

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135504.9 + phase: 0 /pseudo
(1303 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
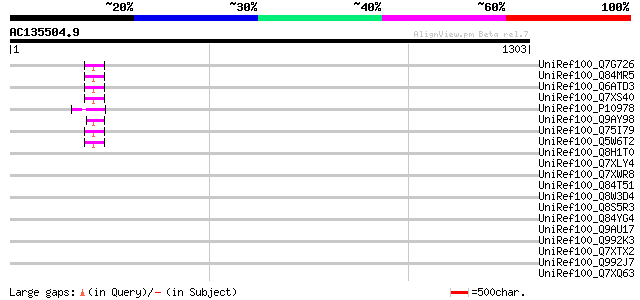
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7G726 Putative polyprotein [Oryza sativa] 53 7e-05
UniRef100_Q84MR5 Putative copia-type pol polyprotein [Oryza sativa] 52 9e-05
UniRef100_Q6ATD3 Putative polyprotein [Oryza sativa] 51 2e-04
UniRef100_Q7XS40 OSJNBa0069D17.3 protein [Oryza sativa] 51 3e-04
UniRef100_P10978 Retrovirus-related Pol polyprotein from transpo... 50 5e-04
UniRef100_Q9AY98 Putative copia-type pol polyprotein [Oryza sativa] 50 6e-04
UniRef100_Q75I79 Putative copia-type pol polyprotein [Oryza sativa] 50 6e-04
UniRef100_Q5W6T2 Putative polyprotein [Oryza sativa] 49 8e-04
UniRef100_Q8H1T0 Gag polyprotein [Triticum aestivum] 47 0.004
UniRef100_Q7XLY4 OSJNBa0042I15.6 protein [Oryza sativa] 47 0.004
UniRef100_Q7XWR8 OSJNBa0091C12.14 protein [Oryza sativa] 46 0.007
UniRef100_Q84T51 Putative copia-type pol polyprotein [Oryza sativa] 46 0.007
UniRef100_Q8W3D4 Putative gag-pol polyprotein [Oryza sativa] 46 0.007
UniRef100_Q8S5R3 Putative copia-type pol polyprotein [Oryza sativa] 46 0.007
UniRef100_Q84YG4 Zinc finger protein [Triticum aestivum] 46 0.009
UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense] 46 0.009
UniRef100_Q992K3 Gag protein [Equine infectious anemia virus] 45 0.011
UniRef100_Q7XTX2 OSJNBa0019K04.26 protein [Oryza sativa] 45 0.011
UniRef100_Q992J7 Gag protein [Equine infectious anemia virus] 45 0.019
UniRef100_Q7XQ63 OSJNBa0072K14.13 protein [Oryza sativa] 44 0.033
>UniRef100_Q7G726 Putative polyprotein [Oryza sativa]
Length = 688
Score = 52.8 bits (125), Expect = 7e-05
Identities = 27/56 (48%), Positives = 30/56 (53%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RKED KG CFNC K GHFIADCP + K K KK
Sbjct: 209 RFKHFLRKSGYGKGRKEDDKGKRQSKRACFNCGKYGHFIADCPKSNEAKVKGGKKK 264
>UniRef100_Q84MR5 Putative copia-type pol polyprotein [Oryza sativa]
Length = 1896
Score = 52.4 bits (124), Expect = 9e-05
Identities = 26/56 (46%), Positives = 30/56 (53%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L R GY RK+D KG CFNC + GHFIADCP + K K KK
Sbjct: 288 RFKHFLRRSGYGKERKDDDKGKRQSKRACFNCGEYGHFIADCPKTNEAKGKGGKKK 343
>UniRef100_Q6ATD3 Putative polyprotein [Oryza sativa]
Length = 1362
Score = 51.2 bits (121), Expect = 2e-04
Identities = 25/56 (44%), Positives = 31/56 (54%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RK+D KG CFNC + GHFIADCP + K+K KK
Sbjct: 289 RFKHFLRKSGYGKGRKDDDKGKRQSKRACFNCGEYGHFIADCPKSNEAKAKGGKKK 344
>UniRef100_Q7XS40 OSJNBa0069D17.3 protein [Oryza sativa]
Length = 1156
Score = 50.8 bits (120), Expect = 3e-04
Identities = 25/56 (44%), Positives = 31/56 (54%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RK+D KG CFNC + GHFIADCP + K+K KK
Sbjct: 197 RFKHFLRKSGYGKGRKDDDKGKRQSKRACFNCGEYGHFIADCPRSNEAKAKGGKKK 252
>UniRef100_P10978 Retrovirus-related Pol polyprotein from transposon TNT 1-94
[Contains: Protease (EC 3.4.23.-); Reverse transcriptase
(EC 2.7.7.49); Endonuclease] [Nicotiana tabacum]
Length = 1328
Score = 50.1 bits (118), Expect = 5e-04
Identities = 26/85 (30%), Positives = 44/85 (51%), Gaps = 7/85 (8%)
Query: 156 IRLEPFNL*YSLFFLGKGKIEKKPLESKFGGRFRYGKVLGRGGYKNSRKEDQKGCFNCKK 215
+R +P N +L G+G+ ++ + YG+ RG KN K + C+NC +
Sbjct: 185 MRKKPENQGQALITEGRGRSYQRSSNN-------YGRSGARGKSKNRSKSRVRNCYNCNQ 237
Query: 216 PGHFIADCPDLQKEKSKSRPKKQSE 240
PGHF DCP+ +K K ++ +K +
Sbjct: 238 PGHFKRDCPNPRKGKGETSGQKNDD 262
>UniRef100_Q9AY98 Putative copia-type pol polyprotein [Oryza sativa]
Length = 1778
Score = 49.7 bits (117), Expect = 6e-04
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 7/51 (13%)
Query: 194 LGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
L + GY RK+D KG CFNC + GHFIADCP + K+K KK
Sbjct: 283 LRKSGYGKGRKDDDKGKRQSRRACFNCGEYGHFIADCPKSNEAKAKGGKKK 333
>UniRef100_Q75I79 Putative copia-type pol polyprotein [Oryza sativa]
Length = 1177
Score = 49.7 bits (117), Expect = 6e-04
Identities = 24/56 (42%), Positives = 30/56 (52%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RK+D KG CFNC + GHFIADCP + K+ KK
Sbjct: 97 RFKHFLRKSGYGKGRKDDDKGKRQSKRACFNCGEYGHFIADCPKSNESKANGGKKK 152
>UniRef100_Q5W6T2 Putative polyprotein [Oryza sativa]
Length = 2049
Score = 49.3 bits (116), Expect = 8e-04
Identities = 25/56 (44%), Positives = 30/56 (52%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RK D KG CFNC + GHFIADCP + K+K KK
Sbjct: 270 RFKHFLRKSGYGKGRKVDDKGKRQSKRACFNCGEFGHFIADCPKSNEAKAKGGKKK 325
>UniRef100_Q8H1T0 Gag polyprotein [Triticum aestivum]
Length = 196
Score = 47.0 bits (110), Expect = 0.004
Identities = 24/60 (40%), Positives = 33/60 (55%), Gaps = 3/60 (5%)
Query: 183 KFGGRFRYGKVLGRGGYKN---SRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKKQS 239
KF R R+GK ++ SR ++ C CKKPGH+I DCP +KE +K + K S
Sbjct: 80 KFSRRGRFGKSSRNDDVRSDSSSRDYKKRTCHKCKKPGHYIQDCPLWEKESNKKKYKDYS 139
>UniRef100_Q7XLY4 OSJNBa0042I15.6 protein [Oryza sativa]
Length = 1510
Score = 47.0 bits (110), Expect = 0.004
Identities = 24/56 (42%), Positives = 30/56 (52%), Gaps = 7/56 (12%)
Query: 189 RYGKVLGRGGYKNSRKEDQKG-------CFNCKKPGHFIADCPDLQKEKSKSRPKK 237
R+ L + GY RK+D KG CFNC + GHFIAD P + K+K KK
Sbjct: 188 RFKHFLRKSGYGKGRKDDDKGKKQSKRACFNCGEYGHFIADFPKSNEAKAKGGKKK 243
>UniRef100_Q7XWR8 OSJNBa0091C12.14 protein [Oryza sativa]
Length = 393
Score = 46.2 bits (108), Expect = 0.007
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 11/63 (17%)
Query: 189 RYGKVLGRGGY-----------KNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKK 237
++GK + R G+ K+S + + C+ CK+PGHFIADCP L+ KK
Sbjct: 279 KFGKFMRRSGFFKGNSSKHHSSKSSGRHSARVCYVCKEPGHFIADCPHLKDGSHIKEDKK 338
Query: 238 QSE 240
+ E
Sbjct: 339 KDE 341
>UniRef100_Q84T51 Putative copia-type pol polyprotein [Oryza sativa]
Length = 2027
Score = 46.2 bits (108), Expect = 0.007
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 11/63 (17%)
Query: 189 RYGKVLGRGGY-----------KNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKK 237
++GK + R G+ K+S + + C+ CK+PGHFIADCP L+ KK
Sbjct: 279 KFGKFMRRSGFFKGGSSKHYSNKSSGRHSARMCYVCKEPGHFIADCPHLKDGSHIKEDKK 338
Query: 238 QSE 240
+ E
Sbjct: 339 KGE 341
>UniRef100_Q8W3D4 Putative gag-pol polyprotein [Oryza sativa]
Length = 491
Score = 46.2 bits (108), Expect = 0.007
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 11/63 (17%)
Query: 189 RYGKVLGRGGY-----------KNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKK 237
++GK + R G+ K+S + + C+ CK+PGHFIADCP L+ KK
Sbjct: 210 KFGKFMRRSGFFKGGSSKHYSNKSSGRHSARVCYVCKEPGHFIADCPHLKDGSHIKEDKK 269
Query: 238 QSE 240
+ E
Sbjct: 270 KDE 272
>UniRef100_Q8S5R3 Putative copia-type pol polyprotein [Oryza sativa]
Length = 1866
Score = 46.2 bits (108), Expect = 0.007
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 11/63 (17%)
Query: 189 RYGKVLGRGGY-----------KNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKK 237
++GK + R G+ K+S + + C+ CK+PGHFIADCP L+ KK
Sbjct: 144 KFGKFMRRSGFFKGGSSKHYSNKSSGRHSARVCYVCKEPGHFIADCPHLKDGSHIKEDKK 203
Query: 238 QSE 240
+ E
Sbjct: 204 KDE 206
>UniRef100_Q84YG4 Zinc finger protein [Triticum aestivum]
Length = 269
Score = 45.8 bits (107), Expect = 0.009
Identities = 29/90 (32%), Positives = 39/90 (43%), Gaps = 9/90 (10%)
Query: 183 KFGGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPDLQKE--------KSKSR 234
KF R +GK L R +S ++ C CKKPGH+I DCP +KE S
Sbjct: 80 KFSRRGHFGKSL-RSNDSSSSDYKKRLCHKCKKPGHYIQDCPQWEKESKTKYKDYSSDDA 138
Query: 235 PKKQSELVKTAGVPKGKRETKVMVPRQRVF 264
KK+S ++ K K + R F
Sbjct: 139 KKKKSSKYSSSKSSKSSSHKKSSSKKARAF 168
>UniRef100_Q9AU17 Polyprotein-like [Lycopersicon chilense]
Length = 1328
Score = 45.8 bits (107), Expect = 0.009
Identities = 19/51 (37%), Positives = 29/51 (56%)
Query: 190 YGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKKQSE 240
YG+ RG K K + C+NC +PGHF DCP+ ++ K +S +K +
Sbjct: 213 YGRSGARGKSKVRSKSKARNCYNCDQPGHFKRDCPNPKRGKGESSGQKNDD 263
>UniRef100_Q992K3 Gag protein [Equine infectious anemia virus]
Length = 400
Score = 45.4 bits (106), Expect = 0.011
Identities = 23/64 (35%), Positives = 38/64 (58%), Gaps = 3/64 (4%)
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADC---PDLQKEKSKSRPKKQSELVKTAGVPKGK 251
G+ G+ +S+ + K CF CK+PGHF C P K+ ++ RP+KQ+ V+ + K +
Sbjct: 303 GKPGHFSSQCKAPKVCFKCKQPGHFSKQCRNAPKNGKQGAQGRPQKQTFPVQKESMNKTQ 362
Query: 252 RETK 255
+E K
Sbjct: 363 KEEK 366
>UniRef100_Q7XTX2 OSJNBa0019K04.26 protein [Oryza sativa]
Length = 559
Score = 45.4 bits (106), Expect = 0.011
Identities = 18/43 (41%), Positives = 26/43 (59%)
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKK 237
GR +++ ++ CFNC + GHFIADCP + K+K KK
Sbjct: 290 GRKDDDKGKRQSERACFNCGEYGHFIADCPKTNEAKAKGDKKK 332
>UniRef100_Q992J7 Gag protein [Equine infectious anemia virus]
Length = 400
Score = 44.7 bits (104), Expect = 0.019
Identities = 22/64 (34%), Positives = 38/64 (59%), Gaps = 3/64 (4%)
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADC---PDLQKEKSKSRPKKQSELVKTAGVPKGK 251
G+ G+ +S+ + K CF CK+PGHF C P ++ ++ RP+KQ+ V+ + K +
Sbjct: 303 GKPGHFSSQCKAPKICFKCKQPGHFSKQCRNAPKNGRQGAQGRPQKQTFPVQKGSMDKTQ 362
Query: 252 RETK 255
+E K
Sbjct: 363 KEEK 366
>UniRef100_Q7XQ63 OSJNBa0072K14.13 protein [Oryza sativa]
Length = 901
Score = 43.9 bits (102), Expect = 0.033
Identities = 17/41 (41%), Positives = 26/41 (62%)
Query: 200 KNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKSRPKKQSE 240
K+S + + C+ CK+P HFIADCP + +EK K K + +
Sbjct: 216 KSSGRHSARVCYMCKEPEHFIADCPHINEEKKKIEKKVEKK 256
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.382 0.170 0.717
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,532,064,328
Number of Sequences: 2790947
Number of extensions: 50127679
Number of successful extensions: 307511
Number of sequences better than 10.0: 324
Number of HSP's better than 10.0 without gapping: 233
Number of HSP's successfully gapped in prelim test: 98
Number of HSP's that attempted gapping in prelim test: 306918
Number of HSP's gapped (non-prelim): 642
length of query: 1303
length of database: 848,049,833
effective HSP length: 139
effective length of query: 1164
effective length of database: 460,108,200
effective search space: 535565944800
effective search space used: 535565944800
T: 11
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 81 (35.8 bits)
Medicago: description of AC135504.9


