

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135504.16 - phase: 0 /pseudo
(343 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
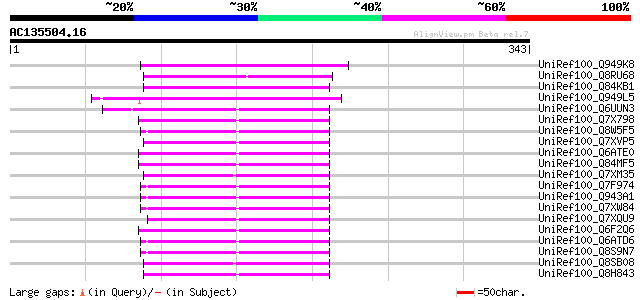
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q949K8 Gag polyprotein [Cicer arietinum] 102 2e-20
UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type... 95 2e-18
UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo] 92 3e-17
UniRef100_Q949L5 Gag polyprotein [Cicer arietinum] 92 3e-17
UniRef100_Q6UUN3 Putative gag-pol polyprotein [Oryza sativa] 91 6e-17
UniRef100_Q7X798 OSJNBb0108J11.19 protein [Oryza sativa] 90 8e-17
UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa] 89 2e-16
UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa] 87 7e-16
UniRef100_Q6ATE0 Putative polyprotein [Oryza sativa] 87 7e-16
UniRef100_Q84MF5 Putative retrotransposon gag protein [Oryza sat... 87 9e-16
UniRef100_Q7XM35 OSJNBb0022P19.7 protein [Oryza sativa] 87 9e-16
UniRef100_Q7F974 OSJNBb0003A12.3 protein [Oryza sativa] 87 9e-16
UniRef100_Q943A1 Putative polyprotein [Oryza sativa] 86 1e-15
UniRef100_Q7XW84 OSJNBa0019J05.10 protein [Oryza sativa] 86 1e-15
UniRef100_Q7XQU9 OSJNBa0086B14.15 protein [Oryza sativa] 86 1e-15
UniRef100_Q6F2Q6 Putative polyprotein [Oryza sativa] 86 1e-15
UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa] 86 1e-15
UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa] 86 1e-15
UniRef100_Q8SB08 Putative polyprotein [Oryza sativa] 86 2e-15
UniRef100_Q8H843 Putative polyprotein [Oryza sativa] 85 3e-15
>UniRef100_Q949K8 Gag polyprotein [Cicer arietinum]
Length = 277
Score = 102 bits (253), Expect = 2e-20
Identities = 51/138 (36%), Positives = 80/138 (57%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
R L F R +PP FKG + +W++EVE+IF ++ C KV + T+ML +A+ WW
Sbjct: 54 RGLVDFRRYNPPKFKGDEGSEKADQWIQEVEKIFDMINCQAGVKVSYATYMLLGDAEYWW 113
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
S ++ V W F+R+FL++YFP R K +FL+L G ++V +YAAKF L
Sbjct: 114 RSARLLMGAAHEEVNWESFKRKFLDKYFPMSTRTKLGDDFLKLHQGSLTVGEYAAKFESL 173
Query: 207 AKFYPYYTAEIVEFSKCY 224
++ + ++ EI E C+
Sbjct: 174 SRHFRFFREEIDEPFMCH 191
>UniRef100_Q8RU68 Putative 22 kDa kafirin cluster; Ty3-Gypsy type [Oryza sativa]
Length = 1230
Score = 95.1 bits (235), Expect = 2e-18
Identities = 45/125 (36%), Positives = 71/125 (56%), Gaps = 1/125 (0%)
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVS 148
L F + PPTF G +P ++W+ +E+ F M C++ +K+ + T+ML A +WW +
Sbjct: 71 LGEFQKLKPPTFSGTANPLEAEEWIVAMEKSFEAMGCTDKEKIIYATYMLQSSAFEWWDA 130
Query: 149 LLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
+ +TW +F+ F +YFPE V+ E EFLELK G+ SV +Y +F LA+
Sbjct: 131 HKKSYSER-IFITWELFKEAFYKKYFPESVKRMKEKEFLELKQGNKSVAEYEIEFSRLAR 189
Query: 209 FYPYY 213
F P +
Sbjct: 190 FAPEF 194
>UniRef100_Q84KB1 Gag-protease polyprotein [Cucumis melo]
Length = 429
Score = 91.7 bits (226), Expect = 3e-17
Identities = 47/124 (37%), Positives = 67/124 (53%), Gaps = 1/124 (0%)
Query: 89 LETFLRNHPPTFKGRY-DPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWV 147
L F + +P TF G DP Q WL +E IFR M+C E QKV+ ML + WW
Sbjct: 60 LRDFRKYNPTTFDGSLEDPTRAQMWLSSLETIFRYMKCPEDQKVQCAVFMLTDRGTAWWE 119
Query: 148 SLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELA 207
+ +L D + +TW F+ F ++F +R EFL L+ GDM+V +Y A+F L+
Sbjct: 120 TTERMLGGDVSQITWQQFKESFYAKFFSASLRDAKRQEFLNLEQGDMTVEQYDAEFDMLS 179
Query: 208 KFYP 211
+F P
Sbjct: 180 RFAP 183
>UniRef100_Q949L5 Gag polyprotein [Cicer arietinum]
Length = 209
Score = 91.7 bits (226), Expect = 3e-17
Identities = 59/176 (33%), Positives = 90/176 (50%), Gaps = 12/176 (6%)
Query: 55 AGRNDPAIAAALEAVA*AVGQQPNAAAGND-----------GVRMLETFLRNHPPTFKGR 103
A RND +A A+ +A +V Q A D R LE F R +PP FKG
Sbjct: 13 ANRNDQ-MAEAMNNMAASVAAQTAAKTQRDLEKRGREIRAAESRGLEDFRRYNPPKFKGD 71
Query: 104 YDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDGAVVTWA 163
+ +W++EVE+I +++C V + T+ML +A+ W S ++ V W
Sbjct: 72 ESSEKADQWIQEVEKIIDMIKCQAGVTVSYATYMLLGDAEYWRRSGRLLMGAAHEEVNWE 131
Query: 164 VFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAKFYPYYTAEIVE 219
F+R+FL++YFP R K +FL+L G M+V +YAAKF L++ + + E E
Sbjct: 132 SFKRKFLDKYFPMSTRTKLGDDFLKLHQGSMTVGEYAAKFGSLSRHFRFVREETDE 187
>UniRef100_Q6UUN3 Putative gag-pol polyprotein [Oryza sativa]
Length = 964
Score = 90.5 bits (223), Expect = 6e-17
Identities = 50/150 (33%), Positives = 72/150 (47%), Gaps = 2/150 (1%)
Query: 62 IAAALEAVA*AVGQQPNAAAGNDGVRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFR 121
IAA+ A A + Q G+ L F R PP F DP WL+ +ER
Sbjct: 22 IAASAAATATHIAQGAGGG-GHHAAGGLAEFQRTQPPVFTRSDDPLDADDWLRTIERKLT 80
Query: 122 VMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGK 181
+++C + +K L A DWW + L ++ DG VVTWA FR F + P+ +
Sbjct: 81 LIRCPDAEKTNLAAEQLQAAAGDWWENFL-AMQPDGRVVTWAQFRDAFRAAHVPKGIMDL 139
Query: 182 NEIEFLELKHGDMSVTKYAAKFVELAKFYP 211
+ EFL L G+ SV +Y +F LA++ P
Sbjct: 140 KQCEFLSLTQGNKSVMEYLCEFNHLARYAP 169
>UniRef100_Q7X798 OSJNBb0108J11.19 protein [Oryza sativa]
Length = 1516
Score = 90.1 bits (222), Expect = 8e-17
Identities = 46/126 (36%), Positives = 66/126 (51%), Gaps = 1/126 (0%)
Query: 86 VRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDW 145
V L FLR PPTF +P WL+ +E+ ++QC++ +KV F TH L A W
Sbjct: 78 VSKLLDFLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAW 137
Query: 146 WVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVE 205
W + + D VTWA F + F PE + + + EF L+ G MSVT+Y +F
Sbjct: 138 WDNYV-ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTMSVTEYLHEFNR 196
Query: 206 LAKFYP 211
LA++ P
Sbjct: 197 LARYAP 202
>UniRef100_Q8W5F5 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 89.0 bits (219), Expect = 2e-16
Identities = 47/125 (37%), Positives = 67/125 (53%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 92 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWW 150
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + G+ + EF L+ G +V +Y +F L
Sbjct: 151 DNYM-VTRPTGAEVTWTEFRHSFNKAQVPEGIVGQKKREFRSLQQGTKTVIEYLHEFNRL 209
Query: 207 AKFYP 211
A++ P
Sbjct: 210 ARYAP 214
>UniRef100_Q7XVP5 OSJNBa0023J03.1 protein [Oryza sativa]
Length = 1827
Score = 87.0 bits (214), Expect = 7e-16
Identities = 45/123 (36%), Positives = 64/123 (51%), Gaps = 1/123 (0%)
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVS 148
L FLR PPTF +P WL +E+ ++QC++ +KV F TH L A WW +
Sbjct: 398 LPEFLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDN 457
Query: 149 LLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
+ G VTWA F R F P+ V + + EF L G+M+VT+Y +F LA+
Sbjct: 458 HM-ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNMTVTEYLHEFNRLAR 516
Query: 209 FYP 211
+ P
Sbjct: 517 YAP 519
>UniRef100_Q6ATE0 Putative polyprotein [Oryza sativa]
Length = 1495
Score = 87.0 bits (214), Expect = 7e-16
Identities = 44/126 (34%), Positives = 65/126 (50%), Gaps = 1/126 (0%)
Query: 86 VRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDW 145
V L FLR PPTF +P WL+ +E+ ++QC++ +KV F TH L A W
Sbjct: 78 VSKLLDFLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAW 137
Query: 146 WVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVE 205
W + + V D VTW F + F PE + + + EF L+ G +VT+Y +F
Sbjct: 138 WDNYV-VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
Query: 206 LAKFYP 211
LA++ P
Sbjct: 197 LARYAP 202
>UniRef100_Q84MF5 Putative retrotransposon gag protein [Oryza sativa]
Length = 486
Score = 86.7 bits (213), Expect = 9e-16
Identities = 44/126 (34%), Positives = 65/126 (50%), Gaps = 1/126 (0%)
Query: 86 VRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDW 145
V L FLR PPTF +P WL+ +E+ ++QC++ +KV F TH L A W
Sbjct: 78 VSKLLDFLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASVW 137
Query: 146 WVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVE 205
W + + V D VTW F + F PE + + + EF L+ G +VT+Y +F
Sbjct: 138 WDNYV-VTRPDATKVTWVEFCQNFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
Query: 206 LAKFYP 211
LA++ P
Sbjct: 197 LARYAP 202
>UniRef100_Q7XM35 OSJNBb0022P19.7 protein [Oryza sativa]
Length = 449
Score = 86.7 bits (213), Expect = 9e-16
Identities = 43/123 (34%), Positives = 64/123 (51%), Gaps = 1/123 (0%)
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVS 148
L FL+ PPTF DP WL+ +E+ ++QC++ ++V F TH L A +WW S
Sbjct: 76 LSEFLKTEPPTFAIAVDPMEASDWLRTIEKKLGLIQCTDQERVGFATHQLVGPASEWWDS 135
Query: 149 LLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
+G +TW F F + P + + EF LK GD++VTKY +F LA+
Sbjct: 136 -YEASRPEGHTITWNEFSSVFRRSHVPAGMITLRKREFRYLKQGDLTVTKYLHEFYRLAR 194
Query: 209 FYP 211
+ P
Sbjct: 195 YAP 197
>UniRef100_Q7F974 OSJNBb0003A12.3 protein [Oryza sativa]
Length = 1471
Score = 86.7 bits (213), Expect = 9e-16
Identities = 46/125 (36%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 66 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWW 124
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + + + EF L+ G +V +Y +F L
Sbjct: 125 DNYM-VTRPTGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNRL 183
Query: 207 AKFYP 211
A++ P
Sbjct: 184 ARYAP 188
>UniRef100_Q943A1 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 86.3 bits (212), Expect = 1e-15
Identities = 46/125 (36%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 92 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWW 150
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + + + EF L+ G +V +Y +F L
Sbjct: 151 DNYM-VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNRL 209
Query: 207 AKFYP 211
A++ P
Sbjct: 210 ARYAP 214
>UniRef100_Q7XW84 OSJNBa0019J05.10 protein [Oryza sativa]
Length = 1525
Score = 86.3 bits (212), Expect = 1e-15
Identities = 46/125 (36%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 92 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWW 150
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + + + EF L+ G +V +Y +F L
Sbjct: 151 DNYM-VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNRL 209
Query: 207 AKFYP 211
A++ P
Sbjct: 210 ARYAP 214
>UniRef100_Q7XQU9 OSJNBa0086B14.15 protein [Oryza sativa]
Length = 1516
Score = 86.3 bits (212), Expect = 1e-15
Identities = 42/120 (35%), Positives = 63/120 (52%), Gaps = 1/120 (0%)
Query: 92 FLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLP 151
FLR PPTF +P WL+ +E+ ++QC++ +KV F TH L A WW + +
Sbjct: 84 FLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDNYV- 142
Query: 152 VLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAKFYP 211
V D VTW F + F PE + + + EF L+ G +VT+Y +F LA++ P
Sbjct: 143 VTRPDATEVTWVEFCQNFRKALIPEGIMAQQKREFRALQQGTRTVTEYLHEFNRLARYAP 202
>UniRef100_Q6F2Q6 Putative polyprotein [Oryza sativa]
Length = 1516
Score = 86.3 bits (212), Expect = 1e-15
Identities = 44/126 (34%), Positives = 65/126 (50%), Gaps = 1/126 (0%)
Query: 86 VRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDW 145
V L FLR PPTF +P WL+ +E+ ++QC++ +KV F TH L A W
Sbjct: 78 VSKLLDFLRIQPPTFSSTTNPMEANDWLRAIEKKLNLLQCNDREKVAFATHQLQGPASAW 137
Query: 146 WVSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVE 205
W + + D VTWA F + F PE + + + EF L+ G +VT+Y +F
Sbjct: 138 WDNYV-ATRLDATEVTWAEFCQSFRKAQIPEGIMAQQKREFRALQQGTRTVTEYLHEFNR 196
Query: 206 LAKFYP 211
LA++ P
Sbjct: 197 LARYAP 202
>UniRef100_Q6ATD6 Putative polyprotein [Oryza sativa]
Length = 1524
Score = 86.3 bits (212), Expect = 1e-15
Identities = 46/125 (36%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 92 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASIWW 150
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + + + EF L+ G +V +Y +F L
Sbjct: 151 DNYM-VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFRSLQQGTRTVIEYLHEFNRL 209
Query: 207 AKFYP 211
A++ P
Sbjct: 210 ARYAP 214
>UniRef100_Q8S9N7 Similar to polyprotein [Oryza sativa]
Length = 1524
Score = 85.9 bits (211), Expect = 1e-15
Identities = 46/125 (36%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
++LE FLR PPTF +P WL +E+ ++QC+E +KV F TH L A WW
Sbjct: 92 KLLE-FLRVKPPTFSSTTNPIEAHDWLHAIEKKLNLLQCNEQEKVAFATHQLQGPASVWW 150
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVEL 206
+ + V GA VTW FR F PE + + + EF L+ G +V +Y +F L
Sbjct: 151 DNYM-VTRPAGAEVTWTEFRHSFNKAQVPEGIVAQKKREFHSLQQGTRTVIEYLHEFNRL 209
Query: 207 AKFYP 211
A++ P
Sbjct: 210 ARYAP 214
>UniRef100_Q8SB08 Putative polyprotein [Oryza sativa]
Length = 501
Score = 85.5 bits (210), Expect = 2e-15
Identities = 42/123 (34%), Positives = 64/123 (51%), Gaps = 1/123 (0%)
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVS 148
L FL+ PPTF DP WL+ +E+ ++QC++ ++V F TH L A +WW +
Sbjct: 128 LSEFLKTEPPTFAIAVDPMEASDWLRTIEKKLGLIQCTDQERVGFATHQLVGPASEWWDN 187
Query: 149 LLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
+ +G +TW F F Y P + + EF LK GD++VT+Y +F LA
Sbjct: 188 -YEASQPEGHTITWNEFSSVFRRSYVPAGMITLRKREFRYLKQGDLTVTEYLHEFYRLAH 246
Query: 209 FYP 211
+ P
Sbjct: 247 YAP 249
>UniRef100_Q8H843 Putative polyprotein [Oryza sativa]
Length = 1796
Score = 85.1 bits (209), Expect = 3e-15
Identities = 44/123 (35%), Positives = 63/123 (50%), Gaps = 1/123 (0%)
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVS 148
L FLR PPTF +P WL +E+ ++QC++ +KV F TH L A WW +
Sbjct: 367 LPEFLRVRPPTFSSTTNPMEANDWLHAIEKKLNLLQCNDQEKVAFATHQLQGPASAWWDN 426
Query: 149 LLPVLEQDGAVVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
+ G VTWA F R F P+ V + + EF L G+ +VT+Y +F LA+
Sbjct: 427 HM-ATRPPGTEVTWAEFCRSFRKAQVPDGVVAQKKREFRALHQGNRTVTEYLQEFNRLAR 485
Query: 209 FYP 211
+ P
Sbjct: 486 YAP 488
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.355 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 487,866,606
Number of Sequences: 2790947
Number of extensions: 17698136
Number of successful extensions: 90621
Number of sequences better than 10.0: 475
Number of HSP's better than 10.0 without gapping: 439
Number of HSP's successfully gapped in prelim test: 36
Number of HSP's that attempted gapping in prelim test: 89498
Number of HSP's gapped (non-prelim): 787
length of query: 343
length of database: 848,049,833
effective HSP length: 128
effective length of query: 215
effective length of database: 490,808,617
effective search space: 105523852655
effective search space used: 105523852655
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 75 (33.5 bits)
Medicago: description of AC135504.16


