

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
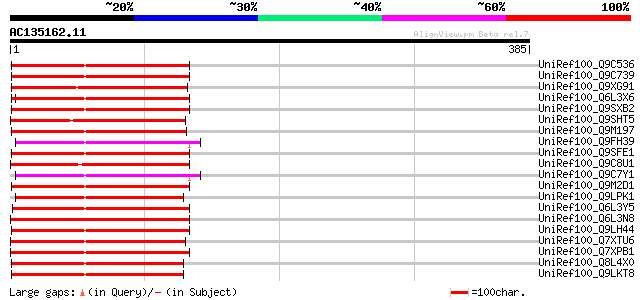
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis t... 126 9e-28
UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis t... 126 9e-28
UniRef100_Q9XG91 Tpv2-1c protein [Phaseolus vulgaris] 125 3e-27
UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum] 125 3e-27
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 124 3e-27
UniRef100_Q9SHT5 Putative retroelement pol polyprotein [Arabidop... 124 4e-27
UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [... 124 6e-27
UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana] 123 8e-27
UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana] 123 8e-27
UniRef100_Q9C8U1 Hypothetical protein T3M13.16 [Arabidopsis thal... 123 8e-27
UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [... 123 8e-27
UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana] 123 1e-26
UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana] 123 1e-26
UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum] 122 1e-26
UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum] 122 2e-26
UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidops... 122 2e-26
UniRef100_Q7XTU6 OSJNBb0034I13.10 protein [Oryza sativa] 119 2e-25
UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa] 117 5e-25
UniRef100_Q8L4X0 Putative pol polyprotein [Oryza sativa] 116 9e-25
UniRef100_Q9LKT8 Hypothetical protein T32B20.i [Arabidopsis thal... 116 9e-25
>UniRef100_Q9C536 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1320
Score = 126 bits (317), Expect = 9e-28
Identities = 61/132 (46%), Positives = 88/132 (66%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL KALYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 942 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1000
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1001 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1060
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1061 KFKMDDSNPVCT 1072
>UniRef100_Q9C739 Copia-type polyprotein, putative [Arabidopsis thaliana]
Length = 1352
Score = 126 bits (317), Expect = 9e-28
Identities = 61/132 (46%), Positives = 88/132 (66%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL KALYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1093 KFKMDDSNPVCT 1104
>UniRef100_Q9XG91 Tpv2-1c protein [Phaseolus vulgaris]
Length = 374
Score = 125 bits (313), Expect = 3e-27
Identities = 61/131 (46%), Positives = 85/131 (64%), Gaps = 1/131 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E + +L KALYGLKQ+ RAWN RI + + F +C E+ +Y KN+ N++ + LYV
Sbjct: 2 EKKVLKLKKALYGLKQAPRAWNTRIDTYFKENGFKQCPYEHALYAKNN-GGNMIFVALYV 60
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI +N IE FKG M ++F MTDLG + +FLG+E + G+ Q+KYA +ILK
Sbjct: 61 DDLIFMGNNNDMIEEFKGTMRREFEMTDLGLMKFFLGLEVRQKETGIFVSQEKYAKEILK 120
Query: 122 RFNMLNCNPAS 132
++ M NCNP S
Sbjct: 121 KYKMENCNPVS 131
>UniRef100_Q6L3X6 Putative polyprotein [Solanum demissum]
Length = 1758
Score = 125 bits (313), Expect = 3e-27
Identities = 64/132 (48%), Positives = 89/132 (66%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED ++ L KALYGLKQS RAW +R+ N L+Q F + ++E +YVK + +L+ + +YV
Sbjct: 835 EDQVYLLTKALYGLKQSPRAWYERMDNHLIQLGFSRSQSEATLYVKVTAGESLI-VSIYV 893
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DD++VT S + I+ FK M K F MTDLG + YFLGME ++S+G+ Q+KY DIL
Sbjct: 894 DDMLVTGSKIELIQRFKDEMEKIFEMTDLGVMKYFLGMEVLQSSDGIFICQQKYILDILN 953
Query: 122 RFNMLNCNPAST 133
RF M +C P ST
Sbjct: 954 RFKMQDCKPVST 965
Score = 84.7 bits (208), Expect = 4e-15
Identities = 46/129 (35%), Positives = 78/129 (59%), Gaps = 3/129 (2%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
I RLHKA Y LKQ+ RA + N L+ F+K +++ +++V++ + +V++ +YVDD+
Sbjct: 1449 ICRLHKAFYRLKQAPRACYNELKNHLLLLGFIKSESDNSLFVRHHSHA-IVYLLVYVDDI 1507
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
IVT +N + + + + + DL LTYFLG+E ++ +G++ Q Y DIL+ N
Sbjct: 1508 IVTGNNTSVVNQVISSLAAR--VKDLEALTYFLGIEVIRSVDGIIMTQSTYKRDILREEN 1565
Query: 125 MLNCNPAST 133
M +C A T
Sbjct: 1566 MADCKLAKT 1574
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 124 bits (312), Expect = 3e-27
Identities = 60/132 (45%), Positives = 87/132 (65%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL K LYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1093 KFKMDDSNPVCT 1104
>UniRef100_Q9SHT5 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1307
Score = 124 bits (311), Expect = 4e-27
Identities = 60/130 (46%), Positives = 86/130 (66%), Gaps = 1/130 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
NE +++LHKALYGLKQ+ RAWN ++ L + FVKC E +VY + ++ L+ + +Y
Sbjct: 916 NEGKVYKLHKALYGLKQAPRAWNTKLNKILQELNFVKCSKEPSVY-RRQEEKKLLIVAIY 974
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDDL+VT S+L I FK M KF M+DLG+LTY+LG+E G++ Q++YA I+
Sbjct: 975 VDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLHRKNGIILRQERYAMKII 1034
Query: 121 KRFNMLNCNP 130
+ M NCNP
Sbjct: 1035 EEAGMSNCNP 1044
>UniRef100_Q9M197 Copia-type reverse transcriptase-like protein [Arabidopsis thaliana]
Length = 1272
Score = 124 bits (310), Expect = 6e-27
Identities = 59/130 (45%), Positives = 87/130 (66%), Gaps = 1/130 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL K LYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPA 131
+F M + NP+
Sbjct: 1093 KFKMDDSNPS 1102
>UniRef100_Q9FH39 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1334
Score = 123 bits (309), Expect = 8e-27
Identities = 65/157 (41%), Positives = 94/157 (59%), Gaps = 21/157 (13%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+++L KALYGLKQ+ RAW RI F ++ F KC E+ ++VK + LV + +YVDDL
Sbjct: 965 VYKLKKALYGLKQAPRAWYSRIEEFFGKEGFEKCYCEHTLFVKKERSDFLV-VSVYVDDL 1023
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I T S++ IE FK MM++F MTDLGK+ YFLG+E + G+ +Q+KYA++I+K++
Sbjct: 1024 IYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYFLGVEVIQDERGIFINQRKYAAEIIKKYG 1083
Query: 125 MLNCNPAS--------------------TLFKQIVGS 141
M CN T FKQ++GS
Sbjct: 1084 MEGCNSVKNPIVPGQKLTKAGAGDAVDPTEFKQLIGS 1120
>UniRef100_Q9SFE1 T26F17.17 [Arabidopsis thaliana]
Length = 1291
Score = 123 bits (309), Expect = 8e-27
Identities = 60/132 (45%), Positives = 87/132 (65%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL KALYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 913 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 971
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + + Q+ YA ++LK
Sbjct: 972 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNRIFITQEGYAKEVLK 1031
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1032 KFKMDDSNPVCT 1043
>UniRef100_Q9C8U1 Hypothetical protein T3M13.16 [Arabidopsis thaliana]
Length = 503
Score = 123 bits (309), Expect = 8e-27
Identities = 63/134 (47%), Positives = 92/134 (68%), Gaps = 3/134 (2%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKN-SKDSNLVHICL 59
+ED +++L+KALYGL+Q+ RAWN ++ + LV+ +F+KC E +VY K SKD L+ I +
Sbjct: 111 SEDKVYKLNKALYGLRQAPRAWNNKLNHILVELQFLKCSKEPSVYRKEVSKD--LLLIAI 168
Query: 60 YVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDI 119
YVDDL VT +NL I FK M KF M++LGKLTY+LG+E + +G+ +Q +YA I
Sbjct: 169 YVDDLFVTGTNLDIINKFKEEMSSKFEMSNLGKLTYYLGIEVIQHKDGIALNQNRYALKI 228
Query: 120 LKRFNMLNCNPAST 133
L+ M++CN T
Sbjct: 229 LEEAGMIDCNRVHT 242
>UniRef100_Q9C7Y1 Copia-type polyprotein, putative; 28768-32772 [Arabidopsis thaliana]
Length = 1334
Score = 123 bits (309), Expect = 8e-27
Identities = 65/157 (41%), Positives = 94/157 (59%), Gaps = 21/157 (13%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+++L KALYGLKQ+ RAW RI F ++ F KC E+ ++VK + LV + +YVDDL
Sbjct: 965 VYKLKKALYGLKQAPRAWYSRIEEFFGKEGFEKCYCEHTLFVKKERSDFLV-VSVYVDDL 1023
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I T S++ IE FK MM++F MTDLGK+ YFLG+E + G+ +Q+KYA++I+K++
Sbjct: 1024 IYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYFLGVEVIQDERGIFINQRKYAAEIIKKYG 1083
Query: 125 MLNCNPAS--------------------TLFKQIVGS 141
M CN T FKQ++GS
Sbjct: 1084 MEGCNSVKNPIVPGQKLTKAGAGDAVDPTEFKQLIGS 1120
>UniRef100_Q9M2D1 Copia-type polyprotein [Arabidopsis thaliana]
Length = 1352
Score = 123 bits (308), Expect = 1e-26
Identities = 59/132 (44%), Positives = 87/132 (65%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL K LYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPAST 133
+F + + NP T
Sbjct: 1093 KFKIDDSNPVCT 1104
>UniRef100_Q9LPK1 F6N18.1 [Arabidopsis thaliana]
Length = 1207
Score = 123 bits (308), Expect = 1e-26
Identities = 59/125 (47%), Positives = 86/125 (68%), Gaps = 1/125 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+++L KALYGLKQ+ RAW RI F ++ F KC E+ ++VK + LV + +YVDDL
Sbjct: 870 VYKLKKALYGLKQAPRAWYSRIEEFFGKEGFEKCYCEHTLFVKKERSDFLV-VSVYVDDL 928
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I T S++ IE FK MM++F MTDLGK+ YFLG+E + G+ +Q+KYA++I+K++
Sbjct: 929 IYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYFLGVEVIQDERGIFINQRKYAAEIIKKYG 988
Query: 125 MLNCN 129
M CN
Sbjct: 989 MEGCN 993
>UniRef100_Q6L3Y5 Putative gag-pol polyprotein [Solanum demissum]
Length = 1133
Score = 122 bits (307), Expect = 1e-26
Identities = 63/131 (48%), Positives = 87/131 (66%), Gaps = 1/131 (0%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D ++ L KALYGLKQ+ RAW RI ++L+ F + E +Y++ D L+ + LYVD
Sbjct: 758 DQVYLLRKALYGLKQAPRAWYARIDSYLLNLGFKRSCNETTLYIRLQNDDFLL-VSLYVD 816
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
DL+VT S+ I+ FK M K F M DLGK+TYFLGME ++S+G+ QKKYA++ILK+
Sbjct: 817 DLLVTGSSTQSIKDFKEQMKKVFEMNDLGKMTYFLGMEVNQSSQGIFVSQKKYATEILKK 876
Query: 123 FNMLNCNPAST 133
F + C P ST
Sbjct: 877 FCLDKCKPVST 887
>UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum]
Length = 1333
Score = 122 bits (306), Expect = 2e-26
Identities = 58/133 (43%), Positives = 85/133 (63%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
NE+ +++L KALYGLKQ+ RAW +I +F F + E +Y+K + +CLY
Sbjct: 954 NENKVYKLRKALYGLKQAPRAWYSKIDSFFQGSGFRRSDNEPTLYLKKQGTDEFLLVCLY 1013
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDD+I S+ + + FK +MM+ F M+DLG L YFLG+E + +G+ QKKYA D+L
Sbjct: 1014 VDDMIYIGSSKSLVNDFKSNMMRNFEMSDLGLLKYFLGLEVIQDKDGIFISQKKYAEDLL 1073
Query: 121 KRFNMLNCNPAST 133
K+F M+NC A+T
Sbjct: 1074 KKFQMMNCEVATT 1086
>UniRef100_Q9LH44 Copia-like retrotransposable element [Arabidopsis thaliana]
Length = 1499
Score = 122 bits (305), Expect = 2e-26
Identities = 63/132 (47%), Positives = 83/132 (62%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E + +LHKALYGLKQ+ RAW RI + ++ F + + YVK + LV + LYV
Sbjct: 977 EGKVLKLHKALYGLKQAPRAWYGRIDGYFIKNGFERSINDAAFYVKKTSKEILV-VSLYV 1035
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DD+IVT SN+ EIE FK M +F MTDLG+L+YFLGME + EG+ Q+ YA +LK
Sbjct: 1036 DDIIVTGSNVKEIERFKEEMKNEFEMTDLGELSYFLGMEVNQDDEGIFLSQENYAKKLLK 1095
Query: 122 RFNMLNCNPAST 133
+F M C ST
Sbjct: 1096 KFGMQECKSVST 1107
>UniRef100_Q7XTU6 OSJNBb0034I13.10 protein [Oryza sativa]
Length = 1425
Score = 119 bits (297), Expect = 2e-25
Identities = 61/130 (46%), Positives = 84/130 (63%), Gaps = 1/130 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
N + +L KALYGL+Q+ RAWN ++ N L+ +F K E VYV+ DS L+ + +Y
Sbjct: 1049 NASKVLKLRKALYGLRQAPRAWNAKLDNTLLSLKFNKSATESAVYVRGVGDSKLI-VGVY 1107
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDDLI+T S EI+AFK M ++FNM+DLG L+Y+LGME + EG+ Q YA IL
Sbjct: 1108 VDDLIITGSQKKEIDAFKLQMKQRFNMSDLGFLSYYLGMEVVQKGEGIFLSQSAYAGKIL 1167
Query: 121 KRFNMLNCNP 130
++ M CNP
Sbjct: 1168 EKTGMEGCNP 1177
>UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa]
Length = 1449
Score = 117 bits (293), Expect = 5e-25
Identities = 57/133 (42%), Positives = 88/133 (65%), Gaps = 1/133 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
+++ ++RLHKALYGL+Q+ RAWN ++ + L+ F + +E+ VY + L+ + +Y
Sbjct: 1069 HKNKVYRLHKALYGLRQAPRAWNAKLDSSLLSLGFHRSSSEHGVYTRTRGGRRLM-VGVY 1127
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDDLI+T + EI +FKG MMK F M+DLG L Y+LG+E T+ S+G+ Q YA IL
Sbjct: 1128 VDDLIITGDHDDEIRSFKGEMMKLFKMSDLGALRYYLGIEVTQDSDGITLGQAAYAGKIL 1187
Query: 121 KRFNMLNCNPAST 133
++ + +CNP T
Sbjct: 1188 EKAGLKDCNPCQT 1200
>UniRef100_Q8L4X0 Putative pol polyprotein [Oryza sativa]
Length = 1426
Score = 116 bits (291), Expect = 9e-25
Identities = 58/128 (45%), Positives = 84/128 (65%), Gaps = 1/128 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E + RL KALYGLKQ+ RAWN ++ N L+ F+K + E VYV+ + S L+ + +YV
Sbjct: 1051 EGQVLRLKKALYGLKQAPRAWNAKLHNTLISLNFIKSETESAVYVRGTGSSRLI-VGVYV 1109
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI++ + +EI+ FK M KKF M+DLG L+Y+LGME + +G+ Q YA+ IL+
Sbjct: 1110 DDLIISGAQASEIDFFKEEMKKKFRMSDLGLLSYYLGMEVVQKDDGVFLSQTAYAAKILE 1169
Query: 122 RFNMLNCN 129
+ M CN
Sbjct: 1170 KTGMEGCN 1177
>UniRef100_Q9LKT8 Hypothetical protein T32B20.i [Arabidopsis thaliana]
Length = 724
Score = 116 bits (291), Expect = 9e-25
Identities = 56/128 (43%), Positives = 86/128 (66%), Gaps = 1/128 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED +++L+KALYGLKQ+ RAWN ++ L++ +FVKC E ++Y K + L+ + +YV
Sbjct: 574 EDQVYKLNKALYGLKQAPRAWNHKLNQILMELKFVKCSKEPSLYRKGENEDLLI-VAVYV 632
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDL+VT S+L I FK + KF M+DLG+LTY+LG+E + G++ Q++Y + IL+
Sbjct: 633 DDLLVTRSSLRMILEFKKEVSTKFEMSDLGRLTYYLGIEVLQHENGIILSQERYVNKILE 692
Query: 122 RFNMLNCN 129
M CN
Sbjct: 693 ETKMDGCN 700
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 455,412,649
Number of Sequences: 2790947
Number of extensions: 15341052
Number of successful extensions: 106982
Number of sequences better than 10.0: 1184
Number of HSP's better than 10.0 without gapping: 1034
Number of HSP's successfully gapped in prelim test: 150
Number of HSP's that attempted gapping in prelim test: 105030
Number of HSP's gapped (non-prelim): 1305
length of query: 385
length of database: 848,049,833
effective HSP length: 129
effective length of query: 256
effective length of database: 488,017,670
effective search space: 124932523520
effective search space used: 124932523520
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 76 (33.9 bits)
Medicago: description of AC135162.11


