

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.1 - phase: 0 /pseudo
(488 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
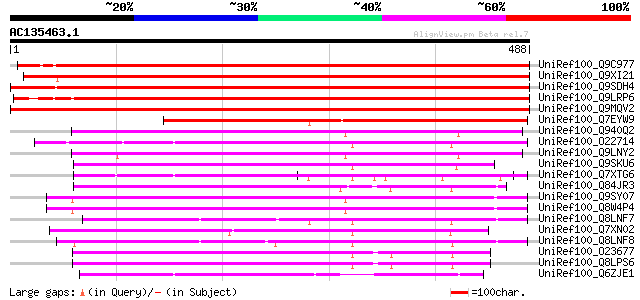
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C977 Hypothetical protein F5I6.2 [Arabidopsis thaliana] 561 e-158
UniRef100_Q9XI21 F9L1.43 [Arabidopsis thaliana] 550 e-155
UniRef100_Q9SDH4 EST C93513(C53164) corresponds to a region of t... 504 e-141
UniRef100_Q9LRP6 DNA-binding protein [Arabidopsis thaliana] 492 e-138
UniRef100_Q9MQV2 DNA-binding protein [Triticum aestivum] 489 e-137
UniRef100_Q7EYW9 Putative DNA-binding protein [Oryza sativa] 327 6e-88
UniRef100_Q940Q2 At1g07590/F22G5_2 [Arabidopsis thaliana] 179 1e-43
UniRef100_O22714 F8A5.28 protein [Arabidopsis thaliana] 170 7e-41
UniRef100_Q9LNY2 F22G5.3 [Arabidopsis thaliana] 167 6e-40
UniRef100_Q9SKU6 Hypothetical protein At2g20710 [Arabidopsis tha... 166 1e-39
UniRef100_Q7XTG6 OSJNBb0026L04.10 protein [Oryza sativa] 162 2e-38
UniRef100_Q84JR3 Hypothetical protein At4g21705 [Arabidopsis tha... 161 3e-38
UniRef100_Q9SY07 Hypothetical protein T5J8.14 [Arabidopsis thali... 158 3e-37
UniRef100_Q8W4P4 Hypothetical protein At4g02820; T5J8.14 [Arabid... 157 7e-37
UniRef100_Q8LNF7 Putative leaf protein [Oryza sativa] 154 4e-36
UniRef100_Q7XN02 OSJNBb0038F03.9 protein [Oryza sativa] 153 1e-35
UniRef100_Q8LNF8 Putative leaf protein [Oryza sativa] 148 3e-34
UniRef100_O23677 T7I23.8 protein [Arabidopsis thaliana] 139 1e-31
UniRef100_Q8LPS6 At1g02150/T7I23.8 [Arabidopsis thaliana] 139 2e-31
UniRef100_Q6ZJE1 Putative DNA-binding protein [Oryza sativa] 134 5e-30
>UniRef100_Q9C977 Hypothetical protein F5I6.2 [Arabidopsis thaliana]
Length = 596
Score = 561 bits (1445), Expect = e-158
Identities = 280/482 (58%), Positives = 367/482 (76%), Gaps = 5/482 (1%)
Query: 8 TEPDADGDSRDTAEIDINELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVDSALD 67
T D D E + EL+L ETD S K +++SELFK IVS GLS+ SALD
Sbjct: 94 TSSDEDEGKLSADEEEEEELDL--IETDVSRKTV--EKKQSELFKTIVSAPGLSIGSALD 149
Query: 68 KWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKL 127
KWVE+G E++R EI A+ LRRR+MYGRALQ +WLE+NKK+E TE++YAS+LDL K+
Sbjct: 150 KWVEEGNEITRVEIAKAMLQLRRRRMYGRALQMSEWLEANKKIEMTERDYASRLDLTVKI 209
Query: 128 RGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQ 187
RGL K E ++ +P SF+GE+LYRTLLANC + N++K+E FNKM++LGFP++ F C+Q
Sbjct: 210 RGLEKGEACMQKIPKSFKGEVLYRTLLANCVAAGNVKKSELVFNKMKDLGFPLSGFTCDQ 269
Query: 188 LLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGC 247
+LL++K+ID+KKIADVLL+MEKEN+KPS TYKILIDVKG +NDI GM QI+ETMK EG
Sbjct: 270 MLLLHKRIDRKKIADVLLLMEKENIKPSLLTYKILIDVKGATNDISGMEQILETMKDEGV 329
Query: 248 ELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVER 307
ELD T+A ARHY+ AGL +K E +LKE+EGE+L+ N LL +YA LGR DEV+R
Sbjct: 330 ELDFQTQALTARHYSGAGLKDKAEKVLKEMEGESLEANRRAFKDLLSIYASLGREDEVKR 389
Query: 308 IWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFE-MMSNKWKLTARNYESLLKIYIRHK 366
IWK+CESKP E+ LAAI+A+G+L K++EAEA+FE ++ + ++ Y LL++Y+ HK
Sbjct: 390 IWKICESKPYFEESLAAIQAFGKLNKVQEAEAIFEKIVKMDRRASSSTYSVLLRVYVDHK 449
Query: 367 MLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTF 426
ML+KGKDL+K M +SGC I TTWDAL+ LYV+AGEVEKAD++L KA +Q+ K M +F
Sbjct: 450 MLSKGKDLVKRMAESGCRIEATTWDALIKLYVEAGEVEKADSLLDKASKQSHTKLMMNSF 509
Query: 427 MTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADN 486
M IM++Y+KRGDVHN EKIF ++R+A Y SR+ F AL QAY NAK PAYG+R+R+KADN
Sbjct: 510 MYIMDEYSKRGDVHNTEKIFLKMREAGYTSRLRQFQALMQAYINAKSPAYGMRDRLKADN 569
Query: 487 LF 488
+F
Sbjct: 570 IF 571
>UniRef100_Q9XI21 F9L1.43 [Arabidopsis thaliana]
Length = 623
Score = 550 bits (1417), Expect = e-155
Identities = 269/493 (54%), Positives = 364/493 (73%), Gaps = 18/493 (3%)
Query: 14 GDSRDTAEIDINELELSDTETDSSDKKSFS-----------------SRRRSELFKAIVS 56
GD D + +++ +T +DS D + FS S+R SE+FKAIVS
Sbjct: 106 GDDDDLEDKNVDLATPDETSSDSEDGEEFSGDEGDIEGAELELHVPESKRPSEMFKAIVS 165
Query: 57 VSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKE 116
VSGLSV SALDKWVE+GK+ +R+E A+ LR+R+M+GRALQ +WL+ NK+ E E++
Sbjct: 166 VSGLSVGSALDKWVEQGKDTNRKEFESAMLQLRKRRMFGRALQMTEWLDENKQFEMEERD 225
Query: 117 YASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMREL 176
YA +LDLI+K+RG K E Y++ +P SFRGEL+YRTLLAN + N+R E FNKM++L
Sbjct: 226 YACRLDLISKVRGWYKGEAYIKTIPESFRGELVYRTLLANHVATSNVRTAEAVFNKMKDL 285
Query: 177 GFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMS 236
GFP++ F CNQ+L++YK++DKKKIADVLL++EKEN+KP+ TYKILID KG SNDI GM
Sbjct: 286 GFPLSTFTCNQMLILYKRVDKKKIADVLLLLEKENLKPNLNTYKILIDTKGSSNDITGME 345
Query: 237 QIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLY 296
QIVETMK+EG ELD RA +ARHYA+AGL EK E +LKE+EGE+L+EN +C LL +Y
Sbjct: 346 QIVETMKSEGVELDLRARALIARHYASAGLKEKAEKVLKEMEGESLEENRHMCKDLLSVY 405
Query: 297 AILGRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFE-MMSNKWKLTARNY 355
L R DEV R+WK+CE PR + LAAI A+G++ K+++AEAVFE ++ ++++ Y
Sbjct: 406 GYLQREDEVRRVWKICEENPRYNEVLAAILAFGKIDKVKDAEAVFEKVLKMSHRVSSNVY 465
Query: 356 ESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQ 415
LL++Y+ HKM+++GKDL+K M DSGC IG TWDA++ LYV+AGEVEKA++ L KA+Q
Sbjct: 466 SVLLRVYVDHKMVSEGKDLVKQMSDSGCNIGALTWDAVIKLYVEAGEVEKAESSLSKAIQ 525
Query: 416 QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPA 475
++KP+ ++FM +M +Y +RGDVHN EKIF R++QA Y SR + L QAY NAK PA
Sbjct: 526 SKQIKPLMSSFMYLMHEYVRRGDVHNTEKIFQRMKQAGYQSRFWAYQTLIQAYVNAKAPA 585
Query: 476 YGIRERMKADNLF 488
YG++ERMKADN+F
Sbjct: 586 YGMKERMKADNIF 598
>UniRef100_Q9SDH4 EST C93513(C53164) corresponds to a region of the predicted gene
[Oryza sativa]
Length = 591
Score = 504 bits (1299), Expect = e-141
Identities = 241/490 (49%), Positives = 354/490 (72%), Gaps = 3/490 (0%)
Query: 1 MEIEDRLTEPDADGDSRDTAEID-INELELSDTETDSSDKKSFSSRRRSELFKAIVSVSG 59
+E+ + D D S ++++ D I+ + LS E D+ ++ + +S L KA++
Sbjct: 77 LEVPPEADKKDLDLTSDESSDEDTIDAIGLSQVEADAKPEEPIK-KSQSTLLKALLVSPR 135
Query: 60 LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYAS 119
+ V A KW+ G L R E+ L SLR+RK+Y +ALQ L+++E +K + E++YAS
Sbjct: 136 VDVAGATKKWLNDGNTLERSELFYVLLSLRKRKLYTKALQLLEYVEESKLFDLGERDYAS 195
Query: 120 KLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFP 179
++DL+AK+ G+ KAEKY+E+VP S RGE++YRTLLANC ++ N++KTE+ FNKM++LGFP
Sbjct: 196 RVDLVAKVHGIYKAEKYIENVPASHRGEVVYRTLLANCVAIANVKKTEQVFNKMKDLGFP 255
Query: 180 VTAFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
VT F+CNQLLL+YK++DKKK+ DVL MMEKENVKPS +TYK+L+D KG + DI+ M +++
Sbjct: 256 VTVFSCNQLLLLYKRVDKKKLGDVLTMMEKENVKPSLFTYKLLVDTKGAARDIEDMEKVI 315
Query: 240 ETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAIL 299
+ M+A+G E D L +A++ARHY G EK EAIL++IEG+++ EN C +L LYA L
Sbjct: 316 QAMQADGIEPDLLIQATIARHYIFGGYREKAEAILEQIEGDDINENRSACKFVLPLYAFL 375
Query: 300 GRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWK-LTARNYESL 358
G+ +VERIWKVCE R+++C++AIEA+G+L +E+AE +FE M WK L+ Y ++
Sbjct: 376 GKKADVERIWKVCEVNARLDECMSAIEAFGKLGDVEKAEEIFENMFKTWKTLSFEYYNAM 435
Query: 359 LKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNK 418
LK+Y K+ +KGK+L K MGD GC +GP+T D+LV LY AGEVEKAD++L K +NK
Sbjct: 436 LKVYANKKLFDKGKELAKRMGDDGCRLGPSTLDSLVKLYSDAGEVEKADSILHKLSYKNK 495
Query: 419 MKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGI 478
+KP++TT++ +++ Y+K+GDVHNAEK+F ++RQ Y RI + L +AY NAK P YG
Sbjct: 496 IKPLYTTYLMLLDSYSKKGDVHNAEKLFSKVRQMGYTGRIRQYQLLLEAYLNAKTPPYGF 555
Query: 479 RERMKADNLF 488
+ERMKAD++F
Sbjct: 556 KERMKADDIF 565
>UniRef100_Q9LRP6 DNA-binding protein [Arabidopsis thaliana]
Length = 610
Score = 492 bits (1267), Expect = e-138
Identities = 243/486 (50%), Positives = 351/486 (72%), Gaps = 12/486 (2%)
Query: 4 EDRLTEPDADGDSRDTAEIDINELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVD 63
+D L EP+ D+ D LE+ + + K + R +SEL+++IV+ SV
Sbjct: 110 DDSLFEPELGSDNDD--------LEIEEKHSKDGGKPT-KKRGQSELYESIVAYK--SVK 158
Query: 64 SALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDL 123
L+KWV++GK+LS+ E+ LA+++LR+RK Y LQ +WL +N + EFTE YAS+LDL
Sbjct: 159 HVLEKWVKEGKDLSQAEVTLAIHNLRKRKSYAMCLQLWEWLGANTQFEFTEANYASQLDL 218
Query: 124 IAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAF 183
+AK+ L KAE +L+ +P S RGE++YRTLLANC ++ K E+ FNKM+EL FP + F
Sbjct: 219 VAKVHSLQKAEIFLKDIPESSRGEVVYRTLLANCVLKHHVNKAEDIFNKMKELKFPTSVF 278
Query: 184 ACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMK 243
ACNQLLL+Y D+KKI+DVLL+ME+EN+KPS TY LI+ KGL+ DI GM +IVET+K
Sbjct: 279 ACNQLLLLYSMHDRKKISDVLLLMERENIKPSRATYHFLINSKGLAGDITGMEKIVETIK 338
Query: 244 AEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRAD 303
EG ELD ++ LA++Y AGL E+ + ++KEIEG+ L++ WVC +LL LYA +G +D
Sbjct: 339 EEGIELDPELQSILAKYYIRAGLKERAQDLMKEIEGKGLQQTPWVCRSLLPLYADIGDSD 398
Query: 304 EVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARNYESLLKIY 362
V R+ + + PR ++C++AI+AWG+LK++EEAEAVFE + K+K+ Y +L++IY
Sbjct: 399 NVRRLSRFVDQNPRYDNCISAIKAWGKLKEVEEAEAVFERLVEKYKIFPMMPYFALMEIY 458
Query: 363 IRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPM 422
+KML KG+DL+K MG++G IGP+TW ALV LY++AGEV KA+ +L +A + NKM+PM
Sbjct: 459 TENKMLAKGRDLVKRMGNAGIAIGPSTWHALVKLYIKAGEVGKAELILNRATKDNKMRPM 518
Query: 423 FTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERM 482
FTT+M I+E+YAKRGDVHN EK+F ++++A+Y +++ + + AY NAK PAYG+ ERM
Sbjct: 519 FTTYMAILEEYAKRGDVHNTEKVFMKMKRASYAAQLMQYETVLLAYINAKTPAYGMIERM 578
Query: 483 KADNLF 488
KADN+F
Sbjct: 579 KADNVF 584
>UniRef100_Q9MQV2 DNA-binding protein [Triticum aestivum]
Length = 612
Score = 489 bits (1258), Expect = e-137
Identities = 238/491 (48%), Positives = 347/491 (70%), Gaps = 3/491 (0%)
Query: 1 MEIEDRLTEPDADGDSRDTAEID-INELELSDTETDSS-DKKSFSSRRRSELFKAIVSVS 58
+E+ + D + S ++++ D ++ + LS + D+ DK+ +S L K ++
Sbjct: 97 LEVPPEADKKDVELPSEESSDDDAVDGIGLSGDDADAKPDKEPMKKASQSPLLKVMLEAP 156
Query: 59 GLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYA 118
V L KW++ G R +I + +LR+RK Y +ALQ L+WLE +K ++ E++YA
Sbjct: 157 RNGVSGTLKKWLDGGNTFDRSDIFYVIMNLRKRKFYFKALQLLEWLEDSKVIDLGERDYA 216
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
S+LDL+AK+ G+ KAEKY++ +P S RGE++YRTLLANC S N++K+EE FNKM++LGF
Sbjct: 217 SRLDLVAKVHGVYKAEKYIDSIPISHRGEIVYRTLLANCVSEANVKKSEEVFNKMKDLGF 276
Query: 179 PVTAFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQI 238
PVT FA NQLLL+YK++DKKKIADVL MEKENVKPS +TYK+L+D KG DI GM ++
Sbjct: 277 PVTVFAINQLLLLYKRVDKKKIADVLAKMEKENVKPSLFTYKLLVDTKGAIRDIAGMEKV 336
Query: 239 VETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAI 298
VE+M+AEG E D L +A++A+HY AG EK EAIL+ +EG ++K N C LL LYA
Sbjct: 337 VESMQAEGVEPDLLFQATIAKHYIFAGHREKAEAILESMEGGDIKGNRNACKILLPLYAF 396
Query: 299 LGRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWK-LTARNYES 357
LG+ D+VER+W+VCE+ PR+++CL+AIE++GRL +E AE VFE M WK L+++ Y +
Sbjct: 397 LGKKDDVERMWQVCEANPRLDECLSAIESFGRLGDVERAEKVFEDMFATWKTLSSKFYNA 456
Query: 358 LLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQN 417
L+K+Y + KGK+L K M + GC +G +T D+LV LYV AGEV+KA+++L K + N
Sbjct: 457 LMKVYADQNLFKKGKELAKRMDEDGCRLGISTIDSLVKLYVGAGEVDKAESILHKLSKSN 516
Query: 418 KMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYG 477
KMKP +++++ +++ Y+K+GD+HN+EK+F +LRQ Y R+ + L AY +AK P YG
Sbjct: 517 KMKPQYSSYLMLLDTYSKKGDIHNSEKVFDQLRQMGYNGRVRQYQLLLNAYVHAKTPVYG 576
Query: 478 IRERMKADNLF 488
RERMKADN+F
Sbjct: 577 FRERMKADNIF 587
>UniRef100_Q7EYW9 Putative DNA-binding protein [Oryza sativa]
Length = 411
Score = 327 bits (837), Expect = 6e-88
Identities = 163/347 (46%), Positives = 240/347 (68%), Gaps = 5/347 (1%)
Query: 145 RGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVL 204
R E+LY TL+ NC +++K EE F ++++L +T CNQ++L+YK+I K+A VL
Sbjct: 44 RNEVLYETLIVNCVLAGDIQKAEEVFKEIKDLCLRLTVTLCNQMILLYKRIAPGKVASVL 103
Query: 205 LMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAA 264
++MEKENVKPS++TY++LID+KG SND+ G+ ++ MKA G E T+ +AR Y
Sbjct: 104 MLMEKENVKPSAFTYRLLIDLKGRSNDLAGIEVVLNEMKAYGIEPSTSTQTMVARFYIHG 163
Query: 265 GLTEKTEAILKEIEGE--NLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCL 322
GLTEK EA++KE+E + N K+ V +LL LYA L + ++V RIW++C ++P +ED L
Sbjct: 164 GLTEKAEAVVKEMEAQLSNSKDGRHVIKSLLHLYAALNKPNDVARIWEMC-TEPMLEDFL 222
Query: 323 AAIEAWGRLKKIEEAEAVFEMMSN-KWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDS 381
+AI+AWG L IE+AE FE M+N KL+++ Y ++L +Y ++K+L+KGK ++ M
Sbjct: 223 SAIKAWGELGLIEKAEETFEAMANAPEKLSSKYYNAMLNVYAQNKLLSKGKQFVERMCRD 282
Query: 382 GCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQN-KMKPMFTTFMTIMEQYAKRGDVH 440
GC GP TWDAL++LYV +GEVEKAD+ L ++N KP+FT++ +M+ YAKRGD+H
Sbjct: 283 GCPNGPLTWDALINLYVNSGEVEKADSFLLNVAEENPDRKPLFTSYFFLMKGYAKRGDIH 342
Query: 441 NAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
N EKIF RL+ Y R + L +AY NAK+PA+G ERM+ DN+
Sbjct: 343 NTEKIFDRLKNVGYAPRPLHYAVLLEAYVNAKVPAHGFLERMRGDNV 389
>UniRef100_Q940Q2 At1g07590/F22G5_2 [Arabidopsis thaliana]
Length = 534
Score = 179 bits (455), Expect = 1e-43
Identities = 126/434 (29%), Positives = 212/434 (48%), Gaps = 10/434 (2%)
Query: 59 GLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYA 118
G++V SAL W+ G + ++ A+N LR+ RAL+ ++W+ + E EY+
Sbjct: 78 GVTVGSALQSWMGDGFPVHGGDVYHAINRLRKLGRNKRALELMEWIIRERPYRLGELEYS 137
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
L+ KL G+ + EK VP F+ ELLY L+ C +R E KMRELG+
Sbjct: 138 YLLEFTVKLHGVSQGEKLFTRVPQEFQNELLYNNLVIACLDQGVIRLALEYMKKMRELGY 197
Query: 179 PVTAFACNQLLLIYKKIDKKK-IADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQ 237
+ N+L++ ++K IA L +M+ + P TY IL+ ++ ++IDG+ +
Sbjct: 198 RTSHLVYNRLIIRNSAPGRRKLIAKDLALMKADKATPHVSTYHILMKLEANEHNIDGVLK 257
Query: 238 IVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYA 297
+ MK G E + ++ LA +A A L EA +EIE +N L+ LY
Sbjct: 258 AFDGMKKAGVEPNEVSYCILAMAHAVARLYTVAEAYTEEIEKSITGDNWSTLDILMILYG 317
Query: 298 ILGRADEVERIWKVCES--KPRVEDCLAAIEAWGRLKKIEEAEAVF-EMMSNKWKLTARN 354
LG+ E+ R W V R + L A EA+ R+ ++ AE ++ EM + K
Sbjct: 318 RLGKEKELARTWNVIRGFHHVRSKSYLLATEAFARVGNLDRAEELWLEMKNVKGLKETEQ 377
Query: 355 YESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL 414
+ SLL +Y + ++ K + + M +G T+ L +A +++A ++ L
Sbjct: 378 FNSLLSVYCKDGLIEKAIGVFREMTGNGFKPNSITYRHLALGCAKAKLMKEALKNIEMGL 437
Query: 415 QQNKMK------PMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAY 468
K P T ++I+E +A++GDV N+EK+F ++ A Y ++AL +AY
Sbjct: 438 NLKTSKSIGSSTPWLETTLSIIECFAEKGDVENSEKLFEEVKNAKYNRYAFVYNALFKAY 497
Query: 469 KNAKLPAYGIRERM 482
AK+ + +RM
Sbjct: 498 VKAKVYDPNLFKRM 511
>UniRef100_O22714 F8A5.28 protein [Arabidopsis thaliana]
Length = 491
Score = 170 bits (431), Expect = 7e-41
Identities = 131/476 (27%), Positives = 227/476 (47%), Gaps = 16/476 (3%)
Query: 24 INELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVDSALDKWVEKGKELSRQEIGL 83
+ L S T S KK + LFK + + V L+++++ K + + E+G
Sbjct: 3 MRHLSRSRDVTKRSTKKYIEEPLYNRLFKD--GGTEVKVRQQLNQFLKGTKHVFKWEVGD 60
Query: 84 ALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNS 143
+ LR R +Y AL+ + +E + + T + A LDL+AK R + E Y +P +
Sbjct: 61 TIKKLRNRGLYYPALKLSEVMEE-RGMNKTVSDQAIHLDLVAKAREITAGENYFVDLPET 119
Query: 144 FRGELLYRTLLANCASLENL-RKTEETFNKMRELGFPVTAFACNQLLLIYKKI-DKKKIA 201
+ EL Y +LL NC E L K E NKM+EL ++ + N L+ +Y K + +K+
Sbjct: 120 SKTELTYGSLL-NCYCKELLTEKAEGLLNKMKELNITPSSMSYNSLMTLYTKTGETEKVP 178
Query: 202 DVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEG-CELDHLTRASLARH 260
++ ++ ENV P SYTY + + +NDI G+ +++E M +G D T +++A
Sbjct: 179 AMIQELKAENVMPDSYTYNVWMRALAATNDISGVERVIEEMNRDGRVAPDWTTYSNMASI 238
Query: 261 YAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCE-SKPRVE 319
Y AGL++K E L+E+E +N + + L+ LY LG+ EV RIW+ + P+
Sbjct: 239 YVDAGLSQKAEKALQELEMKNTQRDFTAYQFLITLYGRLGKLTEVYRIWRSLRLAIPKTS 298
Query: 320 DC--LAAIEAWGRLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIK 376
+ L I+ +L + AE +F E +N R L+ Y + ++ K +L +
Sbjct: 299 NVAYLNMIQVLVKLNDLPGAETLFKEWQANCSTYDIRIVNVLIGAYAQEGLIQKANELKE 358
Query: 377 TMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL-----QQNKMKPMFTTFMTIME 431
G + TW+ + YV++G++ +A + KA+ K P T +M
Sbjct: 359 KAPRRGGKLNAKTWEIFMDYYVKSGDMARALECMSKAVSIGKGDGGKWLPSPETVRALMS 418
Query: 432 QYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ ++ DV+ AE + L+ F L + Y A +R R+K +N+
Sbjct: 419 YFEQKKDVNGAENLLEILKNGTDNIGAEIFEPLIRTYAAAGKSHPAMRRRLKMENV 474
>UniRef100_Q9LNY2 F22G5.3 [Arabidopsis thaliana]
Length = 555
Score = 167 bits (423), Expect = 6e-40
Identities = 126/455 (27%), Positives = 212/455 (45%), Gaps = 31/455 (6%)
Query: 59 GLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQA------------------ 100
G++V SAL W+ G + ++ A+N LR+ RAL+
Sbjct: 78 GVTVGSALQSWMGDGFPVHGGDVYHAINRLRKLGRNKRALEMSNLGMLLDRNEFLLMKIL 137
Query: 101 ---LDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANC 157
++W+ + E EY+ L+ KL G+ + EK VP F+ ELLY L+ C
Sbjct: 138 LNLMEWIIRERPYRLGELEYSYLLEFTVKLHGVSQGEKLFTRVPQEFQNELLYNNLVIAC 197
Query: 158 ASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKK-IADVLLMMEKENVKPSS 216
+R E KMRELG+ + N+L++ ++K IA L +M+ + P
Sbjct: 198 LDQGVIRLALEYMKKMRELGYRTSHLVYNRLIIRNSAPGRRKLIAKDLALMKADKATPHV 257
Query: 217 YTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKE 276
TY IL+ ++ ++IDG+ + + MK G E + ++ LA +A A L EA +E
Sbjct: 258 STYHILMKLEANEHNIDGVLKAFDGMKKAGVEPNEVSYCILAMAHAVARLYTVAEAYTEE 317
Query: 277 IEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCES--KPRVEDCLAAIEAWGRLKKI 334
IE +N L+ LY LG+ E+ R W V R + L A EA+ R+ +
Sbjct: 318 IEKSITGDNWSTLDILMILYGRLGKEKELARTWNVIRGFHHVRSKSYLLATEAFARVGNL 377
Query: 335 EEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDAL 393
+ AE ++ EM + K + SLL +Y + ++ K + + M +G T+ L
Sbjct: 378 DRAEELWLEMKNVKGLKETEQFNSLLSVYCKDGLIEKAIGVFREMTGNGFKPNSITYRHL 437
Query: 394 VSLYVQAGEVEKADTVLQKALQQNKMK------PMFTTFMTIMEQYAKRGDVHNAEKIFY 447
+A +++A ++ L K P T ++I+E +A++GDV N+EK+F
Sbjct: 438 ALGCAKAKLMKEALKNIEMGLNLKTSKSIGSSTPWLETTLSIIECFAEKGDVENSEKLFE 497
Query: 448 RLRQANYISRISPFHALAQAYKNAKLPAYGIRERM 482
++ A Y ++AL +AY AK+ + +RM
Sbjct: 498 EVKNAKYNRYAFVYNALFKAYVKAKVYDPNLFKRM 532
>UniRef100_Q9SKU6 Hypothetical protein At2g20710 [Arabidopsis thaliana]
Length = 490
Score = 166 bits (420), Expect = 1e-39
Identities = 117/404 (28%), Positives = 196/404 (47%), Gaps = 8/404 (1%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
S+ LD W+++G + E+ + LR+ + ALQ DW+ ++ E +E + A +
Sbjct: 53 SIIKVLDGWLDQGNLVKTSELHSIIKMLRKFSRFSHALQISDWMSEHRVHEISEGDVAIR 112
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPV 180
LDLIAK+ GL +AEK+ E +P R LY LL AS + L K E+ F +M+ELGF
Sbjct: 113 LDLIAKVGGLGEAEKFFETIPMERRNYHLYGALLNCYASKKVLHKAEQVFQEMKELGFLK 172
Query: 181 TAFACNQLLLIYKKIDKKKIADVLLM-MEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
N +L +Y + K + + LL ME E VKP +T + + +D++GM + +
Sbjct: 173 GCLPYNVMLNLYVRTGKYTMVEKLLREMEDETVKPDIFTVNTRLHAYSVVSDVEGMEKFL 232
Query: 240 ETMKA-EGCELDHLTRASLARHYAAAGLTEKT-EAILKEIEGENLKENMWVCPTLLRLYA 297
+A +G LD T A A Y AGLTEK E + K + N ++ L+ Y
Sbjct: 233 MRCEADQGLHLDWRTYADTANGYIKAGLTEKALEMLRKSEQMVNAQKRKHAYEVLMSFYG 292
Query: 298 ILGRADEVERIWKVCESKPRVEDC--LAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARN 354
G+ +EV R+W + + + ++ I A ++ IEE E + E L R
Sbjct: 293 AAGKKEEVYRLWSLYKELDGFYNTGYISVISALLKMDDIEEVEKIMEEWEAGHSLFDIRI 352
Query: 355 YESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL 414
L+ Y + M+ K ++++ + +TW+ L Y AG++EKA ++A+
Sbjct: 353 PHLLITGYCKKGMMEKAEEVVNILVQKWRVEDTSTWERLALGYKMAGKMEKAVEKWKRAI 412
Query: 415 QQNK--MKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYIS 456
+ +K +P M+ ++ + D+ KI L + +IS
Sbjct: 413 EVSKPGWRPHQVVLMSCVDYLEGQRDMEGLRKILRLLSERGHIS 456
>UniRef100_Q7XTG6 OSJNBb0026L04.10 protein [Oryza sativa]
Length = 524
Score = 162 bits (410), Expect = 2e-38
Identities = 133/476 (27%), Positives = 221/476 (45%), Gaps = 51/476 (10%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
+V +D +++ K + E+G+ + LR++ +Y AL+ L + + + + T + A +
Sbjct: 34 AVREEVDGFLDSRKRAFKWEVGVCVRRLRKQALYRPALK-LSEVMARRGMNPTVSDQAIR 92
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENL-RKTEETFNKMRELGFP 179
LDL+AK RG+ AEKY +P + + L Y LL NC + + K E KM+EL F
Sbjct: 93 LDLVAKSRGIAAAEKYFLDLPETSKTHLTYGALL-NCYCKDLMTEKAEALMGKMKELNFA 151
Query: 180 VTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQI 238
TA N L+ +Y K+++ +K+ V+ M+ ++V P YTY + + DI G+ ++
Sbjct: 152 FTAMCYNSLMTLYTKVNQHEKVPSVIQDMKADDVLPDIYTYNVWMRALAARVDIKGVERV 211
Query: 239 VETMKAEG-CELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYA 297
+E MK +G D T ++LA Y AGL EK EA LKE+E N ++ L+ LYA
Sbjct: 212 IEEMKRDGRVTPDWTTYSNLASIYVDAGLFEKAEAALKELEKWNTSNDLEAYQFLITLYA 271
Query: 298 ILGRADEVERIWK-VCESKPRVEDC--LAAIEAWGRLKKIEEAEAVF------------- 341
EV R+W+ + ++PR + L I+A LK + AEA F
Sbjct: 272 RTQNLVEVHRVWRSLKRNQPRRANMSYLNMIQALANLKDLPGAEACFKEWEAQYINPPKT 331
Query: 342 --------EMMSNKWKLTA-----------------RNYESLLKIYIRHKMLNKGKDLIK 376
E SN+ + A R +++K YI M +K + K
Sbjct: 332 NTKAPGTAETSSNESDVKATKDKGTDGELKHPKYDIRVANAMIKAYITEGMFDKAVAVKK 391
Query: 377 TMGDSGCTIGPTTWDALVSLYVQAGEVEK----ADTVLQKALQQNKM-KPMFTTFMTIME 431
G + TW+ + Y++ G+++ AD ++K ++ P T+M+
Sbjct: 392 RAKMRGGRLNAKTWEIFMEHYLKEGDLKMVHWCADRAIKKGHSAGRIWVPPHEVTETLMD 451
Query: 432 QYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ K DV AEK L++ F L + Y A G+R R+K +N+
Sbjct: 452 YFEKNKDVDGAEKFVEVLKKVQKDLGTVVFEPLVRTYAAAGKKLPGMRHRLKIENV 507
Score = 49.7 bits (117), Expect = 2e-04
Identities = 48/213 (22%), Positives = 92/213 (42%), Gaps = 15/213 (7%)
Query: 271 EAILKEIEG------ENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAA 324
+A+ +E++G K + VC LR A+ A ++ + P V D
Sbjct: 33 QAVREEVDGFLDSRKRAFKWEVGVCVRRLRKQALYRPALKLSEVMARRGMNPTVSDQAIR 92
Query: 325 IEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCT 384
++ + + I AE F + K T Y +LL Y + M K + L+ M +
Sbjct: 93 LDLVAKSRGIAAAEKYFLDLPETSK-THLTYGALLNCYCKDLMTEKAEALMGKMKELNFA 151
Query: 385 IGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEK 444
+++L++LY + + EK +V+Q ++ + + P T+ M A R D+ E+
Sbjct: 152 FTAMCYNSLMTLYTKVNQHEKVPSVIQD-MKADDVLPDIYTYNVWMRALAARVDIKGVER 210
Query: 445 IFYRLRQANYISRISP----FHALAQAYKNAKL 473
+ +++ R++P + LA Y +A L
Sbjct: 211 VIEEMKRD---GRVTPDWTTYSNLASIYVDAGL 240
>UniRef100_Q84JR3 Hypothetical protein At4g21705 [Arabidopsis thaliana]
Length = 492
Score = 161 bits (408), Expect = 3e-38
Identities = 113/422 (26%), Positives = 206/422 (48%), Gaps = 21/422 (4%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
SV L WV+ GK++S E+ ++ LRRRK + AL+ W+ F+ E+A
Sbjct: 40 SVYPELQNWVQCGKKVSVAELIRIVHDLRRRKRFLHALEVSKWMNETGVCVFSPTEHAVH 99
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPV 180
LDLI ++ G AE+Y E++ ++ + Y LL +N+ K+ F KM+E+GF
Sbjct: 100 LDLIGRVYGFVTAEEYFENLKEQYKNDKTYGALLNCYVRQQNVEKSLLHFEKMKEMGFVT 159
Query: 181 TAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
++ N ++ +Y I + +K+ VL M++ENV P +Y+Y+I I+ G D++ + +
Sbjct: 160 SSLTYNNIMCLYTNIGQHEKVPKVLEEMKEENVAPDNYSYRICINAFGAMYDLERIGGTL 219
Query: 240 ETM-KAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAI 298
M + + +D T A A+ Y G ++ +LK E K++ L+ LYA
Sbjct: 220 RDMERRQDITMDWNTYAVAAKFYIDGGDCDRAVELLKMSENRLEKKDGEGYNHLITLYAR 279
Query: 299 LGRADEVERIW----KVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARN 354
LG+ EV R+W VC+ + +D L +++ ++ + EAE V +WK +
Sbjct: 280 LGKKIEVLRLWDLEKDVCKRRIN-QDYLTVLQSLVKIDALVEAEEVL----TEWKSSGNC 334
Query: 355 YE-----SLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTV 409
Y+ ++++ YI M K + +++ + G P +W+ + + Y + G +E A
Sbjct: 335 YDFRVPNTVIRGYIGKSMEEKAEAMLEDLARRGKATTPESWELVATAYAEKGTLENAFKC 394
Query: 410 LQKAL----QQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALA 465
++ AL K +P T +++ G + E LR +++ +HAL
Sbjct: 395 MKTALGVEVGSRKWRPGLTLVTSVLSWVGDEGSLKEVESFVASLRNCIGVNK-QMYHALV 453
Query: 466 QA 467
+A
Sbjct: 454 KA 455
>UniRef100_Q9SY07 Hypothetical protein T5J8.14 [Arabidopsis thaliana]
Length = 532
Score = 158 bits (400), Expect = 3e-37
Identities = 124/460 (26%), Positives = 213/460 (45%), Gaps = 8/460 (1%)
Query: 35 DSSDKKSFSSRRRSELFKAIVSV--SGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRK 92
+S++KK R L ++S+ + S + KW E+G + + E+ + LR+ K
Sbjct: 48 ESANKKETVVGGRDTLGGRLLSLVYTKRSAVVTIRKWKEEGHSVRKYELNRIVRELRKIK 107
Query: 93 MYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRT 152
Y AL+ +W+ + ++ +YA LDLI+K+RGL AEK+ E +P+ RG +
Sbjct: 108 RYKHALEICEWMVVQEDIKLQAGDYAVHLDLISKIRGLNSAEKFFEDMPDQMRGHAACTS 167
Query: 153 LLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENV 212
LL + + K E F KM E GF + N +L +Y + + VL+ K
Sbjct: 168 LLHSYVQNKLSDKAEALFEKMGECGFLKSCLPYNHMLSMYISRGQFEKVPVLIKELKIRT 227
Query: 213 KPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEA 272
P TY + + ND++G ++ K E D +T + L YA EK
Sbjct: 228 SPDIVTYNLWLTAFASGNDVEGAEKVYLKAKEEKLNPDWVTYSVLTNLYAKTDNVEKARL 287
Query: 273 ILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCES---KPRVEDCLAAIEAWG 329
LKE+E K+N +L+ L+A LG D V WK +S K + L+ I A
Sbjct: 288 ALKEMEKLVSKKNRVAYASLISLHANLGDKDGVNLTWKKVKSSFKKMNDAEYLSMISAVV 347
Query: 330 RLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPT 388
+L + E+A+ ++ E S AR +L Y+ + G+ + + + G +
Sbjct: 348 KLGEFEQAKGLYDEWESVSGTGDARIPNLILAEYMNRDEVLLGEKFYERIVEKGINPSYS 407
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMT-IMEQYAKRGDVHNAEKIFY 447
TW+ L Y++ ++EK KA+ K + + ++ ++G+V AEK+
Sbjct: 408 TWEILTWAYLKRKDMEKVLDCFGKAIDSVKKWTVNVRLVKGACKELEEQGNVKGAEKLMT 467
Query: 448 RLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
L++A Y++ +++L + Y A A + ERM DN+
Sbjct: 468 LLQKAGYVN-TQLYNSLLRTYAKAGEMALIVEERMAKDNV 506
>UniRef100_Q8W4P4 Hypothetical protein At4g02820; T5J8.14 [Arabidopsis thaliana]
Length = 532
Score = 157 bits (397), Expect = 7e-37
Identities = 123/460 (26%), Positives = 213/460 (45%), Gaps = 8/460 (1%)
Query: 35 DSSDKKSFSSRRRSELFKAIVSV--SGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRK 92
+S++KK R L ++S+ + S + KW E+G + + E+ + LR+ K
Sbjct: 48 ESANKKETVVGGRDTLGGRLLSLVYTKRSAVVTIRKWKEEGHSVRKYELNRIVRELRKIK 107
Query: 93 MYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRT 152
Y AL+ +W+ + ++ +YA LDLI+K+RGL AEK+ E +P+ RG +
Sbjct: 108 RYKHALEICEWMVVQEDIKLQAGDYAVHLDLISKIRGLNSAEKFFEDMPDQMRGHAACTS 167
Query: 153 LLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENV 212
LL + + K E F KM E GF + N +L +Y + + VL+ K
Sbjct: 168 LLHSYVQNKLSDKAEALFEKMGECGFLKSCLPYNHMLSMYISRGQFEKVPVLIKELKIRT 227
Query: 213 KPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEA 272
P TY + + ND++G ++ K E D +T + L YA EK
Sbjct: 228 SPDIVTYNLWLTAFASGNDVEGAEKVYLKAKEEKLNPDWVTYSVLTNLYAKTDNVEKARL 287
Query: 273 ILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCES---KPRVEDCLAAIEAWG 329
L+E+E K+N +L+ L+A LG D V WK +S K + L+ I A
Sbjct: 288 ALREMEKLVSKKNRVAYASLISLHANLGDKDGVNLTWKKVKSSFKKMNDAEYLSMISAVV 347
Query: 330 RLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPT 388
+L + E+A+ ++ E S AR +L Y+ + G+ + + + G +
Sbjct: 348 KLGEFEQAKGLYDEWESVSGTGDARIPNLILAEYMNRDEVLLGEKFYERIVEKGINPSYS 407
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMT-IMEQYAKRGDVHNAEKIFY 447
TW+ L Y++ ++EK KA+ K + + ++ ++G+V AEK+
Sbjct: 408 TWEILTWAYLKRKDMEKVLDCFGKAIDSVKKWTVNVRLVKGACKELEEQGNVKGAEKLMT 467
Query: 448 RLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
L++A Y++ +++L + Y A A + ERM DN+
Sbjct: 468 LLQKAGYVN-TQLYNSLLRTYAKAGEMALIVEERMAKDNV 506
>UniRef100_Q8LNF7 Putative leaf protein [Oryza sativa]
Length = 513
Score = 154 bits (390), Expect = 4e-36
Identities = 115/429 (26%), Positives = 203/429 (46%), Gaps = 13/429 (3%)
Query: 69 WVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLR 128
W+ + + +S+ E+ + LRRR + +AL+ W+ + L + + A +L+LI+K+
Sbjct: 73 WLAEERPVSKPELQSLVKYLRRRCRFSQALELSMWMTERRHLHLSPGDVAYRLELISKVH 132
Query: 129 GLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQL 188
GL KA +Y + VPN R Y +LL A E + K EE F MR +G ++A N +
Sbjct: 133 GLDKAVEYFDAVPNQLRELQCYGSLLRCYAEAERVEKAEELFENMRGMGM-ANSYAYNAM 191
Query: 189 LLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGC 247
+ +Y +I + +++ + ME+ + P +T L+ D++ + +++E
Sbjct: 192 MNLYSQIGQVERVDSMYKAMEEGGIVPDIFTIDNLVSAYADVEDVEAIEKVLEKASCNNL 251
Query: 248 ELDHLTRASLARHYAAAGLTEKTEAILKEIEGE--NLKENMWVCPTLLRLYAILGRADEV 305
H + A + + + AG+ E+ +E E K+ LL +YA L EV
Sbjct: 252 MSWH-SFAIVGKVFMKAGMQERALQAFQESEKRITARKDGRVAYGFLLTMYADLQMDSEV 310
Query: 306 ERIWKVCESKPRVEDC----LAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARNYESLLK 360
+RIW V SK C + I ++ I AE +E +K +R LL
Sbjct: 311 DRIWDVYRSKVPASACNTMYMCRISVLLKMNDIVGAEKAYEEWESKHVYHDSRLINILLT 370
Query: 361 IYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL--QQNK 418
Y + ++ K + L+ G T+ TW L Y + G+V KA + +KAL N+
Sbjct: 371 AYCKEGLMEKAEALVDQFIKKGRTLFSNTWYKLAGGYFKVGQVSKAADLTKKALASASNE 430
Query: 419 MKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGI 478
KP + + +A++ +V AE++ L++ ++R +H L + Y NA PA +
Sbjct: 431 WKPDLANVLMSINYFAEQKNVEAAEEMASLLQRLVPLTR-DVYHGLLKTYVNAGEPASDL 489
Query: 479 RERMKADNL 487
+RMK D +
Sbjct: 490 LDRMKKDGI 498
>UniRef100_Q7XN02 OSJNBb0038F03.9 protein [Oryza sativa]
Length = 511
Score = 153 bits (386), Expect = 1e-35
Identities = 120/425 (28%), Positives = 198/425 (46%), Gaps = 13/425 (3%)
Query: 38 DKKSFSSRRRSELFKAI-VSVSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGR 96
D + ++ R S L++ I V G V L W E + L + E+ LR+ + +
Sbjct: 66 DYERRAALRWSSLYRRIAVGHGGRPVGRTLGAWDEGERRLDKWELCRIARELRKFRRFNL 125
Query: 97 ALQALDWL-ESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLA 155
ALQ DW+ E + + + A +LDLIAK+RG+ AE+Y E +P+ + + Y +LL
Sbjct: 126 ALQVYDWMTERRDRFSLSSSDMAIQLDLIAKVRGVSHAEEYFEELPDPLKDKRTYGSLLN 185
Query: 156 NCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLL--MMEKENVK 213
A KTE TF +MR+ GF N L+ Y ++ + +L+ MME+ NV
Sbjct: 186 VYAQAMMKEKTESTFEQMRKKGFATDTLPFNVLMNFYVDAEEAEKVSILIDEMMER-NVA 244
Query: 214 PSSYTYKILIDVKGLSNDIDGMSQIVETM-KAEGCELDHLTRASLARHYAAAGLTEKTEA 272
TY I I D D M Q+ M + E + T +LA + G +EK E
Sbjct: 245 FDVCTYNIWIKSCAAMQDADAMEQVFNQMIRDETVVANWTTYTTLASMHIKLGNSEKAEE 304
Query: 273 ILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESK-PRVEDC--LAAIEAWG 329
LKE E L+ LY+ LG+ +EV R+W ++ P + + + A
Sbjct: 305 SLKEAEKRTTGREKKCFHYLMTLYSHLGKKEEVYRVWNWYKATFPTIHNLGYQEVLSALV 364
Query: 330 RLKKIEEAEAVFEMMSNK-WKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPT 388
RL IE AE ++E ++K + LL Y R + K + + + G P
Sbjct: 365 RLGDIEGAELLYEEWASKSSSFDPKTMNILLAWYAREGFVTKAEQTLNRFVEKGGNPKPN 424
Query: 389 TWDALVSLYVQAGEVEKADTVLQKA---LQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKI 445
TW+ L + Y++ G+ +A + L+KA +K +P T +++ + ++ D +A+++
Sbjct: 425 TWEILGTAYLKDGQSSEALSCLEKATAVASPSKWRPRPTNVESLLANFKEKNDAESADRL 484
Query: 446 FYRLR 450
LR
Sbjct: 485 MNVLR 489
>UniRef100_Q8LNF8 Putative leaf protein [Oryza sativa]
Length = 545
Score = 148 bits (374), Expect = 3e-34
Identities = 116/457 (25%), Positives = 222/457 (48%), Gaps = 18/457 (3%)
Query: 45 RRRSELFKAIVSVSG--LSVDSALDKW-VEKGKELSRQEIGLALNSLRRRKMYGRALQAL 101
R++ LF+ + + + L + L++W + + + +++ EI + L RR+ + +ALQ
Sbjct: 78 RKKDSLFRRVAAAADPRLPLSPVLEQWCLAEERPIAKPEIQSIIKYLCRRRRFSQALQLS 137
Query: 102 DWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLE 161
W+ L + + A +L+LI K+ GL +A +Y + +P+ + + Y +LL A +
Sbjct: 138 MWMTERLHLHLSPGDVAYRLELITKVHGLDRAVEYFDSMPDQLKQQQCYGSLLKCYAEAK 197
Query: 162 NLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYK 220
+ K EE F KMR +G +++A N ++ +Y + + +++ + ME+ + +T
Sbjct: 198 CVEKAEELFEKMRGMGM-ASSYAYNVMMRLYLQDGQVERVHSMHRTMEESGIVADVFTTD 256
Query: 221 ILIDVKGLSNDIDGMSQIVETMKAEGCE--LDHLTRASLARHYAAAGLTEKTEAILKEIE 278
L+ ++ DI+ + +++E KA+ C + + A++ + +G+ E+ +E E
Sbjct: 257 TLVAAYVVAEDIEAIEKVLE--KADTCNDLMTWHSYATIGKVLMQSGMEERALQAFQESE 314
Query: 279 GENLKE-NMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDC----LAAIEAWGRLKK 333
+ K+ N LL +YA LG EV+RIW V +SK C + I ++
Sbjct: 315 KKIAKKSNRVAYGFLLTMYADLGMNSEVDRIWDVYKSKVPASACNSMYMCRISVLLKMND 374
Query: 334 IEEAEAVFEMMSNKWKL-TARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDA 392
I AE +E +K +R LL Y + ++ K + L+ G T TW
Sbjct: 375 IVGAEKAYEEWESKHVYHDSRLINLLLTAYCKEGLMEKAEALVDQFVKKGRTPFGNTWYK 434
Query: 393 LVSLYVQAGEVEKADTVLQKALQ--QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLR 450
L Y + G+ KA + +KAL N+ P T + + +A++ +V AE++ L+
Sbjct: 435 LAGGYFKVGQASKAADLTKKALASGSNEWTPDLTNVLMSLNYFAEQKNVEAAEEMASLLQ 494
Query: 451 QANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ +R +H L + Y NA P + +RMK D +
Sbjct: 495 RLITPTR-DIYHGLLKTYVNAGKPVSDLLDRMKKDGM 530
>UniRef100_O23677 T7I23.8 protein [Arabidopsis thaliana]
Length = 524
Score = 139 bits (351), Expect = 1e-31
Identities = 100/407 (24%), Positives = 194/407 (47%), Gaps = 18/407 (4%)
Query: 60 LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESN-KKLEFTEKEYA 118
L S L++W + G++L++ E+ + LR+ K +AL+ DW+ + ++ + + A
Sbjct: 81 LGAASVLNQWEKAGRKLTKWELCRVVKELRKYKRANQALEVYDWMNNRGERFRLSASDAA 140
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
+LDLI K+RG+P AE++ +P +F+ +Y +LL ++ K E N MR+ G+
Sbjct: 141 IQLDLIGKVRGIPDAEEFFLQLPENFKDRRVYGSLLNAYVRAKSREKAEALLNTMRDKGY 200
Query: 179 PVTAFACNQLLLIYKKI-DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQ 237
+ N ++ +Y + + K+ ++ M++++++ Y+Y I + G ++ M
Sbjct: 201 ALHPLPFNVMMTLYMNLREYDKVDAMVFEMKQKDIRLDIYSYNIWLSSCGSLGSVEKMEL 260
Query: 238 IVETMKAE-GCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLY 296
+ + MK++ + T +++A Y G TEK E L+++E N LL LY
Sbjct: 261 VYQQMKSDVSIYPNWTTFSTMATMYIKMGETEKAEDALRKVEARITGRNRIPYHYLLSLY 320
Query: 297 AILGRADEVERIWKVCES-KPRVEDC--LAAIEAWGRLKKIEEAEAVFEMMSNKWKLTAR 353
LG E+ R+W V +S P + + A + + R+ IE AE V+E +W
Sbjct: 321 GSLGNKKELYRVWHVYKSVVPSIPNLGYHALVSSLVRMGDIEGAEKVYE----EWLPVKS 376
Query: 354 NYES-----LLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADT 408
+Y+ L+ Y+++ L + L M + G +TW+ L + + + +A T
Sbjct: 377 SYDPRIPNLLMNAYVKNDQLETAEGLFDHMVEMGGKPSSSTWEILAVGHTRKRCISEALT 436
Query: 409 VLQKALQ---QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQA 452
L+ A + +P + + DV + E + LRQ+
Sbjct: 437 CLRNAFSAEGSSNWRPKVLMLSGFFKLCEEESDVTSKEAVLELLRQS 483
>UniRef100_Q8LPS6 At1g02150/T7I23.8 [Arabidopsis thaliana]
Length = 524
Score = 139 bits (349), Expect = 2e-31
Identities = 99/407 (24%), Positives = 194/407 (47%), Gaps = 18/407 (4%)
Query: 60 LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESN-KKLEFTEKEYA 118
L S L++W + G++L++ E+ + LR+ K +A++ DW+ + ++ + + A
Sbjct: 81 LGAASVLNQWEKAGRKLTKWELCRVVKELRKYKRANQAIEVYDWMNNRGERFRLSASDAA 140
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
+LDLI K+RG+P AE++ +P +F+ +Y +LL ++ K E N MR+ G+
Sbjct: 141 IQLDLIGKVRGIPDAEEFFLQLPENFKDRRVYGSLLNAYVRAKSREKAEALLNTMRDKGY 200
Query: 179 PVTAFACNQLLLIYKKI-DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQ 237
+ N ++ +Y + + K+ ++ M++++++ Y+Y I + G ++ M
Sbjct: 201 ALHPLPFNVMMTLYMNLREYDKVDAMVFEMKQKDIRLDIYSYNIWLSSCGSLGSVEKMEL 260
Query: 238 IVETMKAE-GCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLY 296
+ + MK++ + T +++A Y G TEK E L+++E N LL LY
Sbjct: 261 VYQQMKSDVSIYPNWTTFSTMATMYIKMGETEKAEDALRKVEARITGRNRIPYHYLLSLY 320
Query: 297 AILGRADEVERIWKVCES-KPRVEDC--LAAIEAWGRLKKIEEAEAVFEMMSNKWKLTAR 353
LG E+ R+W V +S P + + A + + R+ IE AE V+E +W
Sbjct: 321 GSLGNKKELYRVWHVYKSVVPSIPNLGYHALVSSLVRMGDIEGAEKVYE----EWLPVKS 376
Query: 354 NYES-----LLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADT 408
+Y+ L+ Y+++ L + L M + G +TW+ L + + + +A T
Sbjct: 377 SYDPRIPNLLMNAYVKNDQLETAEGLFDHMVEMGGKPSSSTWEILAVGHTRKRCISEALT 436
Query: 409 VLQKALQ---QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQA 452
L+ A + +P + + DV + E + LRQ+
Sbjct: 437 CLRNAFSAEGSSNWRPKVLMLSGFFKLCEEESDVTSKEAVLELLRQS 483
>UniRef100_Q6ZJE1 Putative DNA-binding protein [Oryza sativa]
Length = 506
Score = 134 bits (338), Expect = 5e-30
Identities = 101/385 (26%), Positives = 186/385 (48%), Gaps = 39/385 (10%)
Query: 66 LDKWVEK-GKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLI 124
+++W K G L E+ + LR+R+ + +AL+ +W+ + ++F K++A LDLI
Sbjct: 83 IERWAAKPGNRLRHVELERIVKELRKRRRHRQALEVSEWMNAKGHVKFLPKDHAVHLDLI 142
Query: 125 AKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENL-RKTEETFNKMRELGFPVTAF 183
++ G AE Y ++P+ + E Y LL NC + E L K+ F KM+ELGF +
Sbjct: 143 GEIHGSSAAETYFNNLPDKDKTEKPYGALL-NCYTRELLVEKSLAHFQKMKELGFVFSTL 201
Query: 184 ACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETM 242
N ++ +Y + + +K+ V+ M+ + P +++Y+I I+ G D GM +E M
Sbjct: 202 PYNNIMGLYTNLGQHEKVPSVIAEMKSNGIVPDNFSYRICINSYGTRADFFGMENTLEEM 261
Query: 243 KAE-GCELDHLTRASLARHYAAAGLTEKTEAILKEIEGE-NLKENMWVCPTLLRLYAILG 300
+ E +D T A +A +Y + EK + LK+ E + N+K++ L+ LY LG
Sbjct: 262 ECEPKIVVDWNTYAVVASNYIKGNIREKAFSALKKAEAKINIKDSD-SYNHLISLYGHLG 320
Query: 301 RADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLK 360
EV R+W + MSN + ++Y ++L
Sbjct: 321 DKSEVNRLWAL-------------------------------QMSNCNRHINKDYTTMLA 349
Query: 361 IYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMK 420
+ ++ + + + L+K SG + L++ Y Q ++KA+ +L L++ KM
Sbjct: 350 VLVKLNEIEEAEVLLKEWESSGNAFDFQVPNVLLTGYRQKDLLDKAEALLDDFLKKGKMP 409
Query: 421 PMFTTFMTIMEQYAKRGDVHNAEKI 445
P T++ + YA++GD A ++
Sbjct: 410 PS-TSWAIVAAGYAEKGDAAKAYEL 433
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.317 0.133 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 752,565,847
Number of Sequences: 2790947
Number of extensions: 30024543
Number of successful extensions: 110836
Number of sequences better than 10.0: 733
Number of HSP's better than 10.0 without gapping: 313
Number of HSP's successfully gapped in prelim test: 429
Number of HSP's that attempted gapping in prelim test: 106593
Number of HSP's gapped (non-prelim): 2653
length of query: 488
length of database: 848,049,833
effective HSP length: 131
effective length of query: 357
effective length of database: 482,435,776
effective search space: 172229572032
effective search space used: 172229572032
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Medicago: description of AC135463.1


