

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
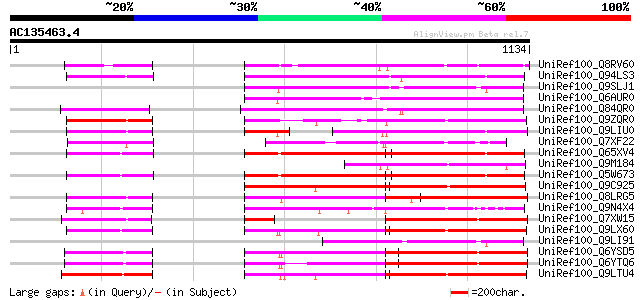
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 517 e-145
UniRef100_Q94LS3 Putative helicase [Oryza sativa] 470 e-131
UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana] 426 e-117
UniRef100_Q6AUR0 Hypothetical protein OSJNBa0077L08.8 [Oryza sat... 404 e-111
UniRef100_Q84QR0 Helicase-like protein [Oryza sativa] 358 7e-97
UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana] 355 4e-96
UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II B... 326 3e-87
UniRef100_Q7XF22 Helicase-like protein [Oryza sativa] 324 1e-86
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 306 3e-81
UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thali... 305 4e-81
UniRef100_Q5W673 Putative helicase [Oryza sativa] 304 9e-81
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 286 2e-75
UniRef100_Q8LRG5 Putative helicase [Oryza sativa] 286 2e-75
UniRef100_Q9N4X4 Hypothetical protein Y46B2A.2 [Caenorhabditis e... 285 7e-75
UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa] 281 1e-73
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 280 2e-73
UniRef100_Q9LI91 Similarity to unknown protein [Arabidopsis thal... 276 3e-72
UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa] 275 4e-72
UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa] 274 1e-71
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 271 6e-71
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 517 bits (1331), Expect = e-145
Identities = 280/673 (41%), Positives = 405/673 (59%), Gaps = 73/673 (10%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP++ RRD G V KNGI LDNR V+PYNP L++KY AHIN+E+ ++S +KYLFKYI K
Sbjct: 480 FPIYKRRDDGRFVEKNGILLDNRFVIPYNPTLLLKYQAHINVEWVSQSKIVKYLFKYINK 539
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
G D+VTA ++ P+ EI+ +YDCRY+SP E+ WRIFGFD+H H
Sbjct: 540 GNDKVTARIKEGTGPN--EIKRWYDCRYISPCEAAWRIFGFDLH-----------HSSTS 586
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQ---SWNLTYVQFPRKFTYNVENR 690
Q I++ D ++VL + ++ + FLA++ N+ ++ + TY++FP F + +
Sbjct: 587 QNIIYDDDEDVEDVLNKEENQTSQFLAFLKTNKKIAEDPEARKFTYIEFPSHFVWKKKQM 646
Query: 691 SWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAA 750
W L + SVGR+ V P + + +R+LLN+ G T +EDI+TV G +Y ++++AC A
Sbjct: 647 EWTLRQRSVSVGRVYHVTPSAGQRFYLRILLNKVKGPTCYEDIKTVGGILYPTFKDACYA 706
Query: 751 LGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEG---- 806
L LL +D+++ID I E GSG +R++F LL + +S P +VW TW+ LS+
Sbjct: 707 LSLLEDDQEYIDTIIEASQWGSGRYMRRLFAQLLTSESLSTPYHVWLNTWNQLSDDQIQN 766
Query: 807 -ILYERRRTLNSPG*LLNS----------------------------------------- 824
L E + L + G L +
Sbjct: 767 QALLEIEKLLRNSGSTLRNYESMPYPENNNFPAAMNTLIQDELCYDRHACAQDHLRLFSK 826
Query: 825 ---DQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
+Q Y++I+ V + +FFV+G+GG GKT+LW LS + RS G+IVLNVASS +
Sbjct: 827 LTVEQRKIYDQIIEAVYSNSGGVFFVNGFGGTGKTYLWKTLSTYIRSRGEIVLNVASSGM 886
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
A+LLL GRTAHS+F+IPL + E S C+I + + A LL A LIIWDEAPM+ R +EA
Sbjct: 887 AALLLDGGRTAHSRFAIPLQVNETSTCSISPDSDLASLLLRAKLIIWDEAPMLHRHCYEA 946
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT++DI V ++ PFGGKT+ GGDFRQILPV+ K R I+ A + SS LW
Sbjct: 947 LDRTLKDI----VQADNHKPFGGKTITFGGDFRQILPVITKGSREQIIHASLTSSRLWNS 1002
Query: 1002 CRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNS 1061
C+VL LTKN+RL +D +++ F+ WIL +GDGKL NDGEA I+IP+D+ L +S
Sbjct: 1003 CKVLTLTKNMRLTADPTEKD--NIKEFSDWILKLGDGKLSEPNDGEAAIDIPEDMLLLDS 1060
Query: 1062 GNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCD 1121
+ + I YPNLL N+ + +FF++RAIL PT + V +VN++++ L+PGE KEY S D
Sbjct: 1061 LHPIDSIANCVYPNLLQNLNDQTFFRERAILCPTNDDVSEVNNHIMDLLPGEVKEYFSSD 1120
Query: 1122 TVLKCDEEVGIDR 1134
+ CD + ++R
Sbjct: 1121 KI--CDFDTSVER 1131
Score = 127 bits (318), Expect = 3e-27
Identities = 72/196 (36%), Positives = 119/196 (59%), Gaps = 16/196 (8%)
Query: 119 SDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVP-VKLRLFRNRNFDS-RVY 176
++ L +V+ + ++D+CN VK RM ++ + ++++P +KL+L R+ +S + Y
Sbjct: 99 TESKRLREDIVEAIKCMLDQCNPYVKVLRMAKERFDHSIEMPNLKLKLVSGRSTNSGKTY 158
Query: 177 NVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRY 236
N+P +SEV L VGDFD+ RDI+V K R +H YP+LFP GED Y
Sbjct: 159 NLPTVSEVGGLFVGDFDNSVGPRDIIVETKT---RELH----------YPLLFPYGEDGY 205
Query: 237 EEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVE 296
++L +TT S R +VS+REF AY++ +R +E+ + +LFQQF+VD +M+E
Sbjct: 206 HIDLELKKKTTN-SENPRTKVSMREFFAYKIMERKDESPYLMHTGKLFQQFLVDGLTMIE 264
Query: 297 NQRLSYVRKNQSQIRS 312
+R+SY++ NQ +R+
Sbjct: 265 AERISYLQFNQKTLRA 280
>UniRef100_Q94LS3 Putative helicase [Oryza sativa]
Length = 1501
Score = 470 bits (1210), Expect = e-131
Identities = 252/626 (40%), Positives = 372/626 (59%), Gaps = 16/626 (2%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP++ RRD G + K + L+N VVPYN L+ K+ H+N+E+ N+S SIKYLFK I
Sbjct: 702 FPIYRRRDDGRQIKKGRVNLNNGFVVPYNKDLLAKFQVHMNVEWFNRSRSIKYLFKSICN 761
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
D+ TA +E ++E + DEI+ Y C+Y + +E+ WRIF F +H + P V+RL+FH E
Sbjct: 762 EDDQATAVVEETDEKNNDEIKRYLGCKYTTATEACWRIFKFPLHYQEPPVERLSFHEENE 821
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVENRSWH 693
Q ++F DS+ ++ R R TMF WM+ N+ + + LTY +FP K+T++ + W
Sbjct: 822 QHVIFPDSTDLQEIVRRPRSGVTMFTEWMETNKRHEDARELTYSEFPTKWTWDKNVKKWV 881
Query: 694 L*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGL 753
K G +GR+ P + E Y +R++LN GCT+FEDIRTV+G V++SY+ AC ALG
Sbjct: 882 RRKGGMKIGRIYNAHPASGERYYLRVILNTAKGCTTFEDIRTVNGTVHSSYKSACHALGF 941
Query: 754 LANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRR 813
L +D ++I+ I E SG +R++F ++L ++DP +W +W LS+ I + +
Sbjct: 942 LNDDSEWIECIKEASCWASGMKLRQLFATVLCHCEVTDPKRLWESSWEKLSKDIQHTQSW 1001
Query: 814 TLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGI-------------GKTFLWN 860
LN P L ++ + N + Y GI KT+LW
Sbjct: 1002 ALNFPTSCLTPSHRRKC-ALIEIEKNMRQAGKSLKEYAGIEPPNMAKLSEIENSKTYLWK 1060
Query: 861 ALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLL 920
A++ RSEGKIVL VASS +A+LLL GRTAHS F+IP+ L +E C I++ + A LL
Sbjct: 1061 AITTRLRSEGKIVLAVASSGVAALLLQGGRTAHSAFNIPINLTDEYTCFIKQGSHIADLL 1120
Query: 921 TMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVV 980
SLI+WDEAPM +R FEA D+++RD+ + + PFGG TVVLGGDFRQILP+V
Sbjct: 1121 MKTSLILWDEAPMANRNCFEALDKSLRDVQRCRNENSCQKPFGGMTVVLGGDFRQILPIV 1180
Query: 981 PKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDASLRL-FARWILDVGDGK 1039
PK R V+A I S LW+ V LTKN+RL + + Q + FA WIL +G+G
Sbjct: 1181 PKGRREHTVNATIKCSYLWQHFEVFNLTKNMRLNYVSKDQTEHQKSAEFAEWILQIGNGD 1240
Query: 1040 LGIDNDGEAVIEIPDDICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELV 1099
I D + + +P D+ L+ + I+ESTYP+L N + ++ ++RAIL P E V
Sbjct: 1241 T-ISLDEKGWVRMPSDLLLQKGDDPKAQIIESTYPDLQDNCCKQNYLEERAILCPVNENV 1299
Query: 1100 EKVNDYVLSLVPGEEKEYLSCDTVLK 1125
++N+Y++ + G++ YLS D+V K
Sbjct: 1300 NELNEYIMDQIQGDKVTYLSRDSVSK 1325
Score = 129 bits (325), Expect = 4e-28
Identities = 76/191 (39%), Positives = 113/191 (58%), Gaps = 2/191 (1%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISE 183
LD+ + L+K++DE N L + FRM RD + + LRL NR+ D R N+P SE
Sbjct: 305 LDQKTIASLLKMLDENNTLAQTFRMARDRFKEDDYHNYTLRLLDNRDQDGRQDNMPSASE 364
Query: 184 VVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLN 243
V LIV D GRDIV+ KD +RI ETH K + +QYP+LFP GED Y IK +
Sbjct: 365 VALLIVKDPTKKSYGRDIVLEYKDMRPKRISETHPKLMAMQYPLLFPYGEDGYRPGIKYS 424
Query: 244 HRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYV 303
+ + KK V++ E+ AYRLQ+R +++ + L Q++VD ++ +E RL+++
Sbjct: 425 GKEGVRNDKK--CVTMLEYYAYRLQQRQDQSMLPLMCGNLSMQYMVDAHACIEQIRLNWI 482
Query: 304 RKNQSQIRSGL 314
R+NQ +R+ L
Sbjct: 483 RQNQGVLRTEL 493
Score = 36.2 bits (82), Expect = 5.9
Identities = 16/34 (47%), Positives = 21/34 (61%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
L++LD R K P I+ +ICAE+PD PE F
Sbjct: 623 LIFLDKRGKSLEPSQIDELICAEIPDRDKDPETF 656
>UniRef100_Q9SLJ1 F20D21.24 protein [Arabidopsis thaliana]
Length = 1250
Score = 426 bits (1096), Expect = e-117
Identities = 246/628 (39%), Positives = 365/628 (57%), Gaps = 36/628 (5%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F L+ RR+ VLK +LDNR VVP+N +++ KY AHIN+E+CNKS++IKYLFKYITK
Sbjct: 452 FILYRRRNDQRYVLKGQTRLDNRFVVPHNLEILKKYKAHINVEWCNKSSAIKYLFKYITK 511
Query: 574 GVDRVTATLETS------------EEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYP 621
GVD+ T ++ EE +EI Y DCRYLS E++WRIF F++H P
Sbjct: 512 GVDKATFIIQKGNSVNGQGSGNGFEEKPRNEINEYLDCRYLSACEAMWRIFMFNIHHHNP 571
Query: 622 AVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPR 681
V+RL HL GEQ +F + +NV R H+RTM + + N+ + L YVQ P
Sbjct: 572 PVQRLPLHLPGEQSTIFEEEENLENVEYRYGHERTMLTEYFELNKICEDARKLKYVQVPT 631
Query: 682 KFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVY 741
F ++ N+ + K +++GR+ + P +LY +R+LLN+ G TSF+ ++TV G V+
Sbjct: 632 MFVWDSTNKMYTRRKQRENIGRIVNILPTAGDLYYLRILLNKVKGATSFDYLKTVGGVVH 691
Query: 742 NSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWH 801
S++ AC GLL D+++ D + E + +R +F +L+ +S+PL +W+ W
Sbjct: 692 ESFKAACHTRGLLDGDKEWHDAMDEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHCWE 751
Query: 802 ILSEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNA 861
+++ + ++++ LN P L +++L Y ++ + H + Y
Sbjct: 752 SMADDVFRKQQKVLNFPQLELKAEELEKY-TLIEIETLLRQHEKSLSDY---------PE 801
Query: 862 LSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLT 921
+ +S GK V+ VASS+IA+LLLP GRTAHS+F IP+ + E+S C I+ A +L+
Sbjct: 802 MPQPEKSNGKNVMPVASSAIAALLLPGGRTAHSRFKIPINVHEDSICDIKIGSMLANVLS 861
Query: 922 MASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVP 981
LIIWDEAPM R FEA DRT+RDI+S + A FGGKTV+LGGDFRQILPV+P
Sbjct: 862 KVDLIIWDEAPMAHRHTFEAVDRTLRDILSVGDEKALTKTFGGKTVLLGGDFRQILPVIP 921
Query: 982 KRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKL- 1040
+ R + V A IN S LW C L++N+R+Q P + FA WIL VGDG+
Sbjct: 922 QGTRQETVSAAINRSYLWESCHKYLLSQNMRVQ-PEEIK-------FAEWILQVGDGEAP 973
Query: 1041 ----GIDNDGEA-VIEIPDDICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPT 1095
GID+D E I I ++ L + N + + S +P+ + ++ + A+L P
Sbjct: 974 RKTHGIDDDQEEDNIIIDKNLLLPETENPLEVLCRSVFPDFTNTFQDLENLKGTAVLTPR 1033
Query: 1096 LELVEKVNDYVLSLVPGEEKEYLSCDTV 1123
E V+++NDY+LS VPG KEY S D++
Sbjct: 1034 NETVDEINDYLLSKVPGLAKEYFSADSI 1061
Score = 42.0 bits (97), Expect = 0.11
Identities = 29/91 (31%), Positives = 43/91 (46%), Gaps = 8/91 (8%)
Query: 228 LFPLGEDRYEE------KIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGM 281
+F DRYE IKL + G K+ + E +Q R E I +
Sbjct: 153 VFRKARDRYEAGDCPEFSIKLIGQKKKG--KQYDMPTTDEIAGLIIQTRLTEGMTIVRSK 210
Query: 282 RLFQQFIVDLYSMVENQRLSYVRKNQSQIRS 312
RL Q+IVD Y+ +E +RL + R NQ ++R+
Sbjct: 211 RLLHQYIVDAYTSIEQERLRWYRLNQKKLRA 241
Score = 35.4 bits (80), Expect = 10.0
Identities = 20/66 (30%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISE 183
L+ +V+DL+ +D N L K FR RD EA ++L + + Y++P E
Sbjct: 132 LNEKIVKDLITTVDTFNCLAKVFRKARDRYEAGDCPEFSIKLIGQKK-KGKQYDMPTTDE 190
Query: 184 VVALIV 189
+ LI+
Sbjct: 191 IAGLII 196
>UniRef100_Q6AUR0 Hypothetical protein OSJNBa0077L08.8 [Oryza sativa]
Length = 807
Score = 404 bits (1038), Expect = e-111
Identities = 231/621 (37%), Positives = 349/621 (56%), Gaps = 34/621 (5%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F + R + G TV KNG+ +DNR VVP+N L++K+ AHIN+E N KYLFKY+TK
Sbjct: 25 FTQYARPNNGRTVTKNGVDVDNRFVVPHNVDLVVKFQAHINLEKVNYDGMHKYLFKYVTK 84
Query: 574 GVDRVTATLET-----SEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTF 628
G D + + S +++EI Y +CR ++P+++ WR+ FD+H P+V+RL
Sbjct: 85 GFDCSRVGIHSNSGSQSSSETINEIDNYLECRCVTPNDAAWRLQQFDIHHTDPSVERLLV 144
Query: 629 HLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVE 688
HL E ++F + + V+ T AW++ANR + LTY++FP +T++ +
Sbjct: 145 HLPFENNVIFTEDDNLEEVIEDPNSSTTKLTAWLEANRENPSARQLTYIEFPEHWTWHNQ 204
Query: 689 NRSWHL*KSGQS-VGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREA 747
+ W + Q +GR+ +V P E Y +R+LL+ G SF +IRT+ GH Y ++R A
Sbjct: 205 GKYWDGRRGSQCRIGRIAYVNPSQGEAYYLRMLLHIVKGPRSFAEIRTISGHEYPTFRAA 264
Query: 748 CAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGI 807
C ALGLL +D++ + L +I GS + + L+L D
Sbjct: 265 CEALGLLGDDQECLPLPDDI-----GSVLAQ--NRLILDELSYD---------------- 301
Query: 808 LYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFR 867
+Y T++ LN+ Q + I N N FFV GYGG KTFLW L R
Sbjct: 302 VYNMPSTIDEAISGLNNSQKEVFNAIYNSAINNEGRTFFVYGYGGTRKTFLWTTLLNSIR 361
Query: 868 SEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLII 927
S+GKI L VASS IASLLLP GRT HS+F IPL + + S C+I+K N A+L+ SLI+
Sbjct: 362 SQGKIALVVASSGIASLLLPGGRTPHSRFKIPLEISQNSMCSIKKNTNLAELIQKTSLIV 421
Query: 928 WDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRAD 987
WDEAP+ + FE+ DRT+RDI+S + + FGG TVVLGGDFRQ LPV+ +
Sbjct: 422 WDEAPVNHKYCFESLDRTLRDILSETNPNSLDKQFGGITVVLGGDFRQTLPVIQNATKQQ 481
Query: 988 IVDACINSSMLWRRCRVLKLTKNIRLQFPT-NGQDDASLRLFARWILDVGDGKLGIDNDG 1046
I+ +CI +S LW +C +++LT+N+RL + + QD LR F+ W+L +G+G +
Sbjct: 482 ILRSCIVNSYLWNKCILIELTENMRLTSASISAQDREELRNFSNWLLKIGNGTEPFVDVP 541
Query: 1047 EAV----IEIPDDICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKV 1102
E + IE+P + L ++ ++ Y +S+ +RAILAPT E+V ++
Sbjct: 542 EQLTNMFIEMPQSLLLSPDCRNLDGLISFVYNLGCQPSNLTSYLCERAILAPTNEVVSEI 601
Query: 1103 NDYVLSLVPGEEKEYLSCDTV 1123
N+ +++ + E Y S D++
Sbjct: 602 NNRMIAQLEASEMSYYSSDSI 622
>UniRef100_Q84QR0 Helicase-like protein [Oryza sativa]
Length = 1330
Score = 358 bits (918), Expect = 7e-97
Identities = 214/641 (33%), Positives = 346/641 (53%), Gaps = 29/641 (4%)
Query: 505 CQHDVEFKCFPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSI 564
C+H + R V KNG+ +DNR +VP+N L++KY HIN+E N
Sbjct: 512 CEHTYILQNGFTQYARPNNQIVTKNGVDIDNRVIVPHNVDLVVKYQDHINLESVNHDGMH 571
Query: 565 KYLFKYITKGVDRVTATLE-TSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAV 623
KYLFKY+TK D A + S +++EI Y +CR ++P++ WR+ FD+H P+V
Sbjct: 572 KYLFKYVTKVYDCSRAGIRRNSANETINEIDNYLECRCVTPNDVAWRLLQFDIHHTDPSV 631
Query: 624 KRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKF 683
+RL HL E +++ + + V+ ++++ AW++AN + Q+ TY++FP F
Sbjct: 632 ERLPVHLPLENNVVYTEDDDLEEVIENPGNQKSKLTAWLEANSQFPQAREHTYIEFPEYF 691
Query: 684 TYNVENRSWHL*KSGQS-VGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYN 742
T++ + W + + + +GR+ V P E Y +R+LL+ G +F +IR + G +
Sbjct: 692 TWHASEKYWDIRRGCYNKIGRIAHVDPTKGEKYYLRMLLHIVKGPKTFSEIRNISGQQHP 751
Query: 743 SYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHI 802
++R AC ALGLL +D+++ +++ V +R++F ++LL +++P ++ +
Sbjct: 752 TFRAACEALGLLGDDQEWSHALNDAVHWALPYQLRQLFVTILLFCEVTNPQRLFTEHAQH 811
Query: 803 LSEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYG----------G 852
+SE Y + L+ S+ A + N + ELD + GY
Sbjct: 812 MSEDFRYRTNQNLSQ------SNSSFADSFVGNALLFELDKLLRNAGYSLSHFNLPLPDD 865
Query: 853 IGKT-----FLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESG 907
IG L + LSY S N + IA+LLLP GRT HS+F IPL + E S
Sbjct: 866 IGSASADNRLLLDELSYDITSIASTSANDINR-IAALLLPGGRTPHSRFKIPLDIRENSM 924
Query: 908 CTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTV 967
C+I+K + A+L+ SLI+WDEAP+ + FEA D T+R I+S++ A + FGG TV
Sbjct: 925 CSIKKNTHLAELIQQTSLIVWDEAPVNHKYCFEALDHTLRYILSDIRPNAQHRQFGGITV 984
Query: 968 VLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPT-NGQDDASLR 1026
GGDFRQ LPV+ R I+ A I +S LW +C VL+LT+N+RL + D LR
Sbjct: 985 AFGGDFRQTLPVIQNATRHQILRASIVNSYLWHQCVVLQLTENMRLSSQNLSPSDKEELR 1044
Query: 1027 LFARWILDVGDG---KLGIDNDGEAV-IEIPDDICLKNSGNHVGDIVESTYPNLLSNMKE 1082
+FA W+L VG+G + I+N+ IEIP + L + ++ ++ Y
Sbjct: 1045 VFADWVLRVGNGTEPHISIENETNGTFIEIPQSLLLPSDSRNLDSLISFVYDLGYEPSNI 1104
Query: 1083 SSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTV 1123
+++F DRAIL+PT ++V ++N+ +++ V E Y S DT+
Sbjct: 1105 TTYFCDRAILSPTNKVVSEINNKIIAQVTAAEISYYSSDTI 1145
Score = 119 bits (297), Expect = 7e-25
Identities = 64/193 (33%), Positives = 107/193 (55%), Gaps = 1/193 (0%)
Query: 112 LNYHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNF 171
+N S D + +V+ L+++ D N +VK FR R+ N K+RLF + +
Sbjct: 138 INIASSSRDSFHANEEIVRSLIEMFDTHNPIVKLFRTARERLSENESDHYKIRLFGSTDA 197
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPL 231
+Y+ P +EVV L+VGD + GRDI+++ L+RI E H K++ +QYP+LFP
Sbjct: 198 HGDIYSAPVAAEVVGLVVGDIGVTDIGRDIIIQHHSSQLQRIDEKHRKFMAMQYPILFPY 257
Query: 232 GEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDL 291
GED + E + ++T + S +R + ++ EF AY + R + + + RL Q ++VD
Sbjct: 258 GEDGFHES-NMYNQTASSSALRRNKATMVEFFAYIMHDRAGQFNTPLRCGRLTQSYLVDG 316
Query: 292 YSMVENQRLSYVR 304
Y VE +R+ + R
Sbjct: 317 YCCVEGERIQHYR 329
>UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana]
Length = 1265
Score = 355 bits (911), Expect = 4e-96
Identities = 228/664 (34%), Positives = 322/664 (48%), Gaps = 145/664 (21%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR V K GI+ DNR V+PYN K ++Y+AHIN+E+CN+++SIKYLFKYI K
Sbjct: 542 YPIYRRRKIDDYVEKGGIKCDNRYVMPYNKKFSLRYNAHINVEWCNQNDSIKYLFKYINK 601
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
G D+V +E +++ +
Sbjct: 602 GPDKVIFIVEPTQQATAG------------------------------------------ 619
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNY-----SQSWNLTYVQFPRKFTYNVE 688
DS P K+ W D RN ++ Y + P FT++ E
Sbjct: 620 ------DSETPQQEQRSAEKKKNEIKDWFDCRRNAVGKNGKRARECLYAEIPAYFTWDGE 673
Query: 689 NRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREAC 748
N+++ G S+GR+ +V ++ Y +R+LLN +
Sbjct: 674 NKAFKKRTRGFSIGRIHYVSRKMEDDYFLRVLLNISV----------------------- 710
Query: 749 AALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGIL 808
F L E F LLL+ +S P +VW+QTWHIL+E IL
Sbjct: 711 ----------LFWRLSQEF------------FAMLLLSDSLSRPAHVWSQTWHILAEDIL 748
Query: 809 YERRRTLNSPG*L----------------------------------------LNSDQLS 828
++R +P + L +Q
Sbjct: 749 KKKRDEFKNPEDIDEFPKPTIDGIDNSNRLIVEELRYNRESNLKEKHEEWKQMLTPEQRG 808
Query: 829 AYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPR 888
Y +I V N L +FFV G+GG GKTF+W LS R +IVLNVASS IASLLL
Sbjct: 809 VYNEITEAVFNNLGGVFFVYGFGGTGKTFIWKTLSATIRYRDQIVLNVASSGIASLLLEG 868
Query: 889 GRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRD 948
GRTAHS+F IPL E S C I+ + + A L+ ASL+IWDEAPM+SR FEA D++ D
Sbjct: 869 GRTAHSRFGIPLNPDEFSVCKIKPKSDLANLVKKASLVIWDEAPMMSRFCFEALDKSFSD 928
Query: 949 IMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLT 1008
I+ N N FGGK VV GGDFRQ+ PV+ GRA+IV + +N+S LW C+VLKLT
Sbjct: 929 IIKN----TDNTVFGGKVVVFGGDFRQVFPVINGAGRAEIVMSSLNASYLWDNCKVLKLT 984
Query: 1009 KNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGD 1067
KN RL + +A ++ F+ W+L VGDG++ NDG A+I+IP+D+ + N+ +
Sbjct: 985 KNTRLLANNLSETEAKEIQEFSDWLLAVGDGRINESNDGVAIIDIPEDLLITNADKPIES 1044
Query: 1068 IVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVLK 1125
I Y P +L + + FFQ RAILA E V +N+Y+L + EE+ YLS D++
Sbjct: 1045 ITNEIYGDPKILHEITDPKFFQGRAILASKNEDVNTINEYLLDQLHAEERIYLSADSIDP 1104
Query: 1126 CDEE 1129
D +
Sbjct: 1105 TDSD 1108
Score = 139 bits (350), Expect = 5e-31
Identities = 71/190 (37%), Positives = 122/190 (63%), Gaps = 3/190 (1%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-FDSRVYNVPEIS 182
LD++L++ ++K+++ CN V+KFR R+ + N + P +R+ +R D R Y++ S
Sbjct: 143 LDKNLIEVIIKMLNRCNPYVRKFRTARERIQTNDEEPFHMRIIADRQGVDGRTYSMHTTS 202
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIK 241
EV ALI GDF RDIV+ +K G L+RI++ H Y+ LQYP++F GED + I+
Sbjct: 203 EVAALIPGDFRHGMPDRDIVIEKKSNGHLKRINQIHISYLALQYPLIFCYGEDGFRPGIE 262
Query: 242 LNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLS 301
++ + K+ +S+R++ A+R+Q+R+ E + + RLFQQF+ D Y+ +E+ RL+
Sbjct: 263 KCFKSKSKKKNKKC-ISMRQWFAFRIQEREVECQTLLRSKRLFQQFLCDAYTTIESNRLN 321
Query: 302 YVRKNQSQIR 311
Y++ NQS++R
Sbjct: 322 YIKFNQSKLR 331
>UniRef100_Q9LIU0 Similar to Arabidopsis thaliana chromosome II BAC T13P21 genomic
sequence [Oryza sativa]
Length = 1278
Score = 326 bits (835), Expect = 3e-87
Identities = 193/467 (41%), Positives = 266/467 (56%), Gaps = 45/467 (9%)
Query: 706 FVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLANDRQFIDLIS 765
++ P ELY +R+LL G F DIRT +G VY ++R+AC A GLL ND ++ L
Sbjct: 607 YISPVAGELYYLRMLLMIVKGVMCFADIRTYEGVVYPTFRQACEARGLLENDNEWHLLFD 666
Query: 766 EIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERR------------- 812
E +V S +R++F ++++ + + +++++ W ++ I + R
Sbjct: 667 EAIVSASSYQLRQLFVTVVMFCSIGNVRSLFDKYWTYFTDDIQHRVRKMFGGNIDDYDLP 726
Query: 813 ----RTLNSPG*L-----------------------LNSDQLSAYEKIVNVVDNELDHMF 845
RT + G LNSDQ ++ I+ V+ F
Sbjct: 727 QSTARTDDDSGNRMVSEELALDSVALAAHADSIIPKLNSDQKRVFDTIMCRVNESKPGFF 786
Query: 846 FVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEE 905
FV G+GG GKTFL NAL RSE KIVL VASS +ASLLLPRGRTAHS+F IP+ + E
Sbjct: 787 FVYGHGGTGKTFLCNALISKVRSEKKIVLAVASSGVASLLLPRGRTAHSRFKIPIDINEN 846
Query: 906 SGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGK 965
S CTI++ A+L+ SLIIWDEAPM R FEA DRT+RD++S +PFGGK
Sbjct: 847 SLCTIKRGTMLAELIQKTSLIIWDEAPMTHRRCFEALDRTLRDLLSEHAPSNGLVPFGGK 906
Query: 966 TVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIR-LQFPTNGQDDAS 1024
VVLGGDFRQILPVV K RA I+DA I +S LW +LKLT N+R LQ Q
Sbjct: 907 VVVLGGDFRQILPVVRKGSRASIIDASITNSPLWSHAVLLKLTVNMRLLQCNLGEQQQQE 966
Query: 1025 LRLFARWILDVGDGKLGIDNDGEAV-IEIPDDICLKNSGNHVGDIVESTYPNLLSNMKES 1083
L FA+W+L +GDGK +++ EA I+IPDD+ +K +G+ + IV +P S +
Sbjct: 967 LEKFAQWVLALGDGK---NDESEATWIDIPDDLLIKTTGDKIHSIVNEVFPGFASKYTDP 1023
Query: 1084 SFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVLKCDEEV 1130
S+ AI+ P V+ +ND ++ +VPGE KEYLSCDT+ K E +
Sbjct: 1024 SYLASCAIVCPNNSTVDDINDRMVDMVPGEVKEYLSCDTISKSSEHI 1070
Score = 112 bits (281), Expect = 5e-23
Identities = 58/107 (54%), Positives = 71/107 (66%), Gaps = 10/107 (9%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F +++RR+ G V+KNGI+LDNR VVPYN KL+ KY AHIN+E+CNKSN IKYLFKY+TK
Sbjct: 495 FTVYMRRNNGRYVVKNGIKLDNRWVVPYNMKLLKKYQAHINVEWCNKSNMIKYLFKYVTK 554
Query: 574 GVDRVTATLET----------SEEPSVDEIQ*YYDCRYLSPSESIWR 610
G DR ET S +EI Y + R+LS ESIWR
Sbjct: 555 GSDRTKVYFETTGNTANKPTDSNAAPRNEIDEYINARFLSTCESIWR 601
Score = 110 bits (274), Expect = 3e-22
Identities = 67/188 (35%), Positives = 98/188 (51%), Gaps = 1/188 (0%)
Query: 125 DRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISEV 184
D +V +L ++DE N LVK FR R + + D + LRL + D YN+P E+
Sbjct: 102 DPFIVTELGAMLDEHNDLVKSFRFARKRLKDHGDEKIALRLLGCNSKDEVQYNLPTSGEI 161
Query: 185 VALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNH 244
+IVGD+ + + D+VV+ D LRR+ H Y+ LQYP+LFP GE + IK
Sbjct: 162 AGIIVGDYSNDKYTYDVVVQSCDSRLRRVSALHPSYMALQYPLLFPYGERGFHLGIKYTD 221
Query: 245 RTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYVR 304
+ +R V++ E+ YRL R N+ + RL VD+YS VE RL ++
Sbjct: 222 FPSITGTSRRY-VTMLEYYRYRLHYRLNKPNPYTCCGRLSDSICVDMYSTVEGSRLKFIA 280
Query: 305 KNQSQIRS 312
NQ +RS
Sbjct: 281 DNQPDLRS 288
>UniRef100_Q7XF22 Helicase-like protein [Oryza sativa]
Length = 1336
Score = 324 bits (830), Expect = 1e-86
Identities = 202/546 (36%), Positives = 295/546 (53%), Gaps = 60/546 (10%)
Query: 560 KSNSIKYLFKYITKGVDRVTATL-ETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHC 618
+ N +KYLFKY+ KG DR + ++ E +DEIQ Y D R+++P E++WRI+GFD+
Sbjct: 714 RRNDVKYLFKYLYKGHDRASISINEADSNGEIDEIQRYRDARWVTPPEALWRIYGFDICH 773
Query: 619 RYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQ 678
P+V++L HL + F +VL + R+M A+ +ANR + + ++ Y
Sbjct: 774 ISPSVRQLQLHLPNMHMLAFDADKDLRDVLDKEDAGRSMLTAYFEANRQHVWARDILYRD 833
Query: 679 FPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDG 738
FP FT+ E Y +R+LLN G TSFED+RTVDG
Sbjct: 834 FPMWFTWQTAEG----------------------ERYYLRVLLNHVAGSTSFEDLRTVDG 871
Query: 739 HVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQ 798
V S+R A GL+ D + + E V +S+ ++F ++L+ SD +W++
Sbjct: 872 VVMPSFRAAAERRGLIEADNTLDECLMEAAVFQMPASLCRLFATILVYCEPSDVRGLWDK 931
Query: 799 TWHILSEGILYERRRTL--------NSPG*---------LLNSDQLSAYEKIVNVVDNEL 841
+S+ Y R T +S G LN +Q SA+ KI+N V +
Sbjct: 932 HLDAMSDD--YRRNNTCPHVVQQMESSIGVETDDMNLSDQLNDEQRSAFNKIMNAVGSAQ 989
Query: 842 DHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLM 901
+FFVDG GG GKTFL+ AL R +G I + A+S +A+ ++P GRTAHS+F IPL
Sbjct: 990 GGVFFVDGPGGTGKTFLYRALLATVRGKGDIAVATATSGVAASIMPGGRTAHSRFKIPLN 1049
Query: 902 LLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLP 961
+ E S C+ K+ A+LL MASLIIWDEA M R A EA D +MRDIM G P
Sbjct: 1050 IDEGSYCSFTKQSGTAKLLQMASLIIWDEASMTKRQAVEALDMSMRDIM-----GCPRSP 1104
Query: 962 FGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQD 1021
FGGKT+V GGDFRQ+LPV+ K R+ I DA + S LW LKL +N+R Q T
Sbjct: 1105 FGGKTIVFGGDFRQVLPVIRKGTRSQITDATLRRSYLWDCMVQLKLVRNMRAQSDT---- 1160
Query: 1022 DASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNH--VGDIVESTYPNLLSN 1079
FA ++L VG+G ++ +G +I +P DIC+ GN + ++++ +PNL N
Sbjct: 1161 -----WFADYLLRVGNGTEEVNKEG--LIGLPSDICVSCKGNETDLERLIDTVFPNLNDN 1213
Query: 1080 MKESSF 1085
+ + ++
Sbjct: 1214 LTDPNY 1219
Score = 90.1 bits (222), Expect = 3e-16
Identities = 60/197 (30%), Positives = 96/197 (48%), Gaps = 9/197 (4%)
Query: 127 SLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISEVVA 186
SL QD+++ + + R +A+ ++ L + D R YNVP +EV A
Sbjct: 306 SLDQDVIRTVSDVLRNNPYSETFRSLGQADDLANYRVTLNLDHRLDQRRYNVPVTTEVAA 365
Query: 187 LIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNHRT 246
+ V + + + + D + I + Y PL YP+ FP GE + + + + T
Sbjct: 366 VWVEGNERRKFEPSVTIYGNDCTRKCIQPFYGCYDPLSYPLFFPRGESGWHQGLPKDKIT 425
Query: 247 TTGSVKK---------RVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVEN 297
+ ++ R+RVS+R++ Y+ Q R + I G RLFQQF VD+Y VE+
Sbjct: 426 MKAANERHGDDPDCNSRIRVSVRDYYCYKFQMRRGIFNPILHGGRLFQQFAVDMYIKVES 485
Query: 298 QRLSYVRKNQSQIRSGL 314
RL YVR NQ +IR+ L
Sbjct: 486 SRLDYVRNNQKEIRADL 502
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 306 bits (783), Expect = 3e-81
Identities = 157/305 (51%), Positives = 209/305 (68%), Gaps = 2/305 (0%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LN++Q A++ I+ + L + FVDGYGG GKT+LW A++ RSEGKIVL VAS I
Sbjct: 1047 LNAEQKVAFDSIIESTNKGLGKLMFVDGYGGTGKTYLWRAITTKLRSEGKIVLTVASCGI 1106
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
A+LLL GRTAHS+F IPL++ EES C I++ + A+LL SLI+WDEAPM +R+ FEA
Sbjct: 1107 AALLLHGGRTAHSRFHIPLIVTEESTCDIKQGSHLAELLKKTSLILWDEAPMANRICFEA 1166
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DR++RDI+ + + S PFGG TVVLGGDFRQILPVV K R IV+A I S LW+
Sbjct: 1167 LDRSLRDILRSKGEDNSTKPFGGMTVVLGGDFRQILPVVRKGRRTQIVNASIKRSYLWQH 1226
Query: 1002 CRVLKLTKNIRLQFPTNGQDDASLRL-FARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
+ KLT+N+RL + +D+ FA+WIL++GDGK DGE IEIPDD+ LK
Sbjct: 1227 FHIFKLTRNMRLSCISRDEDEQKRTADFAQWILNIGDGKT-TSADGEEWIEIPDDLILKK 1285
Query: 1061 SGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSC 1120
G+ +IV+S YPNL+ N K+ F + RAIL P E K+N+++++++ GEE YLSC
Sbjct: 1286 GGDPKEEIVKSIYPNLVQNYKKRDFLEQRAILCPRNETARKINEFIMNMIEGEEITYLSC 1345
Query: 1121 DTVLK 1125
DTV K
Sbjct: 1346 DTVCK 1350
Score = 270 bits (690), Expect = 2e-70
Identities = 132/320 (41%), Positives = 201/320 (62%), Gaps = 4/320 (1%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP + RRD G + K ++LDNR VVPYN L++KY AHIN+E CN+S SIKYLFKY+ K
Sbjct: 675 FPTYRRRDNGRYIEKGNVKLDNRYVVPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHK 734
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
G D+ TA +E+ DEI+ Y +C Y+S ++ WRIF F++H RYP+V+RL FHL E
Sbjct: 735 GDDQATALIESDH----DEIKKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENE 790
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVENRSWH 693
Q+++F DS+ ++ + R T F WM+ N+ ++ + TY +FP K+ + + + W+
Sbjct: 791 QQVIFPDSADLRKIVRKERIGVTKFTQWMETNKINDEARDFTYAEFPSKWVWKNKLKQWN 850
Query: 694 L*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGL 753
K G+ +GR+ + P + + Y +R+LLN G +FE+IRTVDG V+ S++ AC ALG
Sbjct: 851 KRKKGKMIGRIYYAHPASGDKYYLRMLLNTVKGPRTFEEIRTVDGVVHPSFKSACEALGF 910
Query: 754 LANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRR 813
L +DR++++ I E SG+ +R +FT++L ++DP +W W L E I Y++R+
Sbjct: 911 LDDDREWVECIREASNYASGNQLRHLFTTILCHCEVTDPKRIWESCWEDLGEDIEYKQRK 970
Query: 814 TLNSPG*LLNSDQLSAYEKI 833
LN P L Q + I
Sbjct: 971 NLNYPTLRLTEQQKKGHALI 990
Score = 142 bits (357), Expect = 8e-32
Identities = 81/192 (42%), Positives = 115/192 (59%), Gaps = 4/192 (2%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRD-FREANVDVPVKLRLFRNRNFDSRVYNVPEIS 182
+D +V L ++D N+L + FRM RD F+E + V LRL R R D R +N+P S
Sbjct: 278 IDSHIVLGLKNMLDRENVLAQTFRMARDRFKEGDYH-NVSLRLIRKRGGDGRQHNMPSAS 336
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKL 242
EV ALIV D + GRDI+V KD G RRI E H K++ +QYP+LFP GED + KI
Sbjct: 337 EVAALIVNDTSENQKGRDIIVHYKDTGPRRISENHPKFMAMQYPLLFPYGEDGFTNKIL- 395
Query: 243 NHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSY 302
+R GS KR +++ E+ AYR+Q+R N+ + +L QFIVD + + RL +
Sbjct: 396 -YRDNHGSKCKRKHLTMLEYYAYRIQQRKNQCMHLLMCEKLTLQFIVDALACIIQYRLDW 454
Query: 303 VRKNQSQIRSGL 314
+RK+Q +R+ L
Sbjct: 455 IRKHQGNLRTEL 466
>UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thaliana]
Length = 830
Score = 305 bits (782), Expect = 4e-81
Identities = 174/434 (40%), Positives = 255/434 (58%), Gaps = 41/434 (9%)
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+D++TV G V+ S+R+A ALGLL +D+++I+ I + S +R++F +LL+ ++
Sbjct: 53 KDLKTVKGVVHKSFRDAVFALGLLDDDKEYINAIKDANFWCSAKYVRRLFVIMLLSESLT 112
Query: 791 DPLNVWNQTWHILSEGI--------------LYERRRTLNSPG*LLNS------------ 824
P VW++TW ILS+ I L E R L G L+
Sbjct: 113 KPEMVWDETWRILSKDIEHLQLSDEERQQYCLQEIARLLTKNGVSLSKWNRCHKFQMNTM 172
Query: 825 ------DQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVAS 878
Y++I++VV ++ +FFV G+GG GKTFLW LS RS+G I LNVAS
Sbjct: 173 VDYGDFRAKKIYDEIMDVVLHDRGGVFFVYGFGGTGKTFLWKLLSAAVRSKGDISLNVAS 232
Query: 879 SSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLA 938
S IA+L L GRTAHS+F IP+ E S C I + + +L+ A LIIWDEAPM+S+
Sbjct: 233 SGIAALRLDGGRTAHSRFDIPINPNESSTCNISRGSDLGELVKEAKLIIWDEAPMMSKHC 292
Query: 939 FEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSML 998
FE+ DRT++DI++N D P GGK +V GGDFRQ+LPV+ GR +IV A +NSS +
Sbjct: 293 FESLDRTLKDIVNNPGD----KPLGGKVIVFGGDFRQVLPVINGAGREEIVFAALNSSYI 348
Query: 999 WRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICL 1058
W +VL+LTKN+RL + + + F++WILDVGDGK+ NDG A+I+IP++ +
Sbjct: 349 WEHSKVLELTKNMRLLADISEHEKRDIEDFSKWILDVGDGKISQPNDGIALIDIPEEFLI 408
Query: 1059 KNSGNHVGDIVESTYPNLLSNMKESS-----FFQDRAILAPTLELVEKVNDYVLSLVPGE 1113
+ V I+E+ Y N K+ +Q RAIL PT E V +N++++ ++ GE
Sbjct: 409 NGDNDPVESIIEAVYGNTFMEEKDPKKTDYPQYQGRAILCPTNEDVNSINEHMMRMLDGE 468
Query: 1114 EKEYLSCDTVLKCD 1127
E+ YLS D++ D
Sbjct: 469 ERIYLSSDSIDPAD 482
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 304 bits (779), Expect = 9e-81
Identities = 156/305 (51%), Positives = 209/305 (68%), Gaps = 2/305 (0%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LN++Q A++ I+ + L + FVDGYGG GKT+LW A++ RSEGKIVL VAS I
Sbjct: 1156 LNAEQKVAFDSIIESTNKGLGKLMFVDGYGGTGKTYLWRAITTKLRSEGKIVLTVASCGI 1215
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
A+LLL GRTAHS+F IPL++ EES C I++ + A+LL SLI+WDEAPM +R+ FEA
Sbjct: 1216 AALLLHGGRTAHSRFHIPLIVTEESTCDIKQGSHLAELLKKTSLILWDEAPMANRICFEA 1275
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DR++RDI+ + + S PFGG TVVLGGDFRQILPVV K R IV+A I S LW+
Sbjct: 1276 LDRSLRDILRSKGEDNSTKPFGGMTVVLGGDFRQILPVVRKGRRTQIVNASIKRSYLWQH 1335
Query: 1002 CRVLKLTKNIRLQFPTNGQDDASLRL-FARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
+ KLT+N+RL + +D+ FA+WIL++GDGK DGE IEIPDD+ LK
Sbjct: 1336 FHIFKLTRNMRLSCISRDEDEQKRTADFAQWILNIGDGKT-TSADGEEWIEIPDDLILKK 1394
Query: 1061 SGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSC 1120
G+ +IV+S YPNL+ N K+ F + RAIL P E ++N+++++++ GEE YLSC
Sbjct: 1395 GGDPKEEIVKSIYPNLVQNYKKRDFLEQRAILCPRNETAREINEFIMNMIEGEEITYLSC 1454
Query: 1121 DTVLK 1125
DTV K
Sbjct: 1455 DTVCK 1459
Score = 273 bits (698), Expect = 2e-71
Identities = 134/320 (41%), Positives = 203/320 (62%), Gaps = 4/320 (1%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP + RRD G + K ++LDNR VVPYN L++KY AHIN+E CN+S SIKYLFKY+ K
Sbjct: 784 FPTYRRRDNGRYIEKGNVKLDNRYVVPYNRDLLVKYQAHINVERCNRSKSIKYLFKYMHK 843
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
G D+ TA +E+ DEI+ Y +C Y+S ++ WRIF F++H RYP+V+RL FHL E
Sbjct: 844 GDDQATALIESDH----DEIKKYLECTYISGHDACWRIFQFEMHYRYPSVERLPFHLENE 899
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVENRSWH 693
Q+++F DS+ ++ + R T F WM+ N+ ++ +LTY +FP K+ + + + W+
Sbjct: 900 QQVIFPDSADLRKIVRKERIGVTKFTQWMETNKINDEARDLTYAEFPSKWVWKNKLKQWN 959
Query: 694 L*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGL 753
K G+ +GR+ + P + + Y +R+LLN G +FE+IRTVDG V+ S++ AC ALG
Sbjct: 960 KRKKGKMIGRIYYAHPASGDKYYLRMLLNTVKGPRTFEEIRTVDGVVHPSFKSACEALGF 1019
Query: 754 LANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRR 813
L +DR++++ I E SG+ +R +FT++L ++DP +W W LSE I Y++R+
Sbjct: 1020 LDDDREWVECIREASNYASGNQLRHLFTTILCHCEVTDPKRIWESCWEDLSEDIEYKQRK 1079
Query: 814 TLNSPG*LLNSDQLSAYEKI 833
LN P L Q + I
Sbjct: 1080 NLNYPTLRLTEQQKKGHALI 1099
Score = 142 bits (357), Expect = 8e-32
Identities = 81/192 (42%), Positives = 115/192 (59%), Gaps = 4/192 (2%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRD-FREANVDVPVKLRLFRNRNFDSRVYNVPEIS 182
+D +V L ++D N+L + FRM RD F+E + V LRL R R D R +N+P S
Sbjct: 387 IDSHIVLGLKNMLDRENVLAQTFRMARDRFKEGDYH-NVSLRLIRKRGGDGRQHNMPSAS 445
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKL 242
EV ALIV D + GRDI+V KD G RRI E H K++ +QYP+LFP GED + KI
Sbjct: 446 EVAALIVNDTSENQKGRDIIVHYKDTGPRRISENHPKFMAMQYPLLFPYGEDGFTNKIL- 504
Query: 243 NHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSY 302
+R GS KR +++ E+ AYR+Q+R N+ + +L QFIVD + + RL +
Sbjct: 505 -YRDNHGSKCKRKHLTMLEYYAYRIQQRKNQCMHLLMCEKLTLQFIVDALACIIQYRLDW 563
Query: 303 VRKNQSQIRSGL 314
+RK+Q +R+ L
Sbjct: 564 IRKHQGNLRTEL 575
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 286 bits (733), Expect = 2e-75
Identities = 145/307 (47%), Positives = 204/307 (66%), Gaps = 4/307 (1%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSS 880
++ S+Q Y++I++ V ++ +FFV G+GG GKTFLW LS RS+G I LNVASS
Sbjct: 543 MVTSEQKKIYDEIMDAVLHDRGGVFFVYGFGGTGKTFLWKLLSAAIRSKGDISLNVASSG 602
Query: 881 IASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFE 940
IA+LLL GRT HS+F IP+ E S C I + + +L+ A+LIIWDE PM+S+ FE
Sbjct: 603 IAALLLDGGRTTHSRFGIPINPNESSTCNISRGSDLGELVKEANLIIWDETPMMSKHCFE 662
Query: 941 AFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWR 1000
+ DRT+RDIM+N D PFGGK +V GGDFRQ+LPV+ GR +IV A +NSS +W
Sbjct: 663 SLDRTLRDIMNNPGDK----PFGGKGIVFGGDFRQVLPVINGAGREEIVFAALNSSYIWE 718
Query: 1001 RCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
C+VL+LTKN+RL + + + F++WILDVGDGK+ NDG A+I+IP++ +
Sbjct: 719 HCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGKISQPNDGIALIDIPEEFLING 778
Query: 1061 SGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSC 1120
+ V I+E+ Y N K+ FFQ RAIL PT E V +N++++S++ GEE+ YLS
Sbjct: 779 DNDPVESIIEAVYGNTFMEEKDPKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSS 838
Query: 1121 DTVLKCD 1127
D++ D
Sbjct: 839 DSIDPAD 845
Score = 257 bits (656), Expect = 2e-66
Identities = 129/337 (38%), Positives = 204/337 (60%), Gaps = 20/337 (5%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP++ RRDTG+ V KNG Q DNR V+PYN K+ ++Y AHIN+E CN+S SIKYLFKY+ K
Sbjct: 149 FPVYRRRDTGIYVEKNGFQCDNRYVIPYNEKVSLRYQAHINVELCNQSGSIKYLFKYVHK 208
Query: 574 GVDRVTATLETSEEPSV----DEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFH 629
G DRVT T+E +++ + DE++ Y+DCRY+S E++WRIF F +H R V +L FH
Sbjct: 209 GHDRVTVTVEPNDQDTAKKEKDEVKDYFDCRYVSACEAMWRIFKFPIHYRTTPVVKLFFH 268
Query: 630 LHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRN----------------YSQSWN 673
G+Q + ++ ++V+ R + T FLAW N+
Sbjct: 269 EEGKQPVYYKPGETTESVMDRLSSEATQFLAWFQLNKKPPSRTIRANAKKLPKAAPDPTK 328
Query: 674 LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDI 733
L + + P FT+N + + + + + G ++GR+ FVP ++ Y +R+LLN + G TS++D+
Sbjct: 329 LLFEEIPNHFTWNSKEKKFMIRERGFAIGRINFVPRTIEDAYYLRILLNIKRGVTSYKDL 388
Query: 734 RTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPL 793
+TV G V+ S+R+A ALGLL +D+++I+ I + S +R++F +LL+ ++ P
Sbjct: 389 KTVKGVVHKSFRDAVFALGLLDDDKEYINGIKDAKFWCSAKYVRRLFVIMLLSESLTKPE 448
Query: 794 NVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
VW++TW ILSE I +R+ P L+ ++ Y
Sbjct: 449 MVWDETWRILSEDIERRKRKEWKRPDLQLSDEERQQY 485
>UniRef100_Q8LRG5 Putative helicase [Oryza sativa]
Length = 1453
Score = 286 bits (732), Expect = 2e-75
Identities = 156/314 (49%), Positives = 203/314 (63%), Gaps = 5/314 (1%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LNS+Q + ++ IV+ V FFV G+GG GKTFLWN L RSEG IVL VASS +
Sbjct: 955 LNSEQQNFFDTIVSRVSESRPGFFFVYGHGGTGKTFLWNVLISKIRSEGNIVLAVASSGV 1014
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLLPRGRTAHS+F IP+ + E S C+I++ A+L+ SLIIWDEAPM R FEA
Sbjct: 1015 ASLLLPRGRTAHSRFKIPIDIDENSICSIKRGTMLAELIQKTSLIIWDEAPMTHRRCFEA 1074
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT+RD++S S LPFGGK VVLGGDFRQILPV+ K R IVDA I +S LW+
Sbjct: 1075 LDRTLRDLLSEHNPSNSVLPFGGKFVVLGGDFRQILPVIKKGTRNSIVDASITNSPLWQH 1134
Query: 1002 CRVLKLTKNIRL-QFPTNGQDDASLRLFARWILDVGDGKL----GIDNDGEAVIEIPDDI 1056
+LKLT N+RL Q + L FARW+L +GDG L ID I+IPDD+
Sbjct: 1135 VVLLKLTVNMRLFQSGLSEGRRHDLEQFARWVLALGDGMLPVSKRIDESEATWIDIPDDL 1194
Query: 1057 CLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKE 1116
++ S + + IV +P + +SS+ RAI+ P V+++NDY+++++PGE KE
Sbjct: 1195 LIRASDDKIYSIVNEVFPCYVHRYTDSSYLASRAIVCPNNSTVDEINDYMVAMIPGEMKE 1254
Query: 1117 YLSCDTVLKCDEEV 1130
YLSCDT+ K E +
Sbjct: 1255 YLSCDTISKTSEHI 1268
Score = 237 bits (604), Expect = 2e-60
Identities = 139/400 (34%), Positives = 219/400 (54%), Gaps = 24/400 (6%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F ++ RR+ G V+KNGI+LDNR VVPYN KL+ KY AHIN+E CNKSN IKYLFKYITK
Sbjct: 570 FTVYRRRNDGRYVVKNGIKLDNRWVVPYNMKLLKKYQAHINVESCNKSNMIKYLFKYITK 629
Query: 574 GVDRVTATLETS---EEPSVD-------EIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAV 623
G DR ET+ +VD EI Y + R+LS E+ WR F FD+H R PAV
Sbjct: 630 GGDRTKLYFETTGNTPNKTVDGTVLPPNEIDEYINARFLSTCEAFWRAFEFDIHYRVPAV 689
Query: 624 KRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKF 683
+RL HL + ++ + +L K+TM W + N+ + + LTY FP+++
Sbjct: 690 ERLPIHLPNMNFVQYKKGTDLKKLLDSPAAKKTMLTEWFECNKKHPNARTLTYCDFPKQW 749
Query: 684 TYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNS 743
T++ R W + +GR+ +V P ELY +R+LL G S+ D+RT +G VY +
Sbjct: 750 TWDNSARCWRPRTPVEKIGRIYYVSPAAGELYYLRMLLMTVKGAKSYADVRTFEGTVYPT 809
Query: 744 YREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHIL 803
+R+AC + GLL ND + L E +V S +R++F ++++ + + +++++ W
Sbjct: 810 FRQACESRGLLENDNDWHLLFDEAIVSASSLQLRQLFVTVVMFCSVGNVRSLFDKYWLYF 869
Query: 804 SEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALS 863
++ I + R L++P ++ D+L ++++ EL H F + G I L +
Sbjct: 870 TDDIQHRLRTALSNPAYVVPHDRL------LSLLIKEL-HSAFANSGGNIDDYDLPRSTI 922
Query: 864 YHFRSEGKIVLN-------VASSSIASLLLPRGRTAHSQF 896
+ G ++N A ++ ASL++PR + F
Sbjct: 923 HSDDEFGNRMVNEELALDTAALAAHASLMIPRLNSEQQNF 962
Score = 103 bits (257), Expect = 3e-20
Identities = 64/188 (34%), Positives = 94/188 (49%), Gaps = 1/188 (0%)
Query: 125 DRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISEV 184
D S+V L ++D+ N LVK FR RD + + + LRL D YN+P E+
Sbjct: 173 DPSIVTGLGAMLDQHNDLVKSFRYARDRLNEHGNEQIALRLLGCNAKDEVQYNLPTSGEI 232
Query: 185 VALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNH 244
+IVGD + D+VV+ D LR++ H Y+ LQYP+LFP GE + IK
Sbjct: 233 AGIIVGDSSNDAYTYDVVVQSSDNRLRQVSALHPSYMALQYPLLFPYGERGFHLGIKYTD 292
Query: 245 RTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYVR 304
+ +R V++ E+ YR R N+ + RL VD YS VE RL ++
Sbjct: 293 FPSIAGTSRRY-VTMLEYYRYRFHYRLNKPNPYTCCGRLSDSICVDAYSTVEGSRLKFIH 351
Query: 305 KNQSQIRS 312
+Q ++RS
Sbjct: 352 DHQPELRS 359
>UniRef100_Q9N4X4 Hypothetical protein Y46B2A.2 [Caenorhabditis elegans]
Length = 1365
Score = 285 bits (728), Expect = 7e-75
Identities = 214/654 (32%), Positives = 330/654 (49%), Gaps = 75/654 (11%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P + RRD G V LDNR VVPYN +L+++Y+AH+N+E C ++KYLFKY+ K
Sbjct: 578 YPGYRRRDDGRYVDYGTQHLDNRRVVPYNKRLLLRYNAHMNVEICGFIEAVKYLFKYVYK 637
Query: 574 GVDRVTATL---ETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHL 630
G DR + + VDEI+ + D RY+ E+I I GF + + + RL HL
Sbjct: 638 GHDRAALNIIQNVRGDGNVVDEIREHLDARYVCAPEAIHHILGFKLEKKSDTIYRLAVHL 697
Query: 631 HGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDAN---RNYSQSWNLT----------YV 677
G Q I FR +S L + T AW N ++ ++S N+ Y+
Sbjct: 698 EGFQTIYFR-ASVTTQQLESSSQTDTTLTAWFKINQKSKDIAESGNIPSTFVDSRQFFYM 756
Query: 678 QFPRKFTYNVENRSWHL*KSG-QSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTV 736
P FT+ V+ W + G + +GR+ VPP E Y +R+LL G TSFED+RTV
Sbjct: 757 DMPTHFTF-VKKDGWKVRGRGTRQIGRMYTVPPYETERYALRILLLNIKGATSFEDLRTV 815
Query: 737 DGH-----VYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSD 791
VY +Y EA A GLL +D +++ + E +++R +F +++L + + D
Sbjct: 816 LDENNVPVVYATYVEAAKAQGLLNDDSEYLKSLKEWAGCSVPAALRSMFVAIILFNEVHD 875
Query: 792 PLNVWNQTWHILSEGILYE-RRRTLNSPG-------------------*LLNSDQLSAYE 831
N W + G ++ + ++N P LN Q A +
Sbjct: 876 ----LNALWDAVKVGKRFDVTKPSINPPPIDLDTVNPAQCASEGNRLLATLNDQQKRAAD 931
Query: 832 KIVNVVDN-ELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGR 890
+I+ +D+ L +F++DG GG GKT+L+ L G V A + IA+ LLP GR
Sbjct: 932 QILAALDDASLPRLFYLDGPGGSGKTYLYITLYNICVGRGLKVACTAWTGIAANLLPLGR 991
Query: 891 TAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIM 950
T+ S F + + +S +++ +AQ L + IWDEA M+ + A + D +RD+
Sbjct: 992 TSASLFKLDIRNQCKSSLH-QRQLKEAQELAENDVFIWDEASMVPKTALDTVDVLLRDLT 1050
Query: 951 SNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKN 1010
+ PFGGK ++LGGDFRQILPVV + RAD VDACI S LW ++L L N
Sbjct: 1051 K------IDQPFGGKILILGGDFRQILPVVERSSRADQVDACIKRSPLWTEFQILHLISN 1104
Query: 1011 IRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVE 1070
+R+ T+G D + +++L+VGDG ND ++ + +P + + +IVE
Sbjct: 1105 MRV---TSGDSD-----WIQFLLNVGDGSA---NDSDSKVTLPLSVMCDH------NIVE 1147
Query: 1071 STYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVL 1124
+ ++ +S D IL P V ++ND V + + GEE+ YLS D V+
Sbjct: 1148 EVFGAVID--PTTSDPCDNVILTPKNVDVAQLNDDVHNRMVGEERIYLSRDEVI 1199
Score = 85.5 bits (210), Expect = 8e-15
Identities = 57/197 (28%), Positives = 98/197 (48%), Gaps = 14/197 (7%)
Query: 125 DRSLVQDLMKLMDECNLLVKKFRMVRDFREAN-------VDVPVKLRLFRNRNF--DSRV 175
D +++ +L L+ N + ++M+ + E V P +RL + + D R
Sbjct: 181 DPAVMAELSSLLLRTNPYAQAYKMMAEVEEKENSEAAKEVRHPGCVRLIFDISTTKDPRR 240
Query: 176 YNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDR 235
YN+P+ +EV + VG+ D R + V + GGL+ I + PL YPVLFP G D
Sbjct: 241 YNLPQANEVAVVFVGEDDDVPTTRSLAVHPRGGGLKTIRDIDRICDPLTYPVLFPNGTDG 300
Query: 236 YEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMV 295
+ ++ S KK+ R++ + + +Y L +R + + G LFQQF VD + +
Sbjct: 301 WHPDLE-----KRPSEKKQGRITQKMYYSYLLMERSGVFNPLHHGRALFQQFAVDSWVKI 355
Query: 296 ENQRLSYVRKNQSQIRS 312
E RL+Y R +Q +++
Sbjct: 356 EQNRLNYHRTHQVDLKA 372
>UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa]
Length = 2052
Score = 281 bits (718), Expect = 1e-73
Identities = 150/315 (47%), Positives = 205/315 (64%), Gaps = 7/315 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LNSDQ ++ I++ V E FFV G+GG GKTFLWNA+ RSE KIVL +ASS +
Sbjct: 932 LNSDQRKVFDTIIDRVSFEKPGFFFVYGHGGTGKTFLWNAIILKIRSEQKIVLAIASSGV 991
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLLPRGRTAHS+F IP+ + E S C+I + A+L+ LIIWDEAPM RL FEA
Sbjct: 992 ASLLLPRGRTAHSRFKIPIDISENSICSIRRGTILAELIQKTLLIIWDEAPMTHRLCFEA 1051
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT+RD++S + +PFGGK +VLGGDFRQILPV+ K RA I DA I +S LWR
Sbjct: 1052 LDRTLRDLLSEHDPANAIVPFGGKVIVLGGDFRQILPVIQKGSRASIDDASITNSPLWRH 1111
Query: 1002 CRVLKLTKNIRL--QFPTNGQDDASLRLFARWILDVGDGKLGI---DNDGEAV-IEIPDD 1055
++L L N+RL T + D L FA+W+L +G+G + + + E +EIP D
Sbjct: 1112 VKLLSLKINMRLLRSGLTQTKKD-ELDNFAKWVLHIGNGDVPATQRERETEPTWVEIPQD 1170
Query: 1056 ICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEK 1115
+ +K G+ + +++ +P+LL N + ++ RAI+ P V+ +N+YV+ L+PGEEK
Sbjct: 1171 LLIKTDGDKIPALIDEVFPDLLHNHTDPTYLSCRAIVCPNNGTVDDINNYVVGLLPGEEK 1230
Query: 1116 EYLSCDTVLKCDEEV 1130
EYLSCDT+ K E +
Sbjct: 1231 EYLSCDTIAKSSEHI 1245
Score = 99.4 bits (246), Expect = 6e-19
Identities = 60/197 (30%), Positives = 96/197 (48%), Gaps = 1/197 (0%)
Query: 114 YHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDS 173
+ E S D+ +V+ L ++DE N LVK FR ++ E + LRL D
Sbjct: 365 FDKESSSDGPPDKQIVEKLGSMLDEHNELVKSFRYAKERLEEHGYQDTALRLMGCNAKDE 424
Query: 174 RVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGE 233
YN+P E+ ++VGD+ D++V+ KD LR++ H Y+ L YP+LFP G+
Sbjct: 425 VQYNLPSNGEIAGIVVGDYSKDAYIYDVLVKSKDNRLRQVSALHPSYMALPYPLLFPYGD 484
Query: 234 DRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYS 293
+ IK + +R V++ E+ YR+ R N+ + RL +VD YS
Sbjct: 485 RGFHLGIKYTDFRSLAPTARRY-VTMLEYYIYRMHYRLNKPNPYTCCGRLSDSLVVDAYS 543
Query: 294 MVENQRLSYVRKNQSQI 310
VE RL ++ +Q +
Sbjct: 544 TVEGSRLQFIADHQPDL 560
Score = 95.1 bits (235), Expect = 1e-17
Identities = 42/64 (65%), Positives = 54/64 (83%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F ++ RR+ G V+KNG++LDN++VVPYN KL+ KY AHIN+E+CNKSN IKYLFKY+TK
Sbjct: 753 FTVYKRRNDGRYVVKNGLKLDNKNVVPYNMKLLKKYQAHINVEWCNKSNMIKYLFKYVTK 812
Query: 574 GVDR 577
G DR
Sbjct: 813 GSDR 816
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 280 bits (715), Expect = 2e-73
Identities = 149/312 (47%), Positives = 208/312 (65%), Gaps = 7/312 (2%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSS 880
+LN++Q Y++I V N+L +FF+ G+GG GKTF+W L+ RS G+IVLNVASS
Sbjct: 1271 MLNTEQRGIYDEITGAVFNDLGGVFFIYGFGGTGKTFIWKTLAAAVRSRGQIVLNVASSG 1330
Query: 881 IASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFE 940
IASLLL GRTAHS+F+IPL E S C I + + A L+ ASLIIWDEAPM+S+ FE
Sbjct: 1331 IASLLLEGGRTAHSRFAIPLNPDEFSVCKITPKSDLANLIKEASLIIWDEAPMMSKFCFE 1390
Query: 941 AFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWR 1000
+ D++ DI++N N FGGK VV GGDFRQ+LPV+ GR +IV + +N+S LW
Sbjct: 1391 SLDKSFYDILNN----KDNKVFGGKVVVFGGDFRQVLPVINGAGRVEIVMSSLNASYLWD 1446
Query: 1001 RCRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLK 1059
C+VLKLTKN+RL ++A ++ F+ W+L VGDG++ NDGEA+I+IP+++ +K
Sbjct: 1447 HCKVLKLTKNMRLLSGGLSSEEAKEIQQFSDWLLAVGDGRINEPNDGEALIDIPEELLIK 1506
Query: 1060 NSGNHVGDIVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEY 1117
+GN + I + Y P+ L + + FFQ RAILAPT E V +N Y+L + EE+ Y
Sbjct: 1507 EAGNPIEAISKEIYGDPSELHMINDPKFFQRRAILAPTNEDVNTINQYMLEHLKSEERIY 1566
Query: 1118 LSCDTVLKCDEE 1129
LS D++ D +
Sbjct: 1567 LSADSIDPTDSD 1578
Score = 236 bits (602), Expect = 3e-60
Identities = 132/340 (38%), Positives = 200/340 (58%), Gaps = 23/340 (6%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR T + K G + DN VVPYN KL ++Y AHIN+E+CN+S SIKYLFKYI K
Sbjct: 873 YPIYRRRMTEDYIEKGGFKCDNGYVVPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINK 932
Query: 574 GVDRVTATLE-----------TSEEP------SVDEIQ*YYDCRYLSPSESIWRIFGFDV 616
G DRV +E TS EP DEI+ ++DCRY+S SE++WRI+ F +
Sbjct: 933 GADRVVFIVEPVNQDKTTENATSGEPPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPL 992
Query: 617 HCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWN--- 673
R AV+RL+FH G+Q + + + ++VL R ++ +MF+AW+ N+N N
Sbjct: 993 QDRSTAVQRLSFHDEGKQPVYAKPDADIEDVLERISNEDSMFMAWLTLNKNNDVGKNGKR 1052
Query: 674 ---LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSF 730
L Y Q P FT++ +N+ W G S+GR+ +V + Y +R+LLN G S+
Sbjct: 1053 ARELLYSQIPAYFTWDGKNKQWVKRIRGFSLGRINYVCRKMEVEYYLRVLLNIVKGPMSY 1112
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+DI+T +G VY S++EAC A G+L +D+ +ID + E G +R F LLL+ ++
Sbjct: 1113 DDIKTFNGVVYPSFKEACFARGILDDDQVYIDGLHEASQFCFGDYLRNFFAMLLLSDSLA 1172
Query: 791 DPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
P +VW++TWH+L+E I ++R +P L ++ Y
Sbjct: 1173 RPEHVWSETWHLLAEDIENKKREDFKNPDLKLTLAEIRNY 1212
Score = 126 bits (317), Expect = 3e-27
Identities = 72/192 (37%), Positives = 116/192 (59%), Gaps = 7/192 (3%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-FDSRVYNVPEIS 182
L + +++ L++++++ N V KFR R+ + + D P +R+ +R D R Y++P S
Sbjct: 474 LKKEVIEALIEMLNKVNPYVDKFRQARERIQDDNDEPFHMRIVADRKGVDRRTYSMPTSS 533
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIK 241
EV ALI G F RDIV+ EK G L RI + H Y+ LQYP++ GED Y I+
Sbjct: 534 EVAALIPGGFQPSMFDRDIVLEEKTTGHLTRISQIHISYLALQYPLILCYGEDGYTPGIE 593
Query: 242 LNHRTTTGSVKKRVR--VSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQR 299
+ S KK+ + +S+R++ A+R+Q+R NE + + RLFQQF+ D Y+ +E+ R
Sbjct: 594 ---KCLPNSAKKKKKKCISMRQWFAFRIQERPNECKTLTRSKRLFQQFLCDAYTTIESNR 650
Query: 300 LSYVRKNQSQIR 311
LSY++ QS++R
Sbjct: 651 LSYIKFKQSKLR 662
>UniRef100_Q9LI91 Similarity to unknown protein [Arabidopsis thaliana]
Length = 619
Score = 276 bits (706), Expect = 3e-72
Identities = 166/447 (37%), Positives = 255/447 (56%), Gaps = 24/447 (5%)
Query: 683 FTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYN 742
F + N+ + K +++GR+ + P +LY +R+LLN+ G TSF+ ++TV G V+
Sbjct: 2 FVLDSTNKMYTRRKQRENIGRIVNILPTAGDLYYLRILLNKVKGATSFDYLKTVGGVVHE 61
Query: 743 SYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHI 802
S++ AC A GLL D+++ D + E + +R +F +L+ +S+PL +W+ W
Sbjct: 62 SFKAACHARGLLDGDKEWHDAMDEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHCWES 121
Query: 803 LSEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNAL 862
+++ +L +++R LN P L + +L Y ++ + H + Y + +
Sbjct: 122 MADDVLRKQQRVLNFPQLELKAKELEKYT-LIEIETLLRQHEKSLSDYPEMPQPE----- 175
Query: 863 SYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTM 922
+S GK V+ VASS+IA+LLLP GRTAHS F IP+ + E+ C I+ A +L+
Sbjct: 176 ----KSNGKNVMPVASSAIAALLLPGGRTAHSWFKIPINVHEDFICDIKIGSMLANVLSK 231
Query: 923 ASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPK 982
LIIWDEAPM R FEA DRT+RDI+S + A GGKTV+LGGDFRQILPV+P+
Sbjct: 232 VDLIIWDEAPMAHRHTFEAVDRTLRDILSVGDEKALTKTLGGKTVLLGGDFRQILPVIPQ 291
Query: 983 RGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKL-- 1040
R R + V A IN S LW C L++N+R+Q P + FA WIL +GDG+
Sbjct: 292 RTRQETVSAAINRSYLWESCHKYLLSQNMRVQ-PEEIK-------FAEWILQIGDGEAPR 343
Query: 1041 ---GIDNDGEA-VIEIPDDICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTL 1096
GID+D E I I ++ L + N + + +S P+ + ++ + A+L P
Sbjct: 344 KTHGIDDDQEEDNIIIDKNLLLPETENPLEVLCQSVSPDFTNTFQDLENLKGTAVLTPRN 403
Query: 1097 ELVEKVNDYVLSLVPGEEKEYLSCDTV 1123
E V+++NDY+LS VPG KEY S D++
Sbjct: 404 ETVDEINDYLLSKVPGLAKEYFSADSI 430
>UniRef100_Q6YSD5 Helicase-like protein [Oryza sativa]
Length = 1516
Score = 275 bits (704), Expect = 4e-72
Identities = 146/316 (46%), Positives = 203/316 (64%), Gaps = 9/316 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LN+DQ++ + I + FFV G+GG GKTFLWN + RS+ KIVL VASS +
Sbjct: 991 LNNDQITVFSTICSRAIANEPGFFFVSGHGGTGKTFLWNTIIAKLRSQNKIVLAVASSGV 1050
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLLPRGRTAHS+F IP+ + E S C I++ A+LL +LIIWDEAPM R FEA
Sbjct: 1051 ASLLLPRGRTAHSRFKIPIDIDETSICNIKRGTMLAELLAETALIIWDEAPMTHRRCFEA 1110
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT+RDI+S S +PFGGK +VLGGDF+QILPV+PK R I++A I +S LW+
Sbjct: 1111 LDRTLRDILSETCPSNSIVPFGGKPIVLGGDFKQILPVIPKGSRQAIINASITNSELWKH 1170
Query: 1002 CRVLKLTKNIRL---QFPTNGQDDASLRLFARWILDVGDGKLGID-NDGE---AVIEIPD 1054
+L L N+RL P N + + L F++W+L +G+G L + +GE A I IPD
Sbjct: 1171 VALLSLNINMRLLNPMLPDNQKKE--LHDFSQWVLAIGNGTLPMTAKEGENYPAWITIPD 1228
Query: 1055 DICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEE 1114
D+ + SG+ + IV Y + L+ ++ + RAI+ PT V+++NDY++ LVPG+
Sbjct: 1229 DLLVMTSGDKIAAIVHEVYSDFLTCYRDIEYLASRAIVCPTNTTVDEINDYIIGLVPGDS 1288
Query: 1115 KEYLSCDTVLKCDEEV 1130
+ YLSCDT+ K E++
Sbjct: 1289 RVYLSCDTISKSSEQI 1304
Score = 246 bits (627), Expect = 4e-63
Identities = 134/346 (38%), Positives = 207/346 (59%), Gaps = 16/346 (4%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F ++ RR+ G +++KNGI LDNRSVVPYN L+ KY AHIN+E+CNKSN IKYLFKYITK
Sbjct: 606 FTIYRRRNDGRSIMKNGILLDNRSVVPYNMALLKKYEAHINVEWCNKSNLIKYLFKYITK 665
Query: 574 GVDRVTATLETSEE-----PSVD-----EIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAV 623
G DR ET+ + P+ D EI Y D R+LS E++ R+F FD+H R P V
Sbjct: 666 GHDRARIYFETTGKTQNASPNHDLAPRNEILEYMDARFLSTYEALHRLFEFDIHYRVPPV 725
Query: 624 KRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKF 683
+RL HL G+ + + + VL KR+M W + N+ S++ +LTY +FP+++
Sbjct: 726 ERLVVHLPGKNFVRYEKGADLRAVLESPGAKRSMLTEWFETNKKNSKAHSLTYCEFPKEW 785
Query: 684 TYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNS 743
T+ +++WH +GR+ +V P ELY +R+LL G S+ D+RT DG VY +
Sbjct: 786 TWEPSSKTWHERTPAPKIGRIYYVHPTAGELYYLRMLLMIVKGAQSYADVRTYDGVVYGT 845
Query: 744 YREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHIL 803
YREAC A GLL D ++ L E +V S + +R++F ++LL + D +++++ W +
Sbjct: 846 YREACEARGLLEGDNEWHLLFDEAIVTASSAQLRQLFVTVLLYCSVGDVRSLFDKYWLYM 905
Query: 804 SEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDG 849
++ I ++ L++P ++ D L +N++ +EL +F G
Sbjct: 906 TDDIHNRLKKALDNPHCVIPHDHL------LNMLLHELIAVFANSG 945
Score = 103 bits (257), Expect = 3e-20
Identities = 67/197 (34%), Positives = 105/197 (53%), Gaps = 7/197 (3%)
Query: 119 SDGNGLDRS---LVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRV 175
+DG+ D++ +++ L ++D N LV+ FR R+ + + V LRL D
Sbjct: 203 NDGDNSDKADPEIIRALSSMLDAENTLVQSFRYARERVIQHGNQQVTLRLLGCNAKDDVQ 262
Query: 176 YNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDR 235
YN+P SE+ A+IVGDF + E D++V +K GL +I H Y+ LQYP+LFP GE
Sbjct: 263 YNLPTNSEIAAIIVGDFSAKEYKFDVLVYDKGRGLCQISPLHPSYMALQYPLLFPYGERG 322
Query: 236 YEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMV 295
+ IK ++ G + V++ E+ Y + R NE + RL Q VD++S +
Sbjct: 323 FHLGIKYSNYDGIG----KKYVTMPEYYRYEMHYRLNEPNPFTCYGRLSDQIDVDIFSTI 378
Query: 296 ENQRLSYVRKNQSQIRS 312
E RL Y +Q ++RS
Sbjct: 379 ETNRLQYFIDHQKELRS 395
>UniRef100_Q6YTQ6 Helicase-like protein [Oryza sativa]
Length = 1430
Score = 274 bits (701), Expect = 1e-71
Identities = 145/316 (45%), Positives = 202/316 (63%), Gaps = 9/316 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LN+DQ++ + I + FFV G+GG GKTFLWN + RS+ KIVL VASS +
Sbjct: 905 LNNDQITVFNTICSRAIANEPGFFFVSGHGGTGKTFLWNTIIAKLRSQNKIVLAVASSGV 964
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLLPRGRT HS+F IP+ + E S C I++ A+LL +LIIWDEAPM R FEA
Sbjct: 965 ASLLLPRGRTTHSRFKIPIDIDETSICNIKRGTMLAELLAETALIIWDEAPMTHRRCFEA 1024
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT+RDI+S S +PFGGK +VLGGDF+QILPV+PK R I++A I +S LW+
Sbjct: 1025 LDRTLRDILSETCPSNSIIPFGGKPIVLGGDFKQILPVIPKGSRQAIINASITNSELWKH 1084
Query: 1002 CRVLKLTKNIRL---QFPTNGQDDASLRLFARWILDVGDGKLGID-NDGE---AVIEIPD 1054
+L L N+RL P N + + L F++W+L +G+G L + +GE A I IPD
Sbjct: 1085 VALLSLNINMRLLNPMLPDNQKKE--LHDFSQWVLAIGNGTLPMTAKEGENYPAWITIPD 1142
Query: 1055 DICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEE 1114
D+ + SG+ + IV Y + L+ ++ + RAI+ PT V+++NDY++ LVPG+
Sbjct: 1143 DLLVMTSGDKIAAIVHEVYSDFLTCYRDIEYLASRAIVCPTNTTVDEINDYIIGLVPGDS 1202
Query: 1115 KEYLSCDTVLKCDEEV 1130
+ YLSCDT+ K E++
Sbjct: 1203 RVYLSCDTISKSSEQI 1218
Score = 197 bits (502), Expect = 1e-48
Identities = 121/346 (34%), Positives = 185/346 (52%), Gaps = 58/346 (16%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F ++ RR+ G +++KNGI LDNRSVVPYN L+ KY AHIN+E+CNKSN IKYLFKYITK
Sbjct: 562 FTIYRRRNDGRSIMKNGILLDNRSVVPYNMALLKKYEAHINVEWCNKSNLIKYLFKYITK 621
Query: 574 GVDRVTATLETSEE-----PSVD-----EIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAV 623
G DR ET+ + P+ D EI Y D R+LS
Sbjct: 622 GHDRARIYFETTGKTQNASPNHDLAPRNEILEYMDARFLS-------------------- 661
Query: 624 KRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKF 683
TF L KR+M W + N+ S++ +LTY +FP+++
Sbjct: 662 ---TF-------------------LESPGAKRSMLTEWFETNKKNSKAHSLTYCEFPKEW 699
Query: 684 TYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNS 743
T+ +++WH +GR+ +V P ELY +R+LL G S+ D+RT DG VY +
Sbjct: 700 TWEPSSKTWHERTPAPKIGRIYYVHPTAGELYYLRMLLMIVKGAQSYADVRTYDGVVYGT 759
Query: 744 YREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHIL 803
YREAC A GLL D ++ L E +V S + +R++F ++LL + D +++++ W +
Sbjct: 760 YREACEARGLLEGDNEWHLLFDEAIVTASSAQLRQLFVTVLLYCSVGDVRSLFDKYWLYM 819
Query: 804 SEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDG 849
++ I ++ L++P ++ D L +N++ +EL +F G
Sbjct: 820 TDDIHNRLKKALDNPHCVIPHDHL------LNMLLHELIAVFANSG 859
Score = 107 bits (268), Expect = 2e-21
Identities = 68/197 (34%), Positives = 107/197 (53%), Gaps = 7/197 (3%)
Query: 119 SDGNGLDRS---LVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRV 175
+DG+ D++ +++ L ++D N LV+ FR R+ + + V LRL D
Sbjct: 226 NDGDNSDKADPEIIRALSSMLDAENTLVQSFRYARERVIQHGNQQVTLRLLGCNAKDDVQ 285
Query: 176 YNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDR 235
YN+P SE+ A+IVGDF + E D++V +K GLR+I H Y+ LQYP+LFP GE
Sbjct: 286 YNLPTNSEIAAIIVGDFSAKEYKFDVLVYDKGRGLRQISPLHPSYMALQYPLLFPYGERG 345
Query: 236 YEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMV 295
+ IK ++ G + V++ E+ Y + R NE++ RL Q VD++S +
Sbjct: 346 FHLGIKYSNYDGIG----KKYVTMPEYYRYEMHYRLNESNPFTCYGRLSDQIDVDIFSTI 401
Query: 296 ENQRLSYVRKNQSQIRS 312
E RL Y +Q ++RS
Sbjct: 402 ETNRLQYFIDHQKELRS 418
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 271 bits (694), Expect = 6e-71
Identities = 147/311 (47%), Positives = 202/311 (64%), Gaps = 7/311 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
L +Q Y++I N V N+L +FFV G+GG GKTF+W L+ RS+G+I LNVASS I
Sbjct: 949 LTPEQRGIYDQITNAVFNDLGGVFFVYGFGGTGKTFIWKTLAAAVRSKGQICLNVASSGI 1008
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLL GRTAHS+FSIPL E S C I+ + + A L+ ASLIIWDEAPM+S+ FEA
Sbjct: 1009 ASLLLEGGRTAHSRFSIPLNPDEFSVCKIKPKSDLADLIKEASLIIWDEAPMMSKFCFEA 1068
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
D++ DI+ V N FGGK +V GGDFRQ+LPV+ GRA+IV + +N+S LW
Sbjct: 1069 LDKSFSDIIKRV----DNKVFGGKVMVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWDH 1124
Query: 1002 CRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
C+VL+LTKN+RL D+A ++ F+ W+L VGDG++ NDGE +I+IP+++ ++
Sbjct: 1125 CKVLRLTKNMRLLNNDLSVDEAKEIQEFSDWLLAVGDGRVNEPNDGEVIIDIPEELLIQE 1184
Query: 1061 SGNHVGDIVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYL 1118
+ N + I Y P L + + FFQ RAILAP E V +N Y+L + EE+ YL
Sbjct: 1185 ADNPIEAISREIYGDPTKLHEISDPKFFQRRAILAPKNEDVNTINQYMLEHLDSEERIYL 1244
Query: 1119 SCDTVLKCDEE 1129
S D++ D +
Sbjct: 1245 SADSIDPSDSD 1255
Score = 226 bits (576), Expect = 3e-57
Identities = 129/349 (36%), Positives = 201/349 (56%), Gaps = 32/349 (9%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR T + K G++ DNR VVPYN KL ++Y AHIN+E+CN++ SIKYLFKYI K
Sbjct: 541 YPIYRRRLTDDYIEKGGVKCDNRYVVPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYINK 600
Query: 574 GVDRVTATLETSEEPSV-----------------DEIQ*YYDC---------RYLSPSES 607
G DRV +E +E + DEI+ ++DC RY+S SE+
Sbjct: 601 GPDRVVFIVEPIKEATSSDTTAPVVESDTTEKKKDEIKDWFDCSSYISFSPARYVSASEA 660
Query: 608 IWRIFGFDVHCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANR- 666
IWRIF F + R V++L+FH G+Q F ++ +VL R ++ + FLAW+ NR
Sbjct: 661 IWRIFKFPIQHRSTPVQKLSFHDKGKQPAYFDAKAKMADVLERVSNEDSQFLAWLTLNRK 720
Query: 667 -----NYSQSWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLL 721
N ++ + Y + P FT++ EN+ + G S+GR+ +V ++ Y +R+LL
Sbjct: 721 NAVGKNGKRARDCLYAEIPAYFTWDGENKQFKKRTRGFSLGRINYVSRKMEDEYYLRVLL 780
Query: 722 NRQIGCTSFEDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFT 781
N G S++DI+TV+G VY SY+ AC A G+L +D+ +I+ + E G +R F+
Sbjct: 781 NIVRGPQSYDDIKTVNGVVYPSYKLACFARGILDDDQVYINGLIEASQFCFGDYLRNFFS 840
Query: 782 SLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
+LL+ ++ P +VW++TWH+LSE IL ++R + L Q+ Y
Sbjct: 841 MMLLSDSLARPEHVWSETWHLLSEDILIKKRDEFKNQELTLTEAQIQNY 889
Score = 144 bits (364), Expect = 1e-32
Identities = 76/201 (37%), Positives = 127/201 (62%), Gaps = 4/201 (1%)
Query: 113 NYHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-F 171
N S L++ L+ ++K++++ N V+KFR R+ ++ D P +R+ +R
Sbjct: 132 NNGSSTKGKKNLNKQLIDAIIKMLNQVNPYVEKFRSARERIDSTNDEPFHMRIVSDRKGT 191
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFP 230
D R+YN+P EV ALI GDF S RDI++ +K G L+RI + H Y+ LQYP++F
Sbjct: 192 DGRLYNMPTAGEVAALIPGDFVSQMPVRDIILEKKSTGRLKRISQIHISYLALQYPLIFC 251
Query: 231 LGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVD 290
GED Y I+ +++ G KK+ +S+R++ A+R+Q+R++E+ + Q RLFQQF+ D
Sbjct: 252 YGEDGYTPGIEKCYKS--GYTKKKKCISMRQWYAFRIQEREDESHTLLQSKRLFQQFLCD 309
Query: 291 LYSMVENQRLSYVRKNQSQIR 311
Y+ +E+ RL+Y++ NQS++R
Sbjct: 310 AYTTIESNRLAYIKFNQSKLR 330
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,807,975,267
Number of Sequences: 2790947
Number of extensions: 74908906
Number of successful extensions: 267479
Number of sequences better than 10.0: 237
Number of HSP's better than 10.0 without gapping: 133
Number of HSP's successfully gapped in prelim test: 105
Number of HSP's that attempted gapping in prelim test: 264144
Number of HSP's gapped (non-prelim): 2136
length of query: 1134
length of database: 848,049,833
effective HSP length: 138
effective length of query: 996
effective length of database: 462,899,147
effective search space: 461047550412
effective search space used: 461047550412
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 80 (35.4 bits)
Medicago: description of AC135463.4


