

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
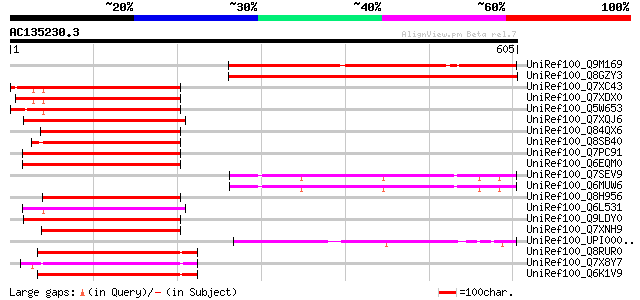
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M169 Hypothetical protein T6D9_70 [Arabidopsis thali... 483 e-135
UniRef100_Q8GZY3 Hypothetical protein OSJNBa0013D02.19 [Oryza sa... 446 e-123
UniRef100_Q7XC43 Hypothetical protein [Oryza sativa] 208 4e-52
UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza... 207 7e-52
UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sat... 201 4e-50
UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa] 195 3e-48
UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sat... 192 3e-47
UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa] 187 6e-46
UniRef100_Q7PC91 Putative transposase [Oryza sativa] 184 7e-45
UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa] 184 7e-45
UniRef100_Q7SEV9 Hypothetical protein [Neurospora crassa] 184 7e-45
UniRef100_Q6MUW6 Hypothetical protein 7C14.110 [Neurospora crassa] 184 7e-45
UniRef100_Q8H956 B protein [Oryza sativa] 181 4e-44
UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa] 181 4e-44
UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana] 180 9e-44
UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa] 180 9e-44
UniRef100_UPI000042E7CD UPI000042E7CD UniRef100 entry 173 2e-41
UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa] 171 7e-41
UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa] 171 7e-41
UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa] 171 7e-41
>UniRef100_Q9M169 Hypothetical protein T6D9_70 [Arabidopsis thaliana]
Length = 376
Score = 483 bits (1243), Expect = e-135
Identities = 242/343 (70%), Positives = 282/343 (81%), Gaps = 10/343 (2%)
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
R SFGIAFDIDGVILLG++PVGGSP+ALR+LY+ G LK P++FLTNGGG+PE+KRASE+
Sbjct: 36 RSSFGIAFDIDGVILLGSSPVGGSPSALRRLYDDSGALKIPFLFLTNGGGLPESKRASEM 95
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 381
S LLG+ VS QV+Q HSPFR+LVNRFE++L+VAAGKGEPA VMS YGFKNVIS+D YAS
Sbjct: 96 SHLLGVQVSPLQVIQAHSPFRKLVNRFENELVVAAGKGEPAAVMSNYGFKNVISMDEYAS 155
Query: 382 RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLC 441
F+NIDPLAPYKK + E + DV S+RVQAAFIVSDPVDWSRD+QVLC
Sbjct: 156 YFDNIDPLAPYKK-----LMFRQDGHKELRSREDVLSQRVQAAFIVSDPVDWSRDIQVLC 210
Query: 442 DILKTGGLPGRNVGTQPHLYFANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHSLEY 501
DIL+TGGLPG+ +G QPHLY ANDDL+YQT+FP+ERLGMGAFRIALESIFNR H LEY
Sbjct: 211 DILRTGGLPGKEIGPQPHLYIANDDLDYQTEFPTERLGMGAFRIALESIFNRIHEKPLEY 270
Query: 502 TCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRGARQT 561
T FGKP+P VFKNAE VL++ + Y N + + F+TLYMIGDNP +DIRGARQ
Sbjct: 271 TSFGKPNPFVFKNAEDVLKE----IATSPYSSN-QGSHHFKTLYMIGDNPKIDIRGARQA 325
Query: 562 GHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKE 604
G PWFSILTRTGVFKGK+NH +FPADLVVDTVEEAVD+IL +E
Sbjct: 326 GTPWFSILTRTGVFKGKDNHSEFPADLVVDTVEEAVDFILTRE 368
>UniRef100_Q8GZY3 Hypothetical protein OSJNBa0013D02.19 [Oryza sativa]
Length = 389
Score = 446 bits (1146), Expect = e-123
Identities = 214/344 (62%), Positives = 263/344 (76%)
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
RPSFGIAFDIDGVIL G +P+GGSP A+R+LY+ DGTLK P++FLTNGGG+PE KRA EL
Sbjct: 44 RPSFGIAFDIDGVILRGRSPIGGSPQAIRRLYSEDGTLKIPFLFLTNGGGVPEHKRAQEL 103
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 381
SELLG+N+S +QV+ G SP+++LVNRFE+ LI+A GKGEPA VM +YGF+ V+SID Y+S
Sbjct: 104 SELLGVNISPAQVVHGSSPYKELVNRFENDLIIAVGKGEPAAVMVDYGFRKVLSIDEYSS 163
Query: 382 RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLC 441
F +IDPLAP+KKW + N ++ P DVF ERV+ F+VSDPVDW RD+QVLC
Sbjct: 164 YFGDIDPLAPFKKWIVQQPDNINLMSEKVHPSYDVFEERVKGVFVVSDPVDWGRDLQVLC 223
Query: 442 DILKTGGLPGRNVGTQPHLYFANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHSLEY 501
DIL TGGLPG G QP LYFA+DDLEYQ FPSERLGMGAFRIALESIFN+ + H L+Y
Sbjct: 224 DILSTGGLPGSGRGDQPPLYFASDDLEYQAAFPSERLGMGAFRIALESIFNQVNDHRLKY 283
Query: 502 TCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRGARQT 561
+GKP+P VFKNA +L+K + S F T+YM+GDNP VDI GA +
Sbjct: 284 ISYGKPNPFVFKNAANILEKLAICMHPSSLPTKEVEEHRFSTIYMVGDNPKVDINGALKA 343
Query: 562 GHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKEC 605
G PW +LTRTGVF+GK+N ++PADLVVDTVE+A++ IL KEC
Sbjct: 344 GPPWSPVLTRTGVFRGKDNDPQYPADLVVDTVEDAINCILEKEC 387
>UniRef100_Q7XC43 Hypothetical protein [Oryza sativa]
Length = 513
Score = 208 bits (529), Expect = 4e-52
Identities = 103/212 (48%), Positives = 138/212 (64%), Gaps = 10/212 (4%)
Query: 1 MDPFDIEAYFQKRDVEDTYIVNRFIQR-----RKKLEEGSASRT---RKYFNRDHVAANQ 52
+DP +I Y + + ++N F R + +LE G++ RT RKY NR+H A+
Sbjct: 9 IDPAEI--YTTDMFMAEHSVLNSFAGRIDRRIKARLEGGTSRRTSGPRKYINRNHEGAHD 66
Query: 53 RLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAK 112
+L DYFA +P Y A FRRR+RM++HVFL IV +L +YFT +VD G SPL K
Sbjct: 67 QLFADYFAEDPLYSAATFRRRFRMRRHVFLHIVDELGKWSSYFTHRVDCTGCLGHSPLQK 126
Query: 113 CTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQ 172
CT A+RMLAYG AAD +DEY+K+ ST LECL F +G++ ++ YLR PT +D R+
Sbjct: 127 CTAAIRMLAYGTAADTLDEYLKVPQSTALECLENFVEGVVEVFSSRYLRRPTAEDLERLL 186
Query: 173 HVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
V E+RGFP M+GSIDCMHW WKN P AW+G+
Sbjct: 187 QVGESRGFPGMLGSIDCMHWRWKNCPTAWKGQ 218
>UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza sativa]
Length = 308
Score = 207 bits (527), Expect = 7e-52
Identities = 99/206 (48%), Positives = 135/206 (65%), Gaps = 8/206 (3%)
Query: 7 EAYFQKRDVEDTYIVNRFIQR-----RKKLEEGSASRT---RKYFNRDHVAANQRLIDDY 58
E Y + + ++N F R + +LE G++ RT RKY NR+H A+ +L DY
Sbjct: 13 EVYTTDMFMAEHSVLNSFAGRIDKRIKARLEGGTSRRTSGPRKYINRNHEGAHDQLFADY 72
Query: 59 FANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMR 118
FA +P Y A FRRR+RM++HVFL IV +L +YFT +++ + G SPL KCT A+R
Sbjct: 73 FAKDPLYSAATFRRRFRMRRHVFLHIVDELGKWSSYFTHRINCTGRLGHSPLQKCTAAIR 132
Query: 119 MLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETR 178
MLAYG AAD +DEY+K+ ST LECL F +G++ ++ YLR PT +D R+ V E+R
Sbjct: 133 MLAYGTAADTLDEYLKVPQSTALECLENFVEGVVEVFSSRYLRRPTAEDLERLLQVGESR 192
Query: 179 GFPWMIGSIDCMHWEWKNYPKAWEGK 204
GFP M+GSIDCMHW WKN P AW+G+
Sbjct: 193 GFPRMLGSIDCMHWRWKNCPTAWKGQ 218
>UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sativa]
Length = 444
Score = 201 bits (512), Expect = 4e-50
Identities = 102/209 (48%), Positives = 138/209 (65%), Gaps = 6/209 (2%)
Query: 1 MDPFDI-EAYFQKRDVEDTYIVNRFIQRRK-KLEEGSASR---TRKYFNRDHVAANQRLI 55
+DP + + Y +++V ++ +R I++ K + G R T K RDHV A+ RL+
Sbjct: 15 LDPSKVLDKYISEQNVLGSF-ASRIIEKMKGRFGAGRLKRQGGTIKTIRRDHVDAHSRLV 73
Query: 56 DDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTT 115
DYFA P Y + MFR R+RM K +FLRIV LS YFTQ+ D + +SPL KCT
Sbjct: 74 ADYFAEHPLYPERMFRTRFRMHKPLFLRIVEALSQWSPYFTQRGDCSGHTSLSPLQKCTA 133
Query: 116 AMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVS 175
A+RMLAYG ADA+DEY+KIG ST LECL F +G+I ++ YLR PT++D I HV+
Sbjct: 134 ALRMLAYGTPADALDEYLKIGKSTALECLEMFSRGVIVVFGGTYLRRPTREDVEHILHVN 193
Query: 176 ETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
E+RGFP M+GSIDCMHW W++ P+AW G+
Sbjct: 194 ESRGFPGMLGSIDCMHWRWESCPRAWRGQ 222
>UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa]
Length = 454
Score = 195 bits (496), Expect = 3e-48
Identities = 96/194 (49%), Positives = 121/194 (61%)
Query: 17 DTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRM 76
+T I + KK + G ++ RKY R+ A RL YF P Y+D FR RYRM
Sbjct: 44 ETQIEEDISYQIKKQQRGPPTQQRKYIRRERQDAEDRLKAYYFLENPMYNDTQFRWRYRM 103
Query: 77 QKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIG 136
+KH+FL IV LS F Q+ DA K G SPL KCT A+RMLAYG +AD +DE ++I
Sbjct: 104 KKHLFLHIVQTLSHWSPVFQQRKDAFGKVGFSPLLKCTAALRMLAYGTSADILDENLQIA 163
Query: 137 SSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKN 196
+T +E L+ FCKGII + YLR PT +D R+ HV E RGFP M+GSIDCMHWEWKN
Sbjct: 164 ETTVIESLVNFCKGIIDCFGPKYLRRPTAEDIQRLLHVGEARGFPGMLGSIDCMHWEWKN 223
Query: 197 YPKAWEGKLAFANH 210
P AW+G+ +H
Sbjct: 224 CPVAWKGQYTRGDH 237
>UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sativa]
Length = 626
Score = 192 bits (487), Expect = 3e-47
Identities = 88/168 (52%), Positives = 119/168 (70%)
Query: 37 SRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFT 96
S TR+Y R+ A+N L+ +YF+ P Y D MFRRR+RM+K +FLRIV LS YFT
Sbjct: 238 SGTRRYIPRNREASNADLVANYFSESPIYTDKMFRRRFRMRKPLFLRIVSALSEWSPYFT 297
Query: 97 QQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYE 156
++DA + G SPL KCT A+RMLAYG AD +DE +KIG +T LECL +F +G+I ++
Sbjct: 298 NRLDATGRAGHSPLQKCTAAIRMLAYGTPADQLDEVLKIGPNTALECLGKFAEGVIEIFR 357
Query: 157 QVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
+ YLRAP D+ R+ V+++RGFP M+G+IDCMHW WKN P +W G+
Sbjct: 358 KEYLRAPRSDEVERLLQVADSRGFPGMLGNIDCMHWAWKNCPVSWCGQ 405
>UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa]
Length = 915
Score = 187 bits (476), Expect = 6e-46
Identities = 91/177 (51%), Positives = 117/177 (65%), Gaps = 4/177 (2%)
Query: 27 RRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVG 86
RR++ + G R+Y R ++ L+ +YF+ P Y D FRRR+RM K +FLRIV
Sbjct: 519 RRRQRQSGP----RRYIPRPREKGHEDLVANYFSANPIYTDEQFRRRFRMNKPLFLRIVN 574
Query: 87 DLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLR 146
LS+ D +FTQ+VDA + SPL KCT A+RML YG ADA+DE +KI +ST+LECL +
Sbjct: 575 ALSNWDQFFTQRVDATGRDSHSPLQKCTAAIRMLGYGTPADALDEVLKIAASTSLECLGK 634
Query: 147 FCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEG 203
F GII + YLR PT D+ +I +E RGFP MIGSIDCMHW+WKN PK W G
Sbjct: 635 FAVGIIECFGSEYLRPPTSDELEKILQENEARGFPGMIGSIDCMHWQWKNCPKGWAG 691
>UniRef100_Q7PC91 Putative transposase [Oryza sativa]
Length = 482
Score = 184 bits (467), Expect = 7e-45
Identities = 87/189 (46%), Positives = 119/189 (62%)
Query: 16 EDTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYR 75
E T +V I+ + RK+ R A+Q+L++DYF+ P Y +FRRR+R
Sbjct: 67 EATVVVQSTIEGLQNEASDHRHHPRKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFR 126
Query: 76 MQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKI 135
M + +FLRIV L YFTQ+VDA N+KG+SPL KCT A+R LA G AD +DEY+KI
Sbjct: 127 MSRPLFLRIVEALGQWSVYFTQRVDAVNRKGLSPLQKCTAAIRQLATGSGADELDEYLKI 186
Query: 136 GSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWK 195
G +T +E + F KG+ ++ + YLR PT +D R+ + E RGFP M GSIDCMHW W+
Sbjct: 187 GETTAMEAMKNFVKGLQDVFGERYLRRPTMEDTERLLQLGEKRGFPGMFGSIDCMHWHWE 246
Query: 196 NYPKAWEGK 204
P AW+G+
Sbjct: 247 RCPVAWKGQ 255
>UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa]
Length = 447
Score = 184 bits (467), Expect = 7e-45
Identities = 87/189 (46%), Positives = 119/189 (62%)
Query: 16 EDTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYR 75
E T +V I+ + RK+ R A+Q+L++DYF+ P Y +FRRR+R
Sbjct: 32 EATVVVQSTIEGLQNEASDHRHHPRKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFR 91
Query: 76 MQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKI 135
M + +FLRIV L YFTQ+VDA N+KG+SPL KCT A+R LA G AD +DEY+KI
Sbjct: 92 MSRPLFLRIVEALGQWSVYFTQRVDAVNRKGLSPLQKCTAAIRQLATGSGADELDEYLKI 151
Query: 136 GSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWK 195
G +T +E + F KG+ ++ + YLR PT +D R+ + E RGFP M GSIDCMHW W+
Sbjct: 152 GETTAMEAMKNFVKGLQDVFGERYLRRPTMEDTERLLQLGEKRGFPGMFGSIDCMHWHWE 211
Query: 196 NYPKAWEGK 204
P AW+G+
Sbjct: 212 RCPVAWKGQ 220
>UniRef100_Q7SEV9 Hypothetical protein [Neurospora crassa]
Length = 419
Score = 184 bits (467), Expect = 7e-45
Identities = 127/375 (33%), Positives = 185/375 (48%), Gaps = 40/375 (10%)
Query: 263 PSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELS 322
P F AFDIDGV+L TP+ G+P AL+ L + D P++ LTNGGG E +R +LS
Sbjct: 45 PPFAFAFDIDGVLLHVATPIPGAPEALKFLNDND----IPFILLTNGGGKHETERVKDLS 100
Query: 323 ELLGLNVSASQVLQGHSPFRQLVN-----RFEDKLIVAAGKGEPALVMSEYGFKNVISID 377
+ LG+ ++ +Q H+PFRQLV+ R + L+ A + L+ EYGF+NV++
Sbjct: 101 QKLGVELTTDNFVQSHTPFRQLVDGPDSLREKTILVTGANAEKCRLIAEEYGFRNVVTPA 160
Query: 378 AYASRFENIDPLAPY--KKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSR 435
+ P P +T P F + ++ A F+ +DP DW+
Sbjct: 161 DILKAHPEVFPFDPLLDTVYTATARPLPRPIFTPGSGMRLSDALKIDAMFVFNDPRDWAY 220
Query: 436 DVQVLCDIL------------KTG--GLP--GRNVGTQPHLYFANDDLEYQTKFPSERLG 479
D+Q++ D+L K G LP G QP LYF+N DL + T F R G
Sbjct: 221 DIQLIVDLLLSQQGYVGTYSPKNGDAALPSCGWQQDGQPPLYFSNADLLWSTGFHLPRFG 280
Query: 480 MGAFRIALESIFNR-THHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNA 538
GAF+ A+ ++ R T+ H L+ GKP+ ++ AE VL H V H
Sbjct: 281 QGAFQAAVAGVWRRLTNGHELQRRVIGKPYSETYQFAERVLTTHRHEVLRRR---GHHEP 337
Query: 539 QPFQTLYMIGDNPAVDIRGAR----QTGHPWFSILTRTGVFKGKENHDK-----FPADLV 589
+++YM+GDNP DI GA + G W SIL RTGV+ + +K F ++
Sbjct: 338 GTLKSVYMVGDNPESDIAGANDFHSEAGTEWCSILVRTGVWSPEHAGEKALQGRFKPKVI 397
Query: 590 VDTVEEAVDYILAKE 604
VD EAV + L +E
Sbjct: 398 VDDAREAVRWALKRE 412
>UniRef100_Q6MUW6 Hypothetical protein 7C14.110 [Neurospora crassa]
Length = 458
Score = 184 bits (467), Expect = 7e-45
Identities = 127/375 (33%), Positives = 185/375 (48%), Gaps = 40/375 (10%)
Query: 263 PSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASELS 322
P F AFDIDGV+L TP+ G+P AL+ L + D P++ LTNGGG E +R +LS
Sbjct: 84 PPFAFAFDIDGVLLHVATPIPGAPEALKFLNDND----IPFILLTNGGGKHETERVKDLS 139
Query: 323 ELLGLNVSASQVLQGHSPFRQLVN-----RFEDKLIVAAGKGEPALVMSEYGFKNVISID 377
+ LG+ ++ +Q H+PFRQLV+ R + L+ A + L+ EYGF+NV++
Sbjct: 140 QKLGVELTTDNFVQSHTPFRQLVDGPDSLREKTILVTGANAEKCRLIAEEYGFRNVVTPA 199
Query: 378 AYASRFENIDPLAPY--KKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSR 435
+ P P +T P F + ++ A F+ +DP DW+
Sbjct: 200 DILKAHPEVFPFDPLLDTVYTATARPLPRPIFTPGSGMRLSDALKIDAMFVFNDPRDWAY 259
Query: 436 DVQVLCDIL------------KTG--GLP--GRNVGTQPHLYFANDDLEYQTKFPSERLG 479
D+Q++ D+L K G LP G QP LYF+N DL + T F R G
Sbjct: 260 DIQLIVDLLLSQQGYVGTYSPKNGDAALPSCGWQQDGQPPLYFSNADLLWSTGFHLPRFG 319
Query: 480 MGAFRIALESIFNR-THHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNA 538
GAF+ A+ ++ R T+ H L+ GKP+ ++ AE VL H V H
Sbjct: 320 QGAFQAAVAGVWRRLTNGHELQRRVIGKPYSETYQFAERVLTTHRHEVLRRR---GHHEP 376
Query: 539 QPFQTLYMIGDNPAVDIRGAR----QTGHPWFSILTRTGVFKGKENHDK-----FPADLV 589
+++YM+GDNP DI GA + G W SIL RTGV+ + +K F ++
Sbjct: 377 GTLKSVYMVGDNPESDIAGANDFHSEAGTEWCSILVRTGVWSPEHAGEKALQGRFKPKVI 436
Query: 590 VDTVEEAVDYILAKE 604
VD EAV + L +E
Sbjct: 437 VDDAREAVRWALKRE 451
>UniRef100_Q8H956 B protein [Oryza sativa]
Length = 455
Score = 181 bits (460), Expect = 4e-44
Identities = 82/165 (49%), Positives = 113/165 (67%)
Query: 40 RKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQV 99
RK+ R A+Q L++DYF+ P Y +FRRR+RM + +FLRIV L +YFTQ+V
Sbjct: 57 RKHIKRPREEAHQNLVNDYFSENPLYPSNIFRRRFRMYRPLFLRIVDALGQWSDYFTQRV 116
Query: 100 DAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVY 159
DAA ++G+SPL KCT A+R LA G AD +DEY+KIG +T ++ + F KGI ++ + Y
Sbjct: 117 DAAGRQGLSPLQKCTAAIRQLATGSGADELDEYLKIGETTAMDAMKNFVKGIREVFGERY 176
Query: 160 LRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
LR PT +D R+ + E RGFP M GSIDCMHW+W+ P AW+G+
Sbjct: 177 LRRPTVEDTERLLELGERRGFPGMFGSIDCMHWQWERCPTAWKGQ 221
>UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa]
Length = 443
Score = 181 bits (460), Expect = 4e-44
Identities = 91/203 (44%), Positives = 119/203 (57%), Gaps = 9/203 (4%)
Query: 16 EDTYIVNRFIQRRKKLEEGSASRT---------RKYFNRDHVAANQRLIDDYFANEPTYD 66
ED I + K +G AS R+Y R+ + L+ +YF+ P Y
Sbjct: 26 EDEIIEELLAEEFKAAGDGEASSVSRRRRQSGPRRYIPRNRERGHDDLVANYFSANPIYT 85
Query: 67 DAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAA 126
D MFRRR+RM++ +FLRIV +LS YFTQ+VDA + +SPL KC A+RML YG AA
Sbjct: 86 DEMFRRRFRMRRPLFLRIVHELSEWSPYFTQRVDATGRNSLSPLQKCIAAIRMLGYGTAA 145
Query: 127 DAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGS 186
D +DE ++I +ST LECL +F +G+I + YLR P D+ +I +E RGFP M GS
Sbjct: 146 DQLDEVLRIAASTCLECLGKFAEGVIEKFGDEYLRLPRADELEKILQENEARGFPGMYGS 205
Query: 187 IDCMHWEWKNYPKAWEGKLAFAN 209
IDCMHW WKN PK W G N
Sbjct: 206 IDCMHWPWKNCPKGWAGMFTSGN 228
>UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana]
Length = 415
Score = 180 bits (457), Expect = 9e-44
Identities = 87/188 (46%), Positives = 120/188 (63%)
Query: 17 DTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRM 76
D N F++ E +R R YF+R + L +DYF++ + FRRR+RM
Sbjct: 22 DQQFENVFVELSNSEEVAKKNRKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRM 81
Query: 77 QKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIG 136
+K +FLRIV LSS +F + DA + G SP+ KCT A+R+LAYG A+DAVDEY+++G
Sbjct: 82 KKPLFLRIVDRLSSELMFFQHRRDATGRFGHSPIQKCTAAIRLLAYGYASDAVDEYLRMG 141
Query: 137 SSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKN 196
+T + CL F KGII + YLRAPT + R+ ++ + RGFP MIGS+DCMHWEWKN
Sbjct: 142 ETTAMSCLENFTKGIISFFGDEYLRAPTATNLRRLLNIGKIRGFPGMIGSLDCMHWEWKN 201
Query: 197 YPKAWEGK 204
P AW+G+
Sbjct: 202 CPTAWKGQ 209
>UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa]
Length = 450
Score = 180 bits (457), Expect = 9e-44
Identities = 83/166 (50%), Positives = 109/166 (65%)
Query: 39 TRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQ 98
TR+Y +RDH + RL YF+ P Y D FRRR+RM++H+FLRIV L YF +
Sbjct: 64 TRRYISRDHEDDHNRLFAKYFSESPLYTDDQFRRRFRMRRHLFLRIVQALGVWSPYFRLR 123
Query: 99 VDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQV 158
DA K G+SPL KCT AMRMLAYG AD +DE + ST +EC++ F +G+ L+ +
Sbjct: 124 RDAFGKVGLSPLQKCTAAMRMLAYGTPADLMDETFGVAESTAMECMINFVQGVRHLFGEQ 183
Query: 159 YLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
YLR PT +D R+ E GFP M+GSIDCMHWEW++ P AW+G+
Sbjct: 184 YLRRPTVEDIQRLLQFGEAHGFPGMLGSIDCMHWEWQSCPVAWKGQ 229
>UniRef100_UPI000042E7CD UPI000042E7CD UniRef100 entry
Length = 350
Score = 173 bits (438), Expect = 2e-41
Identities = 122/355 (34%), Positives = 188/355 (52%), Gaps = 45/355 (12%)
Query: 268 AFDIDGVILLGNTPVGGSPAALRKLYNYDGTLK--FPYVFLTNGGGIPEAKRASELSELL 325
AFDIDGV+ G+ + + ++ L DG L P++ +TNGGG+ + +R S LS L
Sbjct: 19 AFDIDGVLKQGHNVLPEAKRTMKLLTGEDGRLPKPIPFLLITNGGGVLDHERLSLLSSEL 78
Query: 326 GLNVSASQVLQGHSPFRQLVNRFEDK-LIVAAGKGEPALVMSE-YGFKNV-ISIDAYASR 382
G+ ++ Q++Q H+P R ++++DK ++V GKGE ++E YG KN I D A
Sbjct: 79 GVQLTPDQLVQSHTPMRDYAHKYKDKHVLVIGGKGESCRKVAESYGMKNAHIPQDVIA-- 136
Query: 383 FENIDPLAPYKKWTTKL-ATTQNPKFDES--GPQIDVFSERVQAAFIVSDPVDWSRDVQV 439
W + + T+ K +E+ PQ D S + A F++ D DW RD +
Sbjct: 137 ------------WKSSIWDRTELAKEEEAFVRPQ-DFSSIQFSAIFVMHDSHDWGRDTTL 183
Query: 440 LCDILKTGG------LPGRNVGTQP-HLYFANDDLEYQTKFPSERLGMGAFRIALESIFN 492
+ D+L + GR G + L +N D+E+++ +P RLG GAFRI LE+++
Sbjct: 184 ILDLLNSENGYLGTRTEGRKNGEEAVELIMSNPDVEWRSDWPIPRLGQGAFRIGLEAVYK 243
Query: 493 RTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPA 552
T L YT +GKP + + +E+ L++++ V DS H +YM+GDNPA
Sbjct: 244 ATTGLDLTYTQYGKPFKATYDFSELSLRRYLASVGRDSSVPLH--------VYMVGDNPA 295
Query: 553 VDIRGARQTGHPWFSILTRTGVFKGKENHDKFPA---DLVVDTVEEAVDYILAKE 604
DI GA H W SIL RTGVF + H + PA + D VE+ V++ + KE
Sbjct: 296 SDIAGA--NAHGWSSILVRTGVF--HDTHGEKPAYQPTAIADNVEKGVEWAIGKE 346
>UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa]
Length = 626
Score = 171 bits (432), Expect = 7e-41
Identities = 83/191 (43%), Positives = 118/191 (61%), Gaps = 2/191 (1%)
Query: 34 GSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 93
G + + +R+ A + RL DYF+N PTY +FRRR RM + +FLRI+ + D+
Sbjct: 243 GGSVMGHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDD 302
Query: 94 YFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIR 153
YF Q+ +AA G S K T AMR LAYG+AADA+DEY+ I ST +E L RF K +++
Sbjct: 303 YFVQKRNAAGLIGFSCHQKVTAAMRQLAYGIAADALDEYLGIAESTAIESLRRFVKAVVQ 362
Query: 154 LYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGKLAFANHVRA 213
++E YLR+P ++D R+ + E RGFP M+GSIDCMHW+WKN P G + HV
Sbjct: 363 VFEHEYLRSPNENDTTRLLELGEDRGFPGMLGSIDCMHWKWKNCPTELHG--MYQGHVHE 420
Query: 214 RSRVASRITSR 224
+ + + S+
Sbjct: 421 PTIILEAVASK 431
>UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa]
Length = 440
Score = 171 bits (432), Expect = 7e-41
Identities = 84/216 (38%), Positives = 127/216 (57%), Gaps = 8/216 (3%)
Query: 14 DVEDTYIVNRFI-----QRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDA 68
D +D YI + +R K+ GS R+ +RD A N R++ DYFA+ P Y
Sbjct: 30 DDDDYYIAAALLTDIEHKRTKRKHHGSVPG-REIIHRDRFAGNLRIVADYFADPPIYSAK 88
Query: 69 MFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADA 128
+FRRR+RM + +FLRIV + + D+YF Q+ +A G + L K A+RMLAY + AD+
Sbjct: 89 LFRRRFRMSRELFLRIVASVEAHDDYFRQRPNAVGLLGATALQKVYGAIRMLAYDIPADS 148
Query: 129 VDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSID 188
+DE ++I ST +E F K ++ ++ YLRAPT +D R+ ++ RGFP M+G ID
Sbjct: 149 LDEVVRISESTMIEAFKHFVKAVVDVFADQYLRAPTAEDTARLMAINTPRGFPGMLGCID 208
Query: 189 CMHWEWKNYPKAWEGKLAFANHVRARSRVASRITSR 224
CMHW WKN P W+G+ ++ HV + + + S+
Sbjct: 209 CMHWRWKNCPTGWKGQ--YSGHVDGPTMILEAVASK 242
>UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa]
Length = 916
Score = 171 bits (432), Expect = 7e-41
Identities = 83/191 (43%), Positives = 118/191 (61%), Gaps = 2/191 (1%)
Query: 34 GSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDN 93
G + + +R+ A + RL DYF+N PTY +FRRR RM + +FLRI+ + D+
Sbjct: 533 GGSVMGHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDD 592
Query: 94 YFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIR 153
YF Q+ +AA G S K T AMR LAYG+AADA+DEY+ I ST +E L RF K +++
Sbjct: 593 YFVQKRNAAGLIGFSCHQKVTAAMRQLAYGIAADALDEYLGIAESTAIESLRRFVKAVVQ 652
Query: 154 LYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGKLAFANHVRA 213
++E YLR+P ++D R+ + E RGFP M+GSIDCMHW+WKN P G + HV
Sbjct: 653 VFEHEYLRSPNENDTTRLLELGEDRGFPGMLGSIDCMHWKWKNCPTELHG--MYQGHVHE 710
Query: 214 RSRVASRITSR 224
+ + + S+
Sbjct: 711 PTIILEAVASK 721
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,021,479,746
Number of Sequences: 2790947
Number of extensions: 43862579
Number of successful extensions: 85058
Number of sequences better than 10.0: 130
Number of HSP's better than 10.0 without gapping: 95
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 84714
Number of HSP's gapped (non-prelim): 143
length of query: 605
length of database: 848,049,833
effective HSP length: 133
effective length of query: 472
effective length of database: 476,853,882
effective search space: 225075032304
effective search space used: 225075032304
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 78 (34.7 bits)
Medicago: description of AC135230.3


