

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135320.7 + phase: 0
(190 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
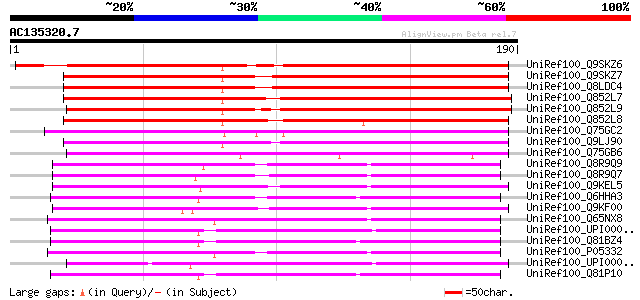
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SKZ6 Putative alanine acetyl transferase [Arabidopsi... 171 7e-42
UniRef100_Q9SKZ7 Putative alanine acetyl transferase [Arabidopsi... 164 1e-39
UniRef100_Q8LDC4 Putative alanine acetyl transferase [Arabidopsi... 163 2e-39
UniRef100_Q852L7 Putative acetyltransferase [Oryza sativa] 156 2e-37
UniRef100_Q852L9 Putative acetyltransferase [Oryza sativa] 137 2e-31
UniRef100_Q852L8 Putative acetyltransferase [Oryza sativa] 133 3e-30
UniRef100_Q75GC2 Putative acetyltransferase [Oryza sativa] 127 2e-28
UniRef100_Q9LJ90 Alanine acetyl transferase-like protein [Arabid... 120 2e-26
UniRef100_Q75GB6 Hypothetical protein OSJNBb0031A14.8 [Oryza sat... 117 2e-25
UniRef100_Q8R9Q9 Acetyltransferases, including N-acetylases of r... 102 4e-21
UniRef100_Q8R9Q7 Acetyltransferases, including N-acetylases of r... 102 5e-21
UniRef100_Q9KEL5 BH0837 protein [Bacillus halodurans] 89 6e-17
UniRef100_Q6HHA3 Ribosomal-protein-alanine acetyltransferase [Ba... 81 2e-14
UniRef100_Q9KF00 Ribosomal-protein-alanine N-acetyltransferase [... 80 3e-14
UniRef100_Q65NX8 YnaD [Bacillus licheniformis] 80 4e-14
UniRef100_UPI00003CBF74 UPI00003CBF74 UniRef100 entry 79 5e-14
UniRef100_Q81BZ4 Ribosomal-protein-alanine acetyltransferase [Ba... 79 8e-14
UniRef100_P05332 Hypothetical P20 protein [Bacillus licheniformis] 79 8e-14
UniRef100_UPI00002E060B UPI00002E060B UniRef100 entry 78 1e-13
UniRef100_Q81P10 Acetyltransferase, GNAT family [Bacillus anthra... 78 1e-13
>UniRef100_Q9SKZ6 Putative alanine acetyl transferase [Arabidopsis thaliana]
Length = 188
Score = 171 bits (434), Expect = 7e-42
Identities = 90/186 (48%), Positives = 124/186 (66%), Gaps = 15/186 (8%)
Query: 3 ITSISSKPGAKEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINF 62
++++SS P K I LRP+ LSD+DD M+W TD V +FC+WE YTS++ I +
Sbjct: 14 LSTVSSSPPEK--------IHLRPMTLSDVDDFMVWATDSNVTRFCTWEPYTSREAAIAY 65
Query: 63 IENIATKFLWCKAICI-NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCV 121
+ + W +AIC+ NDR IG +S++ P D+ R E+GYVLGSKYWGKG+AT
Sbjct: 66 LNDALLPHPWLRAICLDNDRPIGSISVT---PVDEIRG---EIGYVLGSKYWGKGIATEA 119
Query: 122 VKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIIS 181
V+ V F E ++RLEALVDV+N GSQ+VLEK GF KEGV+RK++ +KG RDM++
Sbjct: 120 VRLVAGEIFKEKPEMQRLEALVDVDNVGSQKVLEKVGFVKEGVMRKFMYLKGNVRDMVMF 179
Query: 182 SVLFTD 187
S L +D
Sbjct: 180 SFLPSD 185
>UniRef100_Q9SKZ7 Putative alanine acetyl transferase [Arabidopsis thaliana]
Length = 183
Score = 164 bits (414), Expect = 1e-39
Identities = 84/168 (50%), Positives = 112/168 (66%), Gaps = 7/168 (4%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-N 79
+I+LRP+ LSD+DD M+W TD KVA+FC+WE TS+D+ I +I + W +AIC+ +
Sbjct: 19 RISLRPMTLSDVDDYMVWATDPKVARFCTWEPCTSRDEAIKYITDRVLTHPWLRAICLED 78
Query: 80 DRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERL 139
DR IG + + + N E+GYVL KYWGKG AT V+ V F E +ERL
Sbjct: 79 DRPIGYILIMAVD------NIRKEIGYVLARKYWGKGFATEAVRLVTAEVFEEFPEIERL 132
Query: 140 EALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
EALVDV+N GSQRVLEK GF +EGV+RK++ +KG RD ++ S L TD
Sbjct: 133 EALVDVDNVGSQRVLEKVGFTREGVMRKFICIKGSVRDTVMFSFLSTD 180
>UniRef100_Q8LDC4 Putative alanine acetyl transferase [Arabidopsis thaliana]
Length = 183
Score = 163 bits (413), Expect = 2e-39
Identities = 84/168 (50%), Positives = 112/168 (66%), Gaps = 7/168 (4%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-N 79
+I+LRP+ LSD+DD M+W TD KVA+FC+WE TS+D+ I +I + W +AIC+ +
Sbjct: 19 RISLRPMTLSDVDDYMVWATDPKVARFCTWEPCTSRDEAIKYITDRVLTHPWLQAICLED 78
Query: 80 DRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERL 139
DR IG + + + N E+GYVL KYWGKG AT V+ V F E +ERL
Sbjct: 79 DRPIGYILIMAVD------NIRKEIGYVLARKYWGKGFATEAVRLVTAEVFEEFPEIERL 132
Query: 140 EALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
EALVDV+N GSQRVLEK GF +EGV+RK++ +KG RD ++ S L TD
Sbjct: 133 EALVDVDNVGSQRVLEKVGFTREGVMRKFICIKGSVRDTVMFSFLSTD 180
>UniRef100_Q852L7 Putative acetyltransferase [Oryza sativa]
Length = 193
Score = 156 bits (395), Expect = 2e-37
Identities = 81/178 (45%), Positives = 114/178 (63%), Gaps = 15/178 (8%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-- 78
+++LR +L+D+D +M+W +D +VA C WE Y S + + ++ + W +AIC+
Sbjct: 18 EVSLRRFDLADVDAMMVWASDPQVAAVCRWEPYESTEPLLAYLRDTVLPHPWFRAICVAA 77
Query: 79 --------NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAF 130
DR +G VS+S ++ + AELGYV+ +WGKGVAT VK+VV F
Sbjct: 78 AFDGDGGGEDRPVGAVSVSPTADACR-----AELGYVVARAHWGKGVATAAVKRVVAAVF 132
Query: 131 CELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTDP 188
E+ LER+EALVDV NA SQRVLEKAGF++E VLR Y V+KG+ RDM+I S + TDP
Sbjct: 133 GEVEGLERVEALVDVRNAASQRVLEKAGFRREAVLRSYCVLKGEVRDMVIYSFISTDP 190
>UniRef100_Q852L9 Putative acetyltransferase [Oryza sativa]
Length = 169
Score = 137 bits (344), Expect = 2e-31
Identities = 75/168 (44%), Positives = 105/168 (61%), Gaps = 6/168 (3%)
Query: 22 ITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-ND 80
+TLR L+D D +M W +D +V F +WE Y S D FI + W +AIC+ D
Sbjct: 4 VTLRRFELADADAMMAWASDPEVTAFMTWEPYESVDSLRAFIRDTVLPHPWFRAICLAGD 63
Query: 81 RAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLE 140
G VS++ ++ D+ R AE+ + +WGKGVAT +++ + AF +L +ER+E
Sbjct: 64 GDGGAVSVTPTA--DRCR---AEVAVAVARAHWGKGVATAALRRALAAAFADLDGVERVE 118
Query: 141 ALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTDP 188
ALVDV NA S+R LEKAGFQ+E VLR Y V+KG+ RDM+I S + TDP
Sbjct: 119 ALVDVGNAASRRALEKAGFQQEAVLRSYCVVKGQLRDMVIYSFISTDP 166
>UniRef100_Q852L8 Putative acetyltransferase [Oryza sativa]
Length = 177
Score = 133 bits (334), Expect = 3e-30
Identities = 72/177 (40%), Positives = 108/177 (60%), Gaps = 15/177 (8%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-- 78
++TLR LSD+D +M W +D VA FC WE Y S + + ++ + W +AIC+
Sbjct: 2 EVTLRRFELSDVDAMMAWASDPAVAAFCRWEPYQSTEPLLAYLRDTVLPHPWFRAICLAT 61
Query: 79 -------NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFC 131
+ R +G VSL+ ++ + ELGYV+ +WGKGVAT V++ V
Sbjct: 62 GAGAGDGDGRPVGAVSLAPTADACRG-----ELGYVVARAHWGKGVATAAVRRAVAAVLG 116
Query: 132 -ELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
E+S L R+EALVDV+N SQRV+EKAGF++EGVLR++ KG+ RD+++ S + +D
Sbjct: 117 GEVSGLARVEALVDVDNRASQRVVEKAGFRREGVLRRHYWHKGRVRDLVMYSFVSSD 173
>UniRef100_Q75GC2 Putative acetyltransferase [Oryza sativa]
Length = 203
Score = 127 bits (319), Expect = 2e-28
Identities = 70/187 (37%), Positives = 107/187 (56%), Gaps = 13/187 (6%)
Query: 14 EERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWC 73
E + ++TLR +D + L W +D +V +F + Y+ D+ +I + W
Sbjct: 2 EAEAEAAEVTLREFTEADAEALFAWASDPRVVRFQRRDAYSHVDEARRYIVDKVLPHPWY 61
Query: 74 KAICIN--DRAIGCVSLSSS---------SPGDKSRNKC--AELGYVLGSKYWGKGVATC 120
+AIC+ DR +G +S+ + S + R+ C A +GY + +WG+GVAT
Sbjct: 62 RAICVAGADRPVGSISVKPADDLPLPEPESETGRLRSGCCRASVGYRVAHAHWGRGVATR 121
Query: 121 VVKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMII 180
V+ V + E +LERLEA+ DVEN SQRVLEKAGF +EGVLR+Y+V+KG+ RDM++
Sbjct: 122 AVRAVAEAVLAEWPWLERLEAVADVENPASQRVLEKAGFAREGVLRRYVVLKGRPRDMVM 181
Query: 181 SSVLFTD 187
S + D
Sbjct: 182 FSRVRAD 188
>UniRef100_Q9LJ90 Alanine acetyl transferase-like protein [Arabidopsis thaliana]
Length = 175
Score = 120 bits (300), Expect = 2e-26
Identities = 66/169 (39%), Positives = 99/169 (58%), Gaps = 5/169 (2%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-- 78
+I LRP NLSD +D+ W D+ V ++ W+ S ++ I N A W ++I +
Sbjct: 7 RIFLRPFNLSDAEDVFKWAGDDDVTRYLRWDSVNSLEEAKQHILNKAIPHPWRRSISLLQ 66
Query: 79 NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLER 138
+ +IG VS+ S + R A+L Y + ++WG+G+AT V+ V+ A + + R
Sbjct: 67 DGHSIGYVSVKPDSGDGRCR---ADLAYAVAKEFWGRGIATAAVRMAVEQALEDFPEVVR 123
Query: 139 LEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
L+A+V+VEN SQRVLEKAGF+KEG+L KY KG RDM + S + D
Sbjct: 124 LQAVVEVENKASQRVLEKAGFRKEGLLEKYGFSKGVIRDMFLYSYVKDD 172
>UniRef100_Q75GB6 Hypothetical protein OSJNBb0031A14.8 [Oryza sativa]
Length = 243
Score = 117 bits (292), Expect = 2e-25
Identities = 75/200 (37%), Positives = 99/200 (49%), Gaps = 34/200 (17%)
Query: 22 ITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICINDR 81
+TLRPL L+D DD M W +DE+V +F L +++ + I + W +AIC++D
Sbjct: 21 VTLRPLGLADADDFMAWASDERVMRFLRRPLCAAREQAVAQIRDTVLGHPWFRAICVDDD 80
Query: 82 AIGC----VSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVV--------------- 122
G V S P A LGY L WG+G+AT +
Sbjct: 81 DAGAGRRPVGQVSVWPYADEGGHRANLGYALSHGLWGRGIATAAITMLTAKCINPLFKTL 140
Query: 123 --------------KQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKY 168
QVV F EL LERLEA+ DVEN SQR LEKAGF+KEGVLR+Y
Sbjct: 141 VTVDRRVHARRRPGMQVVARVFDELPGLERLEAVTDVENVRSQRALEKAGFRKEGVLRRY 200
Query: 169 LVMK-GKSRDMIISSVLFTD 187
+V + G+ D +I S L +D
Sbjct: 201 IVRRSGEVMDAVIYSFLASD 220
>UniRef100_Q8R9Q9 Acetyltransferases, including N-acetylases of ribosomal proteins
[Thermoanaerobacter tengcongensis]
Length = 185
Score = 102 bits (255), Expect = 4e-21
Identities = 56/174 (32%), Positives = 101/174 (57%), Gaps = 11/174 (6%)
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFL----- 71
++ +++ L+ ++L D +D+ + +D +V K+ SWE + + +D + FI + +++
Sbjct: 7 LETSRLILKKISLEDAEDMFEYASDPEVTKYVSWEYHKNIEDSLKFINLLLSRYEKGEPS 66
Query: 72 -WCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAF 130
W + N + IG +N E+GYVLG KYW KG+ T V++V++ F
Sbjct: 67 DWGLYLKENGKLIGTCGFLFID----EKNMVGEVGYVLGRKYWNKGIMTEAVRKVIEFGF 122
Query: 131 CELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
+L+ L R++A VEN S+RV++K G + EGVLR+ + +KG+ DM + S+L
Sbjct: 123 EKLN-LNRIQARCKVENIASERVMQKVGMKFEGVLREAVFVKGRFWDMKMYSIL 175
>UniRef100_Q8R9Q7 Acetyltransferases, including N-acetylases of ribosomal proteins
[Thermoanaerobacter tengcongensis]
Length = 187
Score = 102 bits (254), Expect = 5e-21
Identities = 55/174 (31%), Positives = 101/174 (57%), Gaps = 12/174 (6%)
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIAT------KF 70
++ +++ L+ ++L D +D+ + +D +V K+ SWE + + +D +N I+ I +
Sbjct: 7 LETSRLILKKISLEDAEDMFEYASDPEVTKYVSWEYHKNIEDSLNLIKRILSISDKEEVA 66
Query: 71 LWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAF 130
+W + N++ IG L+ +N E+ + LG KYW KG+ T V++V++ F
Sbjct: 67 IWGLYLKENEKLIGTCELNIDR-----KNMIGEIAFALGKKYWNKGIMTEAVREVIEFGF 121
Query: 131 CELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
+L+ L R++A VEN S+RV++K G + EG+LR+ L KG+ DM + S+L
Sbjct: 122 EKLN-LNRIQARCMVENIASERVMQKVGMKFEGILREALFAKGRFWDMKMYSIL 174
>UniRef100_Q9KEL5 BH0837 protein [Bacillus halodurans]
Length = 197
Score = 89.0 bits (219), Expect = 6e-17
Identities = 57/177 (32%), Positives = 88/177 (49%), Gaps = 11/177 (6%)
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKF------ 70
++ ++ LR D + + ++E+V K+ WE + S D F+ K+
Sbjct: 13 LETERLRLRKFYKDDAAAIYDYASNEQVTKYVLWETHQSIKDSEAFLAFALNKYDEKDVS 72
Query: 71 LWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAF 130
W + N+R IG V P DK+ AELGYVL YWG+G+ T V +V+ F
Sbjct: 73 PWAIELKRNERMIGTVDFVWWKPKDKT----AELGYVLSEPYWGQGIMTEAVNALVEFGF 128
Query: 131 CELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
+ LER++A EN S RV+EKAG EG R+ + +KG RD + +++ D
Sbjct: 129 NNME-LERIQAKCFAENISSARVMEKAGLIYEGTHRRAIYVKGAHRDFKVYAIIRED 184
>UniRef100_Q6HHA3 Ribosomal-protein-alanine acetyltransferase [Bacillus
thuringiensis]
Length = 180
Score = 80.9 bits (198), Expect = 2e-14
Identities = 51/175 (29%), Positives = 91/175 (51%), Gaps = 11/175 (6%)
Query: 16 RIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATK------ 69
+++ ++ LR L L D + + + + E V ++ + + + + I+ +
Sbjct: 5 KLETERLQLRELTLLDAETMFHYFSKESVIRYFGMDSFENIEQAKTTIQTFKNRYEEGNV 64
Query: 70 FLWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVA 129
F W D+ IG + + +K AE+GY L YWG+G A+ ++ ++
Sbjct: 65 FRWGIEKKGTDQLIGTCGFHLIN----NHHKRAEIGYELDDTYWGQGYASEALQAILAYG 120
Query: 130 FCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F E L R+ A+V VEN SQ++L KAGFQ+EG+LRK+++ G + D I+ S+L
Sbjct: 121 F-ETLQLIRIAAVVYVENKASQKLLNKAGFQEEGLLRKHMIQNGVAHDTILYSLL 174
>UniRef100_Q9KF00 Ribosomal-protein-alanine N-acetyltransferase [Bacillus halodurans]
Length = 190
Score = 80.1 bits (196), Expect = 3e-14
Identities = 52/177 (29%), Positives = 92/177 (51%), Gaps = 11/177 (6%)
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFI---ENIA---TKF 70
++ ++ LR + D ++ + +D++V K+ E + + +D + I E+I T
Sbjct: 11 LETKRLILRKITTDDARSILSYLSDKEVMKYFGLEPFQTLEDALGEIAWYESILHEQTGI 70
Query: 71 LWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAF 130
W + D IG P ++ AE+G+ L YWG+G+A+ ++ V++ F
Sbjct: 71 RWGITLKGQDEVIGSCGFHQWVP----KHHRAEIGFELSKLYWGQGIASEAIRAVIQYGF 126
Query: 131 CELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
L L+R++AL++ N SQR++EK GF EG+LR Y GK D+ + S+L D
Sbjct: 127 EHLE-LQRIQALIEPPNIPSQRLVEKQGFISEGLLRSYEYTCGKFDDLYMYSLLKRD 182
>UniRef100_Q65NX8 YnaD [Bacillus licheniformis]
Length = 178
Score = 79.7 bits (195), Expect = 4e-14
Identities = 57/172 (33%), Positives = 89/172 (51%), Gaps = 3/172 (1%)
Query: 15 ERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCK 74
E + ++TLR + L D D L + +D +V K+ + +T + I+ I L +
Sbjct: 2 ETLYTERLTLRKMELEDADVLCQYWSDPEVTKYMNIAPFTDVSQARDMIQMINDLSLEGQ 61
Query: 75 A--ICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCE 132
A I + G V + N AE+GY LG +WGKG A+ VK+++ F
Sbjct: 62 ANRFSIIAKETGEVIGTCGFNMIDQENGRAEIGYDLGRNHWGKGFASEAVKKLIDYGFTS 121
Query: 133 LSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
L+ L R+EA V+ EN S ++L FQKEG+LR+Y KG+ D+ + S+L
Sbjct: 122 LN-LNRIEAKVEPENTPSIKLLNSLSFQKEGLLREYEKAKGRLIDVYMFSLL 172
>UniRef100_UPI00003CBF74 UPI00003CBF74 UniRef100 entry
Length = 180
Score = 79.3 bits (194), Expect = 5e-14
Identities = 50/175 (28%), Positives = 90/175 (50%), Gaps = 11/175 (6%)
Query: 16 RIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATK------ 69
+++ ++ LR L L D + + + + E V ++ + + + + I+ +
Sbjct: 5 KLETERLQLRELTLLDAETMFRYFSKESVIRYFGMDSFENIEQAKTTIQTFKNRYEEGSV 64
Query: 70 FLWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVA 129
F W I + G + + +K AE+GY L YWG+G AT ++ ++
Sbjct: 65 FRWG----IEKKGTGQLIGTCGFHLINHHHKRAEIGYELDDTYWGQGYATEALQAILTYG 120
Query: 130 FCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F L + R+ A+V VEN SQ++L KAGFQ+EG+LRK+++ G + D I+ S+L
Sbjct: 121 FDTLQLI-RIAAVVYVENKASQKLLSKAGFQEEGLLRKHMIQNGVAHDTILYSLL 174
>UniRef100_Q81BZ4 Ribosomal-protein-alanine acetyltransferase [Bacillus cereus]
Length = 180
Score = 78.6 bits (192), Expect = 8e-14
Identities = 50/175 (28%), Positives = 92/175 (52%), Gaps = 11/175 (6%)
Query: 16 RIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATK------ 69
+++ ++ LR L L D + + + + E V ++ + + + + I+ +
Sbjct: 5 KLETERLFLRELTLLDAETMFQYFSKESVIRYFGMDSFQNIEQAKTTIQTFKNRYEEGSV 64
Query: 70 FLWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVA 129
F W I + G + + + +K AE+GY L YWG+G A+ ++ ++
Sbjct: 65 FRWG----IEKKGTGQLIGTCGFHLINNHHKRAEIGYELDDTYWGQGYASEALQAILAYG 120
Query: 130 FCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F E+ L R+ A+V VEN SQ++L KAGFQ+EG+LRK+++ G + D I+ S+L
Sbjct: 121 F-EMLQLIRIAAVVYVENKASQKLLIKAGFQEEGLLRKHMIQNGVAHDTILYSLL 174
>UniRef100_P05332 Hypothetical P20 protein [Bacillus licheniformis]
Length = 178
Score = 78.6 bits (192), Expect = 8e-14
Identities = 56/176 (31%), Positives = 88/176 (49%), Gaps = 11/176 (6%)
Query: 15 ERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCK 74
E + ++TLR + L D D L + +D +V K+ + +T + I+ I L +
Sbjct: 2 ETLYTERLTLRKMELEDADVLCQYWSDPEVTKYMNITPFTDVSQARDMIQMINDLSLEGQ 61
Query: 75 A------ICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKV 128
A + D IG + N AE+GY LG +WGKG A+ V++++
Sbjct: 62 ANRFSIIVKETDEVIGTCGFNMID----QENGRAEIGYDLGRNHWGKGFASEAVQKLIDY 117
Query: 129 AFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F L+ L R+EA V+ EN S ++L FQKEG+LR Y KG+ D+ + S+L
Sbjct: 118 GFTSLN-LNRIEAKVEPENTPSIKLLNSLSFQKEGLLRDYEKAKGRLIDVYMFSLL 172
>UniRef100_UPI00002E060B UPI00002E060B UniRef100 entry
Length = 178
Score = 78.2 bits (191), Expect = 1e-13
Identities = 51/167 (30%), Positives = 91/167 (53%), Gaps = 3/167 (1%)
Query: 22 ITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENI-ATKFLWCKAICIND 80
++LR + +DL + T+ V + SW L S DD + + I +T + + + D
Sbjct: 14 LSLRQIERADLGAWYAYLTNPDVYRHTSWNL-RSPDDLLPLFDAIDSTDPDSIRRLAVID 72
Query: 81 RAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLE 140
A G ++ + + N+ AE+ Y L +WG+G+A+ + + V AF + ++ R++
Sbjct: 73 DASGALAGTIGLHTVSTVNRSAEIAYDLAPSHWGRGIASALCEAVTAWAFADGGFM-RMQ 131
Query: 141 ALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
A+V NAGS RVL+K G++ EG+LR Y +++G D + + L TD
Sbjct: 132 AVVLTGNAGSARVLQKCGYRYEGLLRAYKMVRGTPGDFAMYARLATD 178
>UniRef100_Q81P10 Acetyltransferase, GNAT family [Bacillus anthracis]
Length = 180
Score = 78.2 bits (191), Expect = 1e-13
Identities = 50/175 (28%), Positives = 90/175 (50%), Gaps = 11/175 (6%)
Query: 16 RIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATK------ 69
+++ ++ LR L L D + + + + E V ++ + + + + I+ +
Sbjct: 5 KLETERLQLRELTLLDAETMFHYFSKESVIRYFGMDSFENIEQAKTTIQTFKNRYEEGSV 64
Query: 70 FLWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVA 129
F W I + G + + +K AE+GY L YWG+G A+ ++ ++
Sbjct: 65 FRWG----IEKKGTGQLIGTCGFHLINHHHKRAEIGYELDDTYWGQGYASEALQAILTYG 120
Query: 130 FCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVL 184
F E L R+ A+V VEN SQ++L KAGFQ+EG+LRK+++ G + D I+ S+L
Sbjct: 121 F-ETLQLIRIAAVVYVENKASQKLLSKAGFQEEGLLRKHMIQNGVAHDTILYSLL 174
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 304,758,536
Number of Sequences: 2790947
Number of extensions: 11561462
Number of successful extensions: 23748
Number of sequences better than 10.0: 764
Number of HSP's better than 10.0 without gapping: 318
Number of HSP's successfully gapped in prelim test: 446
Number of HSP's that attempted gapping in prelim test: 23046
Number of HSP's gapped (non-prelim): 780
length of query: 190
length of database: 848,049,833
effective HSP length: 120
effective length of query: 70
effective length of database: 513,136,193
effective search space: 35919533510
effective search space used: 35919533510
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC135320.7


