

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136142.6 + phase: 0
(147 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
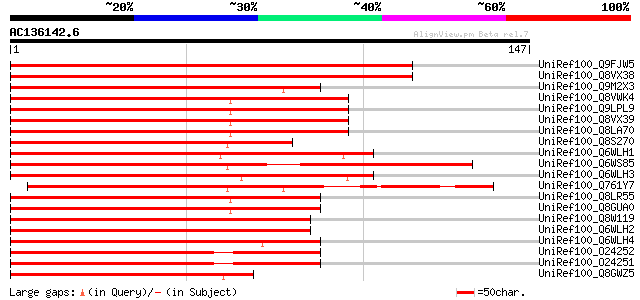
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJW5 Similarity to MYB-like DNA-binding protein [Ara... 152 2e-36
UniRef100_Q8VX38 Telomere repeat binding factor 2 [Arabidopsis t... 152 2e-36
UniRef100_Q9M2X3 MYB-like protein [Arabidopsis thaliana] 132 1e-30
UniRef100_Q8VWK4 Telomere repeat binding factor 1 [Arabidopsis t... 130 8e-30
UniRef100_Q9LPL9 F2J10.16 protein [Arabidopsis thaliana] 130 8e-30
UniRef100_Q8VX39 Telomere repeat binding factor 1 [Arabidopsis t... 130 8e-30
UniRef100_Q8LA70 DNA-binding protein PcMYB1, putative [Arabidops... 130 8e-30
UniRef100_Q8S270 Putative single myb histone 6 [Oryza sativa] 124 4e-28
UniRef100_Q6WLH1 Single myb histone 6 [Zea mays] 120 6e-27
UniRef100_Q6WS85 Single myb histone 1 [Zea mays] 118 3e-26
UniRef100_Q6WLH3 Single myb histone 5 [Zea mays] 114 4e-25
UniRef100_Q761Y7 BRI1-KD interacting protein 127 [Oryza sativa] 107 7e-23
UniRef100_Q8LR55 Histone H1-like protein [Oryza sativa] 105 2e-22
UniRef100_Q8GUA0 MYB28 protein [Oryza sativa] 105 2e-22
UniRef100_Q8W119 Histone H1-like protein [Zea mays] 103 1e-21
UniRef100_Q6WLH2 Single myb histone 4 [Zea mays] 103 1e-21
UniRef100_Q6WLH4 Single myb histone 3 [Zea mays] 100 7e-21
UniRef100_O24252 DNA-binding protein PcMYB1 [Petroselinum crispum] 95 4e-19
UniRef100_O24251 DNA-binding protein PcMYB1 [Petroselinum crispum] 95 4e-19
UniRef100_Q8GWZ5 Putative myb-related DNA-binding protein [Arabi... 88 3e-17
>UniRef100_Q9FJW5 Similarity to MYB-like DNA-binding protein [Arabidopsis thaliana]
Length = 299
Score = 152 bits (383), Expect = 2e-36
Identities = 76/114 (66%), Positives = 86/114 (74%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEEAALKAGV+KHG GKWRTIL D EFS IL++RSNVDLKDKWRNI+VTA
Sbjct: 1 MGAPKQKWTPEEEAALKAGVLKHGTGKWRTILSDTEFSLILKSRSNVDLKDKWRNISVTA 60
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQLALGKVVQREDFLDIKPLTISGGTFQSPK 114
+WGSR+KAKLALK +PP K D+N AL V D KP + G SP+
Sbjct: 61 LWGSRKKAKLALKRTPPGTKQDDNNTALTIVALTNDDERAKPTSPGGSGGGSPR 114
>UniRef100_Q8VX38 Telomere repeat binding factor 2 [Arabidopsis thaliana]
Length = 299
Score = 152 bits (383), Expect = 2e-36
Identities = 76/114 (66%), Positives = 86/114 (74%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEEAALKAGV+KHG GKWRTIL D EFS IL++RSNVDLKDKWRNI+VTA
Sbjct: 1 MGAPKQKWTPEEEAALKAGVLKHGTGKWRTILSDSEFSLILKSRSNVDLKDKWRNISVTA 60
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQLALGKVVQREDFLDIKPLTISGGTFQSPK 114
+WGSR+KAKLALK +PP K D+N AL V D KP + G SP+
Sbjct: 61 LWGSRKKAKLALKRTPPGTKQDDNNTALTIVALTNDDERAKPTSPGGSGGGSPR 114
>UniRef100_Q9M2X3 MYB-like protein [Arabidopsis thaliana]
Length = 295
Score = 132 bits (333), Expect = 1e-30
Identities = 63/89 (70%), Positives = 75/89 (83%), Gaps = 1/89 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPK KWT EEE ALKAGV+KHG GKWRTIL DP +S+IL++RSNVDLKDKWRNI+VTA
Sbjct: 1 MGAPKLKWTPEEETALKAGVLKHGTGKWRTILSDPVYSTILKSRSNVDLKDKWRNISVTA 60
Query: 61 IWGSRQKAKLALKNSP-PAPKTDNNQLAL 88
+WGSR+KAKLALK +P + D+N A+
Sbjct: 61 LWGSRKKAKLALKRTPLSGSRQDDNATAI 89
>UniRef100_Q8VWK4 Telomere repeat binding factor 1 [Arabidopsis thaliana]
Length = 300
Score = 130 bits (326), Expect = 8e-30
Identities = 62/97 (63%), Positives = 77/97 (78%), Gaps = 1/97 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEE+ALK+GV+KHG GKWRTIL DPEFS +L RSNVDLKDKWRN++V A
Sbjct: 1 MGAPKQKWTQEEESALKSGVIKHGPGKWRTILKDPEFSGVLYLRSNVDLKDKWRNMSVMA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLALGKVVQRED 96
WGSR+K++LA+K + PK + N LAL +Q ++
Sbjct: 61 NGWGSREKSRLAVKRTFSLPKQEENSLALTNSLQSDE 97
>UniRef100_Q9LPL9 F2J10.16 protein [Arabidopsis thaliana]
Length = 318
Score = 130 bits (326), Expect = 8e-30
Identities = 62/97 (63%), Positives = 77/97 (78%), Gaps = 1/97 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEE+ALK+GV+KHG GKWRTIL DPEFS +L RSNVDLKDKWRN++V A
Sbjct: 1 MGAPKQKWTQEEESALKSGVIKHGPGKWRTILKDPEFSGVLYLRSNVDLKDKWRNMSVMA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLALGKVVQRED 96
WGSR+K++LA+K + PK + N LAL +Q ++
Sbjct: 61 NGWGSREKSRLAVKRTFSLPKQEENSLALTNSLQSDE 97
>UniRef100_Q8VX39 Telomere repeat binding factor 1 [Arabidopsis thaliana]
Length = 300
Score = 130 bits (326), Expect = 8e-30
Identities = 62/97 (63%), Positives = 77/97 (78%), Gaps = 1/97 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEE+ALK+GV+KHG GKWRTIL DPEFS +L RSNVDLKDKWRN++V A
Sbjct: 1 MGAPKQKWTQEEESALKSGVIKHGPGKWRTILKDPEFSGVLYLRSNVDLKDKWRNMSVMA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLALGKVVQRED 96
WGSR+K++LA+K + PK + N LAL +Q ++
Sbjct: 61 NGWGSREKSRLAVKRTFSLPKQEENSLALTNSLQSDE 97
>UniRef100_Q8LA70 DNA-binding protein PcMYB1, putative [Arabidopsis thaliana]
Length = 300
Score = 130 bits (326), Expect = 8e-30
Identities = 62/97 (63%), Positives = 77/97 (78%), Gaps = 1/97 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT EEE+ALK+GV+KHG GKWRTIL DPEFS +L RSNVDLKDKWRN++V A
Sbjct: 1 MGAPKQKWTQEEESALKSGVIKHGPGKWRTILKDPEFSGVLYLRSNVDLKDKWRNMSVMA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLALGKVVQRED 96
WGSR+K++LA+K + PK + N LAL +Q ++
Sbjct: 61 NGWGSREKSRLAVKRTFSLPKQEENSLALTNSLQSDE 97
>UniRef100_Q8S270 Putative single myb histone 6 [Oryza sativa]
Length = 297
Score = 124 bits (311), Expect = 4e-28
Identities = 59/81 (72%), Positives = 70/81 (85%), Gaps = 1/81 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQ+WTAEEEAALKAGV KHG GKWRTIL DPEF+++LR RSNVDLKDKWRN++VTA
Sbjct: 1 MGAPKQRWTAEEEAALKAGVAKHGTGKWRTILRDPEFTALLRLRSNVDLKDKWRNLSVTA 60
Query: 61 -IWGSRQKAKLALKNSPPAPK 80
+GSR++A++ALK PK
Sbjct: 61 GGYGSRERARVALKGGKRGPK 81
>UniRef100_Q6WLH1 Single myb histone 6 [Zea mays]
Length = 298
Score = 120 bits (301), Expect = 6e-27
Identities = 61/106 (57%), Positives = 77/106 (72%), Gaps = 3/106 (2%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINV-T 59
MGAPKQ+WT+EEEAAL+AG+ +HG GKWRTIL DPEFSS L RSNVDLKDKWRN+NV
Sbjct: 1 MGAPKQRWTSEEEAALRAGIARHGVGKWRTILKDPEFSSTLCYRSNVDLKDKWRNMNVIV 60
Query: 60 AIWGSRQKAKLALKNSPPAPKTDNNQLALGKVVQ--REDFLDIKPL 103
+ SR KAK ALK PK + + +A+ +V ++ +D KP+
Sbjct: 61 STSSSRDKAKSALKRIRTIPKNNEHTMAITRVTSDIDDEIVDEKPI 106
>UniRef100_Q6WS85 Single myb histone 1 [Zea mays]
Length = 299
Score = 118 bits (295), Expect = 3e-26
Identities = 64/132 (48%), Positives = 87/132 (65%), Gaps = 10/132 (7%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQ+WT EEEAALKAGV KHG GKWRTIL D +FS++LR RSNVDLKDKWRN++VTA
Sbjct: 1 MGAPKQRWTPEEEAALKAGVAKHGPGKWRTILRDSDFSALLRLRSNVDLKDKWRNLSVTA 60
Query: 61 -IWGSRQKAKLALKNSPPAPKTDNNQLALGKVVQREDFLDIKPLTISGGTFQSPKPLTIC 119
+GSR+KA++ALK + + K+ +D+K + + T +PL +
Sbjct: 61 GGYGSREKARMALK---------KGRRVVPKLTAEPMDVDVKDMDDAHDTAIDVEPLAMA 111
Query: 120 SGTLQSPNSKEQ 131
+L + S ++
Sbjct: 112 FESLPTEESPDK 123
>UniRef100_Q6WLH3 Single myb histone 5 [Zea mays]
Length = 286
Score = 114 bits (285), Expect = 4e-25
Identities = 55/106 (51%), Positives = 74/106 (68%), Gaps = 3/106 (2%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQ+WT+EEEAAL+AGV +HG G WR IL DPE SS LR RSNVDLKDKWRN+NV
Sbjct: 1 MGAPKQRWTSEEEAALRAGVARHGVGNWRMILNDPELSSTLRYRSNVDLKDKWRNMNVIV 60
Query: 61 IWGS-RQKAKLALKNSPPAPKTDNNQLALGKVVQR--EDFLDIKPL 103
S R + + + + + APK ++ LA+ + ++ +D+KP+
Sbjct: 61 TSSSTRDRGRTSTRRTRAAPKNNDQLLAMSTITSEVDDEIVDVKPI 106
>UniRef100_Q761Y7 BRI1-KD interacting protein 127 [Oryza sativa]
Length = 292
Score = 107 bits (266), Expect = 7e-23
Identities = 63/134 (47%), Positives = 82/134 (61%), Gaps = 17/134 (12%)
Query: 6 QKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA-IWGS 64
QKWTAEEE+ALKAG+ KHGAGKWRTIL DP+FS++LR RSNVDLKDKWRN+NVT G+
Sbjct: 1 QKWTAEEESALKAGIAKHGAGKWRTILKDPDFSNVLRYRSNVDLKDKWRNMNVTVNASGA 60
Query: 65 RQKAKLALKNSP-PAPKTDNNQLALGKVVQREDFLDIKPLTISGGTFQSPKPLTICSGTL 123
R + K + P APK + + + I +T S G +P+ + + T
Sbjct: 61 RDRVKAPVVKKPRSAPKHEGHSTSTA----------IAAVT-SDGDDDVAEPVPLATST- 108
Query: 124 QSPNSKEQVSRFKN 137
+SK +SR N
Sbjct: 109 ---SSKRSLSRLDN 119
>UniRef100_Q8LR55 Histone H1-like protein [Oryza sativa]
Length = 306
Score = 105 bits (263), Expect = 2e-22
Identities = 51/89 (57%), Positives = 65/89 (72%), Gaps = 1/89 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT+EEE AL+ GV+KHG GKWRTI DPEFS +L +RSN+DLKDKWRN++ +A
Sbjct: 1 MGAPKQKWTSEEEEALRRGVLKHGPGKWRTIQKDPEFSPVLSSRSNIDLKDKWRNLSFSA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLAL 88
GSR K K+ P + + ++Q L
Sbjct: 61 SGLGSRDKLKVPRIKGPSSSTSPSSQTPL 89
>UniRef100_Q8GUA0 MYB28 protein [Oryza sativa]
Length = 304
Score = 105 bits (263), Expect = 2e-22
Identities = 51/89 (57%), Positives = 65/89 (72%), Gaps = 1/89 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT+EEE AL+ GV+KHG GKWRTI DPEFS +L +RSN+DLKDKWRN++ +A
Sbjct: 1 MGAPKQKWTSEEEEALRRGVLKHGPGKWRTIQKDPEFSPVLSSRSNIDLKDKWRNLSFSA 60
Query: 61 I-WGSRQKAKLALKNSPPAPKTDNNQLAL 88
GSR K K+ P + + ++Q L
Sbjct: 61 SGLGSRDKLKVPRIKGPSSSTSPSSQTPL 89
>UniRef100_Q8W119 Histone H1-like protein [Zea mays]
Length = 288
Score = 103 bits (256), Expect = 1e-21
Identities = 48/85 (56%), Positives = 62/85 (72%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT+EEE AL+AGV KHGAGKWRTI DPEFS +L +RSN+DLKDKWRN++ +A
Sbjct: 1 MGAPKQKWTSEEEDALRAGVRKHGAGKWRTIQKDPEFSPVLSSRSNIDLKDKWRNLSFSA 60
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQ 85
K ++ P + + ++Q
Sbjct: 61 SGLGSSKLRVPKITGPSSSPSSSSQ 85
>UniRef100_Q6WLH2 Single myb histone 4 [Zea mays]
Length = 288
Score = 103 bits (256), Expect = 1e-21
Identities = 48/85 (56%), Positives = 62/85 (72%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT+EEE AL+AGV KHGAGKWRTI DPEFS +L +RSN+DLKDKWRN++ +A
Sbjct: 1 MGAPKQKWTSEEEDALRAGVRKHGAGKWRTIQKDPEFSPVLSSRSNIDLKDKWRNLSFSA 60
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQ 85
K ++ P + + ++Q
Sbjct: 61 SGLGSSKLRVPKITGPSSSPSSSSQ 85
>UniRef100_Q6WLH4 Single myb histone 3 [Zea mays]
Length = 285
Score = 100 bits (249), Expect = 7e-21
Identities = 49/89 (55%), Positives = 63/89 (70%), Gaps = 1/89 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MGAPKQKWT+EEE AL+ GV KHGAGKWRTI DP+FS IL +RSN+DLKDKWRN++ +A
Sbjct: 1 MGAPKQKWTSEEEDALRRGVRKHGAGKWRTIQKDPQFSPILSSRSNIDLKDKWRNLSFSA 60
Query: 61 IWGSRQKAKL-ALKNSPPAPKTDNNQLAL 88
K ++ + S +P + + L L
Sbjct: 61 SGLGSSKVRVPKITGSSSSPSSSSQALLL 89
>UniRef100_O24252 DNA-binding protein PcMYB1 [Petroselinum crispum]
Length = 307
Score = 94.7 bits (234), Expect = 4e-19
Identities = 47/88 (53%), Positives = 57/88 (64%), Gaps = 5/88 (5%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MG KQKWTAEEE ALKAGV KHG GKW+TIL+DP+F++ L RSN+DLKDKWRN+ +
Sbjct: 12 MGNHKQKWTAEEEEALKAGVKKHGMGKWKTILVDPDFATALTHRSNIDLKDKWRNLGI-- 69
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQLAL 88
S A K+ P N A+
Sbjct: 70 ---SSSAAAQVSKDKSPVLSITNGSAAI 94
>UniRef100_O24251 DNA-binding protein PcMYB1 [Petroselinum crispum]
Length = 307
Score = 94.7 bits (234), Expect = 4e-19
Identities = 47/88 (53%), Positives = 57/88 (64%), Gaps = 5/88 (5%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVTA 60
MG KQKWTAEEE ALKAGV KHG GKW+TIL+DP+F++ L RSN+DLKDKWRN+ +
Sbjct: 12 MGNHKQKWTAEEEEALKAGVKKHGMGKWKTILVDPDFATALTHRSNIDLKDKWRNLGI-- 69
Query: 61 IWGSRQKAKLALKNSPPAPKTDNNQLAL 88
S A K+ P N A+
Sbjct: 70 ---SSSTAAQVSKDKSPVLSITNGSAAI 94
>UniRef100_Q8GWZ5 Putative myb-related DNA-binding protein [Arabidopsis thaliana]
Length = 296
Score = 88.2 bits (217), Expect = 3e-17
Identities = 43/70 (61%), Positives = 51/70 (72%), Gaps = 1/70 (1%)
Query: 1 MGAPKQKWTAEEEAALKAGVVKHGAGKWRTILMDPEFSSILRTRSNVDLKDKWRNINVT- 59
MG K KWTAEEE AL AGV KHG GKW+ IL DPE + L +RSN+DLKDKWRN++V
Sbjct: 1 MGNQKLKWTAEEEEALLAGVRKHGPGKWKNILRDPELAEQLSSRSNIDLKDKWRNLSVAP 60
Query: 60 AIWGSRQKAK 69
I GS+ K +
Sbjct: 61 GIQGSKDKIR 70
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 254,998,822
Number of Sequences: 2790947
Number of extensions: 9754806
Number of successful extensions: 18127
Number of sequences better than 10.0: 600
Number of HSP's better than 10.0 without gapping: 389
Number of HSP's successfully gapped in prelim test: 211
Number of HSP's that attempted gapping in prelim test: 17595
Number of HSP's gapped (non-prelim): 663
length of query: 147
length of database: 848,049,833
effective HSP length: 123
effective length of query: 24
effective length of database: 504,763,352
effective search space: 12114320448
effective search space used: 12114320448
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 67 (30.4 bits)
Medicago: description of AC136142.6


