

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135318.3 + phase: 1 /pseudo
(726 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
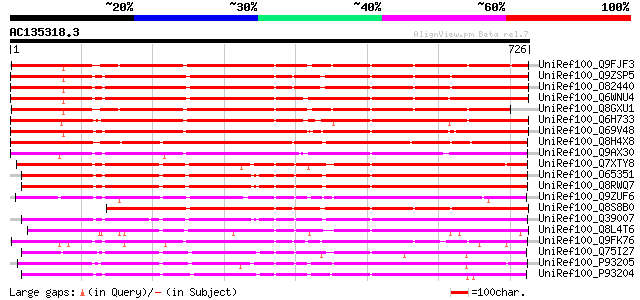
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJF3 Serine protease-like protein [Arabidopsis thali... 705 0.0
UniRef100_Q9ZSP5 Subtilisin-like protease [Arabidopsis thaliana] 697 0.0
UniRef100_O82440 Subtilisin-like protease [Arabidopsis thaliana] 697 0.0
UniRef100_Q6WNU4 Subtilisin-like protease [Glycine max] 692 0.0
UniRef100_Q8GXU1 Putative subtilisin-like protease [Arabidopsis ... 681 0.0
UniRef100_Q6H733 Putative subtilisin-like proteinase AIR3 [Oryza... 645 0.0
UniRef100_Q69V48 Putative subtilisin-like proteinase [Oryza sativa] 603 e-171
UniRef100_Q8H4X8 Putative subtilisin-like serine protease AIR3 [... 598 e-169
UniRef100_Q9AX30 Subtilisin-like protease [Oryza sativa] 548 e-154
UniRef100_Q7XTY8 OSJNBa0019K04.9 protein [Oryza sativa] 548 e-154
UniRef100_O65351 Cucumisin-like serine protease [Arabidopsis tha... 545 e-153
UniRef100_Q8RWQ7 AT5g67360/K8K14_8 [Arabidopsis thaliana] 540 e-152
UniRef100_Q9ZUF6 Subtilisin-like serine protease, putative [Arab... 528 e-148
UniRef100_Q8S8B0 Subtilisin-like serine protease AIR3 [Arabidops... 528 e-148
UniRef100_Q39007 Subtilisin-like protease [Arabidopsis thaliana] 528 e-148
UniRef100_Q8L4T6 Putative subtilisin-like protease [Oryza sativa] 523 e-146
UniRef100_Q9FK76 Subtilisin-like protease [Arabidopsis thaliana] 513 e-144
UniRef100_Q75I27 Putaive subtilisin-like proteinase [Oryza sativa] 508 e-142
UniRef100_P93205 SBT2 protein [Lycopersicon esculentum] 503 e-141
UniRef100_P93204 SBT1 protein [Lycopersicon esculentum] 499 e-139
>UniRef100_Q9FJF3 Serine protease-like protein [Arabidopsis thaliana]
Length = 760
Score = 705 bits (1819), Expect = 0.0
Identities = 378/733 (51%), Positives = 492/733 (66%), Gaps = 38/733 (5%)
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRS 62
L S +GS ENAKEAI YSY + INGFAA+L+E EAA+IAK+P VVSVF +K KLHTT S
Sbjct: 53 LASFVGSHENAKEAIFYSYKRHINGFAAILDENEAAEIAKHPDVVSVFPNKGRKLHTTHS 112
Query: 63 WEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNI 119
W F+ L N + +S W K +GE+TII N+DTGVWPESKSFSD G G +PA+W+G
Sbjct: 113 WNFMLLAKNGVVHKSSLWNKAGYGEDTIIANLDTGVWPESKSFSDEGYGAVPARWKG--- 169
Query: 120 CQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPR--SQQTARDFVGHGTHTLSTAGG 177
K VPCNRKLIGAR+FNK Y G LP S +T RD GHG+HTLSTA G
Sbjct: 170 ------RCHKDVPCNRKLIGARYFNKGYLAYTG-LPSNASYETCRDHDGHGSHTLSTAAG 222
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
NFVPGA++F IGNGT GGSP+ARVA YKVCW D CF AD+L+AI+ AI+DGVD++
Sbjct: 223 NFVPGANVFGIGNGTASGGSPKARVAAYKVCWPPVDGAECFDADILAAIEAAIEDGVDVL 282
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
S S GG ++ + +D I+IG+FHA+ + +V SAGN GP G+V NVAPWV TV A
Sbjct: 283 SASVGG----DAGDYMSDGIAIGSFHAVKNGVTVVCSAGNSGPKSGTVSNVAPWVITVGA 338
Query: 298 STLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
S++DR+F + + + N ++ G SL LP + ++++++ DA +AN DA C+ +L
Sbjct: 339 SSMDREFQAFVELKNGQSFKGTSLSKPLPEEKMYSLISAADANVANGNVTDALLCKKGSL 398
Query: 357 DPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL--STI 414
DP KV GKI+ C R G V +G +A +AGA G++L N + +G ++S+ HVL S I
Sbjct: 399 DPKKVKGKILVCLR-GDNARVDKGMQAAAAGAAGMVLCND-KASGNEIISDAHVLPASQI 456
Query: 415 SYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSIL 474
Y G +L S K +P TLN KPAP MAS+SSRGPN + P IL
Sbjct: 457 DY------KDGETLFSYLSSTKDPKGYIKAPTATLN-TKPAPFMASFSSRGPNTITPGIL 509
Query: 475 KPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPN 534
KPD+TAPGVNI+AA++ ++L +D RR PFN GTSMSCPH++G GL+KTLHP+
Sbjct: 510 KPDITAPGVNIIAAFTEATGPTDLDSDNRR-TPFNTESGTSMSCPHISGVVGLLKTLHPH 568
Query: 535 WSPAAIKSAIMTTATTRDNTNKP-ISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGI 593
WSPAAI+SAIMTT+ TR+N KP + ++F K ANPF+YGSGH++PN A PGLVYDL
Sbjct: 569 WSPAAIRSAIMTTSRTRNNRRKPMVDESFKK--ANPFSYGSGHVQPNKAAHPGLVYDLTT 626
Query: 594 KDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTN 653
DYL+FLCA GYN ++ + +TC +++ D NYPSIT+PNL S+TVTR + N
Sbjct: 627 GDYLDFLCAVGYNNTVVQLFAEDPQYTCRQGANLLDFNYPSITVPNL-TGSITVTRKLKN 685
Query: 654 VGPPSTYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWT 712
VGPP+TY A+ + G +++V P L F K GE K FQ+ ++ VTP Y FGEL WT
Sbjct: 686 VGPPATYNARFREPLGVRVSVEPKQLTFNKTGEVKIFQMTLRPLPVTP-SGYVFGELTWT 744
Query: 713 NGKHIVRSPVTVR 725
+ H VRSP+ V+
Sbjct: 745 DSHHYVRSPIVVQ 757
>UniRef100_Q9ZSP5 Subtilisin-like protease [Arabidopsis thaliana]
Length = 772
Score = 697 bits (1800), Expect = 0.0
Identities = 383/732 (52%), Positives = 489/732 (66%), Gaps = 26/732 (3%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
D LGS GS+E A +AI YSY K INGFAA L+ + A +I+K+P+VVSVF +K KLHTT
Sbjct: 59 DFLGSFTGSRERATDAIFYSYTKHINGFAAHLDHDLAYEISKHPEVVSVFPNKALKLHTT 118
Query: 61 RSWEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+FLGL N +S W+K RFGE+TII N+DTGVWPESKSF D G+GPIP++W+G
Sbjct: 119 RSWDFLGLEHNSYVPSSSIWRKARFGEDTIIANLDTGVWPESKSFRDEGLGPIPSRWKG- 177
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGG 177
ICQ K T CNRKLIGAR+FNK Y G L S + RD GHG+HTLSTA G
Sbjct: 178 -ICQNQKDATFH---CNRKLIGARYFNKGYAAAVGHLNSSFDSPRDLDGHGSHTLSTAAG 233
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
+FVPG SIF GNGT KGGSPRARVA YKVCW C+ ADVL+A D AI DG D+I
Sbjct: 234 DFVPGVSIFGQGNGTAKGGSPRARVAAYKVCWPPVKGNECYDADVLAAFDAAIHDGADVI 293
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
SVS GG ++ F D ++IG+FHA + I++V SAGN GP +V NVAPW TV A
Sbjct: 294 SVSLGGEPTS----FFNDSVAIGSFHAAKKRIVVVCSAGNSGPADSTVSNVAPWQITVGA 349
Query: 298 STLDRDFSSVMTIGN-KTLTGASL-FVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRT 355
ST+DR+F+S + +GN K G SL LP + + I+ S +AK NA+ DA+ C+ +
Sbjct: 350 STMDREFASNLVLGNGKHYKGQSLSSTALPHAKFYPIMASVNAKAKNASALDAQLCKLGS 409
Query: 356 LDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTIS 415
LDP K GKI+ C R G+ V +G+ G G++L N + G LL++PHVL
Sbjct: 410 LDPIKTKGKILVCLR-GQNGRVEKGRAVALGGGIGMVLEN-TYVTGNDLLADPHVLPATQ 467
Query: 416 YPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILK 475
S R + I ++P++T KPAPVMAS+SS+GP+ V P ILK
Sbjct: 468 LTSKDSFAVSRYISQTKKPI-----AHITPSRTDLGLKPAPVMASFSSKGPSIVAPQILK 522
Query: 476 PDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNW 535
PD+TAPGV+++AAY+ S +N D RR FN + GTSMSCPH++G AGL+KT +P+W
Sbjct: 523 PDITAPGVSVIAAYTGAVSPTNEQFDPRR-LLFNAISGTSMSCPHISGIAGLLKTRYPSW 581
Query: 536 SPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKD 595
SPAAI+SAIMTTAT D+ PI +A + A PF++G+GH++PN A++PGLVYDLGIKD
Sbjct: 582 SPAAIRSAIMTTATIMDDIPGPIQNATNMK-ATPFSFGAGHVQPNLAVNPGLVYDLGIKD 640
Query: 596 YLNFLCASGYNQQLISALNFNMTFTCSGTS-SIDDLNYPSITLPNLGLNSVTVTRTVTNV 654
YLNFLC+ GYN IS + N FTCS S+ +LNYPSIT+PNL + VTV+RTV NV
Sbjct: 641 YLNFLCSLGYNASQISVFSGN-NFTCSSPKISLVNLNYPSITVPNLTSSKVTVSRTVKNV 699
Query: 655 GPPSTYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTN 713
G PS Y KV G +AV P+SLNF K+GE+KTF+VI+ + + Y FGEL W++
Sbjct: 700 GRPSMYTVKVNNPQGVYVAVKPTSLNFTKVGEQKTFKVILVKSKGNVAKGYVFGELVWSD 759
Query: 714 GKHIVRSPVTVR 725
KH VRSP+ V+
Sbjct: 760 KKHRVRSPIVVK 771
>UniRef100_O82440 Subtilisin-like protease [Arabidopsis thaliana]
Length = 758
Score = 697 bits (1799), Expect = 0.0
Identities = 383/732 (52%), Positives = 489/732 (66%), Gaps = 26/732 (3%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
D LGS GS+E A +AI YSY K INGFAA L+ + A +I+K+P+VVSVF +K KLHTT
Sbjct: 45 DFLGSFTGSRERATDAIFYSYTKHINGFAAHLDHDLAYEISKHPEVVSVFPNKALKLHTT 104
Query: 61 RSWEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+FLGL N +S W+K RFGE+TII N+DTGVWPESKSF D G+GPIP++W+G
Sbjct: 105 RSWDFLGLEHNSYVPSSSIWRKARFGEDTIIANLDTGVWPESKSFRDEGLGPIPSRWKG- 163
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGG 177
ICQ K T CNRKLIGAR+FNK Y G L S + RD GHG+HTLSTA G
Sbjct: 164 -ICQNQKDATFH---CNRKLIGARYFNKGYAAAVGHLNSSFDSPRDLDGHGSHTLSTAAG 219
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
+FVPG SIF GNGT KGGSPRARVA YKVCW C+ ADVL+A D AI DG D+I
Sbjct: 220 DFVPGVSIFGQGNGTAKGGSPRARVAAYKVCWPPVKGNECYDADVLAAFDAAIHDGADVI 279
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
SVS GG ++ F D ++IG+FHA + I++V SAGN GP +V NVAPW TV A
Sbjct: 280 SVSLGGEPTS----FFNDSVAIGSFHAAKKRIVVVCSAGNSGPADSTVSNVAPWQITVGA 335
Query: 298 STLDRDFSSVMTIGN-KTLTGASL-FVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRT 355
ST+DR+F+S + +GN K G SL LP + + I+ S +AK NA+ DA+ C+ +
Sbjct: 336 STMDREFASNLVLGNGKHYKGQSLSSTALPHAKFYPIMASVNAKAKNASALDAQLCKLGS 395
Query: 356 LDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTIS 415
LDP K GKI+ C R G+ V +G+ G G++L N + G LL++PHVL +
Sbjct: 396 LDPIKTKGKILVCLR-GQNGRVEKGRAVALGGGIGMVLEN-TYVTGNDLLADPHVLPSTQ 453
Query: 416 YPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILK 475
S R + I ++P++T KPAPVMAS+SS+GP+ V P ILK
Sbjct: 454 LTSKDSFAVSRYMTQTKKPI-----AHITPSRTDLGLKPAPVMASFSSKGPSIVAPQILK 508
Query: 476 PDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNW 535
PD+TAPGV+++AAY+ S +N D RR FN + GTSMSCPH++G AGL+KT +P+W
Sbjct: 509 PDITAPGVSVIAAYTGAVSPTNEQFDPRR-LLFNAISGTSMSCPHISGIAGLLKTRYPSW 567
Query: 536 SPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKD 595
SPAAI+SAIMTTATT D+ PI +A + A PF++G+GH++PN A++PGLVYDLGIKD
Sbjct: 568 SPAAIRSAIMTTATTMDDIPGPIQNATNMK-ATPFSFGAGHVQPNLAVNPGLVYDLGIKD 626
Query: 596 YLNFLCASGYNQQLISALNFNMTFTCSGTS-SIDDLNYPSITLPNLGLNSVTVTRTVTNV 654
YLNFLC+ GYN IS + N FTCS S+ +LNYPSIT+PNL + VTV+RTV NV
Sbjct: 627 YLNFLCSLGYNASQISVFSGN-NFTCSSPKISLVNLNYPSITVPNLTSSKVTVSRTVKNV 685
Query: 655 GPPSTYFAKVQLA-GYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTN 713
G PS Y KV G +A+ P+SLNF K+GE KTF+VI+ + + Y FGEL W+
Sbjct: 686 GRPSMYTVKVNNPHGVYVALKPTSLNFTKVGELKTFKVILVKSKGNVAKGYMFGELVWSA 745
Query: 714 GKHIVRSPVTVR 725
KH VRSP+ V+
Sbjct: 746 KKHRVRSPIVVK 757
>UniRef100_Q6WNU4 Subtilisin-like protease [Glycine max]
Length = 773
Score = 692 bits (1785), Expect = 0.0
Identities = 376/730 (51%), Positives = 480/730 (65%), Gaps = 25/730 (3%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
D LGS LGS AK++I YSY + INGFAA L+EE A +IAK+PKV+SVF ++ KLHTT
Sbjct: 58 DFLGSFLGSSNTAKDSIFYSYTRHINGFAATLDEEVAVEIAKHPKVLSVFENRGRKLHTT 117
Query: 61 RSWEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+F+ L N + +S W+K RFGE IIGN+DTGVWPESKSFS++G+GPIP+KWRG
Sbjct: 118 RSWDFMELEHNGVIQSSSIWKKARFGEGVIIGNLDTGVWPESKSFSEQGLGPIPSKWRG- 176
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGG 177
IC +T CNRKLIGAR+FNK Y G L S + RD GHGTHTLSTAGG
Sbjct: 177 -ICDNGIDHTFH---CNRKLIGARYFNKGYASVAGPLNSSFDSPRDNEGHGTHTLSTAGG 232
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
N V S+F G GT KGGSP ARVA YKVCW CF AD+L+A D AI DGVD++
Sbjct: 233 NMVARVSVFGQGQGTAKGGSPMARVAAYKVCWPPVGGEECFDADILAAFDLAIHDGVDVL 292
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
SVS GG SST F D ++IG+FHA R +++V SAGN GP + N+APW TVAA
Sbjct: 293 SVSLGGSSST----FFKDSVAIGSFHAAKRGVVVVCSAGNSGPAEATAENLAPWHVTVAA 348
Query: 298 STLDRDFSSVMTIGNK-TLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
ST+DR F + + +GN T G SL ++ + I+ +TDAKLA+A DA C+ TL
Sbjct: 349 STMDRQFPTYVVLGNDITFKGESLSATKLAHKFYPIIKATDAKLASARAEDAVLCQNGTL 408
Query: 357 DPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISY 416
DP+K GKIV C R G V +G++A AGA G++L N + G ++++PHVL
Sbjct: 409 DPNKAKGKIVVCLR-GINARVDKGEQAFLAGAVGMVLAND-KTTGNEIIADPHVL----- 461
Query: 417 PGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKP 476
P +H T S + ++ KT KPAP MA++SS+GPN + P ILKP
Sbjct: 462 PASHINFTDGSAVFNYINSTKFPVAYITHPKTQLDTKPAPFMAAFSSKGPNTMVPEILKP 521
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWS 536
D+TAPGV+++AAY+ +N + D RR PFN + GTSMSCPHV+G GL++ L+P WS
Sbjct: 522 DITAPGVSVIAAYTEAQGPTNQVFDKRR-IPFNSVSGTSMSCPHVSGIVGLLRALYPTWS 580
Query: 537 PAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDY 596
AAIKSAIMTTATT DN +P+ +A D A PF+YG+GH++PN AMDPGLVYD+ I DY
Sbjct: 581 TAAIKSAIMTTATTLDNEVEPLLNATDGK-ATPFSYGAGHVQPNRAMDPGLVYDITIDDY 639
Query: 597 LNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGP 656
LNFLCA GYN+ IS + C S+ +LNYPSIT+P L SVTVTRT+ NVG
Sbjct: 640 LNFLCALGYNETQISVFT-EGPYKCRKKFSLLNLNYPSITVPKLS-GSVTVTRTLKNVGS 697
Query: 657 PSTYFAKVQLA-GYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGK 715
P TY A VQ G ++V PS L FK +GE+K+F++ +A Y FG+L W++GK
Sbjct: 698 PGTYIAHVQNPYGITVSVKPSILKFKNVGEEKSFKLTFKAMQGKATNNYAFGKLIWSDGK 757
Query: 716 HIVRSPVTVR 725
H V SP+ V+
Sbjct: 758 HYVTSPIVVK 767
>UniRef100_Q8GXU1 Putative subtilisin-like protease [Arabidopsis thaliana]
Length = 756
Score = 681 bits (1758), Expect = 0.0
Identities = 365/708 (51%), Positives = 476/708 (66%), Gaps = 37/708 (5%)
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRS 62
L S +GS ENAKEAI YSY + INGFAA+L+E EAA+IAK+P VVSVF +K KLHTT S
Sbjct: 71 LASFVGSHENAKEAIFYSYKRHINGFAAILDENEAAEIAKHPDVVSVFPNKGRKLHTTHS 130
Query: 63 WEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNI 119
W F+ L N + +S W K +GE+TII N+DTGVWPESKSFSD G G +PA+W+G
Sbjct: 131 WNFMLLAKNGVVHKSSLWNKAGYGEDTIIANLDTGVWPESKSFSDEGYGAVPARWKG--- 187
Query: 120 CQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPR--SQQTARDFVGHGTHTLSTAGG 177
K VPCNRKLIGAR+FNK Y G LP S +T RD GHG+HTLSTA G
Sbjct: 188 ------RCHKDVPCNRKLIGARYFNKGYLAYTG-LPSNASYETCRDHDGHGSHTLSTAAG 240
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
NFVPGA++F IGNGT GGSP+ARVA YKVCW D CF AD+L+AI+ AI+DGVD++
Sbjct: 241 NFVPGANVFGIGNGTASGGSPKARVAAYKVCWPPVDGAECFDADILAAIEAAIEDGVDVL 300
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
S S GG ++ + +D I+IG+FHA+ + +V SAGN GP G+V NVAPWV TV A
Sbjct: 301 SASVGG----DAGDYMSDGIAIGSFHAVKNGVTVVCSAGNSGPKSGTVSNVAPWVITVGA 356
Query: 298 STLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
S++DR+F + + + N ++ G SL LP + ++++++ DA +AN DA C+ +L
Sbjct: 357 SSMDREFQAFVELKNGQSFKGTSLSKPLPEEKMYSLISAADANVANGNVTDALLCKKGSL 416
Query: 357 DPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL--STI 414
DP KV GKI+ C R G V +G +A +AGA G++L N + +G ++S+ HVL S I
Sbjct: 417 DPKKVKGKILVCLR-GDNARVDKGMQAAAAGAAGMVLCND-KASGNEIISDAHVLPASQI 474
Query: 415 SYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSIL 474
Y G +L S K +P TLN KPAP MAS+SSRGPN + P IL
Sbjct: 475 DY------KDGETLFSYLSSTKDPKGYIKAPTATLN-TKPAPFMASFSSRGPNTITPGIL 527
Query: 475 KPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPN 534
KPD+TAPGVNI+AA++ ++L +D RR PFN GTSMSCPH++G GL+KTLHP+
Sbjct: 528 KPDITAPGVNIIAAFTEATGPTDLDSDNRR-TPFNTESGTSMSCPHISGVVGLLKTLHPH 586
Query: 535 WSPAAIKSAIMTTATTRDNTNKP-ISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGI 593
WSPAAI+SAIMTT+ TR+N KP + ++F K ANPF+YGSGH++PN A PGLVYDL
Sbjct: 587 WSPAAIRSAIMTTSRTRNNRRKPMVDESFKK--ANPFSYGSGHVQPNKAAHPGLVYDLTT 644
Query: 594 KDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTN 653
DYL+FLCA GYN ++ + +TC +++ D NYPSIT+PNL S+TVTR + N
Sbjct: 645 GDYLDFLCAVGYNNTVVQLFAEDPQYTCRQGANLLDFNYPSITVPNL-TGSITVTRKLKN 703
Query: 654 VGPPSTYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTP 700
VGPP+TY A+ + G +++V P L F K GE K FQ+ ++ VTP
Sbjct: 704 VGPPATYNARFREPLGVRVSVEPKQLTFNKTGEVKIFQMTLRPLPVTP 751
>UniRef100_Q6H733 Putative subtilisin-like proteinase AIR3 [Oryza sativa]
Length = 799
Score = 645 bits (1663), Expect = 0.0
Identities = 364/742 (49%), Positives = 469/742 (63%), Gaps = 42/742 (5%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+LL +LG KE A+EAI YSY + INGFAA L+ AA+IA+ P VVSVF ++ HKLHTT
Sbjct: 76 ELLAGVLGDKEKAREAIFYSYTRHINGFAANLDAAAAAKIAEKPGVVSVFPNRGHKLHTT 135
Query: 61 RSWEFLGLRG---NDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+FLGL G +AW+K RFGE+TIIGN+DTGVWPES+SF D G+GPIP+ WRG
Sbjct: 136 RSWQFLGLAGVGGAPTGAAWKKARFGEDTIIGNLDTGVWPESESFRDDGLGPIPSWWRGE 195
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQ-QTARDFVGHGTHTLSTAG 176
CQ + CNRKLIGARFFNK Y G L S T RD GHGTHTLSTAG
Sbjct: 196 --CQKGQ---DDAFSCNRKLIGARFFNKGYASAVGNLNTSLFDTPRDTDGHGTHTLSTAG 250
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V GAS+F GNGT GGSP ARVA Y+VC++ + + CF AD+L+A D AI DGV +
Sbjct: 251 GAPVAGASVFGYGNGTASGGSPMARVAAYRVCYTPVNGSECFDADILAAFDAAIHDGVHV 310
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
+SVS GG ++ + F D ++IG+FHA+ I +V SAGN GP PG+V NVAPW+FT A
Sbjct: 311 LSVSLGG----DAGDYFADGLAIGSFHAVRHGIAVVCSAGNSGPAPGTVSNVAPWLFTAA 366
Query: 297 ASTLDRDFSSVMTIGNKTLTGASLFVNL--PPNQDFTIVTSTDAKLANATNRDARFCRPR 354
AST+DR+F + + + L G SL + P + F ++ S+ A N T +++ C
Sbjct: 367 ASTMDREFPAYVVFNDTKLKGQSLSASALSPASSSFPMIDSSLAASPNRTQNESQLCFLG 426
Query: 355 TLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTI 414
+LDP KV GKIV C R G V +G+ L AG G++L N G ++++ HVL
Sbjct: 427 SLDPEKVKGKIVVCLR-GVNPRVEKGEAVLEAGGAGMVLAND-VTTGNEIIADAHVL--- 481
Query: 415 SYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNR------RKPAPVMASYSSRGPNK 468
P H + + + + S K SPA T+ R KPAP MA++SS+GPN
Sbjct: 482 --PATHIKFSDGQI------LFSYLKNTKSPAGTITRPETRLGTKPAPFMAAFSSQGPNT 533
Query: 469 VQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLI 528
V P ILKPD+TAPGV+++AA++ ++ ++L D RR FN GTSMSCPHVAG GL+
Sbjct: 534 VTPGILKPDITAPGVSVVAAWTRASAPTDLAFDKRR-VAFNSESGTSMSCPHVAGVVGLL 592
Query: 529 KTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLV 588
+TL P+WSPAAI+SA+MTTA DN I ++ ANPF +G+GH+ P AM+PGLV
Sbjct: 593 RTLRPDWSPAAIRSALMTTAVEVDNERHAILNS-SFAAANPFGFGAGHVSPARAMNPGLV 651
Query: 589 YDLGIKDYLNFLCASGYNQQLISAL---NFNMTFTC-SGTSSIDDLNYPSITLPNLGLNS 644
YDL DYLNFLC+ YN +++ F C + + DLNYPSIT+ NL +S
Sbjct: 652 YDLAAVDYLNFLCSLSYNATVMAMFAGGGGAAPFRCPASPPKVQDLNYPSITVVNL-TSS 710
Query: 645 VTVTRTVTNVGPPSTYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRK 703
TV RTV NVG P Y A V AG ++ V P +L F GEKKTFQV + T+ +
Sbjct: 711 ATVRRTVKNVGKPGVYKAYVTSPAGVRVTVSPDTLPFLLKGEKKTFQVRFEVTNASLAMD 770
Query: 704 YQFGELRWTNGKHIVRSPVTVR 725
Y FG L WTNGK VRSP+ V+
Sbjct: 771 YSFGALVWTNGKQFVRSPLVVK 792
>UniRef100_Q69V48 Putative subtilisin-like proteinase [Oryza sativa]
Length = 790
Score = 603 bits (1556), Expect = e-171
Identities = 349/738 (47%), Positives = 459/738 (61%), Gaps = 33/738 (4%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLLGS+LG +E A++AI Y Y K INGFAA LE EEAA +A+ P VVSVF + ++HTT
Sbjct: 69 DLLGSVLGDREKARDAIFYLYTKNINGFAARLEAEEAAAVAERPGVVSVFPDRGRRMHTT 128
Query: 61 RSWEFLGLRGNDIN----SAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRG 116
RSW+FLGL D + S W+ R+G+N IIGN+D+GVWPES SF+DR +GPIP W+G
Sbjct: 129 RSWQFLGLERPDGSVPPWSPWEAARYGQNIIIGNLDSGVWPESLSFNDRELGPIPNYWKG 188
Query: 117 GNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNG-KLPRSQQTARDFVGHGTHTLSTA 175
C+ + T K CN KLIGAR+FN Y K G L + +T RD GHGTHTL+TA
Sbjct: 189 A--CRNEHDKTFK---CNSKLIGARYFNNGYAKVIGVPLNDTHKTPRDANGHGTHTLATA 243
Query: 176 GGNFVPGASIFNIGNGTIKGGSPRARVATYKVCW-SLTDATSCFGADVLSAIDQAIDDGV 234
GG+ V GA F +G GT +GGSPRARVA Y+VC+ + +C+ +D+L+A + AI DGV
Sbjct: 244 GGSAVRGAEAFGLGGGTARGGSPRARVAAYRVCYPPFNGSDACYDSDILAAFEAAIADGV 303
Query: 235 DIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFT 294
+IS S G + + D I+IGA HA+ I +V SA N GP PG+V NVAPW+ T
Sbjct: 304 HVISASVG----ADPNDYLEDAIAIGALHAVKAGITVVCSASNFGPDPGTVTNVAPWILT 359
Query: 295 VAASTLDRDFSSVMTIGNKTLTGASLFVN-LPPNQDFTIVTSTDAKLANATNRDARFCRP 353
VAAST+DR F + + + G SL L +T++++ +A + DA C
Sbjct: 360 VAASTMDRAFPAHLVFNRNRVEGQSLSPTWLRGKTFYTMISAANAAVPGYPPADALLCEL 419
Query: 354 RTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLST 413
LD KV GKIV C R G + V +G+E AG +IL N E +G ++++ HVL
Sbjct: 420 GALDGKKVMGKIVVCMRGGNPR-VEKGEEVSRAGGAAMILVND-EASGNDVIADAHVLPA 477
Query: 414 ISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSI 473
+ NH+ G +L + K G K ++ AKT+ KPAPVMA++SS+GPN V P I
Sbjct: 478 VHI--NHA--DGHALLAYINSTK-GAKAFITRAKTVVGVKPAPVMAAFSSQGPNTVNPEI 532
Query: 474 LKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHP 533
LKPDVTAPGV+++AA+S A + L D RR FN GTSMSCP V+G AGLIKTLHP
Sbjct: 533 LKPDVTAPGVSVIAAWSGAAGPTGLPYDQRR-VAFNAQSGTSMSCPQVSGVAGLIKTLHP 591
Query: 534 NWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGI 593
+WSPAAIKSAIMTTAT N +PI ++ + A PF+ G+GH+ P+ AMDPGLVYDL +
Sbjct: 592 DWSPAAIKSAIMTTATELGNDMRPIMNS-SMSPATPFSCGAGHVFPHRAMDPGLVYDLTV 650
Query: 594 KDYLNFLCASGYNQQLISALNFNMTFTCSGTSSID--DLNYPSITLPNLGLNS--VTVTR 649
D+L+FLC GYN ++ N F C +D D NYPSIT +L T R
Sbjct: 651 DDHLSFLCTIGYNATALALFN-GAPFRCP-DDPLDPLDFNYPSITAFDLAPAGPPATARR 708
Query: 650 TVTNVGPPSTYFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFG 707
V NVGPP+TY A V + G ++ V P++L F+ GE +TF V P Y FG
Sbjct: 709 RVRNVGPPATYTAAVVREPEGVQVTVTPTTLTFESTGEVRTFWVKFAVRDPAPAANYAFG 768
Query: 708 ELRWTNGKHIVRSPVTVR 725
+ W++G H VRSP+ V+
Sbjct: 769 AIVWSDGNHQVRSPIVVK 786
>UniRef100_Q8H4X8 Putative subtilisin-like serine protease AIR3 [Oryza sativa]
Length = 762
Score = 598 bits (1543), Expect = e-169
Identities = 346/735 (47%), Positives = 468/735 (63%), Gaps = 33/735 (4%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLLGS+LGSK+ AK+AI+YSY K INGFAA LEEE A QIA++P VV+V S KLHTT
Sbjct: 46 DLLGSVLGSKQLAKDAILYSYTKNINGFAAHLEEEVATQIARHPDVVTVMASTMLKLHTT 105
Query: 61 RSWEFLGL-RGNDI--NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRG-IGPIPAKWRG 116
RSW+F+ + R I +S W+ GRFG++ II N+D+GVWPES SF+D +G +P +W+G
Sbjct: 106 RSWDFMDMERDGQILPDSIWKHGRFGQDVIIANLDSGVWPESNSFTDEEVVGEVPKRWKG 165
Query: 117 GNICQLDKLNTSK-KVPCNRKLIGARFFNKAYQKRN-GKLPRSQQTARDFVGHGTHTLST 174
C +T+K V CN+KLIGAR+FNK N G + +RD GHGTHTLST
Sbjct: 166 S--CS----DTAKYGVSCNKKLIGARYFNKDMLLSNPGAV--DGNWSRDTEGHGTHTLST 217
Query: 175 AGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGV 234
AGG FVP AS+F NGT KGG+PRARVA YKVCWS C ADVL+ + AI DG
Sbjct: 218 AGGRFVPRASLFGYANGTAKGGAPRARVAAYKVCWS----GECAAADVLAGFEAAIHDGA 273
Query: 235 DIISVSAGGPSSTNSEEIFTDE-ISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVF 293
D+ISVS G + + F E +++G+ HA + +V SAGN GP +VVN APWV
Sbjct: 274 DVISVSFGQDAPVATVASFLQEPVTLGSLHAAMNGVSVVCSAGNSGPLEDTVVNAAPWVT 333
Query: 294 TVAASTLDRDFSSVMTIGNKT-LTGASL-FVNLPPNQDFTIVTSTDAKLANATNRDARFC 351
TVAAST+DRDF +V+T+GN +TG SL L Q ++++ ++DA LA++ A C
Sbjct: 334 TVAASTVDRDFPNVVTLGNNAHMTGMSLETTTLHSTQLYSMIKASDAALASSDPAVASTC 393
Query: 352 RPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL 411
P TLDP KV KIV C R G I V +G L+AG G+IL N E++G ++++PHVL
Sbjct: 394 PPGTLDPEKVKNKIVVCVRGGDIPRVTKGMTVLNAGGTGMILAN-GEMDGDDIVADPHVL 452
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQP 471
+ + + + +D + + + +SP+KT K +P +A++SSRGP+ P
Sbjct: 453 PATMITYSEAMSLYKYMDSSKNPVAN-----ISPSKTEVGVKNSPSVAAFSSRGPSGTLP 507
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
+LKPD+ APGV+ILAA++ + S + + D RR + ++ GTSM+CPH++G GL+K
Sbjct: 508 CVLKPDIAAPGVDILAAFTEYVSPTEVPNDERRS-EYAILSGTSMACPHISGVIGLLKAA 566
Query: 532 HPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDL 591
P WSPAA++SAIMTTA T+DNT P+ D D A FA+G+G+I PN A+DPGLVYDL
Sbjct: 567 RPEWSPAAMRSAIMTTARTQDNTGAPMRD-HDGREATAFAFGAGNIHPNRAVDPGLVYDL 625
Query: 592 GIKDYLNFLCASGYNQQLISALNFNMTFTC-SGTSSIDDLNYPSITLPNLGLNSVTVTRT 650
+DY FLC+ G+N ++ L+ FTC ++DLNYPSI +P L S TV R
Sbjct: 626 SKEDYFVFLCSMGFNSSDLAKLSAG-NFTCPEKVPPMEDLNYPSIVVPALRHTS-TVARR 683
Query: 651 VTNVGPPSTYFAKVQLA-GYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGEL 709
+ VG P+TY A + G + V P++L F K GE K F+V ++ + Y FG L
Sbjct: 684 LKCVGRPATYRATWRAPYGVNMTVEPAALEFGKDGEVKEFKVTFKSEKDKLGKGYVFGRL 743
Query: 710 RWTNGKHIVRSPVTV 724
W++G H VRSPV V
Sbjct: 744 VWSDGTHHVRSPVVV 758
>UniRef100_Q9AX30 Subtilisin-like protease [Oryza sativa]
Length = 778
Score = 548 bits (1412), Expect = e-154
Identities = 335/749 (44%), Positives = 453/749 (59%), Gaps = 51/749 (6%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEH-KLHTT 60
LL S+ GS+E A+ +++YSY +NGFAA+L EEEA ++ +VVS F S HTT
Sbjct: 52 LLLSVKGSEEEARASLLYSYKHSLNGFAALLSEEEATALSARTEVVSAFPSNGRWSPHTT 111
Query: 61 RSWEFLGL----RGNDINSAWQKG--RFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKW 114
RSWEF+GL RG D G GE+ I+G +D+G+WPES+SF D G+GP+PA+W
Sbjct: 112 RSWEFVGLEEGVRGPDDTGRLPPGDKAGGEDVIVGVLDSGIWPESRSFGDEGLGPVPARW 171
Query: 115 RGGNICQL-DKLNTSKKVPCNRKLIGARFFNKAYQKRNGKL--PRSQQTARDFVGHGTHT 171
+G +CQ D + S CNRK+IGAR++ KAY+ R G + + ++ RD GHGTHT
Sbjct: 172 KG--VCQGGDSFSPSS---CNRKIIGARYYVKAYEARYGAVNTTNAYRSPRDHDGHGTHT 226
Query: 172 LSTAGGNFVPG-ASIFNIGNGTIKGGSPRARVATYKVCWSLTDAT-----SCFGADVLSA 225
ST G VPG A++ GT GG+P ARVA YKVCW + +CF AD+L+A
Sbjct: 227 ASTVAGRTVPGVAALGGFAPGTASGGAPLARVAVYKVCWPIPGPNPNIENTCFEADMLAA 286
Query: 226 IDQAIDDGVDIISVSAGGPSSTNSEEIFTDE-ISIGAFHALARNILLVASAGNEGPTPGS 284
ID A+ DGVD++SVS G ST F ++ I++GA HA R ++LV S GN GP P +
Sbjct: 287 IDDAVGDGVDVMSVSIG---STGKPLPFAEDGIAVGALHAAMRGVVLVCSGGNSGPKPAT 343
Query: 285 VVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLF-VNLPPNQDFTIVTSTDAKLAN 342
V N+APW+ TVAAS++DR F S + +GN + G ++ LP N+ + +V + DA +
Sbjct: 344 VSNLAPWMLTVAASSIDRAFISPIKLGNGMVIMGQTVTPYQLPGNKPYPLVYAADAVVPG 403
Query: 343 ATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGK 402
+ C P++L P KV GKIV C R ++ V +G E AG +IL N P G+
Sbjct: 404 TPANVSNQCLPKSLAPEKVRGKIVVCLRGTGLR-VEKGLEVKLAGGAAIILGNPPAFGGE 462
Query: 403 TLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYS 462
+ + HVL PG + + I + S + P++T+ KP+PVMA +S
Sbjct: 463 VPV-DAHVL-----PGTAVSSVDVNAIIRYINSSSSPTAVLDPSRTVVDVKPSPVMAQFS 516
Query: 463 SRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVA 522
SRGPN +P+ILKPDVTAPG+NILAA+S +S + L D R +N+M GTSMSCPHV+
Sbjct: 517 SRGPNVNEPNILKPDVTAPGLNILAAWSEASSPTKLDGDNRV-VKYNIMSGTSMSCPHVS 575
Query: 523 GTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSA 582
TA L+K+ HP WS AAI+SAIMTTATT + P+ DA D T+A P YGSGHIRP A
Sbjct: 576 ATAVLLKSAHPGWSSAAIRSAIMTTATTSNAEGGPMMDA-DGTVAGPIDYGSGHIRPKHA 634
Query: 583 MDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSID-DLNYPSITLPNLG 641
+DPGLVYD +DYL F CASG Q + + C T LN+PS+ + L
Sbjct: 635 LDPGLVYDASYQDYLLFACASGGAQ-------LDHSLPCPATPPPPYQLNHPSLAIHGLN 687
Query: 642 LNSVTVTRTVTNVGPPSTYF--AKVQLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVT 699
SVTV RTVTNVG S + A V+ G + V P SL+F + GEKK+F++ ++AT
Sbjct: 688 -GSVTVQRTVTNVGQGSARYSVAVVEPMGVSVKVSPRSLSFARTGEKKSFRIKIEATKGR 746
Query: 700 P--RRKYQF--GELRWTNGKHIVRSPVTV 724
R QF G W++G H+VRSP+ V
Sbjct: 747 GGWRVNGQFVAGSYTWSDGVHVVRSPLVV 775
>UniRef100_Q7XTY8 OSJNBa0019K04.9 protein [Oryza sativa]
Length = 776
Score = 548 bits (1411), Expect = e-154
Identities = 316/731 (43%), Positives = 446/731 (60%), Gaps = 46/731 (6%)
Query: 10 KENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLR 69
+++A IIY+Y +GFAA L+EEEA +A+ V++V +LHTTRS +FLG+
Sbjct: 70 EDDASTRIIYNYETAFHGFAAQLDEEEAELMAEADGVLAVIPETVLQLHTTRSPDFLGIG 129
Query: 70 GNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSK 129
N W + ++G +DTG+WPES SFSD+G+GP+PAKW+G +CQ + T+
Sbjct: 130 PEVSNRIWSDSLADHDVVVGVLDTGIWPESPSFSDKGLGPVPAKWKG--LCQTGRGFTTA 187
Query: 130 KVPCNRKLIGARFFNKAYQKRNGKLPRSQQ--TARDFVGHGTHTLSTAGGNFVPGASIFN 187
CNRK++GAR F Y+ +G + + + + RD GHGTHT +TA G+ V A+++
Sbjct: 188 N--CNRKIVGARIFYNGYEASSGPINETTELKSPRDQDGHGTHTAATAAGSPVQDANLYG 245
Query: 188 IGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSST 247
G +G +PRARVA YKVCW A CF +D+L+A+D+A+ DGVD++S+S GG +S
Sbjct: 246 YAGGVARGMAPRARVAAYKVCW----AGGCFSSDILAAVDRAVSDGVDVLSISLGGGAS- 300
Query: 248 NSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSV 307
+ D +SI +F A+ + + SAGN GP P S+ N++PW+ TV AST+DRDF +
Sbjct: 301 ---RYYLDSLSIASFGAMQMGVFVACSAGNAGPDPISLTNLSPWITTVGASTMDRDFPAT 357
Query: 308 MTIGN-KTLTGASLFV---NLPPNQDFTIVTSTDAKLANATNRDAR-FCRPRTLDPSKVN 362
+T+GN +TG SL+ NL P + + +V N++ D R C TL P V+
Sbjct: 358 VTLGNGANITGVSLYKGLRNLSPQEQYPVVYLG----GNSSMPDPRSLCLEGTLQPHDVS 413
Query: 363 GKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISY---PGN 419
GKIV CDR G V +GQ AG G+IL N NG+ L+++ H+L ++ G
Sbjct: 414 GKIVICDR-GISPRVQKGQVVKEAGGIGMILANTAA-NGEELVADSHLLPAVAVGEAEGI 471
Query: 420 HSRTTGRSLDIIPSDIK-SGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDV 478
+++ +S + + GTKL + +P+PV+A++SSRGPN + ILKPDV
Sbjct: 472 AAKSYSKSAPKPTATLSFGGTKLGI---------RPSPVVAAFSSRGPNILTLEILKPDV 522
Query: 479 TAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPA 538
APGVNILAA+S AS S+L +D+RR FN++ GTSMSCPHVAG A LIK HP+WSPA
Sbjct: 523 VAPGVNILAAWSGDASPSSLSSDSRR-VGFNILSGTSMSCPHVAGVAALIKASHPDWSPA 581
Query: 539 AIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLN 598
IKSA+MTTA DNT +P+ DA + PF +G+GHI P A+ PGLVYD+G DYL
Sbjct: 582 QIKSALMTTAYVHDNTYRPMKDAATGKASTPFEHGAGHIHPVRALTPGLVYDIGQADYLE 641
Query: 599 FLCASGYNQQLISALNFNMTFTCSGT-SSIDDLNYP--SITLPNLGLNSVTVTRTVTNVG 655
FLC + N TC T SS DLNYP S+ + ++TV RTVTNVG
Sbjct: 642 FLCTQHMTPMQLRTFTKNSNMTCRHTFSSASDLNYPAISVVFADQPSKALTVRRTVTNVG 701
Query: 656 PP-STYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTN 713
PP STY KV + G + V P++L+F +K +++V V T+ ++ +FG L W++
Sbjct: 702 PPSSTYHVKVTKFKGADVIVEPNTLHFVSTNQKLSYKVTV--TTKAAQKAPEFGALSWSD 759
Query: 714 GKHIVRSPVTV 724
G HIVRSPV +
Sbjct: 760 GVHIVRSPVVL 770
>UniRef100_O65351 Cucumisin-like serine protease [Arabidopsis thaliana]
Length = 757
Score = 545 bits (1404), Expect = e-153
Identities = 314/715 (43%), Positives = 436/715 (60%), Gaps = 32/715 (4%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L +EEA + P V+SV ++LHTTR+ FLGL + +
Sbjct: 65 LLYTYENAIHGFSTRLTQEEADSLMTQPGVISVLPEHRYELHTTRTPLFLGLDEHTADLF 124
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ G + + ++G +DTGVWPESKS+SD G GPIP+ W+GG C+ T+ CNRK
Sbjct: 125 PEAGSYSD-VVVGVLDTGVWPESKSYSDEGFGPIPSSWKGG--CEAGTNFTASL--CNRK 179
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF + Y+ G + S+++ RD GHGTHT STA G+ V GAS+ +GT +
Sbjct: 180 LIGARFFARGYESTMGPIDESKESRSPRDDDGHGTHTSSTAAGSVVEGASLLGYASGTAR 239
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G +PRARVA YKVCW CF +D+L+AID+AI D V+++S+S GG S + +
Sbjct: 240 GMAPRARVAVYKVCW----LGGCFSSDILAAIDKAIADNVNVLSMSLGGGMS----DYYR 291
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ R IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF ++ +GN K
Sbjct: 292 DGVAIGAFAAMERGILVSCSAGNAGPSSSSLSNVAPWITTVGAGTLDRDFPALAILGNGK 351
Query: 314 TLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
TG SLF P++ + + +A +NATN C TL P KV GKIV CDR G
Sbjct: 352 NFTGVSLFKGEALPDKLLPFIYAGNA--SNATN--GNLCMTGTLIPEKVKGKIVMCDR-G 406
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G+IL N NG+ L+++ H+L + + P
Sbjct: 407 INARVQKGDVVKAAGGVGMILANTAA-NGEELVADAHLLPATTVGEKAGDIIRHYVTTDP 465
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
+ S +S T+ KP+PV+A++SSRGPN + P+ILKPD+ APGVNILAA++
Sbjct: 466 NPTAS-----ISILGTVVGVKPSPVVAAFSSRGPNSITPNILKPDLIAPGVNILAAWTGA 520
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 552
A + L +D+RR FN++ GTSMSCPHV+G A L+K++HP WSPAAI+SA+MTTA
Sbjct: 521 AGPTGLASDSRR-VEFNIISGTSMSCPHVSGLAALLKSVHPEWSPAAIRSALMTTAYKTY 579
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGY-NQQLIS 611
KP+ D + PF +G+GH+ P +A +PGL+YDL +DYL FLCA Y + Q+ S
Sbjct: 580 KDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYDLTTEDYLGFLCALNYTSPQIRS 639
Query: 612 ALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGY 669
N T S + S+ DLNYPS + G+ + TRTVT+VG TY KV + G
Sbjct: 640 VSRRNYTCDPSKSYSVADLNYPSFAVNVDGVGAYKYTRTVTSVGGAGTYSVKVTSETTGV 699
Query: 670 KIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
KI+V P+ LNFK+ EKK++ V S P FG + W++GKH+V SPV +
Sbjct: 700 KISVEPAVLNFKEANEKKSYTVTFTVDSSKPSGSNSFGSIEWSDGKHVVGSPVAI 754
>UniRef100_Q8RWQ7 AT5g67360/K8K14_8 [Arabidopsis thaliana]
Length = 757
Score = 540 bits (1391), Expect = e-152
Identities = 313/715 (43%), Positives = 435/715 (60%), Gaps = 32/715 (4%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L +EEA + P V+SV ++LHTTR+ FLGL + +
Sbjct: 65 LLYTYENAIHGFSTRLTQEEADSLMTQPGVISVLPEHRYELHTTRTPLFLGLDEHTADLF 124
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ G + + ++G +DTGVWPESKS+SD G GPIP+ W+GG C+ T+ CNRK
Sbjct: 125 PEAGSYSD-VVVGVLDTGVWPESKSYSDEGFGPIPSSWKGG--CEAGTNFTASL--CNRK 179
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF + Y+ G + S+++ RD GHGTHT STA G+ V GAS+ +GT +
Sbjct: 180 LIGARFFARGYESTMGPIDESKESRSPRDDDGHGTHTSSTAAGSVVEGASLLGYASGTAR 239
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G +PRARVA YKVCW CF +D+L+AID+AI D V+++S+S GG S + +
Sbjct: 240 GMAPRARVAVYKVCW----LGGCFSSDILAAIDKAIADNVNVLSMSLGGGMS----DYYR 291
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ R IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF ++ +GN K
Sbjct: 292 DGVAIGAFAAMERGILVSCSAGNAGPSSSSLSNVAPWITTVGAGTLDRDFPALAILGNGK 351
Query: 314 TLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
TG SLF P++ + + +A +NATN C TL P KV GKIV CDR G
Sbjct: 352 NFTGVSLFKGEALPDKLLPFIYAGNA--SNATN--GNLCMTGTLIPEKVKGKIVMCDR-G 406
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G+IL N NG+ L+++ H+L + + P
Sbjct: 407 INARVQKGDVVKAAGGVGMILANTAA-NGEELVADAHLLPATTVGEKAGDIIRHYVTTDP 465
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
+ S +S T+ KP+PV+A++SSRGPN + P+ILKPD+ APGVNILAA++
Sbjct: 466 NPTAS-----ISILGTVVGVKPSPVVAAFSSRGPNSITPNILKPDLIAPGVNILAAWTGA 520
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 552
A + L +D+RR FN++ GTSMSCPHV+G A L+K++HP SPAAI+SA+MTTA
Sbjct: 521 AGPTGLASDSRR-VEFNIISGTSMSCPHVSGLAALLKSVHPECSPAAIRSALMTTAYKTY 579
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGY-NQQLIS 611
KP+ D + PF +G+GH+ P +A +PGL+YDL +DYL FLCA Y + Q+ S
Sbjct: 580 KDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYDLTTEDYLGFLCALNYTSPQIRS 639
Query: 612 ALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGY 669
N T S + S+ DLNYPS + G+ + TRTVT+VG TY KV + G
Sbjct: 640 VSRRNYTCDPSKSYSVADLNYPSFAVNVDGVGAYKYTRTVTSVGGAGTYSVKVTSETTGV 699
Query: 670 KIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
KI+V P+ LNFK+ EKK++ V S P FG + W++GKH+V SPV +
Sbjct: 700 KISVEPAVLNFKEANEKKSYTVTFTVDSSKPSGSNSFGSIEWSDGKHVVGSPVAI 754
>UniRef100_Q9ZUF6 Subtilisin-like serine protease, putative [Arabidopsis thaliana]
Length = 754
Score = 528 bits (1360), Expect = e-148
Identities = 319/725 (44%), Positives = 431/725 (59%), Gaps = 40/725 (5%)
Query: 9 SKENAKEAIIYSYNKQINGFAAMLEEEEA-AQIAKNPKVVSVFLSKEHKLHTTRSWEFLG 67
S+ N++ +++Y+Y +GF+A L+ EA + ++ + ++ +F + LHTTR+ EFLG
Sbjct: 52 SQLNSESSLLYTYTTSFHGFSAYLDSTEADSLLSSSNSILDIFEDPLYTLHTTRTPEFLG 111
Query: 68 LRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNT 127
L N G IIG +DTGVWPES+SF D + IP+KW+G C+
Sbjct: 112 L--NSEFGVHDLGSSSNGVIIGVLDTGVWPESRSFDDTDMPEIPSKWKGE--CESGSDFD 167
Query: 128 SKKVPCNRKLIGARFFNKAYQKRNG---KLPRSQQTARDFVGHGTHTLSTAGGNFVPGAS 184
SK CN+KLIGAR F+K +Q +G R + RD GHGTHT +TA G+ V AS
Sbjct: 168 SKL--CNKKLIGARSFSKGFQMASGGGFSSKRESVSPRDVDGHGTHTSTTAAGSAVRNAS 225
Query: 185 IFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGP 244
GT +G + RARVATYKVCWS T CFG+D+L+A+D+AI DGVD++S+S GG
Sbjct: 226 FLGYAAGTARGMATRARVATYKVCWS----TGCFGSDILAAMDRAILDGVDVLSLSLGGG 281
Query: 245 SSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDF 304
S+ + D I+IGAF A+ R + + SAGN GPT SV NVAPWV TV A TLDRDF
Sbjct: 282 SAP----YYRDTIAIGAFSAMERGVFVSCSAGNSGPTRASVANVAPWVMTVGAGTLDRDF 337
Query: 305 SSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNG 363
+ +GN K LTG SL+ + + + + C P +LD S V G
Sbjct: 338 PAFANLGNGKRLTGVSLYSGVGMG-----TKPLELVYNKGNSSSSNLCLPGSLDSSIVRG 392
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
KIV CDR G V +G AG G+I+ N +G+ L+++ H+L I+ +
Sbjct: 393 KIVVCDR-GVNARVEKGAVVRDAGGLGMIMANTAA-SGEELVADSHLLPAIAV----GKK 446
Query: 424 TGRSL-DIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPG 482
TG L + + SD K T L + L+ KP+PV+A++SSRGPN V P ILKPDV PG
Sbjct: 447 TGDLLREYVKSDSKP-TALLVFKGTVLDV-KPSPVVAAFSSRGPNTVTPEILKPDVIGPG 504
Query: 483 VNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKS 542
VNILA +S + L D+RR FN+M GTSMSCPH++G AGL+K HP WSP+AIKS
Sbjct: 505 VNILAGWSDAIGPTGLDKDSRRT-QFNIMSGTSMSCPHISGLAGLLKAAHPEWSPSAIKS 563
Query: 543 AIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCA 602
A+MTTA DNTN P+ DA D +L+NP+A+GSGH+ P A+ PGLVYD+ ++Y+ FLC+
Sbjct: 564 ALMTTAYVLDNTNAPLHDAADNSLSNPYAHGSGHVDPQKALSPGLVYDISTEEYIRFLCS 623
Query: 603 SGYNQQLISALNFNMTFTCSGT-SSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYF 661
Y I A+ + CS S LNYPS ++ G V TR VTNVG S+ +
Sbjct: 624 LDYTVDHIVAIVKRPSVNCSKKFSDPGQLNYPSFSVLFGGKRVVRYTREVTNVGAASSVY 683
Query: 662 AKVQLAG---YKIAVVPSSLNFKKIGEKKTFQV-IVQATSVTPRRKYQFGELRWTNGKHI 717
KV + G I+V PS L+FK +GEKK + V V V+ K +FG + W+N +H
Sbjct: 684 -KVTVNGAPSVGISVKPSKLSFKSVGEKKRYTVTFVSKKGVSMTNKAEFGSITWSNPQHE 742
Query: 718 VRSPV 722
VRSPV
Sbjct: 743 VRSPV 747
>UniRef100_Q8S8B0 Subtilisin-like serine protease AIR3 [Arabidopsis thaliana]
Length = 578
Score = 528 bits (1360), Expect = e-148
Identities = 293/592 (49%), Positives = 382/592 (64%), Gaps = 17/592 (2%)
Query: 136 KLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKG 195
KLIGAR+FNK Y G L S + RD GHG+HTLSTA G+FVPG SIF GNGT KG
Sbjct: 1 KLIGARYFNKGYAAAVGHLNSSFDSPRDLDGHGSHTLSTAAGDFVPGVSIFGQGNGTAKG 60
Query: 196 GSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTD 255
GSPRARVA YKVCW C+ ADVL+A D AI DG D+ISVS GG ++ F D
Sbjct: 61 GSPRARVAAYKVCWPPVKGNECYDADVLAAFDAAIHDGADVISVSLGGEPTS----FFND 116
Query: 256 EISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKTL 315
++IG+FHA + I++V SAGN GP +V NVAPW TV AST+ +++ + +
Sbjct: 117 SVAIGSFHAAKKRIVVVCSAGNSGPADSTVSNVAPWQITVGASTMTVSLLAILFSVMENI 176
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
T S LP + + I+ S +AK NA+ DA+ C+ +LDP K GKI+ C R G+
Sbjct: 177 TSLS-STALPHAKFYPIMASVNAKAKNASALDAQLCKLGSLDPIKTKGKILVCLR-GQNG 234
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDI 435
V +G+ G G++L N + G LL++PHVL S R + I
Sbjct: 235 RVEKGRAVALGGGIGMVLEN-TYVTGNDLLADPHVLPATQLTSKDSFAVSRYISQTKKPI 293
Query: 436 KSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASA 495
++P++T KPAPVMAS+SS+GP+ V P ILKPD+TAPGV+++AAY+ S
Sbjct: 294 -----AHITPSRTDLGLKPAPVMASFSSKGPSIVAPQILKPDITAPGVSVIAAYTGAVSP 348
Query: 496 SNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTN 555
+N D RR FN + GTSMSCPH++G AGL+KT +P+WSPAAI+SAIMTTAT D+
Sbjct: 349 TNEQFDPRR-LLFNAISGTSMSCPHISGIAGLLKTRYPSWSPAAIRSAIMTTATIMDDIP 407
Query: 556 KPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNF 615
PI +A + A PF++G+GH++PN A++PGLVYDLGIKDYLNFLC+ GYN IS +
Sbjct: 408 GPIQNATNMK-ATPFSFGAGHVQPNLAVNPGLVYDLGIKDYLNFLCSLGYNASQISVFSG 466
Query: 616 NMTFTCSGTS-SIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKV-QLAGYKIAV 673
N FTCS S+ +LNYPSIT+PNL + VTV+RTV NVG PS Y KV G +AV
Sbjct: 467 N-NFTCSSPKISLVNLNYPSITVPNLTSSKVTVSRTVKNVGRPSMYTVKVNNPQGVYVAV 525
Query: 674 VPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVR 725
P+SLNF K+GE+KTF+VI+ + + Y FGEL W++ KH VRSP+ V+
Sbjct: 526 KPTSLNFTKVGEQKTFKVILVKSKGNVAKGYVFGELVWSDKKHRVRSPIVVK 577
>UniRef100_Q39007 Subtilisin-like protease [Arabidopsis thaliana]
Length = 746
Score = 528 bits (1360), Expect = e-148
Identities = 309/715 (43%), Positives = 430/715 (59%), Gaps = 34/715 (4%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L +EEA + P V+SV ++LHTTR+ FLGL + +
Sbjct: 56 LLYTYENAIHGFSTRLTQEEADSLMTQPGVISVLPEHRYELHTTRTPLFLGLDEHTADLF 115
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ G + + ++G +DTGVWPESKS+SD G GPIP+ W+GG C+ T+ CNRK
Sbjct: 116 PEAGSYSD-VVVGVLDTGVWPESKSYSDEGFGPIPSSWKGG--CEAGTNFTASL--CNRK 170
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF + Y+ G + S+++ RD GHGTHT STA G+ V GAS+ +GT +
Sbjct: 171 LIGARFFARGYESTMGPIDESKESRSPRDDDGHGTHTSSTAAGSVVEGASLLGYASGTAR 230
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G +A YKVCW CF +D+L+AID+AI D V+++S+S GG S + +
Sbjct: 231 G--MLHALAVYKVCW----LGGCFSSDILAAIDKAIADNVNVLSMSLGGGMS----DYYR 280
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ R IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF ++ +GN K
Sbjct: 281 DGVAIGAFAAMERGILVSCSAGNAGPSSSSLSNVAPWITTVGAGTLDRDFPALAILGNGK 340
Query: 314 TLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
TG SLF P++ + + +A +NATN C TL P KV GKIV CDR G
Sbjct: 341 NFTGVSLFKGEALPDKLLPFIYAGNA--SNATN--GNLCMTGTLIPEKVKGKIVMCDR-G 395
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G+IL N NG+ L+++ H+L + + P
Sbjct: 396 INARVQKGDVVKAAGGVGMILANTAA-NGEELVADAHLLPATTVGEKAGDIIRHYVTTDP 454
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
+ S +S T+ KP+PV+A++SSRGPN + P+ILKPD+ APGVNILAA++
Sbjct: 455 NPTAS-----ISILGTVVGVKPSPVVAAFSSRGPNSITPNILKPDLIAPGVNILAAWTGA 509
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 552
A + L +D+RR FN++ GTSMSCPHV+G A L+K++HP WSPAAI+SA+MTTA
Sbjct: 510 AGPTGLASDSRR-VEFNIISGTSMSCPHVSGLAALLKSVHPEWSPAAIRSALMTTAYKTY 568
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGY-NQQLIS 611
KP+ D + PF +G+GH+ P +A +PGL+YDL +DYL FLCA Y + Q+ S
Sbjct: 569 KDGKPLLDIATGKPSTPFDHGAGHVSPTTATNPGLIYDLTTEDYLGFLCALNYTSPQIRS 628
Query: 612 ALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGY 669
N T S + S+ DLNYPS + G + TRTVT+VG TY KV + G
Sbjct: 629 VSRRNYTCDPSKSYSVADLNYPSFAVNVDGAGAYKYTRTVTSVGGAGTYSVKVTSETTGV 688
Query: 670 KIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
KI+V P+ LNFK+ EKK++ V S P FG + W++GKH+V SPV +
Sbjct: 689 KISVEPAVLNFKEANEKKSYTVTFTVDSSKPSGSNSFGSIEWSDGKHVVGSPVAI 743
>UniRef100_Q8L4T6 Putative subtilisin-like protease [Oryza sativa]
Length = 757
Score = 523 bits (1346), Expect = e-146
Identities = 322/746 (43%), Positives = 431/746 (57%), Gaps = 72/746 (9%)
Query: 25 INGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDI-NSAWQKG-RF 82
I ++E I + P V++V HK+HTTRSW+FL L N AW+ ++
Sbjct: 35 IQVIVVQIDESFVGVIKQLPGVLAVIPDVLHKVHTTRSWDFLELERNGAATGAWKDAAKY 94
Query: 83 GENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLD---KLN---TSKKVPCNR- 135
G + IIGN+DTGVWPES SF D G +P++WRG I D K N + +K+ C
Sbjct: 95 GVDAIIGNVDTGVWPESASFKDDGYS-VPSRWRGKCITGNDTTFKCNNKFSGEKLTCRTC 153
Query: 136 -KLIGARFFNKAYQKRN---GKLPRSQQ---TARDFVGHGTHTLSTAGGNFVPGASIFNI 188
KLIGA FFN + GK P T RD++GHGTHTLSTAGG FVP AS+F
Sbjct: 154 SKLIGAGFFNLGFLASGLLQGKPPSQAAELYTPRDYIGHGTHTLSTAGGGFVPDASVFGH 213
Query: 189 GNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTN 248
G GT KGGSP ARVA YK C+ A C +D+L+A+ A++DGV+++S+S GGP
Sbjct: 214 GKGTAKGGSPLARVAAYKACY----AEGCSSSDILAAMVTAVEDGVNVLSLSVGGP---- 265
Query: 249 SEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVM 308
+++ +D I+IGAF+A+ + +++V SA N GP PGSV NVAPW+ TV AST+DRDF + +
Sbjct: 266 ADDYLSDPIAIGAFYAVQKGVIVVCSASNSGPQPGSVTNVAPWILTVGASTMDRDFPAYV 325
Query: 309 TIG----NKTLTGASLF-VNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNG 363
T G + T+ G SL LP Q + ++ + +A AN + ++ C P +LD KV G
Sbjct: 326 TFGGVTSSMTIKGQSLSNSTLPQGQRYAMINAKNANAANVPSENSTLCFPGSLDSDKVRG 385
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLST--ISYPG--- 418
KIV C R G V +G AG G++L N NG+ ++++PH+++ +SY
Sbjct: 386 KIVVCTR-GVNARVEKGLVVKQAGGVGMVLCNYAG-NGEDVIADPHLIAAAHVSYSQCIN 443
Query: 419 --NHSRTTGRSLD-IIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILK 475
N+ +T + I SD + G KPAPVMA++SSRGPN + P ILK
Sbjct: 444 LFNYLGSTDNPVGYITASDARLGV-------------KPAPVMAAFSSRGPNPITPQILK 490
Query: 476 PDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNW 535
PD+TAPGV+++AAYS S + L D RR P+N+M GTSMSCPHV+G GLIKT +P+W
Sbjct: 491 PDITAPGVSVIAAYSEAVSPTELSFDDRR-VPYNIMSGTSMSCPHVSGIVGLIKTKYPDW 549
Query: 536 SPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKD 595
+PA IKSAIMTTA T DN + I D A PFAYGSGH+R A+DPGLVYD D
Sbjct: 550 TPAMIKSAIMTTAITGDNDSGKIRDE-TGAAATPFAYGSGHVRSVQALDPGLVYDTTSAD 608
Query: 596 YLNFLCASGYNQQLISALNF---NMTFTCSGTSSI---DDLNYPSITLPNLGLNSVTVTR 649
Y +FLCA Q + F CS + +DLNYPSI +P L S TV R
Sbjct: 609 YADFLCALRPTQNPLPLPVFGDDGKPRACSQGAQYGRPEDLNYPSIAVPCLS-GSATVRR 667
Query: 650 TVTNVG-PPSTYFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQF 706
V NVG P Y V LAG K+ V P L+F+ GE++ F V ++ Y F
Sbjct: 668 RVKNVGAAPCRYAVSVTEALAGVKVTVYPPELSFESYGEEREFTVRLEVQDAAAAANYVF 727
Query: 707 GELRWT-------NGKHIVRSPVTVR 725
G + W+ + KH VRSP+ +
Sbjct: 728 GSIEWSEESESDPDRKHRVRSPIVAK 753
>UniRef100_Q9FK76 Subtilisin-like protease [Arabidopsis thaliana]
Length = 791
Score = 513 bits (1322), Expect = e-144
Identities = 313/762 (41%), Positives = 439/762 (57%), Gaps = 64/762 (8%)
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLS--KEHKLHTT 60
L S+ S+E+A+ +++YSY INGFAA L ++A+++ K +VVSVF S ++++ HTT
Sbjct: 51 LQSVKESEEDARASLLYSYKHSINGFAAELTPDQASKLEKLAEVVSVFKSHPRKYEAHTT 110
Query: 61 RSWEFLGL-----------RGNDINSAWQKGR-------FGENTIIGNIDTGVWPESKSF 102
RSWEF+GL R ND + ++ GR G+ I+G +D+GVWPESKSF
Sbjct: 111 RSWEFVGLEEEETDSDVPRRKNDADDRFRVGRNFLKKAKHGDGIIVGVLDSGVWPESKSF 170
Query: 103 SDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQ--- 159
+D+G+GP+P W+G ICQ S CNRK+IGAR++ K Y++ G +
Sbjct: 171 NDKGMGPVPKSWKG--ICQTGVAFNSSH--CNRKIIGARYYVKGYERYYGAFNATANKDF 226
Query: 160 -TARDFVGHGTHTLSTAGGNFVPGASIFN-IGNGTIKGGSPRARVATYKVCWSLTDATS- 216
+ RD GHG+HT STA G V GAS G+ GG+P AR+A YK CW+ +A
Sbjct: 227 LSPRDPDGHGSHTASTAVGRRVLGASALGGFAKGSASGGAPLARLAIYKACWAKPNAEKV 286
Query: 217 ----CFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLV 272
C D+L+AID AI DGV +IS+S G +T D I++GA HA+ RNI++
Sbjct: 287 EGNICLEEDMLAAIDDAIADGVHVISISIG---TTEPFPFTQDGIAMGALHAVKRNIVVA 343
Query: 273 ASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKTLTGASLFVNLPPNQDFTI 332
ASAGN GP PG++ N+APW+ TV ASTLDR F + +GN ++ +
Sbjct: 344 ASAGNSGPKPGTLSNLAPWIITVGASTLDRAFVGGLVLGNGYTIKTDSITAFKMDKFAPL 403
Query: 333 VTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVI 392
V +++ + + C P +L P V+GK+V C R G + +G E AG G+I
Sbjct: 404 VYASNVVVPGIALNETSQCLPNSLKPELVSGKVVLCLR-GAGSRIGKGMEVKRAGGAGMI 462
Query: 393 LRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRR 452
L N NG + S+ H + T G + L+ I +D K + P KT+ +
Sbjct: 463 LGN-IAANGNEVPSDSHFVPT---AGVTPTVVDKILEYIKTD--KNPKAFIKPGKTVYKY 516
Query: 453 KPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQ 512
+ AP M +SSRGPN V P+ILKPD+TAPG+ ILAA+S S S + D +R +N+
Sbjct: 517 QAAPSMTGFSSRGPNVVDPNILKPDITAPGLYILAAWSGADSPSKMSVD-QRVAGYNIYS 575
Query: 513 GTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAY 572
GTSMSCPHVAG L+K +HP WS AAI+SA+MTTA ++ KPI D ANPFA
Sbjct: 576 GTSMSCPHVAGAIALLKAIHPKWSSAAIRSALMTTAWMTNDKKKPIQDTTGLP-ANPFAL 634
Query: 573 GSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTC-SGTSSIDDLN 631
GSGH RP A DPGLVYD + YL + C+ ++ N + TF C S + N
Sbjct: 635 GSGHFRPTKAADPGLVYDASYRAYLLYGCS-------VNITNIDPTFKCPSKIPPGYNHN 687
Query: 632 YPSITLPNLGLNSVTVTRTVTNVG---PPSTYFAKVQ-LAGYKIAVVPSSLNFKKIGEKK 687
YPSI +PNL +VTV RTVTNVG STY V+ +G + +P+ L+F +IG+K+
Sbjct: 688 YPSIAVPNL-KKTVTVKRTVTNVGTGNSTSTYLFSVKPPSGISVKAIPNILSFNRIGQKQ 746
Query: 688 TFQVIV-----QATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
F++++ Q + T + +YQFG WT+ H+VRSP+ V
Sbjct: 747 RFKIVIKPLKNQVMNATEKGQYQFGWFSWTDKVHVVRSPIAV 788
>UniRef100_Q75I27 Putaive subtilisin-like proteinase [Oryza sativa]
Length = 765
Score = 508 bits (1309), Expect = e-142
Identities = 303/732 (41%), Positives = 431/732 (58%), Gaps = 57/732 (7%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y+ ++GF+A L EA +A V++V ++LHTTR+ EFLG+ GND
Sbjct: 60 MLYAYDTVLHGFSARLTAREARDMAAMDGVLAVNPEARYELHTTRTPEFLGIAGND-GLF 118
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
Q G G+ ++G +DTGVWPES+S+ D G+G +P+ W+G + N+S CNRK
Sbjct: 119 PQSGTAGD-VVVGVLDTGVWPESRSYDDAGLGEVPSWWKGECMAGTG-FNSSA---CNRK 173
Query: 137 LIGARFFNKAYQKRNGKLP--RSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
L+GARFFN+ Y+ G + R ++ RD GHGTHT STA G V GAS+ +GT +
Sbjct: 174 LVGARFFNRGYEAAMGPMDTTRESRSPRDDDGHGTHTSSTAAGAAVSGASLLGFASGTAR 233
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G +PRARVA YKVCW CF +D+L+ +D A+ DG ++S+S GG ++ +
Sbjct: 234 GMAPRARVAVYKVCW----LGGCFSSDILAGMDAAVADGCGVLSLSLGGGAADYAR---- 285
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ +N+L+ SAGN GP ++ NVAPW+ TV A TLDRDF + +++GN K
Sbjct: 286 DSVAIGAFAAMEQNVLVSCSAGNAGPGTSTLSNVAPWITTVGAGTLDRDFPAYVSLGNGK 345
Query: 314 TLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
TG SL+ P+ IV + +A + A N C P TL P KV GKIV CDR G
Sbjct: 346 NYTGVSLYAGKALPSTPLPIVYAANASNSTAGN----LCMPGTLTPEKVAGKIVVCDR-G 400
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G AG G++L N NG+ L+++ H+L + P
Sbjct: 401 VSARVQKGFVVRDAGGAGMVLSNTAT-NGEELVADAHLLPAAGVGAKEGAAIKAYVASDP 459
Query: 433 SD----IKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAA 488
S + +GT++ + +P+PV+A++SSRGPN + P ILKPD+ APGVNILAA
Sbjct: 460 SPTATIVVAGTQVDV---------RPSPVVAAFSSRGPNMLTPEILKPDIIAPGVNILAA 510
Query: 489 YSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTA 548
++ A + + DTRR FN++ GTSMSCPHV+G A L+++ HP WSPAA++SA+MTTA
Sbjct: 511 WTGKAGPTGIAADTRR-VAFNIISGTSMSCPHVSGLAALLRSAHPEWSPAAVRSALMTTA 569
Query: 549 TTR---DNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGY 605
+ P+ DA A PF YG+GH+ P SA+DPGLVYDLG DY++FLCA Y
Sbjct: 570 YSTYAGAGDANPLLDAATGAPATPFDYGAGHVDPASAVDPGLVYDLGTADYVDFLCALNY 629
Query: 606 NQQLISALNFNMTFTCS--GTSSIDDLNYPSITL----------PNLGLNSVTVT--RTV 651
+I+A+ + ++ C+ S+ +LNYPS + + G + TVT RT+
Sbjct: 630 TSTMIAAVARSKSYGCTEGKAYSVYNLNYPSFAVAYSTASSQAAESSGAAATTVTHRRTL 689
Query: 652 TNVGPPSTY-FAKVQLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELR 710
TNVG TY + + G +AV P+ L F GEKK++ V A S P FG L
Sbjct: 690 TNVGAAGTYKVSAAAMPGVAVAVEPTELAFTSAGEKKSYTVSFTAKS-QPSGTAGFGRLV 748
Query: 711 WTNGKHIVRSPV 722
W++GKH V SP+
Sbjct: 749 WSDGKHSVASPM 760
>UniRef100_P93205 SBT2 protein [Lycopersicon esculentum]
Length = 775
Score = 503 bits (1295), Expect = e-141
Identities = 294/726 (40%), Positives = 429/726 (58%), Gaps = 40/726 (5%)
Query: 12 NAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGN 71
+ +E I+YSY +G AA L EEE ++ + V++VF +++LHTTRS FLGL
Sbjct: 71 DGEERILYSYQTAFHGVAAQLSEEEVKKLQERNGVLAVFPEIKYQLHTTRSPLFLGLDRE 130
Query: 72 DINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKV 131
D + W N I+G +DTG+WPES SF+D G+ +P+ W+G +C+ + +K
Sbjct: 131 DSSKLWADRLSDHNVIVGVLDTGIWPESPSFNDSGMTSVPSHWKG--VCETGR--GFEKH 186
Query: 132 PCNRKLIGARFFNKAYQKRNGKLPRSQQ--TARDFVGHGTHTLSTAGGNFVPGASIFNIG 189
C++K++GAR F + Y+ +GK+ + +ARD GHGTHT T G+ V GA++
Sbjct: 187 HCSKKIVGARVFFRGYEAASGKINERGEFKSARDQDGHGTHTAGTVAGSVVRGANLLGYA 246
Query: 190 NGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNS 249
GT +G +P ARVA YKVCW CF +D+LSA+DQA+ DGV+I+S+S GG S+ +
Sbjct: 247 YGTARGMAPGARVAAYKVCW----VGGCFSSDILSAVDQAVADGVNILSISLGGGVSSYN 302
Query: 250 EEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMT 309
D +SI AF A+ + + + SAGN GP P S+ NV+PW+ TV AST+DRDF + +
Sbjct: 303 R----DSLSIAAFGAMEKGVFVSCSAGNGGPDPISLTNVSPWITTVGASTMDRDFPATVE 358
Query: 310 IGN-KTLTGASLF---VNLPPNQDFTIVTSTDAKLANATN-RDARFCRPRTLDPSKVNGK 364
+G K +TGASL+ +NL + + ++ +N++N + C TLD + V GK
Sbjct: 359 LGTGKIVTGASLYKGRMNLSTQKQYPLIYLG----SNSSNLMPSSLCLDGTLDKASVAGK 414
Query: 365 IVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTT 424
IV CDR G V +GQ AG G+IL N NG+ L+++ H+L ++
Sbjct: 415 IVICDR-GISPRVQKGQVVKEAGGVGMILTNTAA-NGEELVADSHLLPAVAV----GERE 468
Query: 425 GRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVN 484
GR++ + + + LR K R P+PV+A++SSRGPN + ILKPD+ APGVN
Sbjct: 469 GRAIKLYAAGRSATATLRFLGTKLGIR--PSPVVAAFSSRGPNFLSLEILKPDMVAPGVN 526
Query: 485 ILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAI 544
ILA ++ S+L D RR FN++ GTSMSCPHV+G A L+K HP+WSPAAIKSA+
Sbjct: 527 ILAGWTGALGPSSLPIDQRRT-NFNILSGTSMSCPHVSGIAALLKARHPDWSPAAIKSAL 585
Query: 545 MTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASG 604
MTTA DNT K + DA T + P+ +G+GH+ P A+DPGL+YD+G +DY FLC
Sbjct: 586 MTTAYVHDNTYKSLKDASSVTPSTPYDHGAGHVNPRKAVDPGLIYDIGAQDYFEFLCTQE 645
Query: 605 YNQQLISALNFNMTFTC-SGTSSIDDLNYPSITL---PNLGLNSVTVTRTVTNVGPP-ST 659
+ + TC ++ DLNYP+I+ L+ +T+ RTVTNVG P S
Sbjct: 646 LSPSQLMVFGKFSNRTCHHSLANPGDLNYPAISAVFPEKTKLSMLTLHRTVTNVGSPISN 705
Query: 660 YFAKVQ-LAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIV 718
Y V G + V P LNF +K +++V + +V+ ++ +FG L W +G H V
Sbjct: 706 YHVVVSAFKGAVVKVEPERLNFTSKNQKLSYKVTFK--TVSRQKAPEFGSLIWKDGTHKV 763
Query: 719 RSPVTV 724
RSP+ +
Sbjct: 764 RSPIAI 769
>UniRef100_P93204 SBT1 protein [Lycopersicon esculentum]
Length = 766
Score = 499 bits (1285), Expect = e-139
Identities = 295/725 (40%), Positives = 420/725 (57%), Gaps = 46/725 (6%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+YN I+G++ L +EA +A+ P ++ V ++LHTTRS FLGL G + S
Sbjct: 64 MLYTYNSVIHGYSTQLTADEAKALAQQPGILLVHEEVIYELHTTRSPTFLGLEGRESRSF 123
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ + IIG +DTGVWPESKSF D G+G +PA W+G CQ K + CNRK
Sbjct: 124 FPQTEARSEVIIGVLDTGVWPESKSFDDTGLGQVPASWKGK--CQTGKNFDASS--CNRK 179
Query: 137 LIGARFFNKAYQKRNGKLPRS--QQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF++ Y+ G + + ++ RD GHGTHT +TA G+ V GAS+ GT +
Sbjct: 180 LIGARFFSQGYEAAFGAIDETIESKSPRDDEGHGTHTATTAAGSVVTGASLLGYATGTAR 239
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G + ARVA YKVCW+ CF +D+L+ +DQA+ DGV+++S+S GG S +I
Sbjct: 240 GMASHARVAAYKVCWT----GGCFSSDILAGMDQAVIDGVNVLSLSLGGTISDYHRDI-- 293
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
++IGAF A ++ I + SAGN GP+ G++ NVAPW+ TV A T+DR+F + + IGN K
Sbjct: 294 --VAIGAFSAASQGIFVSCSAGNGGPSSGTLSNVAPWITTVGAGTMDREFPAYIGIGNGK 351
Query: 314 TLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
L G SL+ P+ +V + + ++ N C +L P KV GKIV CDR G
Sbjct: 352 KLNGVSLYSGKALPSSVMPLVYAGNVSQSSNGN----LCTSGSLIPEKVAGKIVVCDR-G 406
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSL-DII 431
+G AG G+IL N + G L+++ H++ T + +T G + I
Sbjct: 407 MNARAQKGLVVKDAGGIGMILANT-DTYGDELVADAHLIPTAAV----GQTAGNLIKQYI 461
Query: 432 PSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSL 491
S+ + K +P+PV+A++SSRGPN + P +LKPD+ APGVNILA ++
Sbjct: 462 ASNSNPTATIAFGGTKL--GVQPSPVVAAFSSRGPNPITPDVLKPDLIAPGVNILAGWTG 519
Query: 492 FASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTR 551
+ L DTR FN++ GTSMSCPHV+G A L+K HP WSPAAI+SA+MTT+ +
Sbjct: 520 KVGPTGLQEDTRN-VGFNIISGTSMSCPHVSGLAALLKAAHPEWSPAAIRSALMTTSYST 578
Query: 552 DNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLIS 611
K I D + PF YG+GH+ P +A+ PGLVYDL + DY+NFLCA Y+ +I
Sbjct: 579 YKNGKTIEDVATGMSSTPFDYGAGHVNPTAAVSPGLVYDLTVDDYINFLCALDYSPSMIK 638
Query: 612 ALNFNMTFTCSGTSS--IDDLNYPSITLP-------NLGLNSVTV---TRTVTNVGPPST 659
+ +C + DLNYPS ++P + ++ TV TRT+TNVG P+T
Sbjct: 639 VI-AKRDISCDENKEYRVADLNYPSFSIPMETAWGEHADSSTPTVTRYTRTLTNVGNPAT 697
Query: 660 YFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHI 717
Y A V + KI V P +L F + EKKT+ V ATS P F L W++G+H+
Sbjct: 698 YKASVSSETQDVKILVEPQTLTFSRKNEKKTYTVTFTATS-KPSGTTSFARLEWSDGQHV 756
Query: 718 VRSPV 722
V SP+
Sbjct: 757 VASPI 761
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,229,731,534
Number of Sequences: 2790947
Number of extensions: 53134879
Number of successful extensions: 129649
Number of sequences better than 10.0: 869
Number of HSP's better than 10.0 without gapping: 450
Number of HSP's successfully gapped in prelim test: 419
Number of HSP's that attempted gapping in prelim test: 126229
Number of HSP's gapped (non-prelim): 1590
length of query: 726
length of database: 848,049,833
effective HSP length: 135
effective length of query: 591
effective length of database: 471,271,988
effective search space: 278521744908
effective search space used: 278521744908
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 79 (35.0 bits)
Medicago: description of AC135318.3


