

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.11 - phase: 0
(295 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
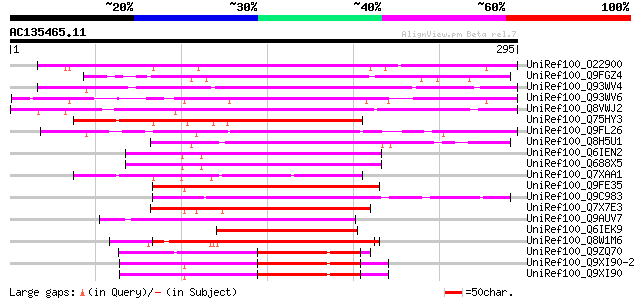
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O22900 Probable WRKY transcription factor 23 [Arabidop... 235 9e-61
UniRef100_Q9FGZ4 Probable WRKY transcription factor 48 [Arabidop... 194 3e-48
UniRef100_Q93WV4 Probable WRKY transcription factor 71 [Arabidop... 173 4e-42
UniRef100_Q93WV6 Probable WRKY transcription factor 68 [Arabidop... 171 2e-41
UniRef100_Q8VWJ2 Probable WRKY transcription factor 28 [Arabidop... 165 2e-39
UniRef100_Q75HY3 Hypothetical protein OSJNBb0035N21.13 [Oryza sa... 157 4e-37
UniRef100_Q9FL26 Probable WRKY transcription factor 8 [Arabidops... 156 7e-37
UniRef100_Q8H5U1 Putative DNA-binding protein WRKY2 [Oryza sativa] 154 2e-36
UniRef100_Q6IEN2 WRKY transcription factor 49 [Oryza sativa] 154 2e-36
UniRef100_Q688X5 WRKY transcription factor [Oryza sativa] 154 2e-36
UniRef100_Q7XAA1 WRKY17 [Oryza sativa] 150 4e-35
UniRef100_Q9FE35 DNA-binding protein WRKY2-like [Oryza sativa] 149 1e-34
UniRef100_Q9C983 Probable WRKY transcription factor 57 [Arabidop... 149 1e-34
UniRef100_Q7X7E3 WRKY3 [Oryza sativa] 146 7e-34
UniRef100_Q9AUV7 Putative DNA binding protein [Oryza sativa] 142 1e-32
UniRef100_Q6IEK9 WRKY transcription factor 72 [Oryza sativa] 124 2e-27
UniRef100_Q8W1M6 WRKY-like drought-induced protein [Retama raetam] 124 3e-27
UniRef100_Q9ZQ70 Probable WRKY transcription factor 3 [Arabidops... 122 9e-27
UniRef100_Q9XI90-2 Splice isoform 2 of Q9XI90 [Arabidopsis thali... 122 1e-26
UniRef100_Q9XI90 Probable WRKY transcription factor 4 [Arabidops... 122 1e-26
>UniRef100_O22900 Probable WRKY transcription factor 23 [Arabidopsis thaliana]
Length = 337
Score = 235 bits (600), Expect = 9e-61
Identities = 152/329 (46%), Positives = 188/329 (56%), Gaps = 51/329 (15%)
Query: 17 SFPSYSFPSVFDLSE----ER---SFMELLGVQQNMNNYSDSLLDLPVVVKEPPLESDGN 69
SF S SV+D + ER FMELL QQ+ + + S + +P ++ +
Sbjct: 10 SFYYPSSQSVWDFGDLAAAERHSLGFMELLSSQQHQDFATVSPHSFLLQTSQPQTQTQPS 69
Query: 70 GKEYSEVLNSQQQ--------------------PATPNSSSISSASSEAINDEHNKTVD- 108
K S ++ + PATPNSSSISSASSEA+N+E KT D
Sbjct: 70 AKLSSSIIQAPPSEQLVTSKVESLCSDHLLINPPATPNSSSISSASSEALNEEKPKTEDN 129
Query: 109 --------QTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKN 160
Q + KQLKAKK NQK+ REAR+AFMTKSEVDHLEDGYRWRKYGQKAVKN
Sbjct: 130 EEEGGEDQQEKSHTKKQLKAKKNNQKRQREARVAFMTKSEVDHLEDGYRWRKYGQKAVKN 189
Query: 161 SPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSR--------SSAV 212
SPFPRSYYRCT+ SCNVKK VERS DP+ VVTTYEG+HTH +P+ SR S+
Sbjct: 190 SPFPRSYYRCTTASCNVKKRVERSFRDPSTVVTTYEGQHTHISPLTSRPISTGGFFGSSG 249
Query: 213 RAGSL----LPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQ 268
A SL P + +T S Q + QY++QQ+QQ L + + + +
Sbjct: 250 AASSLGNGCFGFPIDGST-LISPQFQQLVQYHHQQQQQELMSCFGGVNEYLNSHANEYGD 308
Query: 269 DRPLCNPR--VQDNGLLQDVVPSHMFKEE 295
D + R V+DNGLLQDVVPSHM KEE
Sbjct: 309 DNRVKKSRVLVKDNGLLQDVVPSHMLKEE 337
>UniRef100_Q9FGZ4 Probable WRKY transcription factor 48 [Arabidopsis thaliana]
Length = 399
Score = 194 bits (492), Expect = 3e-48
Identities = 122/279 (43%), Positives = 160/279 (56%), Gaps = 47/279 (16%)
Query: 44 NMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEH 103
N NN + DLP+ P ES SEV+N+ P +PNS+S+SS+S+EA ND +
Sbjct: 122 NNNNNTSFFTDLPL----PQAES-------SEVVNTT--PTSPNSTSVSSSSNEAANDNN 168
Query: 104 N---------KTVDQTNNQ--LNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRK 152
+ + DQ Q QLKAKK NQKK REAR AF+TKS++D+L+DGYRWRK
Sbjct: 169 SGKEVTVKDQEEGDQQQEQKGTKPQLKAKKKNQKKAREARFAFLTKSDIDNLDDGYRWRK 228
Query: 153 YGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAV 212
YGQKAVKNSP+PRSYYRCT+V C VKK VERS DP+IV+TTYEG+HTHP P+ R
Sbjct: 229 YGQKAVKNSPYPRSYYRCTTVGCGVKKRVERSSDDPSIVMTTYEGQHTHPFPMTPRG--- 285
Query: 213 RAGSLLPPPAECTTNFASDQNYDISQ--YYNQQRQQV--LFNTLSSLGFPSKNMNATF-- 266
G L P + AS ++ I Q Y Q Q ++N S ++ + TF
Sbjct: 286 HIGMLTSPILDHGATTASSSSFSIPQPRYLLTQHHQPYNMYNNNSLSMINRRSSDGTFVN 345
Query: 267 --------------SQDRPLCNPRVQDNGLLQDVVPSHM 291
SQ + ++D+GLLQD++PS +
Sbjct: 346 PGPSSSFPGFGYDMSQASTSTSSSIRDHGLLQDILPSQI 384
>UniRef100_Q93WV4 Probable WRKY transcription factor 71 [Arabidopsis thaliana]
Length = 282
Score = 173 bits (439), Expect = 4e-42
Identities = 104/281 (37%), Positives = 148/281 (52%), Gaps = 13/281 (4%)
Query: 17 SFPSYSFPSVFDLSEERSFMELLGVQQ--NMNNYSDSLLDLPVVVKEPPLESDGNGKEYS 74
S F S+ D + M+ +++ + YS + V P L N S
Sbjct: 12 SLEEVHFKSLSDCLQSSLVMDYNSLEKVFKFSPYSSPFQSVSPSVNNPYLNLTSN----S 67
Query: 75 EVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIA 134
V++S P ++ + N+ V +++ QL KQ KK +KK RE R+A
Sbjct: 68 PVVSSSSNEGEPKENTNDKSDQMEDNEGDLHGVGESSKQLTKQ--GKKKGEKKEREVRVA 125
Query: 135 FMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTT 194
FMTKSE+DHLEDGYRWRKYGQKAVKNSP+PRSYYRCT+ CNVKK VERS DP+IV+TT
Sbjct: 126 FMTKSEIDHLEDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSFQDPSIVITT 185
Query: 195 YEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSS 254
YEGKH HP P R + L+ + + +++ + + + ++ S
Sbjct: 186 YEGKHNHPIPSTLRGTVAAEHLLVHRGGGGSLLHSFPRHH--QDFLMMKHSPANYQSVGS 243
Query: 255 LGFPSKNMNATFSQDRPLCNPRVQDNGLLQDVVPSHMFKEE 295
L + + ++++ + N V D GLLQD+VPS K E
Sbjct: 244 LSYEHGHGTSSYNFNN---NQPVVDYGLLQDIVPSMFSKNE 281
>UniRef100_Q93WV6 Probable WRKY transcription factor 68 [Arabidopsis thaliana]
Length = 277
Score = 171 bits (434), Expect = 2e-41
Identities = 124/323 (38%), Positives = 166/323 (51%), Gaps = 76/323 (23%)
Query: 2 MEKKGLNMEDYANIGSFPSYSFPSVF-DLSEER---SFMELLGVQQNMNNYSDSLLDLPV 57
ME G+ M Y ++G Y S F DLS ER FM+LLGV ++ ++
Sbjct: 1 MENVGVGMPFY-DLGQTRVYPLLSDFHDLSAERYPVGFMDLLGVHRHTPTHT-------- 51
Query: 58 VVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAIN--DEHNKTVDQTNNQLN 115
PL P TPNSSS SEA+N DE + ++ ++
Sbjct: 52 -----PL---------------MHFPTTPNSSS-----SEAVNGDDEEEEDGEEQQHKTK 86
Query: 116 KQLKAKKTNQK--KPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV 173
K+ K K ++K K + +++F+T+SEV HL+DGY+WRKYGQK VK+SPFPR+YYRCT+
Sbjct: 87 KRFKFTKMSRKQTKKKVPKVSFITRSEVLHLDDGYKWRKYGQKPVKDSPFPRNYYRCTTT 146
Query: 174 SCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIM---------SRSSAVRAGSLLP--PPA 222
C+VKK VERS SDP+ V+TTYEG+HTHP P++ S SA RA LP PP
Sbjct: 147 WCDVKKRVERSFSDPSSVITTYEGQHTHPRPLLIMPKEGSSPSNGSASRAHIGLPTLPP- 205
Query: 223 ECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPR------ 276
+ Y NQQ+Q + K +N D + R
Sbjct: 206 ------------QLLDYNNQQQQAPSSFGTEYINRQEKGINHDDDDDHVVKKSRTRDLLD 253
Query: 277 ----VQDNGLLQDVVPSHMFKEE 295
V+D+GLLQDVVPSH+ KEE
Sbjct: 254 GAGLVKDHGLLQDVVPSHIIKEE 276
>UniRef100_Q8VWJ2 Probable WRKY transcription factor 28 [Arabidopsis thaliana]
Length = 318
Score = 165 bits (417), Expect = 2e-39
Identities = 112/307 (36%), Positives = 153/307 (49%), Gaps = 23/307 (7%)
Query: 1 MMEKKGLNMEDYANI----GSFPS-YSFPSVFDLSE---ERSFMELLGVQQNMNNYSDSL 52
MM N Y N+ G PS YSF S E + G+ + + +S
Sbjct: 22 MMNLPTSNPSSYGNLPSQNGFNPSTYSFTDCLQSSPAAYESLLQKTFGLSPSSSEVFNSS 81
Query: 53 LDLPVVVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSAS---SEAINDEHNKTVDQ 109
+D + P N N + +P++SS S A ++ + +
Sbjct: 82 ID------QEPNRDVTNDVINGGACNETETRVSPSNSSSSEADHPGEDSGKSRRKRELVG 135
Query: 110 TNNQLNKQL-KAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYY 168
+Q++K++ K KKT KK RE R++FMTKSEVDHLEDGYRWRKYGQKAVKNSP+PRSYY
Sbjct: 136 EEDQISKKVGKTKKTEVKKQREPRVSFMTKSEVDHLEDGYRWRKYGQKAVKNSPYPRSYY 195
Query: 169 RCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNF 228
RCT+ CNVKK VERS DPT+V+TTYEG+H HP P R S+ A ++ +F
Sbjct: 196 RCTTQKCNVKKRVERSFQDPTVVITTYEGQHNHPIPTNLRGSSA-AAAMFSADLMTPRSF 254
Query: 229 ASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQDVVP 288
A D + L G+ S N N + Q + + LL+++ P
Sbjct: 255 AHDMFRTAAYTNGGSVAAALDYGYGQSGYGSVNSNPSSHQ----VYHQGGEYELLREIFP 310
Query: 289 SHMFKEE 295
S FK+E
Sbjct: 311 SIFFKQE 317
>UniRef100_Q75HY3 Hypothetical protein OSJNBb0035N21.13 [Oryza sativa]
Length = 337
Score = 157 bits (396), Expect = 4e-37
Identities = 89/190 (46%), Positives = 117/190 (60%), Gaps = 23/190 (12%)
Query: 38 LLGVQQNMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEVLNSQQQ------PATPNSSSI 91
L+ ++ +Y L PV P L D K + V + Q PATPNSS +
Sbjct: 60 LITPYSSITDYLQGFLQDPVYASSP-LGGDAAVKHETVVDHPSQAGGVAAAPATPNSSVL 118
Query: 92 SSASSEAINDE------HNKTVDQTNNQLNKQ---LKAKKTNQ-------KKPREARIAF 135
SS+S A D+ + D+ +++ + +++ KTN+ KK RE R+AF
Sbjct: 119 SSSSEAAGGDDLRRCKKGRRPEDEEEEEIDDEGSAVQSCKTNKMKNKKGAKKEREPRVAF 178
Query: 136 MTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTY 195
MTKSEVDHLEDGYRWRKYGQKAVKNS +PRSYYRCT+ C VKK VERS DP++V+TTY
Sbjct: 179 MTKSEVDHLEDGYRWRKYGQKAVKNSSYPRSYYRCTAPRCGVKKRVERSEQDPSMVITTY 238
Query: 196 EGKHTHPNPI 205
EG+HTHP+P+
Sbjct: 239 EGQHTHPSPV 248
>UniRef100_Q9FL26 Probable WRKY transcription factor 8 [Arabidopsis thaliana]
Length = 326
Score = 156 bits (394), Expect = 7e-37
Identities = 111/294 (37%), Positives = 153/294 (51%), Gaps = 45/294 (15%)
Query: 19 PSYSFPSVFDLSEERSFMELLGVQQ-----NMNNYSDSLLDLPVVVKEPPLESDGNGKEY 73
P +F F+LS S + Q N NY+ S VV DG
Sbjct: 60 PQGAFELGFELSPSSSDFFNPSLDQENGLYNAYNYNSSQKSHEVV-------GDGCATIK 112
Query: 74 SEVLNSQQQPATPNSSSISSASSEAINDEHNKTV----DQTNNQLNKQLKAKKTNQKKPR 129
SEV S A+P+SS E K + + + K +K KK +KK +
Sbjct: 113 SEVRVS----ASPSSSEADHHPGEDSGKIRKKREVRDGGEDDQRSQKVVKTKKKEEKK-K 167
Query: 130 EARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPT 189
E R++FMTK+EVDHLEDGYRWRKYGQKAVKNSP+PRSYYRCT+ CNVKK VERS DPT
Sbjct: 168 EPRVSFMTKTEVDHLEDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSYQDPT 227
Query: 190 IVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVLF 249
+V+TTYE +H HP P +R +A+ +G+ ASD N S ++ ++
Sbjct: 228 VVITTYESQHNHPIP-TNRRTAMFSGTT-----------ASDYNPSSSPIFS----DLII 271
Query: 250 NT--------LSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQDVVPSHMFKEE 295
NT L + + S N+N ++ Q + + + + LL+++ PS FK+E
Sbjct: 272 NTPRSFSNDDLFRVPYASVNVNPSYHQQQHGFHQQESEFELLKEMFPSVFFKQE 325
>UniRef100_Q8H5U1 Putative DNA-binding protein WRKY2 [Oryza sativa]
Length = 290
Score = 154 bits (390), Expect = 2e-36
Identities = 98/238 (41%), Positives = 122/238 (51%), Gaps = 43/238 (18%)
Query: 83 PATPNSSSISSASSEAINDEHN--KTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSE 140
P TP SS A+S D+ + K + +K L K K+ R+ R AFMTKSE
Sbjct: 63 PETPAGSSADGAASSCSTDDADGGKPAAASTEAASKSLTPGK---KRARQPRFAFMTKSE 119
Query: 141 VDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHT 200
+DHLEDGYRWRKYGQKAVKNSPFPRSYYRCT+ C VKK VERS DP++V+TTYEG+H+
Sbjct: 120 IDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTNSKCTVKKRVERSSDDPSVVITTYEGQHS 179
Query: 201 HPNPIMSRSSAVRAG-----------------------SLLPP----PAECTTNFASDQN 233
H R++A AG +L PP P TT +S
Sbjct: 180 HHTVTFPRAAATAAGFSHIHAMAALAAAPFSAHQQLYSNLQPPPPTMPLAATTPASSSSL 239
Query: 234 YDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQDVVPSHM 291
+ + N + Q V S G+PS S P + D GLL D+VP M
Sbjct: 240 LQLPLHCNHELQVV----ASCGGYPS-------SSSSPPASVLPVDKGLLDDMVPRAM 286
>UniRef100_Q6IEN2 WRKY transcription factor 49 [Oryza sativa]
Length = 418
Score = 154 bits (390), Expect = 2e-36
Identities = 86/180 (47%), Positives = 100/180 (54%), Gaps = 31/180 (17%)
Query: 68 GNGKEYSEVLNSQQQPATPNSSSISSASSEAI------------------------NDEH 103
G G + + P TPNS S+SS SSEA D+
Sbjct: 93 GGGASAAAAAAAVAGPMTPNSMSVSSTSSEACGVGGGAGGDEESAGKCKKEEEGDGGDDD 152
Query: 104 NKTVDQT-------NNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQK 156
K T ++ K K K +K+PR+ R AFMTKSEVDHLEDGYRWRKYGQK
Sbjct: 153 GKEGSSTTKGDGDGEDKNKKGGKGKGKGEKRPRQPRFAFMTKSEVDHLEDGYRWRKYGQK 212
Query: 157 AVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGS 216
AVKNSPFPRSYYRCT+ C VKK VERS D +V+TTYEGKHTHP P R +A G+
Sbjct: 213 AVKNSPFPRSYYRCTTQKCPVKKRVERSYQDAAVVITTYEGKHTHPIPATLRGTAHLLGA 272
>UniRef100_Q688X5 WRKY transcription factor [Oryza sativa]
Length = 419
Score = 154 bits (390), Expect = 2e-36
Identities = 86/180 (47%), Positives = 100/180 (54%), Gaps = 31/180 (17%)
Query: 68 GNGKEYSEVLNSQQQPATPNSSSISSASSEAI------------------------NDEH 103
G G + + P TPNS S+SS SSEA D+
Sbjct: 93 GGGASAAAAAAAVAGPMTPNSMSVSSTSSEACGVGGGAGGDEESAGKCKKEEEGDGGDDD 152
Query: 104 NKTVDQT-------NNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQK 156
K T ++ K K K +K+PR+ R AFMTKSEVDHLEDGYRWRKYGQK
Sbjct: 153 GKEGSSTTKGDGDGEDKNKKGGKGKGKGEKRPRQPRFAFMTKSEVDHLEDGYRWRKYGQK 212
Query: 157 AVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGS 216
AVKNSPFPRSYYRCT+ C VKK VERS D +V+TTYEGKHTHP P R +A G+
Sbjct: 213 AVKNSPFPRSYYRCTTQKCPVKKRVERSYQDAAVVITTYEGKHTHPIPATLRGTAHLLGA 272
>UniRef100_Q7XAA1 WRKY17 [Oryza sativa]
Length = 502
Score = 150 bits (379), Expect = 4e-35
Identities = 89/190 (46%), Positives = 112/190 (58%), Gaps = 25/190 (13%)
Query: 38 LLGVQQNMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEVLNSQQQ------PATPNSSSI 91
L+ ++ +Y L PV P L D K + V + Q PATPNSS +
Sbjct: 60 LITPYSSITDYLQGFLQDPVYASSP-LGGDAAVKHETVVDHPSQAGGVAAAPATPNSSVL 118
Query: 92 SSASSEA-------------INDEHNKTVDQTNNQLNK---QLKAKKTNQKKPREARIAF 135
SS+S A DE + +D + + ++K KK KK RE R+AF
Sbjct: 119 SSSSEAAGGDDLRRCKKGRRPEDEEEEEIDDEGSAVQSCTNKMKNKK-GAKKEREPRVAF 177
Query: 136 MTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTY 195
MTKSEVDHLEDGYRWRKYGQKAVKNS +P SYYRCT+ C VKK VERS DP++V+TTY
Sbjct: 178 MTKSEVDHLEDGYRWRKYGQKAVKNSSYP-SYYRCTAPRCGVKKRVERSEQDPSMVITTY 236
Query: 196 EGKHTHPNPI 205
EG+HTHP+P+
Sbjct: 237 EGQHTHPSPV 246
>UniRef100_Q9FE35 DNA-binding protein WRKY2-like [Oryza sativa]
Length = 379
Score = 149 bits (375), Expect = 1e-34
Identities = 77/147 (52%), Positives = 93/147 (62%), Gaps = 15/147 (10%)
Query: 84 ATPNSS-SISSASSEAIN--------------DEHNKTVDQTNNQLNKQLKAKKTNQKKP 128
ATPNSS S SS+ EA +E + ++ K K KK +K+
Sbjct: 129 ATPNSSASFSSSDGEAEGGKSSRRCKKGQAKAEEEDDKDEEDGENSKKPNKPKKKAEKRQ 188
Query: 129 REARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDP 188
R+ R+AF+TKSEVDHLEDGYRWRKYGQKAVKNSP+PRSYYRCT+ C VKK VERS DP
Sbjct: 189 RQPRVAFLTKSEVDHLEDGYRWRKYGQKAVKNSPYPRSYYRCTTPKCGVKKRVERSYQDP 248
Query: 189 TIVVTTYEGKHTHPNPIMSRSSAVRAG 215
+ V+TTYEG+HTH +P R G
Sbjct: 249 STVITTYEGQHTHHSPASLRGGGGGVG 275
>UniRef100_Q9C983 Probable WRKY transcription factor 57 [Arabidopsis thaliana]
Length = 287
Score = 149 bits (375), Expect = 1e-34
Identities = 89/208 (42%), Positives = 117/208 (55%), Gaps = 10/208 (4%)
Query: 84 ATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDH 143
+T N+ S +S+SSE + + ++T +K KK QK+ R+ R AFMTKS+VD+
Sbjct: 87 STNNNPSATSSSSEDPAENSTASAEKTPPP-ETPVKEKKKAQKRIRQPRFAFMTKSDVDN 145
Query: 144 LEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPN 203
LEDGYRWRKYGQKAVKNSPFPRSYYRCT+ C VKK VERS DP+IV+TTYEG+H H
Sbjct: 146 LEDGYRWRKYGQKAVKNSPFPRSYYRCTNSRCTVKKRVERSSDDPSIVITTYEGQHCHQT 205
Query: 204 PIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMN 263
R + A P T++ YY Q++L PS +
Sbjct: 206 IGFPRGGILTAHD----PHSFTSHHHLPPPLPNPYYY----QELLHQLHRDNNAPSPRLP 257
Query: 264 ATFSQDRPLCNPRVQDNGLLQDVVPSHM 291
++D P + + GLL D+VP M
Sbjct: 258 RPTTEDTPAVS-TPSEEGLLGDIVPQTM 284
>UniRef100_Q7X7E3 WRKY3 [Oryza sativa]
Length = 565
Score = 146 bits (368), Expect = 7e-34
Identities = 80/160 (50%), Positives = 97/160 (60%), Gaps = 32/160 (20%)
Query: 83 PATPNS-SSISSASSEAIN-----------------------DEHNKTV---DQTNNQLN 115
P TPN+ SS+S++SSE + ++ NK ++ N
Sbjct: 261 PLTPNTTSSMSTSSSEGVGGGGGGGAGAGAGEEESPARCKKEEDENKEEGKGEEDEGHKN 320
Query: 116 KQLKAKK-----TNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRC 170
K+ A K +K+ R+ R AFMTKSEVDHLEDGYRWRKYGQKAVKNSP+PRSYYRC
Sbjct: 321 KKGSAAKGGKAGKGEKRARQPRFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPYPRSYYRC 380
Query: 171 TSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSS 210
T+ C VKK VERS DP +V+TTYEGKHTHP P R S
Sbjct: 381 TTQKCPVKKRVERSYQDPAVVITTYEGKHTHPIPATLRGS 420
>UniRef100_Q9AUV7 Putative DNA binding protein [Oryza sativa]
Length = 314
Score = 142 bits (357), Expect = 1e-32
Identities = 73/149 (48%), Positives = 90/149 (59%), Gaps = 3/149 (2%)
Query: 53 LDLPVVVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNN 112
LDLPV V + +E E ++ ++S SS+ A D+ +
Sbjct: 46 LDLPVDVVGAAAREE---EEEEEEAPARSGDGAAAAASSSSSGEPAAPDKRPAAAEAAPA 102
Query: 113 QLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTS 172
K QK+ R+ R AFMTKSE+DHLEDGYRWRKYGQKAVKNSPFPRSYYRCT+
Sbjct: 103 AAATATATAKKGQKRARQPRFAFMTKSEIDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTN 162
Query: 173 VSCNVKKHVERSLSDPTIVVTTYEGKHTH 201
C VKK VERS DP++V+TTYEG+H H
Sbjct: 163 SKCTVKKRVERSSDDPSVVITTYEGQHCH 191
>UniRef100_Q6IEK9 WRKY transcription factor 72 [Oryza sativa]
Length = 245
Score = 124 bits (312), Expect = 2e-27
Identities = 56/82 (68%), Positives = 64/82 (77%)
Query: 121 KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKH 180
+K +KK R R AF T+S+VD L+DGYRWRKYGQKAVKN+ FPRSYYRCT CNVKK
Sbjct: 116 RKKGEKKERRPRFAFQTRSQVDILDDGYRWRKYGQKAVKNNKFPRSYYRCTHQGCNVKKQ 175
Query: 181 VERSLSDPTIVVTTYEGKHTHP 202
V+R D T+VVTTYEG HTHP
Sbjct: 176 VQRLSRDETVVVTTYEGTHTHP 197
>UniRef100_Q8W1M6 WRKY-like drought-induced protein [Retama raetam]
Length = 488
Score = 124 bits (311), Expect = 3e-27
Identities = 70/144 (48%), Positives = 89/144 (61%), Gaps = 14/144 (9%)
Query: 84 ATPNSSSISSASSEAINDEHNKTVDQTNNQLN------KQLK-----AKKTNQKKP-REA 131
A PNS+ SS A ND+ + D+TN++++ K+ K A T+ KP RE
Sbjct: 241 AEPNSTP--DLSSVATNDDSREGADRTNDEVDDDDPFSKRRKMELGFADITHVVKPIREP 298
Query: 132 RIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIV 191
R+ T SEVD L+DGYRWRKYGQK V+ +P PRSYY+CT+ C V+KHVER+ DP V
Sbjct: 299 RVVVKTLSEVDILDDGYRWRKYGQKVVRGNPNPRSYYKCTNAGCPVRKHVERASHDPKAV 358
Query: 192 VTTYEGKHTHPNPIMSRSSAVRAG 215
+TTYEGKH H P SS AG
Sbjct: 359 ITTYEGKHNHDVPAARNSSHDMAG 382
Score = 81.6 bits (200), Expect = 2e-14
Identities = 54/162 (33%), Positives = 83/162 (50%), Gaps = 14/162 (8%)
Query: 59 VKEPPLESDGNGKEYSEVLNS--QQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNK 116
V E ++++G GK S + + + P++ S+ + ++ + V+ +++LN
Sbjct: 69 VSEQSIQAEGPGKAQSFASSPLIECEIDVPSNELSLSSPVQMVSSGASTPVEVDSDELNH 128
Query: 117 Q------LKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRC 170
+ L+A + + K +A S+ DGY WRKYGQK VK S FPRSYY+C
Sbjct: 129 KGNTITVLQASQVEEVKGSGLPVAPERASD-----DGYNWRKYGQKLVKGSEFPRSYYKC 183
Query: 171 TSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAV 212
T +C VKK +E S D I Y+G H HP P SR +V
Sbjct: 184 THPNCEVKKLLECS-HDGQITEIVYKGMHDHPKPQPSRRYSV 224
>UniRef100_Q9ZQ70 Probable WRKY transcription factor 3 [Arabidopsis thaliana]
Length = 513
Score = 122 bits (307), Expect = 9e-27
Identities = 61/148 (41%), Positives = 85/148 (57%), Gaps = 2/148 (1%)
Query: 64 LESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAK-K 122
L +E S+V ++Q +S + +A + ++ + H D ++
Sbjct: 334 LNKSKRDQETSQVTTTEQMSEASDSEEVGNAET-SVGERHEDEPDPKRRNTEVRVSEPVA 392
Query: 123 TNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVE 182
++ + E RI T SEVD L+DGYRWRKYGQK VK +P+PRSYY+CT+ C V+KHVE
Sbjct: 393 SSHRTVTEPRIIVQTTSEVDLLDDGYRWRKYGQKVVKGNPYPRSYYKCTTPDCGVRKHVE 452
Query: 183 RSLSDPTIVVTTYEGKHTHPNPIMSRSS 210
R+ +DP VVTTYEGKH H P SS
Sbjct: 453 RAATDPKAVVTTYEGKHNHDVPAARTSS 480
Score = 78.6 bits (192), Expect = 2e-13
Identities = 36/60 (60%), Positives = 42/60 (70%), Gaps = 1/60 (1%)
Query: 145 EDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNP 204
+DGY WRKYGQK VK S FPRSYY+CT +C VKK VERSL D + Y+G+H H P
Sbjct: 250 DDGYNWRKYGQKQVKGSDFPRSYYKCTHPACPVKKKVERSL-DGQVTEIIYKGQHNHELP 308
>UniRef100_Q9XI90-2 Splice isoform 2 of Q9XI90 [Arabidopsis thaliana]
Length = 487
Score = 122 bits (305), Expect = 1e-26
Identities = 66/164 (40%), Positives = 88/164 (53%), Gaps = 8/164 (4%)
Query: 65 ESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAIN--------DEHNKTVDQTNNQLNK 116
+++ + K E + Q T S +S E N DE+ + + ++
Sbjct: 294 QTNSSNKTKREQHEAVSQATTTEHLSEASDGEEVGNGETDVREKDENEPDPKRRSTEVRI 353
Query: 117 QLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCN 176
A + + E RI T SEVD L+DGYRWRKYGQK VK +P+PRSYY+CT+ C
Sbjct: 354 SEPAPAASHRTVTEPRIIVQTTSEVDLLDDGYRWRKYGQKVVKGNPYPRSYYKCTTPGCG 413
Query: 177 VKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPP 220
V+KHVER+ +DP VVTTYEGKH H P SS A + L P
Sbjct: 414 VRKHVERAATDPKAVVTTYEGKHNHDLPAAKSSSHAAAAAQLRP 457
Score = 79.3 bits (194), Expect = 1e-13
Identities = 36/60 (60%), Positives = 42/60 (70%), Gaps = 1/60 (1%)
Query: 145 EDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNP 204
+DGY WRKYGQK VK S FPRSYY+CT+ C VKK VERSL D + Y+G+H H P
Sbjct: 202 DDGYNWRKYGQKQVKGSEFPRSYYKCTNPGCPVKKKVERSL-DGQVTEIIYKGQHNHEPP 260
>UniRef100_Q9XI90 Probable WRKY transcription factor 4 [Arabidopsis thaliana]
Length = 514
Score = 122 bits (305), Expect = 1e-26
Identities = 66/164 (40%), Positives = 88/164 (53%), Gaps = 8/164 (4%)
Query: 65 ESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAIN--------DEHNKTVDQTNNQLNK 116
+++ + K E + Q T S +S E N DE+ + + ++
Sbjct: 321 QTNSSNKTKREQHEAVSQATTTEHLSEASDGEEVGNGETDVREKDENEPDPKRRSTEVRI 380
Query: 117 QLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCN 176
A + + E RI T SEVD L+DGYRWRKYGQK VK +P+PRSYY+CT+ C
Sbjct: 381 SEPAPAASHRTVTEPRIIVQTTSEVDLLDDGYRWRKYGQKVVKGNPYPRSYYKCTTPGCG 440
Query: 177 VKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPP 220
V+KHVER+ +DP VVTTYEGKH H P SS A + L P
Sbjct: 441 VRKHVERAATDPKAVVTTYEGKHNHDLPAAKSSSHAAAAAQLRP 484
Score = 79.3 bits (194), Expect = 1e-13
Identities = 36/60 (60%), Positives = 42/60 (70%), Gaps = 1/60 (1%)
Query: 145 EDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNP 204
+DGY WRKYGQK VK S FPRSYY+CT+ C VKK VERSL D + Y+G+H H P
Sbjct: 229 DDGYNWRKYGQKQVKGSEFPRSYYKCTNPGCPVKKKVERSL-DGQVTEIIYKGQHNHEPP 287
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.310 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 508,513,056
Number of Sequences: 2790947
Number of extensions: 20822888
Number of successful extensions: 69050
Number of sequences better than 10.0: 537
Number of HSP's better than 10.0 without gapping: 345
Number of HSP's successfully gapped in prelim test: 194
Number of HSP's that attempted gapping in prelim test: 68065
Number of HSP's gapped (non-prelim): 858
length of query: 295
length of database: 848,049,833
effective HSP length: 126
effective length of query: 169
effective length of database: 496,390,511
effective search space: 83889996359
effective search space used: 83889996359
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 74 (33.1 bits)
Medicago: description of AC135465.11


