

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135416.15 - phase: 0 /pseudo
(662 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
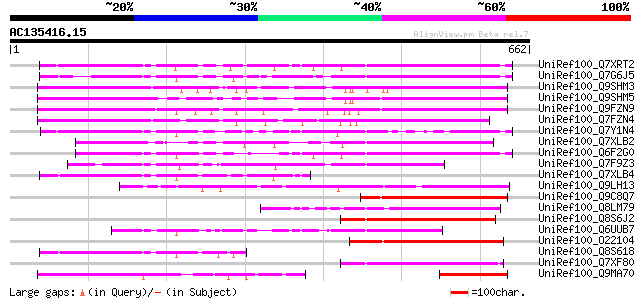
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa] 326 1e-87
UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa] 266 2e-69
UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana] 265 4e-69
UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana] 263 2e-68
UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis ... 258 4e-67
UniRef100_Q7FZN4 T17A2.3 protein [Arabidopsis thaliana] 248 4e-64
UniRef100_Q7Y1N4 Putative retrotransposon gag protein [Oryza sat... 232 2e-59
UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa] 226 2e-57
UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sa... 222 3e-56
UniRef100_Q7F9Z3 OSJNBa0079C19.24 protein [Oryza sativa] 188 4e-46
UniRef100_Q7XLB4 OSJNBa0011K22.15 protein [Oryza sativa] 156 2e-36
UniRef100_Q9LH13 Similarity to Athila retroelement ORF1 protein ... 155 4e-36
UniRef100_Q9C8Q7 Athila ORF 1, putative; 43045-40843 [Arabidopsi... 152 2e-35
UniRef100_Q8LM79 Hypothetical protein OSJNAa0082N11.11 [Oryza sa... 151 7e-35
UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sa... 151 7e-35
UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa] 150 9e-35
UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-... 148 4e-34
UniRef100_Q8S618 Hypothetical protein OSJNBb0048O22.17 [Oryza sa... 148 6e-34
UniRef100_Q7XF80 Hypothetical protein [Oryza sativa] 146 2e-33
UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana] 145 3e-33
>UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa]
Length = 920
Score = 326 bits (836), Expect = 1e-87
Identities = 223/629 (35%), Positives = 326/629 (51%), Gaps = 58/629 (9%)
Query: 38 EAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQ 97
+AVRL LFPFSL RA+ WF++ +I +WD +FL++FFP KT LR +I+RF Q
Sbjct: 298 DAVRLRLFPFSLLRRAKQWFYA-NCAAINTWDKCSTSFLSKFFPIGKTNALRGRISRFQQ 356
Query: 98 KDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNHT 157
E + EAWER +E + CPHHG++ WLI+ FYNGL+ ++ +DAAA GA +K
Sbjct: 357 TRDESIPEAWERLQEYVAACPHHGMDDWLILQNFYNGLTLMSRDHLDAAAGGAFFSKTVQ 416
Query: 158 EAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQ---KFEKL 214
A LIE M N W+ ER T ++ G++ V E LAAK++ L + EK
Sbjct: 417 GAVELIEKMVSNT-GWSKERLQT------RQRGMHTVKETELLAAKLDLLMKCLDDHEKR 469
Query: 215 NVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQGS 274
V + CE+CG +GH G DC E+ Y N RP QG
Sbjct: 470 PQGTVKALDSHVTCEVCGGTGHSGNDC----LETHEEAMYMGKNNEYRP--------QGG 517
Query: 275 YGQVAP----PGYTNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSLSKLTTK 330
G P PG NN + + SL L + Q K L+ + + L + K
Sbjct: 518 EGWNQPRPYYPGGNNNGNFSNQPSLMDL----VFAQAKTTDALRKKLAANDKILENINVK 573
Query: 331 VDSIAT-------HTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSK 383
+D A+ KM++TQ++Q+A V + G PGQP+++ + +V AI G K
Sbjct: 574 LDGFASAFQNQLSFNKMIETQLAQLASLVPANES--GRIPGQPDSSIE-NVKAITTRGGK 630
Query: 384 LEE-----TVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALT-----PLRLTKLNLVTQ 433
A + E+ I+ +K + ++ T P + K ++ Q
Sbjct: 631 STRDPPYPNPAGTNGMSKETPSTDSDDKEIQPDKTVPQEYCDTRLLPFPQQSRKPSVDKQ 690
Query: 434 FTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRE-NSAIIKK 492
F F+ ++QKI I +P +A+ ++P YA++LK+I + K+ + E LT ++ I+ K
Sbjct: 691 FACFVEVIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLPTTEVVKLTEHCSNLILHK 749
Query: 493 PPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLAD 552
P K +DPG I C IG++ +QALCDLGASVS++P +F +L L T L+LAD
Sbjct: 750 LPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKDVFDKLNFTVLAPTPMRLQLAD 809
Query: 553 RSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRI 612
S PVG ED+P KI IP DF+VLD+D E P+ILGRPFL+TAGA +DV G I
Sbjct: 810 SSVHYPVGIAEDVPAKIRDFFIPVDFVVLDMDTGKETPLILGRPFLSTAGANIDVGTGSI 869
Query: 613 IFQVSDELIGFELENYMKGPALYSCNVIK 641
F + + FE + P + C +++
Sbjct: 870 RFHTNGKEEKFEFQ-----PRMEQCTMVR 893
>UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa]
Length = 689
Score = 266 bits (679), Expect = 2e-69
Identities = 197/612 (32%), Positives = 289/612 (47%), Gaps = 78/612 (12%)
Query: 38 EAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQ 97
+ V+L LFPFSL RA+ WF++ ++ +WD AFL++FFP
Sbjct: 75 DVVKLRLFPFSLLGRAKQWFYANRT-AVNTWDKCSTAFLSKFFP---------------- 117
Query: 98 KDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNHT 157
EAWER +E + CPH G++ WLI+ FYNGL+ ++ +DAAA GA +K
Sbjct: 118 ------MEAWERLQEYVAACPHLGMDDWLILQNFYNGLTPMSRDHLDAAARGAFFSKTVQ 171
Query: 158 EAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKF---EKL 214
A LIE M N W+ ER T ++ G++ V E LAAK++ L ++ EK
Sbjct: 172 GAVELIEKMVSN-MGWSEERLQT------RQRGMHTVKETELLAAKLDLLMKRLDDHEKG 224
Query: 215 NVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQGS 274
V + CE+CG +GH G DC E+ Y N G P QG
Sbjct: 225 PQRTVKALDSHVMCEVCGNTGHSGNDC----LETCEEAMYMGNNNGYCP--------QGG 272
Query: 275 YGQVAPPGY----TNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSLSKLTTK 330
G P Y NN + SL+ L + Q K L + N L+ L
Sbjct: 273 QGWNQPRPYYQGGNNNSNFPNQPSLKDL----VFAQAKTTDALSKKLA-ANDKLASLVP- 326
Query: 331 VDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAK 390
A T + Q + V + G P N I G +
Sbjct: 327 ----ANETGRIPGQPDPSIENVKAITMRRGKSTRDPPYPNPVGTNEISKGAPSNDSA--- 379
Query: 391 AKSVKGESVKCLGKKGVIESEKLLDKDKALTPLRLTKLNLVTQFTKFLNILQKICIKIPF 450
K V+ E+ ++ D P R+ K ++ QF +F+ ++QKI I +P
Sbjct: 380 DKEVQPENTV---------PQEYCDTRLLSFPQRMRKPSVDEQFARFVEVIQKIHINVPL 430
Query: 451 AEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRE-NSAIIKKPPTKLRDPGSFAIPCMI 509
+A+ ++P YA++LK+I + K+ + E LT + ++ I+ K P K +DPG I C I
Sbjct: 431 LDAM-QVPTYARYLKDILNNKRPLPTTEVVKLTEQCSNLILHKLPEKKKDPGCPTITCSI 489
Query: 510 GSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDIPVKI 569
G++ +Q C+LGASVS++ +F +L L T L+LAD S P G ED+PVKI
Sbjct: 490 GAQQFDQVSCNLGASVSVVLKDVFDKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKI 549
Query: 570 EGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRIIFQVSDELIGFELENYM 629
IP DF+VLD+D E P+ILGRPFL+TAGA +DV G I F ++ + FE +
Sbjct: 550 RDFFIPVDFVVLDMDTGKETPLILGRPFLSTAGANIDVGTGSIRFHINGKKEKFEFQ--- 606
Query: 630 KGPALYSCNVIK 641
P C++++
Sbjct: 607 --PRTEQCSMVR 616
>UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 265 bits (676), Expect = 4e-69
Identities = 217/708 (30%), Positives = 337/708 (46%), Gaps = 129/708 (18%)
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRF 95
+++ +L LFPFSL D+A W +L SIT+WDD ++ FL++FF ++TA+LR++I+ F
Sbjct: 65 SEDGFKLRLFPFSLGDKAHIWEKNLPHDSITTWDDCKKVFLSKFFSNARTARLRNEISGF 124
Query: 96 NQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKN 155
+QK GE EAWERFK CPHHG K ++ T Y G+ +M +D A+ G NK+
Sbjct: 125 SQKTGESFCEAWERFKGYTNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKD 184
Query: 156 HTEAYSLIEDMAQN--HYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKFEK 213
E + L+E++AQ+ +Y +R V T + E+ N ++ K
Sbjct: 185 VEEGWELVENLAQSDGNYNEDCDRTVRG-TADSDDKHRKEIKALNDKLDRILLSQHKHVH 243
Query: 214 LNVN----AVTPSSASPPCEICGISGHIG--------------VDCQLGSAANIEQLNY- 254
V+ V + E+ I+ + G + + + AN + Y
Sbjct: 244 FLVDDEQYEVQDGEGNQLEEVSYINNNQGGYKGYNYFKTNNPNLSYRSTNVANPQDQVYP 303
Query: 255 -----------AQFNQGTRPNQKFYKNPQGSYGQVAPPGYTNNQR-------VAKKSSLE 296
+NQG P Q+F QG+Y Q PPG+ Q K L+
Sbjct: 304 PQQQQSQNKPFVPYNQGFVPKQQF----QGNY-QQQPPGFAPPQHQGPAAPDADMKQMLQ 358
Query: 297 ILLE---NCIANQNKNLQELKNQTGFLNGSLSKLTTKVDSIATHTKMLKTQISQVAQQVA 353
LL +C K + EL N+ L+ S + L K++++ T + L+ + +
Sbjct: 359 QLLHGQASCSMEMAKKISELHNK---LDCSYNDLNVKMETLDTKVRYLEGHSTSSSATKQ 415
Query: 354 TSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKL 413
TS PG+ NPK + +AI L K T + K+V +S G+ +
Sbjct: 416 TSQ-----LPGKAVQNPKEYAHAITLRSGKALPTREEPKTVTEDSEDQDGE------DLS 464
Query: 414 LDKDKALTPLRLT---KLNLVTQ---------FTK------------------FLNILQK 443
L+KD+A PL L+ L+L Q FT+ + +K
Sbjct: 465 LEKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKEK 524
Query: 444 ICIKIPFAEAL------------SRMPLYAKFLKEIFSKKKAID--------HNETK--- 480
+ + P+ L ++AK +KE+ + +D H K
Sbjct: 525 VFVPPPYKSQLPFPGRHKKALADKYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKDLI 584
Query: 481 -----------ALTRENSAIIKKP--PTKLRDPGSFAIPCMIGSETLNQALCDLGASVSL 527
L+ E SAII+K P KL DPGSF +PC +G N+ LCDLGASVSL
Sbjct: 585 VERIQEVQGMVVLSHECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSL 644
Query: 528 LPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDN 587
+PL + RLG + K +L LADRS P G +E++P++I ++IPTDF+VL++DE+
Sbjct: 645 MPLSVAKRLGFTQYKSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEP 704
Query: 588 ECPIILGRPFLATAGAIVDVQNGRIIFQV-SDELIGFELENYMKGPAL 634
+ P+ILGRPFLATAGA++DV+ G+I + D + F++++ MK P +
Sbjct: 705 KDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTI 752
>UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 263 bits (671), Expect = 2e-68
Identities = 200/634 (31%), Positives = 315/634 (49%), Gaps = 108/634 (17%)
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRF 95
+++ +L LFPFSL D+A W +L SIT+WDD ++AFL++FF ++TA+LR++I+ F
Sbjct: 200 SEDGFKLRLFPFSLGDKAHIWEKNLPHDSITTWDDCKKAFLSKFFSNARTARLRNEISGF 259
Query: 96 NQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKN 155
+QK GE EAWERFK C HHG K ++ T Y G+ +M +D A+ G NK+
Sbjct: 260 SQKTGESFCEAWERFKGYTNQCSHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKD 319
Query: 156 HTEAYSLIEDMAQNH--YQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKFEK 213
E + L+E++AQ++ Y +R V S + +++AL K ++
Sbjct: 320 VEEGWELVENLAQSNGNYNENCDRTVRGTADSDDKH-----------RKEIKALNDKLDR 368
Query: 214 LNVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQG 273
+ + S + + + Q G +E+++Y NQG K Y N +
Sbjct: 369 ILL--------SQHKHVHFLVDDEQYEVQDGEGNQLEEVSYINNNQG---GYKGYNNFK- 416
Query: 274 SYGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSLSKLTTKVDS 333
TNN ++ +S+ +AN + + Q S +K +
Sbjct: 417 ----------TNNPNLSYRSTN-------VANPQDQVYPPQQQQ-----SQNKPFVPYNQ 454
Query: 334 IATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAKAKS 393
T L PG+ NPK + +AI L K T + K+
Sbjct: 455 ATKQTSQL---------------------PGKAVQNPKEYAHAITLHSGKALPTREEPKT 493
Query: 394 VKGESVKCLGKKGVIESEKLLDKDKALTPLRLT---KLNLVTQ---------FTK----- 436
V +S G+ + L KD+A PL L+ L+L Q FT+
Sbjct: 494 VTEDSEDQDGE------DLSLKKDQADKPLDLSLEQPLDLSLQQSLDPPLDSFTRPTTRP 547
Query: 437 -------------FLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALT 483
+ +K+ ++IP +AL+ +P KFLK++ ++ + L+
Sbjct: 548 VIPAASPTAPKPVAVKNKEKVELRIPLVDALALIPDSHKFLKDLIVERIQ-EVQGMVVLS 606
Query: 484 RENSAIIKKP--PTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGEL 541
E SAII+K P KL DPGSF +PC +G N+ LCDLGASVSL+PL + RLG +
Sbjct: 607 HECSAIIQKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQY 666
Query: 542 KLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATA 601
K +L LADRS P G +E++P++I ++IPTDF+VL++DE+ + P+ILGRPFLATA
Sbjct: 667 KSCNISLILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATA 726
Query: 602 GAIVDVQNGRIIFQV-SDELIGFELENYMKGPAL 634
GA++DV+ G+I + D + F++++ MK P +
Sbjct: 727 GAMIDVKKGKIDLNLGKDFRMTFDVKDAMKKPTI 760
>UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1864
Score = 258 bits (659), Expect = 4e-67
Identities = 203/684 (29%), Positives = 329/684 (47%), Gaps = 97/684 (14%)
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRF 95
+++ +L LFPFSL D+A W SL GSITSW+D ++AFLA+FF S+TA+LR+ I+ F
Sbjct: 114 SEDGFKLRLFPFSLGDKAHQWEKSLLQGSITSWNDCKKAFLAKFFSNSRTARLRNDISGF 173
Query: 96 NQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKN 155
Q + E EAWERFK CPHHG K ++ T Y G+ +M +D A+ G +NK+
Sbjct: 174 TQTNNETFCEAWERFKGYQTQCPHHGFSKASLLSTLYRGVLPKIRMLLDTASNGNFLNKD 233
Query: 156 HTEAYSLIEDMAQN--HYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKF-- 211
+ + L+E++AQ+ +Y +R+V + S E E+ N K+ + QK
Sbjct: 234 VEDGWELVENLAQSDGNYNEDYDRSVRTSSDSD-EKHRREMKAMNDKLDKLLLVQQKHIH 292
Query: 212 -----EKLNVNAVTPSSASPPCEICGISG-----------HIGVDCQLGSAANIEQLNYA 255
E V + + G H + + + AN + Y
Sbjct: 293 FLGDDETFQVQDGETMQSEEVSYVQNQGGYNKGFNNFKQNHPNLSYRSTNVANPQDQVYP 352
Query: 256 Q-----------FNQGTR--PNQKFYKNPQGSYG-QVAPPGYTNNQR----VAKKSSLEI 297
+NQG P Q++ QG+Y Q+ PPG+T Q+ S L+
Sbjct: 353 SQQQNQPRPFVPYNQGQGYVPKQQY----QGNYQPQLPPPGFTQQQQQPASTTPDSDLKN 408
Query: 298 LLENCIANQNKNLQELKNQTGFLNG----SLSKLTTKVDSIATHTKMLKTQISQVAQQVA 353
+L+ + Q +L + ++ S + + KV+++ + + ++ Q A
Sbjct: 409 MLQQILQGQATGAMDLSKRMAEIHNKVDCSYNDINIKVEALTSKIRYIEGQTGSTAAPKF 468
Query: 354 TSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAKAKSVKGESVKCLGK---KGVIES 410
T + +N + + +AI L K T +SV G+ + +
Sbjct: 469 TGPSRKSM------SNSEEYAHAITLRSGKELPTKESPNQNTEDSVDQDGEDFCQNGNSA 522
Query: 411 EKLLDKDKALTPLRL----------------TKLNLVT-----------------QFTKF 437
EK +++ P RL TK N+ K+
Sbjct: 523 EKAIEEPILDQPTRLLAPAASPLVEKPAATKTKDNVFVPPPYKPPLPFPGRFKKVMIQKY 582
Query: 438 LNILQK----ICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKP 493
+L+K + + +P + L+ +P K++K++ +++ + L+ E SAII++
Sbjct: 583 KALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITERIK-EVQGMVVLSHECSAIIQQK 641
Query: 494 --PTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLA 551
P KL DPGSF +PC +G N+ LCDLGASVSL+PL + +LG + K +L LA
Sbjct: 642 IIPKKLGDPGSFTLPCALGPLAFNKCLCDLGASVSLMPLSVAKKLGFNKYKPCNISLILA 701
Query: 552 DRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGR 611
DRS P G +ED+PV I +++PTDF+VL++DE+ + P+ILGRPFL T GAI+DV+ G+
Sbjct: 702 DRSVRIPHGLLEDLPVMIGMVEVPTDFVVLEMDEEPKDPLILGRPFLVTVGAIIDVKKGK 761
Query: 612 IIFQVSDEL-IGFELENYMKGPAL 634
I + +L + F++ N MK P +
Sbjct: 762 IDLNLGRDLKMTFDITNTMKKPTI 785
>UniRef100_Q7FZN4 T17A2.3 protein [Arabidopsis thaliana]
Length = 1428
Score = 248 bits (633), Expect = 4e-64
Identities = 184/624 (29%), Positives = 302/624 (47%), Gaps = 66/624 (10%)
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRF 95
+++ +L LFPFSL D+A W +L SIT+WDD ++AFL++FF ++TA+LR++I F
Sbjct: 82 SEDGFKLRLFPFSLGDKAHIWEKNLSHDSITTWDDYKKAFLSKFFSNARTARLRNEIYGF 141
Query: 96 NQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKN 155
+QK GE EAWERFK CPHH K ++ T Y G+ +M +D A+ G NK+
Sbjct: 142 SQKTGESFCEAWERFKGYTNQCPHHSFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKD 201
Query: 156 HTEAYSLIEDMAQN--HYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKFEK 213
E + L+E++AQ+ +Y +R V T + E+ N ++ QK
Sbjct: 202 VEEGWELVENLAQSDGNYNEDCDRTVRG-TADSDDKHRKEIKALNDKLDRILLSQQKHVH 260
Query: 214 LNVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQG 273
V+ ++ G+ + L + + +NQG P Q+F QG
Sbjct: 261 FLVD-------DEQYQVQDGEGNQLEEVYLPQQKQGQNKPFVLYNQGFVPKQQF----QG 309
Query: 274 SYGQVAPPGYTNNQ---RVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSL----SK 326
+Y PPG+ Q A + ++ +++ + Q + E+ + L+ L +
Sbjct: 310 NYQPPPPPGFVTQQNQCHAAPDAKMKQMVKQLLQGQASSSMEIAKKLSELHHKLDCSYND 369
Query: 327 LTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPK--------------- 371
L KV+++ T + L+ Q +T S PG+ NPK
Sbjct: 370 LNAKVEALNTKVRYLE------GQSASTFSPKVTRLPGKSIQNPKEYATAHAITICHDRE 423
Query: 372 ---AHVNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKDKAL-------- 420
HV + + ++E A + V+ V+ G + ++KA
Sbjct: 424 LPTRHVLDLITRDNDVQEGEASTQ-VEASVVEFNHSAGSRHLTQSTSEEKAAIIERMVKR 482
Query: 421 ---TPLRLTKLNLVTQ---FTKFLNI----LQKICIKIPFAEALSRMPLYAKFLKEIFSK 470
TPL L L + ++ ++ L +I +P E L+ + K ++ + +
Sbjct: 483 FKPTPLPLRALPWTFRKAWMERYKSVAAKQLDEIEAVMPLMEVLNLILDPHKVVRNLILE 542
Query: 471 --KKAIDHNETKALTRENSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLL 528
K D ++ T +A + KL DPGSF +PC IG + LCDLGASV+L+
Sbjct: 543 RIKMYHDSDDESDATPSRAADKRIVQEKLEDPGSFTLPCSIGEFAFSDCLCDLGASVNLM 602
Query: 529 PLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNE 588
PL + RL + K + TL LADRS + G ++D+PV I G+++PTDF+VLD++ +++
Sbjct: 603 PLSMARRLEFIQYKPCDLTLILADRSSRKHFGMLKDLPVMINGVEVPTDFVVLDMEVEHK 662
Query: 589 CPIILGRPFLATAGAIVDVQNGRI 612
P+ILGRPFLA+ GA++DV+ G+I
Sbjct: 663 DPLILGRPFLASVGAVIDVKEGKI 686
>UniRef100_Q7Y1N4 Putative retrotransposon gag protein [Oryza sativa]
Length = 697
Score = 232 bits (592), Expect = 2e-59
Identities = 186/615 (30%), Positives = 289/615 (46%), Gaps = 101/615 (16%)
Query: 40 VRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQKD 99
VRL LFPFSL R + WF++ + +WD AFL++FFP KT LR +I+ F Q
Sbjct: 77 VRLRLFPFSLLRRVKQWFYA-NCAAANTWDKCSMAFLSKFFPMGKTNALRGRISSFQQTM 135
Query: 100 GEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNHTEA 159
E + EAWER +E + CPHHG++ WLI+ FYNGL+ ++ +DAAA GA +K A
Sbjct: 136 DESIPEAWERLQEYVAACPHHGMDDWLILQNFYNGLTPMSRDHLDAAAGGAFFSKTVQGA 195
Query: 160 YSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKF---EKLNV 216
IE M N W+ ER T ++ G++ V E L AK++ L ++ EK+
Sbjct: 196 VEFIEKMVSN-MGWSEERLQT------RQRGMHTVKETELLPAKLDLLMKRLDDHEKMPQ 248
Query: 217 NAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQGSYG 276
V + CE+CG +GH G +C E+ Y N G R G
Sbjct: 249 GTVKALDSHVTCEVCGDTGHPGNNC----PETREEAMYMGNNNGYR-----------RQG 293
Query: 277 QVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSLSKLTTKVDSIAT 336
+ ++KK + + + N+N L GF + ++L
Sbjct: 294 DLVFAQAKTTDSLSKKLAAN---DKIVENRNVKLD------GFASAFQNQL--------C 336
Query: 337 HTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAKAKSVKG 396
KM++TQ++Q+A V + G PGQP+++ + +V AI G K +
Sbjct: 337 FNKMMETQLAQLASLVPANE--TGRIPGQPDSSIE-NVKAITTRGGKSTRDPPYPNPTRT 393
Query: 397 ESV-KCLGKKGVIESEKLLDK-------DKALTPL--RLTKLNLVTQFTKFLNILQKICI 446
+ K G+ + E +K D L P R+ K ++ QF +F+ ++QKI I
Sbjct: 394 NGLSKEAPSNGLADKEVQPEKTMPQEYCDIRLLPFPQRMRKPSVDEQFARFVEVIQKIHI 453
Query: 447 KIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDPGSFAIP 506
+P +A+ ++P YA+ LK+I + K+ + E LT +++P T P
Sbjct: 454 NMPLLDAM-QVPTYARCLKDILNNKRPLPTTEVVKLT----GTMQQPDT----------P 498
Query: 507 CMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDIP 566
G E + A P P+ L+L +++++ R +D+P
Sbjct: 499 QAPGEEERSGA-----------PTPM-------RLQLVDSSIRYTARI-------AKDVP 533
Query: 567 VKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRIIFQVSDELIGFELE 626
VKI IP DF+VLD+D E P ILGRPFL+TAGA +DV G I F ++ + FE +
Sbjct: 534 VKIRYFFIPVDFVVLDMDTGKETPFILGRPFLSTAGANIDVGTGSIYFHINGKEEKFEFQ 593
Query: 627 NYMKGPALYSCNVIK 641
P C++++
Sbjct: 594 -----PRTEQCSMVR 603
>UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa]
Length = 551
Score = 226 bits (575), Expect = 2e-57
Identities = 171/550 (31%), Positives = 259/550 (47%), Gaps = 96/550 (17%)
Query: 84 KTAKLRDQITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSV 143
KT LR +I+ F Q E + EAWER +E + CPHHG++ WLI+ FYNGL+ + +
Sbjct: 3 KTNALRGRISSFQQTRDESIPEAWERLQEYMAACPHHGMDDWLILQNFYNGLTPMSHDHL 62
Query: 144 DAAACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAK 203
DAAA GA +K A LIE M + ++ KK G +V + +H+
Sbjct: 63 DAAARGAFFSKTVQGAVDLIEKMLDLLMKRLDDH-------DKKPQGTVKVLD-SHVT-- 112
Query: 204 VEALTQKFEKLNVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRP 263
CE+CG +GH G DC + + Y N G P
Sbjct: 113 ------------------------CEVCGNTGHSGNDC-----LDTREAMYMGNNNGYCP 143
Query: 264 NQKFYKNPQGSYGQVAPPGYTNNQRVAKKSSLEIL---LENCIANQNKNLQELKNQTGFL 320
Q G P Y + + A + L + L++ + Q K + L +
Sbjct: 144 --------QRGQGWNQPCPYVDRKHEAWEICLTPVQPSLKDLVFAQAKTIDALSKKLATN 195
Query: 321 NGSLSKLTTKVDSIAT-------HTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAH 373
+ L + K+D A+ KML+TQ++Q A V +
Sbjct: 196 DKILENINVKLDGFASAFQNQLRFNKMLETQLAQFASLVPANET---------------- 239
Query: 374 VNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALTPL-----RLTKL 428
G + + + V + S E ++ EK++ ++ T L R K
Sbjct: 240 ------GTNGIAKEVPSSDSADKE----------VQLEKIVPQEYCDTWLLSFHQRSRKP 283
Query: 429 NLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRE-NS 487
++ QF +F ++QKI I +P +A+ ++P YA++LK++ + K+ + E LT + +S
Sbjct: 284 SVDEQFARFAEVIQKIHINVPLLDAM-QVPTYARYLKDMLNNKRLLPTMEVVKLTEQCSS 342
Query: 488 AIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETT 547
I+ K P K +DPG I C IG++ +QA CDLGAS+S++P + +L L T
Sbjct: 343 VILHKLPEKKKDPGCPTITCSIGAQQFDQAFCDLGASISVMPKDVPDKLNFTVLTPTLMC 402
Query: 548 LKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDV 607
L+LAD S P G ED+PVKI IP DF+VLD+D E P+ILGRPFL+TAGA +DV
Sbjct: 403 LQLADLSVHYPTGIAEDVPVKIRDFFIPVDFVVLDMDMGKETPLILGRPFLSTAGANIDV 462
Query: 608 QNGRIIFQVS 617
G I F ++
Sbjct: 463 GMGSIRFDIN 472
>UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sativa]
Length = 585
Score = 222 bits (565), Expect = 3e-56
Identities = 174/572 (30%), Positives = 263/572 (45%), Gaps = 92/572 (16%)
Query: 84 KTAKLRDQITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSV 143
KT L +I+ F Q E + EAWE +E + CPHHG++ WLI+ FYN L+ ++ +
Sbjct: 3 KTDALCGRISSFQQTRDESIPEAWELLQEYVAACPHHGMDDWLILQNFYNRLTPMSRDHL 62
Query: 144 DAAACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAK 203
DAAA GA +K LIE M N W+ ER T ++ G++ V E LAAK
Sbjct: 63 DAAAGGAFFSKTVQGTVELIEKMVSN-MGWSKERLQT------RQRGMHTVKETELLAAK 115
Query: 204 VEALTQKF---EKLNVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQG 260
++ L ++ EK V + CE+CG + H G DC E + N G
Sbjct: 116 LDLLMKRLDNHEKRPQGTVKALDSHVTCEVCGGTDHSGNDCP---ETREEAMYMGNNNNG 172
Query: 261 TRPNQKFYKNPQGSYGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFL 320
RP QG G P Y N N+
Sbjct: 173 YRP--------QGVQGWNQPRSYYQG-----------------GNNNE------------ 195
Query: 321 NGSLSKLTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLG 380
TQ++Q+A V + G PGQP ++ + +V AI
Sbjct: 196 ----------------------TQLAQLAYLVPANET--GRIPGQPYSSIE-NVKAITTR 230
Query: 381 GSKLEE-----TVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALT-----PLRLTKLNL 430
G K A + E+ I+ EK + ++ T P K ++
Sbjct: 231 GGKSTRDPPYPNPAGTNGMSKETPSNDSADKEIQPEKTVPQEYCDTRLLPFPQWSRKTSV 290
Query: 431 VTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRE-NSAI 489
F +F+ ++Q+I I + +A+ ++P+YA++LK+I + K+ + E LT ++ I
Sbjct: 291 DELFARFVEVIQRIHINVLLLDAM-QVPIYARYLKDILNNKRPLPTTEVVKLTEHCSNVI 349
Query: 490 IKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLK 549
+ K P K + I C IG++ +QALCDLGASVS++P +F RL L T L+
Sbjct: 350 LHKLPEKKKYSECPTITCSIGAQQFDQALCDLGASVSVMPKDVFDRLNFTVLAPTPMRLQ 409
Query: 550 LADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQN 609
+AD S G ED+P+KI IP DF+VLD+D E P+ILGRPFL+TAGA +DV+
Sbjct: 410 MADSSVRYQAGIAEDVPIKIWDFFIPVDFVVLDMDTRKETPLILGRPFLSTAGANIDVET 469
Query: 610 GRIIFQVSDELIGFELENYMKGPALYSCNVIK 641
I F ++++ FE + P C++++
Sbjct: 470 SSICFHINEKEEKFEFQ-----PRTEQCSMVR 496
>UniRef100_Q7F9Z3 OSJNBa0079C19.24 protein [Oryza sativa]
Length = 461
Score = 188 bits (478), Expect = 4e-46
Identities = 148/492 (30%), Positives = 232/492 (47%), Gaps = 45/492 (9%)
Query: 74 AFLARFFPPSKTAKLRDQITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYN 133
AFL++FFP KT LR +I+ F Q E + EAWER PHHG++ LI+ FYN
Sbjct: 2 AFLSKFFPMGKTNALRGRISNFQQTRDESIPEAWER--------PHHGMDNSLILQNFYN 53
Query: 134 GLSYTTKMSVDAAACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYE 193
GL+ + +DAAA GA +K A LIE M N W+ ER T ++ GI+
Sbjct: 54 GLTTMSHDHLDAAAGGAFFSKTVRGAIDLIEKMVSN-MGWSKERLQT------RQQGIHT 106
Query: 194 VSEYNHLAAKVEALTQKFEKLN---VNAVTPSSASPPCEICGISGHIGVDCQLGSAANIE 250
V E LAAK++ L ++ + + + V CE+CG +GH G DC E
Sbjct: 107 VKETELLAAKLDLLMKRLDDHDNRPQDTVKALDLHVTCEVCGGTGHSGNDCP----ETHE 162
Query: 251 QLNYAQFNQGTRPNQKFYKNPQGSYGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNL 310
+ Y N N ++ PQG G P Y + S + L++ I Q K
Sbjct: 163 EAMYMGKN-----NNEY--RPQGGQGWNQPRPYYQGENNNSNFSNQPSLKDLIFVQAKTT 215
Query: 311 QELKNQTGFLNGSLSKLTTKVDSIATH-------TKMLKTQISQVAQQVATSSQTPGVFP 363
L + + L + ++D + KM++TQ++Q+A V +
Sbjct: 216 DALSKKLAANDKILENINVQLDGFTSSFQNQLIFNKMIETQLAQLASLVPANEIGRIPRR 275
Query: 364 GQPETNPKAHVNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALTPL 423
G T + + N G E + + + + + K + ++ D P
Sbjct: 276 GGKSTRDRPYPNPAGTNGVAREAPSSDSANKEVQPEKAV-------PQEYYDTRLLSFPQ 328
Query: 424 RLTKLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALT 483
R K ++ QF +F+ ++QKI I + +A+ ++P YA++LK+I + K+ + E LT
Sbjct: 329 RSRKPSVDEQFARFVEVIQKIHINVSLFDAM-QVPTYARYLKDILNNKRPLPTTEVIELT 387
Query: 484 RE-NSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELK 542
+ ++ I+ K K + PG I C IG++ NQAL DLG SVS++P+ +F +L L
Sbjct: 388 EQCSNVILHKLSEKRKHPGCPTITCSIGAQQFNQALRDLGVSVSVIPMDVFDKLNFTVLA 447
Query: 543 LTETTLKLADRS 554
T L+LAD S
Sbjct: 448 PTPMRLQLADSS 459
>UniRef100_Q7XLB4 OSJNBa0011K22.15 protein [Oryza sativa]
Length = 406
Score = 156 bits (395), Expect = 2e-36
Identities = 117/356 (32%), Positives = 170/356 (46%), Gaps = 32/356 (8%)
Query: 38 EAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQ 97
+AVRL LF FSL RA+ WF++ +I +WD AFL++FFP KT LR +I+ F Q
Sbjct: 35 DAVRLRLFLFSLLGRAKQWFYANRA-AINTWDKCSTAFLSKFFPMGKTNALRGRISSFQQ 93
Query: 98 KDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNHT 157
E + EAWER +E + CP HG++ WLI+ FYNGL+ ++ +D AA GA +K
Sbjct: 94 TRDESIPEAWERLQEYVAACPLHGMDDWLILLNFYNGLTLMSRDHLDTAAGGAFFSKTVQ 153
Query: 158 EAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKF---EKL 214
A LIE M N W+ ER T + G++ V E LAAK++ L ++ EK
Sbjct: 154 GAVDLIEKMVSN-MGWSEERLQT------HQQGMHIVKETELLAAKLDLLIKRLDDHEKQ 206
Query: 215 NVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQGS 274
V CE+C + H G DC E + N G RP N
Sbjct: 207 PQGTVKALDYHITCEVCSNTSHSGNDC---PETREEAMYMGNNNNGYRPQGGQRWNQPRP 263
Query: 275 YGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQELKNQTGFLNGSLSKLTTKVDSI 334
Y Q G NN + +SSL+ L + Q K L + + L + K+D
Sbjct: 264 YYQ----GGNNNGNFSNQSSLKDL----VFAQAKTTDALSKKLAANDNILENINVKLDGF 315
Query: 335 A-------THTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSK 383
A + KM++T ++Q+A V + + GQP+++ + +V I G K
Sbjct: 316 ASTFQNQLSFNKMIETHLAQLASLVPANESRRVL--GQPDSSVE-NVRVIITRGGK 368
>UniRef100_Q9LH13 Similarity to Athila retroelement ORF1 protein [Arabidopsis
thaliana]
Length = 583
Score = 155 bits (392), Expect = 4e-36
Identities = 125/513 (24%), Positives = 245/513 (47%), Gaps = 37/513 (7%)
Query: 141 MSVDAAACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHL 200
M++D A+ G M K TEA LIE++A ++ + V ++ G E ++ L
Sbjct: 1 MALDTASNGEFMTKTETEATKLIENLAGSN----SNHNVDYDRSNRGGGG--ESKQFAEL 54
Query: 201 AAKVEALTQKFEKLNVNAVTPSSASPPCEICGISGHIGVDCQLG---SAANIEQLNYAQF 257
+AKVE L ++ +K +VN SS + G + ++ + N+E +
Sbjct: 55 SAKVEQLMRRDQK-SVNFCEDSSKGMVHQEFSGDGSEDLQAEINFVNGSTNVENPQDQVY 113
Query: 258 -NQGTRPNQKF----YKNPQGSYGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQE 312
Q QKF ++N GQ PP T + + ++++++ + +Q KN +
Sbjct: 114 PTQAGSQGQKFQPYGFQNKGNYQGQFQPPAGTGHASSFGDNEMKLMMQQVLEDQKKNAAD 173
Query: 313 LKNQTGFLNGSLSKLTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKA 372
+ + ++ + L K ++++H K L+ Q+SQ+ V+ S + G G+ + K
Sbjct: 174 INVK---VDSMYNDLNGKFATLSSHVKTLENQVSQI---VSASMRPDGTHSGKVKPKGKE 227
Query: 373 HVNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKD------KALTPLRLT 426
AI + E VAK V+ L + ++E ++ L + K P R
Sbjct: 228 QCYAIMIQEELREIVVAKQVETNVVVVETLVEDKIVEDDEPLSVEPPPYVPKLPFPGRER 287
Query: 427 KLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTR-- 484
++ ++ +F I++++ +++PF + + +P Y +LK I S K++I+ K +++
Sbjct: 288 QIQRQKEYARFDEIMKQLYVRLPFLQLVLHVPSYRSYLKYILSNKRSIEEG-VKLISKGE 346
Query: 485 ENSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLT 544
E++ +++ + + + + G E +N ++ P + +LG+ K +
Sbjct: 347 EHAQLVESQRQQKEAQLTNVVEMLAGKEMVNTCS-------AIPPATIPKKLGITNFKPS 399
Query: 545 ETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAI 604
+L LADRS P+G E++ ++ IPT+F+VL++D++ P+ LGRPFL T AI
Sbjct: 400 RISLILADRSVQFPMGLAENVHARVGNFYIPTNFVVLELDKEPHDPLNLGRPFLNTVEAI 459
Query: 605 VDVQNGRIIFQVSDELIGFELENYMKGPALYSC 637
+DV+ I Q+ D + F+++ K P + C
Sbjct: 460 IDVRRSTINLQIGDYALEFDIKGTRKNPTIEDC 492
>UniRef100_Q9C8Q7 Athila ORF 1, putative; 43045-40843 [Arabidopsis thaliana]
Length = 530
Score = 152 bits (385), Expect = 2e-35
Identities = 79/190 (41%), Positives = 128/190 (66%), Gaps = 4/190 (2%)
Query: 448 IPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKP--PTKLRDPGSFAI 505
+P + L+ +P K++K++ +++ + L+ E SAII++ KL DPGSF +
Sbjct: 121 MPLVDCLALIPNEHKYVKDLITERIK-EVQGMVVLSHECSAIIQQKIVQEKLEDPGSFTL 179
Query: 506 PCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVEDI 565
PC I T + +LCDLGASVS++PL + +LG + K + TL LADR+ +P G +ED+
Sbjct: 180 PCSIRQLTFSNSLCDLGASVSIMPLSMARKLGFVQYKPCDLTLILADRTSRRPFGLLEDV 239
Query: 566 PVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRIIFQVSDEL-IGFE 624
PV I G+++P DF+VL++DE+++ P+ILGRPFLA+AGA++DV+ G+I + ++ + FE
Sbjct: 240 PVMINGVEVPIDFVVLEMDEESKDPLILGRPFLASAGAVIDVKQGKINLNLGEDFKMKFE 299
Query: 625 LENYMKGPAL 634
+ N MK P +
Sbjct: 300 IRNTMKKPTI 309
>UniRef100_Q8LM79 Hypothetical protein OSJNAa0082N11.11 [Oryza sativa]
Length = 322
Score = 151 bits (381), Expect = 7e-35
Identities = 105/306 (34%), Positives = 164/306 (53%), Gaps = 21/306 (6%)
Query: 321 NGSLSKLTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLG 380
N L + + + KM++TQ++Q+A V + PGQP+++ + ++ AI
Sbjct: 27 NVKLDGFASAFQNQLSFNKMIETQLAQLASLVPANESER--IPGQPDSSIE-NIKAITTR 83
Query: 381 GSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALTPLRLTKLNLVTQFTKFLNI 440
G S + S K+ I+ EK + ++ T + QF + + +
Sbjct: 84 GG------TNGMSKETPSTDLADKE--IQPEKTVPQEYCDT----RGVGNRQQFARSVEV 131
Query: 441 LQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDP 500
+QKI I +P +A+ ++P YA++LK+I + K+ + E ++ I+ K P K +DP
Sbjct: 132 IQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLPTTE-----HCSNVILHKLPEKKKDP 185
Query: 501 GSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVG 560
I C IG++ +QALCDLGASVS++P +F +L L T L+LAD S PVG
Sbjct: 186 RCPTITCSIGAQQFDQALCDLGASVSVMPKDVFDKLNFTVLAPTPMRLQLADSSVHCPVG 245
Query: 561 NVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRIIFQVSDEL 620
ED+PVKI IP DF+VLD+D E +ILG PFL+TAGA +DV G I F ++ +
Sbjct: 246 IEEDVPVKIWDFFIPVDFVVLDMDTGKETSLILGHPFLSTAGANIDVGMGSIRFHINGKE 305
Query: 621 IGFELE 626
FE +
Sbjct: 306 EKFEFQ 311
>UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sativa]
Length = 237
Score = 151 bits (381), Expect = 7e-35
Identities = 88/199 (44%), Positives = 120/199 (60%), Gaps = 2/199 (1%)
Query: 422 PLRLTKLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKA 481
P R K ++ QF F+ ++QKI I +P +A+ ++P YA +LK+I + K+ + E
Sbjct: 36 PQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLPTTEVVK 94
Query: 482 LTRE-NSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGE 540
LT + ++ I+ K P K +DPG I C IG++ +QALCDLGASVS++P +F +L
Sbjct: 95 LTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLNFTV 154
Query: 541 LKLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLAT 600
L T L+LAD S P G ED+PVKI I DF+VLD+D E IILGRPFL T
Sbjct: 155 LAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPFLGT 214
Query: 601 AGAIVDVQNGRIIFQVSDE 619
AGA + V G I FQ E
Sbjct: 215 AGANIHVGTGSIRFQYQRE 233
>UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa]
Length = 450
Score = 150 bits (380), Expect = 9e-35
Identities = 133/431 (30%), Positives = 209/431 (47%), Gaps = 64/431 (14%)
Query: 131 FYNGLSYTTKMSVDAAACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAG 190
FYNGL+ ++ +DAAA GA +K A L+E M N W+ E+ T + G
Sbjct: 74 FYNGLTPMSRDHLDAAAGGAFFSKMVQGAVDLVEKMVSN-MGWSEEQLQTC------QRG 126
Query: 191 IYEVSEYNHLAAKVEALTQKF---EKLNVNAVTPSSASPPCEICGISGHIGVDCQLGSAA 247
++ V E LAAK++ L ++ EK V + CE+C +GH G DC
Sbjct: 127 MHTVKETELLAAKLDLLMKRLDNHEKRPQGTVKALDSHVTCEVCSSTGHSGNDCP----- 181
Query: 248 NIEQLNYAQFNQGTRPNQKFYKNPQGSYG--QVAP--PGYTNNQRVAKKSSLEILLENCI 303
E A + N +Y+ PQG G Q+ P G NN + + SL+ L +
Sbjct: 182 --ETREEAMY----MGNNNWYR-PQGGQGWSQLRPYYQGGNNNGNFSNQPSLKDL----V 230
Query: 304 ANQNKNLQELKNQTGFLNGSLSKLTTKVDSIATHTKMLKTQ-ISQVAQQVATSSQTPGVF 362
+Q K L + +AT+ K+L+ ++Q+A V + G
Sbjct: 231 FSQAKTTDALSKK-----------------LATNDKILENNNLAQLASLVPVNET--GRI 271
Query: 363 PGQPETNPKAHVNAIFLGGSKLEETVAKAKSVKGESVKCLGKKGVIESEKLLDKDKALTP 422
PGQP+++ + +V AI + G+ ++ K + K V ++ D L P
Sbjct: 272 PGQPDSSIE-NVKAITMRGASSSDSADKEDQPE---------KTV--PQEYCDTWLLLFP 319
Query: 423 LRLTKLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKAL 482
R+ K + QF +F+ ++QKI I +P +A+ +MP YA++LK I + K+ + E L
Sbjct: 320 QRMRKPLVDEQFARFVEVIQKIHINVPLLDAM-QMPTYARYLKGILNNKRPLPTTEVVKL 378
Query: 483 TRE-NSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGEL 541
T + ++ I+ K P K +DPG I C IG++ +QALCDLGASVS++P +F +L L
Sbjct: 379 TEQWSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKNVFDKLNFTML 438
Query: 542 KLTETTLKLAD 552
T L+LAD
Sbjct: 439 APTPMHLQLAD 449
>UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-like gene [Vicia
faba]
Length = 238
Score = 148 bits (374), Expect = 4e-34
Identities = 82/199 (41%), Positives = 121/199 (60%), Gaps = 3/199 (1%)
Query: 434 FTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIK-- 491
F FL + +K+ + I EAL +MP YAKF+K+I SKK+ ID + T S+I++
Sbjct: 14 FEIFLEMFKKLELNILVLEALEKMPTYAKFMKDIISKKRTID-TDPIIHTETCSSILQGM 72
Query: 492 KPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLA 551
K P K +D G PC IG + + +LGASVSLLP ++ +LG+ ++ T TL+ A
Sbjct: 73 KIPVKKKDRGFVTTPCTIGDRSFKKYFINLGASVSLLPFYIYKKLGISNVQNTRMTLRFA 132
Query: 552 DRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGR 611
D S P G VED+ VKI+ P DF+VL++ ED E +ILGRPFL T ++D++ G
Sbjct: 133 DHSVKIPYGIVEDVLVKIDKFVFPVDFVVLEMPEDEEITLILGRPFLETRRCLIDIEEGT 192
Query: 612 IIFQVSDELIGFELENYMK 630
+ +V DE + +++N MK
Sbjct: 193 VTLKVYDEELKIDVQNTMK 211
>UniRef100_Q8S618 Hypothetical protein OSJNBb0048O22.17 [Oryza sativa]
Length = 412
Score = 148 bits (373), Expect = 6e-34
Identities = 100/279 (35%), Positives = 141/279 (49%), Gaps = 27/279 (9%)
Query: 38 EAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQ 97
+A++L LFPFSL +A+ WF++ I +WD AF ++FFP KT LR +I+ F Q
Sbjct: 75 DAIKLRLFPFSLIGKAKRWFYANRA-MINTWDKCSTAFPSKFFPMGKTNALRGRISSFQQ 133
Query: 98 KDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNHT 157
E + EAWER +E + +CPHHG++ WLI+ FYNGL+ ++ +DAAA GA +K
Sbjct: 134 TRDESIPEAWERLQEYVAVCPHHGMDDWLILQNFYNGLTLMSRDHLDAAAGGAFFSKTVR 193
Query: 158 EAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQKF---EKL 214
A LIE M N W+ ER T + G++ V E LAAK++ L ++ +K
Sbjct: 194 GAIDLIEKMVSN-MGWSEERLQT------HQQGMHTVKETEMLAAKLDLLMKRLDDHDKR 246
Query: 215 NVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPN---QKFYKN- 270
V + CE+CG +GH G D E + N G RP QK + N
Sbjct: 247 PQGTVKALDSHVTCEVCGSTGHSGNDF---LETREEAMYMGSNNNGYRPQGVIQKIHINV 303
Query: 271 PQGSYGQVAPPGYT-------NNQRVAKKSSLEILLENC 302
P QV P Y NN+R ++ L E C
Sbjct: 304 PLLDAMQV--PTYARYLKDILNNKRPLPTMEVDKLTEQC 340
Score = 36.6 bits (83), Expect = 2.4
Identities = 17/48 (35%), Positives = 31/48 (64%), Gaps = 1/48 (2%)
Query: 440 ILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENS 487
++QKI I +P +A+ ++P YA++LK+I + K+ + E LT + S
Sbjct: 295 VIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLPTMEVDKLTEQCS 341
>UniRef100_Q7XF80 Hypothetical protein [Oryza sativa]
Length = 725
Score = 146 bits (369), Expect = 2e-33
Identities = 88/211 (41%), Positives = 128/211 (59%), Gaps = 4/211 (1%)
Query: 422 PLRLTKLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKA 481
P R K ++ QF F+ ++QKI I +P +A+ ++P YA +LK+I + K+ + E
Sbjct: 67 PQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLPTTEVVK 125
Query: 482 LTRE-NSAIIKKPPTKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGE 540
LT + ++ I+ K P K +DPG I C IG++ +QALCDLGASVS++P +F +L
Sbjct: 126 LTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVFDKLNFTV 185
Query: 541 LKLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLAT 600
L T L+LAD S P G ED+PVKI I DF+VLD+D E IILGRPFL T
Sbjct: 186 LAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIILGRPFLGT 245
Query: 601 AGA-IVDVQNGRIIFQVSDELIGFELENYMK 630
AGA I ++Q + +D L+ F + N+++
Sbjct: 246 AGANIHNIQEVEVEPPKTDSLVKF-MHNFLE 275
>UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana]
Length = 823
Score = 145 bits (367), Expect = 3e-33
Identities = 108/360 (30%), Positives = 172/360 (47%), Gaps = 42/360 (11%)
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRF 95
++++ +L LFPFSL D+A W +L V S+ + DD ++AFLA+FF S+TA+LR++I+ F
Sbjct: 118 SEDSFKLRLFPFSLGDKAHLWEKTLPVDSVDTLDDCKKAFLAKFFSNSRTARLRNEISGF 177
Query: 96 NQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKN 155
NQK+ E EAWERFK CPHHG +K ++ T Y G +M +D + G +NK+
Sbjct: 178 NQKNSESFAEAWERFKGYSTQCPHHGFKKASLLSTLYRGALPKIRMLLDTTSNGNFLNKD 237
Query: 156 HTEAYSLIEDMAQN------HYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVEALTQ 209
E + L+E++AQ+ +Y TN + + K+ + + N E T
Sbjct: 238 VAEGWELVENLAQSDGNYNENYDRTNRGSSDSEDRHKRRTKLLMIRLTNWCLLNREMSTT 297
Query: 210 KFEKLNVNAVTPSSASPPCEICGISGHIGVDCQLGSAANIEQLNYAQFNQGTRPNQKFYK 269
+K N+ G I + + + AN + Y NQ P K +
Sbjct: 298 LQKKSLHNSK--------------KGRILLLRRSNNVANPQDQVYPPQNQ--PPQAKPFI 341
Query: 270 NPQGSYGQ---VAPPGYTN--NQRVAKKSSLEILLE-------NCIANQNKNLQELKNQT 317
Y Q PPG+T Q A+ S ++ LL +C +K L EL T
Sbjct: 342 PYNQGYNQKQNFGPPGFTQQPQQTSAQDSEMKTLLHQLVQGQASCSMTMDKKLAEL---T 398
Query: 318 GFLNGSLSKLTTKVDSIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAI 377
++ S + L K+D++ T K ++ I+ ++ + PG P NPK + +AI
Sbjct: 399 TRIDCSYNDLNIKIDALNTRVKTMEGHIAS-----TSAPKHPGQLPRNSVQNPKEYAHAI 453
Score = 92.0 bits (227), Expect = 5e-17
Identities = 47/87 (54%), Positives = 64/87 (73%), Gaps = 1/87 (1%)
Query: 549 KLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQ 608
KLADRS P G +ED+PVKI I+IPTDF+VL++DE+ + P+ILGRPFLATAGA++DVQ
Sbjct: 580 KLADRSVRVPHGMLEDLPVKIGSIEIPTDFVVLEMDEEPKDPLILGRPFLATAGALIDVQ 639
Query: 609 NGRIIFQVSDEL-IGFELENYMKGPAL 634
G+I + L + F++ MK P +
Sbjct: 640 MGKIDLNLGKNLHMSFDIAKKMKKPTI 666
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,075,698,075
Number of Sequences: 2790947
Number of extensions: 44418749
Number of successful extensions: 122128
Number of sequences better than 10.0: 692
Number of HSP's better than 10.0 without gapping: 466
Number of HSP's successfully gapped in prelim test: 226
Number of HSP's that attempted gapping in prelim test: 121273
Number of HSP's gapped (non-prelim): 920
length of query: 662
length of database: 848,049,833
effective HSP length: 134
effective length of query: 528
effective length of database: 474,062,935
effective search space: 250305229680
effective search space used: 250305229680
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Medicago: description of AC135416.15


