

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.2 + phase: 0
(164 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
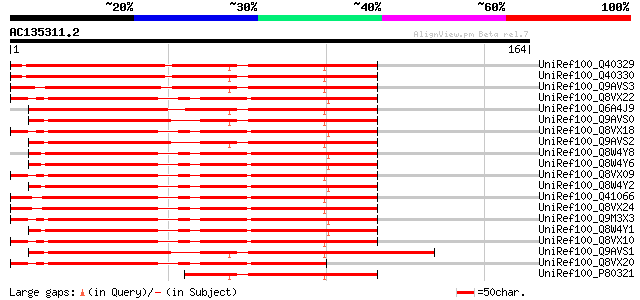
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q40329 Serine proteinase inhibitor [Medicago sativa] 153 2e-36
UniRef100_Q40330 Serine proteinase inhibitor [Medicago sativa] 152 4e-36
UniRef100_Q9AVS3 Trypsin inhibitor precursor [Pisum sativum] 141 5e-33
UniRef100_Q8VX22 Trypsin/chymotrypsin inhibitor precursor [Pisum... 137 1e-31
UniRef100_Q6A4J9 Trypsin inhibitor [Medicago scutellata] 137 1e-31
UniRef100_Q9AVS0 Trypsin inhibitor precursor [Pisum sativum] 135 4e-31
UniRef100_Q8VX18 Trypsin/chymotrypsin inhibitor precursor [Pisum... 135 4e-31
UniRef100_Q9AVS2 Trypsin inhibitor precursor [Pisum sativum] 132 4e-30
UniRef100_Q8W4Y8 Trypsin/chymotrypsin inhibitor [Lens culinaris] 132 4e-30
UniRef100_Q8W4Y6 Trypsin/chymotrypsin inhibitor [Lens ervoides] 131 7e-30
UniRef100_Q8VX09 Trypsin/chymotrypsin inhibitor precursor [Pisum... 130 9e-30
UniRef100_Q8W4Y2 Trypsin/chymotrypsin inhibitor [Lens odemensis] 130 2e-29
UniRef100_Q41066 Seed trypsin/chymotrypsin inhibitor TI5-72 prec... 130 2e-29
UniRef100_Q8VX24 Trypsin/chymotrypsin inhibitor precursor [Pisum... 129 2e-29
UniRef100_Q9M3X3 Seed trypsin/chymotrypsin inhibitor IVA precurs... 129 3e-29
UniRef100_Q8W4Y1 Trypsin/chymotrypsin inhibitor [Lens culinaris ... 128 5e-29
UniRef100_Q8VX10 Trypsin/chymotrypsin inhibitor precursor [Pisum... 128 5e-29
UniRef100_Q9AVS1 Putative trypsin inhibitor precursor [Pisum sat... 125 4e-28
UniRef100_Q8VX20 Trypsin/chymotrypsin inhibitor precursor [Pisum... 108 6e-23
UniRef100_P80321 Bowman-Birk type proteinase inhibitor [Medicago... 79 3e-14
>UniRef100_Q40329 Serine proteinase inhibitor [Medicago sativa]
Length = 113
Score = 153 bits (386), Expect = 2e-36
Identities = 79/119 (66%), Positives = 90/119 (75%), Gaps = 9/119 (7%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELM NKKAM+MKLALLVFLLGFTSTVVDARFD SFI+Q++ NG++ NYD KST T
Sbjct: 1 MELM-MNKKAMMMKLALLVFLLGFTSTVVDARFDQASFISQLLFNGEAA--NYDVKSTTT 57
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACCN C C S+ P QCRC D+G+TCHSACK+C C S P C C DIT+FCY CN
Sbjct: 58 ACCNFCPCTRSIPP---QCRCTDIGETCHSACKSCLCTRSIPPQCRCTDITNFCYPKCN 113
>UniRef100_Q40330 Serine proteinase inhibitor [Medicago sativa]
Length = 113
Score = 152 bits (383), Expect = 4e-36
Identities = 79/119 (66%), Positives = 90/119 (75%), Gaps = 9/119 (7%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELM NKKAM+MKLALLVFLLGFTST VDARFD SFITQ++ NG++ NYD KST T
Sbjct: 1 MELM-MNKKAMMMKLALLVFLLGFTSTGVDARFDRASFITQLLFNGEAA--NYDVKSTTT 57
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACCN C C S+ P QCRC+D+G+TCHSACK+C C S P C C DIT+FCY CN
Sbjct: 58 ACCNFCPCTKSIPP---QCRCSDIGETCHSACKSCICTRSYPPQCRCTDITNFCYPKCN 113
>UniRef100_Q9AVS3 Trypsin inhibitor precursor [Pisum sativum]
Length = 110
Score = 141 bits (356), Expect = 5e-33
Identities = 73/119 (61%), Positives = 85/119 (71%), Gaps = 12/119 (10%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MEL+N K +MKLAL+VFLLGFT+TVVDARFDSTSFITQ++SNGD+ Y KST T
Sbjct: 1 MELINTKK---MMKLALMVFLLGFTATVVDARFDSTSFITQLLSNGDA---GYSIKSTTT 54
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC++C+C S+ P QC C DVGKTCHS C C C S P C C D DFCY+ CN
Sbjct: 55 ACCDSCICTKSIPP---QCHCTDVGKTCHSGCNLCLCTRSFPPQCHCTDTNDFCYQKCN 110
>UniRef100_Q8VX22 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 104
Score = 137 bits (344), Expect = 1e-31
Identities = 74/117 (63%), Positives = 87/117 (74%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLGFTANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRCADVG+TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTASNPPT-CRCADVGETCHSACNSCICAYSNPPKCQCFDTHKFCYKACH 104
>UniRef100_Q6A4J9 Trypsin inhibitor [Medicago scutellata]
Length = 110
Score = 137 bits (344), Expect = 1e-31
Identities = 70/113 (61%), Positives = 81/113 (70%), Gaps = 11/113 (9%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM+MKLALLVFLL F ST VDARFD SFITQ++SNG++ T KST TACC+ C
Sbjct: 6 NKKAMMMKLALLVFLLSFASTTVDARFDRASFITQLLSNGEANT-----KSTTTACCDFC 60
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C S+ P QC+C DV + CHSACK+C C S P C CYDITDFCY C+
Sbjct: 61 PCTRSIPP---QCQCTDVREKCHSACKSCLCTRSFPPQCRCYDITDFCYPSCS 110
>UniRef100_Q9AVS0 Trypsin inhibitor precursor [Pisum sativum]
Length = 104
Score = 135 bits (340), Expect = 4e-31
Identities = 70/113 (61%), Positives = 84/113 (73%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM MKLAL++FLLGFT+TVVDARF STSFITQ++SNGD++ ACC++C
Sbjct: 5 NKKAM-MKLALMLFLLGFTATVVDARFGSTSFITQLLSNGDASNK---------ACCDSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
LC S+ P QC+C D+G+TCHSACK C C S P C+C DITDFCYK CN
Sbjct: 55 LCTRSIPP---QCQCNDIGETCHSACKACLCTRSLPPKCSCTDITDFCYKKCN 104
>UniRef100_Q8VX18 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 104
Score = 135 bits (340), Expect = 4e-31
Identities = 73/117 (62%), Positives = 86/117 (73%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLGFTANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTASNPPT-CRCVDVGETCHSACNSCICAYSNPPKCQCFDTHKFCYKACH 104
>UniRef100_Q9AVS2 Trypsin inhibitor precursor [Pisum sativum]
Length = 104
Score = 132 bits (331), Expect = 4e-30
Identities = 68/113 (60%), Positives = 82/113 (72%), Gaps = 16/113 (14%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM MKLAL++FLLGFT+TVVDARFDS SFITQ++SNG ++ ACC++C
Sbjct: 5 NKKAM-MKLALVLFLLGFTATVVDARFDSNSFITQLLSNGGASNK---------ACCDSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
LC S+ P +CRC D+G+TCHSACK C C S P C C DITDFCY+ CN
Sbjct: 55 LCTRSIPP---RCRCNDIGETCHSACKTCICTRSLPPQCRCIDITDFCYEKCN 104
>UniRef100_Q8W4Y8 Trypsin/chymotrypsin inhibitor [Lens culinaris]
Length = 104
Score = 132 bits (331), Expect = 4e-30
Identities = 68/111 (61%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL++FLLGFT+ VVDARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMLFLLGFTANVVDARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>UniRef100_Q8W4Y6 Trypsin/chymotrypsin inhibitor [Lens ervoides]
Length = 104
Score = 131 bits (329), Expect = 7e-30
Identities = 68/111 (61%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL+VFLLGFT+ VV+ARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMVFLLGFTANVVNARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>UniRef100_Q8VX09 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 114
Score = 130 bits (328), Expect = 9e-30
Identities = 72/117 (61%), Positives = 83/117 (70%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACNSCICALSYPPKCQCFDTQKFCYKACH 104
>UniRef100_Q8W4Y2 Trypsin/chymotrypsin inhibitor [Lens odemensis]
Length = 104
Score = 130 bits (326), Expect = 2e-29
Identities = 67/111 (60%), Positives = 82/111 (73%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKLAL++FLLGFT+ VV+ARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLALMLFLLGFTANVVNARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>UniRef100_Q41066 Seed trypsin/chymotrypsin inhibitor TI5-72 precursor [Pisum
sativum]
Length = 114
Score = 130 bits (326), Expect = 2e-29
Identities = 69/117 (58%), Positives = 82/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK ++MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNKK---VMMKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSDPPT-CRCVDVGETCHSACDSCICALSYPPQCQCFDTHKFCYKACH 104
>UniRef100_Q8VX24 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 104
Score = 129 bits (325), Expect = 2e-29
Identities = 69/117 (58%), Positives = 83/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK ++MKLAL+VFLLGFT+ VV+AR DSTSFITQV+SNG+ D KS
Sbjct: 1 MELMNKK---VMMKLALMVFLLGFTANVVNARLDSTSFITQVLSNGN------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACDSCICALSYPPKCQCFDTHKFCYKACH 104
>UniRef100_Q9M3X3 Seed trypsin/chymotrypsin inhibitor IVA precursor (PSTI IVA)
(TI12-36) [Contains: Seed trypsin/chymotrypsin inhibitor
I (PSTI I)] [Pisum sativum]
Length = 114
Score = 129 bits (324), Expect = 3e-29
Identities = 71/117 (60%), Positives = 84/117 (71%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+ARFDSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARFDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DV +TCHSAC +C C +S+ P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVRETCHSACDSCICAYSNPPKCQCFDTHKFCYKACH 104
>UniRef100_Q8W4Y1 Trypsin/chymotrypsin inhibitor [Lens culinaris subsp. tomentosus]
Length = 104
Score = 128 bits (322), Expect = 5e-29
Identities = 66/111 (59%), Positives = 81/111 (72%), Gaps = 12/111 (10%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKA +MKL L++FLLGFT+ VV+ARFDSTSFITQV+SNGD D KS ACC+TC
Sbjct: 5 NKKA-IMKLVLMLFLLGFTANVVNARFDSTSFITQVLSNGD------DVKS---ACCDTC 54
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCECGWSS-PLCTCYDITDFCYKPCN 116
LC+ + T CRC DV ++CHSAC C C +S+ P C CYD +FCYK C+
Sbjct: 55 LCTRSQPPT-CRCVDVRESCHSACDKCVCAYSNPPQCQCYDTHNFCYKTCH 104
>UniRef100_Q8VX10 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 114
Score = 128 bits (322), Expect = 5e-29
Identities = 71/117 (60%), Positives = 82/117 (69%), Gaps = 14/117 (11%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VFLL F + VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFLLSFAANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P C C+D FCYK C+
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACNSCICALSYPPQCQCFDTQKFCYKACH 104
>UniRef100_Q9AVS1 Putative trypsin inhibitor precursor [Pisum sativum]
Length = 133
Score = 125 bits (314), Expect = 4e-28
Identities = 69/131 (52%), Positives = 86/131 (64%), Gaps = 16/131 (12%)
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
NKKAM MKLAL++FLLGFT+TVVDARFDS SFI Q++S GD++ ACC++C
Sbjct: 5 NKKAM-MKLALMLFLLGFTATVVDARFDSDSFIIQLLSKGDASNK---------ACCDSC 54
Query: 67 LC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCNVWQQQQK 123
LC S+ P +CRC D G+TCHSACK C C S P C C DITDFCY+ N +
Sbjct: 55 LCTKSIPP---RCRCNDTGETCHSACKTCICTRSLPPQCRCIDITDFCYEKRNQPASKPT 111
Query: 124 VQVVVTIVLAQ 134
+ V+ + L Q
Sbjct: 112 INHVMDLNLQQ 122
>UniRef100_Q8VX20 Trypsin/chymotrypsin inhibitor precursor [Pisum sativum]
Length = 90
Score = 108 bits (269), Expect = 6e-23
Identities = 62/100 (62%), Positives = 71/100 (71%), Gaps = 13/100 (13%)
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MELMNK KAM MKLAL+VF L F + VV+AR DSTSFITQV+SNGD D KS
Sbjct: 1 MELMNK--KAM-MKLALMVFPLSFAANVVNARLDSTSFITQVLSNGD------DVKS--- 48
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSSP 100
ACC+TCLC+ + T CRC DVG+TCHSAC +C C S P
Sbjct: 49 ACCDTCLCTKSNPPT-CRCVDVGETCHSACNSCICALSYP 87
>UniRef100_P80321 Bowman-Birk type proteinase inhibitor [Medicago scutellata]
Length = 62
Score = 79.3 bits (194), Expect = 3e-14
Identities = 37/64 (57%), Positives = 43/64 (66%), Gaps = 6/64 (9%)
Query: 56 KSTATACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCY 112
KST TACC+ C C S+ P QC+C DV + CHSACK+C C S P C CYDITDFCY
Sbjct: 2 KSTTTACCDFCPCTRSIPP---QCQCTDVREKCHSACKSCLCTRSFPPQCRCYDITDFCY 58
Query: 113 KPCN 116
C+
Sbjct: 59 PSCS 62
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.131 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 248,365,608
Number of Sequences: 2790947
Number of extensions: 8912990
Number of successful extensions: 36812
Number of sequences better than 10.0: 321
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 250
Number of HSP's that attempted gapping in prelim test: 35361
Number of HSP's gapped (non-prelim): 1000
length of query: 164
length of database: 848,049,833
effective HSP length: 117
effective length of query: 47
effective length of database: 521,509,034
effective search space: 24510924598
effective search space used: 24510924598
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC135311.2


