEL 60
E MP RVM D ++LP G ++I+NGA G++GW A P P+LYKP + P +F+
Sbjct: 390 ETMPTSRVMSDTVILPNGEILIINGAKRGSSGWHLAKEPNFAPLLYKP--NKPLGQRFKE 447
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTL 120
LAP++ PR+YHS A+ LPDG++LVGGSN + Y F +YPTEL ++ + P YL P L +
Sbjct: 448 LAPSTIPRVYHSIAIALPDGKVLVGGSNTNNGYQFNVEYPTELRIEKFSPPYLDPALANM 507
Query: 121 RPVIVAVEVVNSTLSYESLFSVSFLLREVKDV-NRIRVSMVAPSFTTHSFAMNQRLLFLE 179
RP IV + Y +F V L++ + V+M+APSFTTHS +MN RLL L
Sbjct: 508 RPRIVNT-ATPKQIKYGQMFDVKIELKQQNVAKENVMVTMLAPSFTTHSVSMNMRLLMLG 566
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
+ ++ V ++ PPS +APPGYY+LF ++
Sbjct: 567 INNVKNV-----------------------GGDNHQIQAVAPPSGKLAPPGYYLLFAVYN 603
Query: 240 GIPSVATWVHV 250
G+PSV W+ +
Sbjct: 604 GVPSVGEWIQI 614
>UniRef100_Q9M332 Hypothetical protein F5K20_250 [Arabidopsis thaliana]
Length = 545
Score = 181 bits (460), Expect = 1e-44
Identities = 101/250 (40%), Positives = 148/250 (58%), Gaps = 30/250 (12%)
Query: 2 EVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFEL 60
E MP R+M DM+ LPTG ++I+NGA G+ G+E ++P LYP+LY+P D P ++F
Sbjct: 323 EEMPFGRIMGDMVNLPTGEILIINGAQAGSQGFEMGSDPCLYPLLYRP--DQPIGLRFMT 380
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTL 120
L P + PRMYHS+A LLPDGRIL+ GSNPH Y F A++PTEL ++A+ P+YL P+ L
Sbjct: 381 LNPGTVPRMYHSTANLLPDGRILLAGSNPHYFYKFNAEFPTELRIEAFSPEYLSPDRANL 440
Query: 121 RPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEV 180
RP ++ + + Y +F V F+ + V I+++ + F THSF+ QRL+ L V
Sbjct: 441 RP---EIQEIPQIIRYGEVFDV-FVTVPLPVVGIIQMNWGSAPFATHSFSQGQRLVKLTV 496
Query: 181 TALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVG 240
S+ D G G+ Y+ PP+ V+PPGYYM F ++ G
Sbjct: 497 AP------SVPD------------GVGR-----YRIQCTAPPNGAVSPPGYYMAFAVNQG 533
Query: 241 IPSVATWVHV 250
+PS+A W+ +
Sbjct: 534 VPSIARWIRI 543
>UniRef100_Q8S198 Glyoxal oxidase-like [Oryza sativa]
Length = 624
Score = 178 bits (451), Expect = 1e-43
Identities = 101/253 (39%), Positives = 142/253 (55%), Gaps = 27/253 (10%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
++ MP RVM D+L+LPTG++++LNGAA G +GW +L PVLY P L +F +
Sbjct: 396 LDQMPSGRVMGDVLILPTGDLLMLNGAAKGCSGWGFGRQALLSPVLYSPYLRRG-KRFRV 454
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELDT 119
L P++ PRMYHS++ LLPD +LV GSN + Y+F +PTE+ ++ + P YL P+L
Sbjct: 455 LNPSNIPRMYHSTSALLPDATVLVAGSNTNSAYNFSGVDFPTEVRVERFTPPYLSPQLSP 514
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLL-REVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
RP I A V + Y + F+ F + +V+M AP FTTH ++MNQRLL L
Sbjct: 515 NRPAIDAASVPGDGMRYGARFTFRFTTPAQGVGQGDFKVTMYAPPFTTHGYSMNQRLLIL 574
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
VTA + Q Q + TV PP +APPGYYM++V+
Sbjct: 575 PVTAF-----AAQGQR-------------------HTVTVDAPPKPELAPPGYYMVYVVA 610
Query: 239 VGIPSVATWVHVH 251
G+PS A WV +H
Sbjct: 611 KGVPSKAAWVKMH 623
>UniRef100_Q6AT39 Putative glyoxal oxidase [Oryza sativa]
Length = 622
Score = 163 bits (413), Expect = 3e-39
Identities = 95/252 (37%), Positives = 138/252 (54%), Gaps = 27/252 (10%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
+E MPV RVM D+L+LPTG++++LNGAA G++GW A P+L P+LY P +F
Sbjct: 395 VEAMPVGRVMGDLLVLPTGDLLMLNGAAKGSSGWGFARQPILSPILYSPRHPEG-SRFRP 453
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQ-AKYPTELSLDAYYPDYLRPELDT 119
LA ++ RMYHS++ +LPD +LV G N + Y+F +PTE+ ++ + P YL EL
Sbjct: 454 LAASTVARMYHSTSAVLPDATVLVAGGNTNAAYNFSGVDFPTEVRVERFAPPYLSRELTG 513
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLR-EVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
R VI V + Y + F+ F + +RV+M AP FTTH ++MNQRLL L
Sbjct: 514 NRAVIDVASVPAGGMRYGTKFTFRFHTPVAAVEWGDVRVTMYAPPFTTHGYSMNQRLLVL 573
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
V GF + +Y+ TV P +APPGYY+++V+
Sbjct: 574 PVA-----------------GFSAQ-------GQMYELTVDTPRKPELAPPGYYLVYVVS 609
Query: 239 VGIPSVATWVHV 250
+PS A WV +
Sbjct: 610 KDVPSEAAWVKI 621
>UniRef100_UPI000021A206 UPI000021A206 UniRef100 entry
Length = 445
Score = 117 bits (293), Expect = 3e-25
Identities = 86/258 (33%), Positives = 128/258 (49%), Gaps = 34/258 (13%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
M+ MP RVM + +L+ G V +NGA G G+ A P +LY P P +F
Sbjct: 215 MDAMPDGRVMTEGVLMLDGTVFFVNGAHQGAQGFGVADKPAFTSLLYDPA--QPLGQRFT 272
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHR--LYDFQAKYP--TELSLDAYYPDYLRP 115
A ++ PRMYHS +++L D +L+ GSNP + + + A P TE ++ Y P YL
Sbjct: 273 TAATSTIPRMYHSVSIMLEDATVLIAGSNPVQQPILEVSADTPFATEFRVERYTPPYLSN 332
Query: 116 ELDTLRPVIVAVEVVNSTL--SYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQ 173
LRP+ + + N T + S+ +V F L V ++V++ + THS M
Sbjct: 333 GKQNLRPLNMTLSGTNMTPGPAGSSVLNVRFGLPSAT-VKDVKVALYYNGYVTHSVHMGH 391
Query: 174 RLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYM 233
R+++LE T GF GK A ++ V+ PPS N+ PPGYY+
Sbjct: 392 RMVYLEHT-----------------GFAV----GKTAQNLM---VQPPPSNNITPPGYYI 427
Query: 234 LFVIHVGIPSVATWVHVH 251
LFVI GIPSV + ++
Sbjct: 428 LFVIADGIPSVGQQIMIN 445
Score = 37.0 bits (84), Expect = 0.48
Identities = 26/79 (32%), Positives = 38/79 (47%), Gaps = 2/79 (2%)
Query: 7 PRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDN-PFMKFELLAPAS 65
PR P LP G V + +G+ NG A + N Y +L + G+ + F+K +LL A
Sbjct: 50 PRWYPSAQTLPDGRVFVASGSLNGLAPENASNNNPTYEMLDRNGVSSGQFVKMDLLERAQ 109
Query: 66 TPRMYHSSAVLLPDGRILV 84
MY LL DG + +
Sbjct: 110 PYYMY-PFIHLLNDGNLFI 127
>UniRef100_UPI000023EF34 UPI000023EF34 UniRef100 entry
Length = 901
Score = 110 bits (276), Expect = 3e-23
Identities = 81/251 (32%), Positives = 119/251 (47%), Gaps = 41/251 (16%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
+E MP RVMP+M+ LP G +IL GA G G+ A N L V+Y P + P +
Sbjct: 670 IERMPSKRVMPNMVALPDGRYLILGGAQVGRGGFGLADNANLNAVMYDP--EEPLGQRMT 727
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDT 119
+LA + R+YHS AVLL DG++LV GS+P Q K+P E ++ ++PDYL L
Sbjct: 728 VLANTTIARLYHSEAVLLSDGKVLVSGSDPQD----QGKHPQEKRIEYFWPDYL---LSG 780
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLE 179
+ + T F+++ L E +++RVS++A TH +M QR LF E
Sbjct: 781 ATQPNFTISDRDWTYGESYTFTLTSDLEE--GASKLRVSLMASVGATHGVSMGQRTLFPE 838
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
+ S +V PP+ V+PP +Y +FV+
Sbjct: 839 FSC-----------------------------SGKTCSVTAPPNAFVSPPSWYQMFVLDG 869
Query: 240 GIPSVATWVHV 250
PS A WV +
Sbjct: 870 PTPSHAIWVRI 880
>UniRef100_Q6A3Q3 Glyoxal oxidase [Botrytis cinerea]
Length = 656
Score = 104 bits (259), Expect = 2e-21
Identities = 81/254 (31%), Positives = 118/254 (45%), Gaps = 32/254 (12%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
++ MP R M + LLP G V+ LNG G G+ A +P L +LY P +F
Sbjct: 431 LDAMPEGRGMVEGTLLPDGTVVWLNGGNLGAQGFGLAKDPTLEALLYDPTKAKG-QRFST 489
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQA----KYPTELSLDAYYPDYLRPE 116
LA ++ PR+YHS ++LL DG ++V GSNP + Q Y TE ++ Y P YL +
Sbjct: 490 LATSTIPRLYHSVSLLLLDGTLMVAGSNPVEMPKLQPDAADPYVTEFRVENYVPPYLSGD 549
Query: 117 LDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLL 176
RP V + S + S V+F + V++ F THS M R+L
Sbjct: 550 NAKKRPTNVKLS-SGSFKADGSTLDVTFDC--PAGAKAVTVTLYHGGFVTHSVHMGHRML 606
Query: 177 FLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFV 236
L+ T V + Q K TV PP+ NVAPPG Y++++
Sbjct: 607 HLDNTGF--VAGATQQ----------------------KLTVTRPPNNNVAPPGPYVVYI 642
Query: 237 IHVGIPSVATWVHV 250
+ GIP++ +V V
Sbjct: 643 LVDGIPAMGQFVTV 656
>UniRef100_UPI000042F76B UPI000042F76B UniRef100 entry
Length = 631
Score = 97.4 bits (241), Expect = 3e-19
Identities = 76/264 (28%), Positives = 120/264 (44%), Gaps = 54/264 (20%)
Query: 8 RVMPDMLLLPTGNVIILNGAANGTAGW--------ENAANPVLYPVLYKPGLDNPFMKFE 59
R M + + LP G + +NGA GTAG+ E+ A+ LY Y ++
Sbjct: 335 RSMGNFINLPDGRLFYVNGARTGTAGYGTQDWAVGESYADHPLYQSWYFDPSQPSGQRWS 394
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLY-----DFQAKYPTELSLDAYYPDYLR 114
+S PRMYHSSA LLPDG +++ GSNP+ Y + Y T+ ++ +YPDY
Sbjct: 395 KAGVSSIPRMYHSSASLLPDGTVIISGSNPNADYVDAVNNASYTYFTQYQVEIFYPDY-- 452
Query: 115 PELDTLRP--------VIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTT 166
D ++P + + N TLS LF+V ++N+ R ++ F+T
Sbjct: 453 --ADHVKPTPQGMPSNITYGGDYFNITLSASDLFNVPI------NINKTRAVIMRTGFST 504
Query: 167 HSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNV 226
H+ M QR + LE + F ++ G I + + PP+ +
Sbjct: 505 HTMNMGQRHIELETS------------------FTTTDDGGGILH-----VAQLPPNPGI 541
Query: 227 APPGYYMLFVIHVGIPSVATWVHV 250
PG + F++ GIPS A+WV +
Sbjct: 542 LAPGPALFFIVVDGIPSNASWVMI 565
>UniRef100_UPI000042C13A UPI000042C13A UniRef100 entry
Length = 664
Score = 87.8 bits (216), Expect = 2e-16
Identities = 77/251 (30%), Positives = 115/251 (45%), Gaps = 47/251 (18%)
Query: 8 RVMPDMLLLPTGNVIILNGAANGTAGWENAA---------NPVLYPVLYKPGLD-NPFMK 57
R M + +P G + + NG A GTAG+ + P+ P +Y P
Sbjct: 358 RSMGQFIFMPDGKMWMGNGVAMGTAGYGDEGYSIGQSYGQEPLYQPAIYDPSAPAGSRWS 417
Query: 58 FELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYP---DYLR 114
E L ++ RMYHSSA+LL D ILV GSNP++ F+ K+PT S++ +YP + R
Sbjct: 418 REGLGMSTQERMYHSSAILLADSSILVSGSNPNKDATFE-KWPTSYSVEQWYPLWYNEQR 476
Query: 115 PELDTLRPVIVAVEVVNSTLSY-ESLFSVSFL-LREVKDVNRIRVSMVAPSFTTHSFAMN 172
PE + P S+LSY F+VS+ + + +V ++ F+TH+ M
Sbjct: 477 PEPSSSWP---------SSLSYGGEYFNVSYTPSNSSSNSDNTKVVVIRTGFSTHAMNMG 527
Query: 173 QRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYY 232
QR L L T +D+ GE S + PP+ N+ PG
Sbjct: 528 QRYLELNST-------YTKDEASGEVTLHVS---------------QMPPNANIFQPGPA 565
Query: 233 MLFVIHVGIPS 243
M+F++ GIPS
Sbjct: 566 MIFLVVDGIPS 576
>UniRef100_Q7NIL2 Glr2171 protein [Gloeobacter violaceus]
Length = 749
Score = 85.5 bits (210), Expect = 1e-15
Identities = 74/244 (30%), Positives = 112/244 (45%), Gaps = 54/244 (22%)
Query: 8 RVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTP 67
R+ +LLP V++ G+ E+AA L +Y P + +++ A A+
Sbjct: 328 RLHHSAVLLPDRTVLVCGGSGAD----EDAAKAALQAEIYDPVANT----WKVAATATVA 379
Query: 68 RMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTLRPVIVAV 127
R+YHS A+LLPDGR++ GSNP R + EL L+ + P YL RPVI
Sbjct: 380 RLYHSIALLLPDGRVITAGSNPEREVE-------ELRLEVFSPPYL---FRGPRPVI--- 426
Query: 128 EVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEEVV 187
E V + +Y + +V + D+ I S++ P TH+F M+QRL+ + T
Sbjct: 427 ESVAQSWNYGN--AVEIKTPQATDIRWI--SLIRPGTPTHAFDMDQRLVDVPFTL----- 477
Query: 188 NSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVI-HVGIPSVAT 246
N+ T P N+APPG+YMLF+ + +PSVA
Sbjct: 478 -----------------------NTSGGLTATIPSEPNLAPPGWYMLFITDNDKVPSVAA 514
Query: 247 WVHV 250
WV +
Sbjct: 515 WVQL 518
>UniRef100_Q7Z868 Glyoxaloxidase 1 [Ustilago maydis]
Length = 862
Score = 84.3 bits (207), Expect = 3e-15
Identities = 65/212 (30%), Positives = 98/212 (45%), Gaps = 43/212 (20%)
Query: 4 MPVPRVMPDMLLLPTGNVIILNGAANGTAGWEN------------------AANPVLYPV 45
+P R M + LP G +++LNGA GTAG+ N + +P PV
Sbjct: 367 LPEGRSMGQFIHLPDGTMVVLNGANKGTAGYSNQTWNTIQYNGRTVVTEGLSQDPTYVPV 426
Query: 46 LYKPG------LDNPFMKFELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKY 99
+Y P L N +K P++ R+YHSSA+LLPDG ++V GSNPH+ D
Sbjct: 427 IYDPSKPRGQRLSNANLK-----PSTIARLYHSSAILLPDGSVMVAGSNPHQ--DVALDM 479
Query: 100 PTELSLDAYYPDY-----LRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKD--- 151
PT + A+ Y P D+ RP V NS L S F+++ + D
Sbjct: 480 PTGTTPQAFNTTYEVEKWYPPYWDSPRPYPQGVP--NSVLYGGSPFNITVNGTFMGDSAN 537
Query: 152 --VNRIRVSMVAPSFTTHSFAMNQRLLFLEVT 181
+ +++ F+TH+ M QR ++L+ T
Sbjct: 538 AKAANTKFAIIRTGFSTHAMNMGQRAVYLDYT 569
>UniRef100_Q7Z867 Glyoxaloxidase 2 [Ustilago maydis]
Length = 625
Score = 84.0 bits (206), Expect = 3e-15
Identities = 73/260 (28%), Positives = 110/260 (42%), Gaps = 50/260 (19%)
Query: 4 MPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFM------- 56
+P R M + LP G + NG G AG+ N V PV G DNP
Sbjct: 338 LPERRSMGQFINLPDGTLWFGNGVTTGVAGYSTDPNSVGKPVGESYG-DNPSYQPLVYDP 396
Query: 57 ------KFELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYP 110
+++ + + R+YHSSA LLPD ILV GSNP+ + K+ TE ++ +YP
Sbjct: 397 KASRGNRWKRVGSTNIGRLYHSSATLLPDSSILVAGSNPNADVNHHVKWKTEYRIERWYP 456
Query: 111 DYLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFA 170
D+ D RP + + + S+ SY + L + +V ++ F+TH
Sbjct: 457 DF----YDQPRP---SNDGLPSSFSYGGQ-GFTIRLSSAAQAQKAKVVLIRTGFSTHGMN 508
Query: 171 MNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPG 230
M QR++ L+ T GS L ++ PP+ N+ PG
Sbjct: 509 MGQRMIELKSTHR-----------------GSKLYVAQL-----------PPNPNLFAPG 540
Query: 231 YYMLFVIHVGIPSVATWVHV 250
+ FV+ G+PS V V
Sbjct: 541 PALAFVVVDGVPSQGKMVMV 560
>UniRef100_Q01773 Glyoxal oxidase precursor [Phanerochaete chrysosporium]
Length = 559
Score = 80.1 bits (196), Expect = 5e-14
Identities = 59/195 (30%), Positives = 98/195 (50%), Gaps = 25/195 (12%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTA---------GWENAANPVLYPVLYKPGL 51
+E M R+MP+++ +P G ++I NGA G A G NA +PVL P LY P
Sbjct: 320 VEHMLEARMMPELVHVPNGQILITNGAGTGFAALSAVADPVGNSNADHPVLTPSLYTP-- 377
Query: 52 DNPFMKFELLA--PAST-PRMYHSSAVLLPDGRILVGGSNPHRLYDFQA----KYPTELS 104
D P K A P +T PRMYHS+ L G +GG+NP+ + K+P+EL
Sbjct: 378 DAPLGKRISNAGMPTTTIPRMYHSTVTLTQQGNFFIGGNNPNMNFTPPGTPGIKFPSELR 437
Query: 105 LDAYYPDYLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSF 164
++ P P + RP ++ + L + +V + ++++V+++ F
Sbjct: 438 IETLDP----PFMFRSRPALLTMP---EKLKFGQKVTVPITIPSDLKASKVQVALMDLGF 490
Query: 165 TTHSFAMNQRLLFLE 179
++H+F + RL+F+E
Sbjct: 491 SSHAFHSSARLVFME 505
>UniRef100_Q01772 Glyoxal oxidase precursor [Phanerochaete chrysosporium]
Length = 559
Score = 80.1 bits (196), Expect = 5e-14
Identities = 59/195 (30%), Positives = 98/195 (50%), Gaps = 25/195 (12%)
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTA---------GWENAANPVLYPVLYKPGL 51
+E M R+MP+++ +P G ++I NGA G A G NA +PVL P LY P
Sbjct: 320 VEHMLEARMMPELVHVPNGQILITNGAGTGFAALSAVADPVGNSNADHPVLTPSLYTP-- 377
Query: 52 DNPFMKFELLA--PAST-PRMYHSSAVLLPDGRILVGGSNPHRLYDFQA----KYPTELS 104
D P K A P +T PRMYHS+ L G +GG+NP+ + K+P+EL
Sbjct: 378 DAPLGKRISNAGMPTTTIPRMYHSTVTLTQQGNFFIGGNNPNMNFTPPGTPGIKFPSELR 437
Query: 105 LDAYYPDYLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSF 164
++ P P + RP ++ + L + +V + ++++V+++ F
Sbjct: 438 IETLDP----PFMFRSRPALLTMP---EKLKFGQKVTVPITIPSDLKASKVQVALMDLGF 490
Query: 165 TTHSFAMNQRLLFLE 179
++H+F + RL+F+E
Sbjct: 491 SSHAFHSSARLVFME 505
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 443,977,596
Number of Sequences: 2790947
Number of extensions: 18864573
Number of successful extensions: 58157
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 46
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 57953
Number of HSP's gapped (non-prelim): 120
length of query: 251
length of database: 848,049,833
effective HSP length: 124
effective length of query: 127
effective length of database: 501,972,405
effective search space: 63750495435
effective search space used: 63750495435
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC136286.2


