

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.3 - phase: 0
(239 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
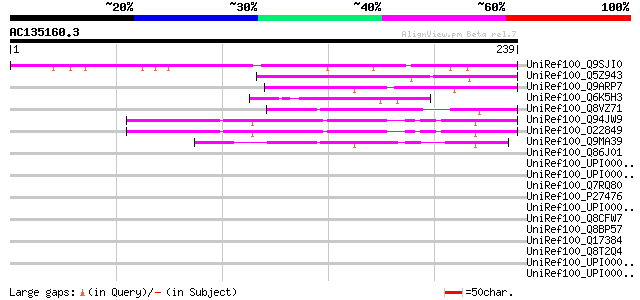
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SJI0 Hypothetical protein At2g42760 [Arabidopsis tha... 150 2e-35
UniRef100_Q5Z943 Hypothetical protein P0659D09.34 [Oryza sativa] 111 2e-23
UniRef100_Q9ARP7 OSJNBa0010K01.19 protein [Oryza sativa] 65 1e-09
UniRef100_Q6K5H3 Hypothetical protein OJ1791_B03.25 [Oryza sativa] 52 1e-05
UniRef100_Q8VZ71 Hypothetical protein At2g31560 [Arabidopsis tha... 52 2e-05
UniRef100_Q94JW9 Hypothetical protein At2g43340; T1O24.8 [Arabid... 48 2e-04
UniRef100_O22849 Hypothetical protein At2g43340 [Arabidopsis tha... 48 2e-04
UniRef100_Q9MA39 T20M3.14 protein [Arabidopsis thaliana] 47 3e-04
UniRef100_Q86J01 Hypothetical protein [Dictyostelium discoideum] 40 0.051
UniRef100_UPI000049A3A5 UPI000049A3A5 UniRef100 entry 40 0.066
UniRef100_UPI00003379F7 UPI00003379F7 UniRef100 entry 38 0.19
UniRef100_Q7RQ80 Subtilisin-like protease 2 [Plasmodium yoelii y... 38 0.19
UniRef100_P27476 Nuclear localization sequence binding protein [... 38 0.19
UniRef100_UPI00002EA5E5 UPI00002EA5E5 UniRef100 entry 38 0.25
UniRef100_Q8CFW7 RIKEN cDNA 5730509K17 gene [Mus musculus] 38 0.25
UniRef100_Q8BP57 Mus musculus 8 days embryo whole body cDNA, RIK... 38 0.25
UniRef100_Q17384 Hum-3 [Caenorhabditis elegans] 38 0.25
UniRef100_Q8T2Q4 Hypothetical protein [Dictyostelium discoideum] 38 0.25
UniRef100_UPI0000499A95 UPI0000499A95 UniRef100 entry 37 0.33
UniRef100_UPI00002E7E5C UPI00002E7E5C UniRef100 entry 37 0.33
>UniRef100_Q9SJI0 Hypothetical protein At2g42760 [Arabidopsis thaliana]
Length = 267
Score = 150 bits (380), Expect = 2e-35
Identities = 114/271 (42%), Positives = 151/271 (55%), Gaps = 37/271 (13%)
Query: 1 MSAEQILTLLDSFWFETTI-------LTNKNPSK--VEEILPQ---DTNLLLVPTPNLDV 48
M+ E++L L + W E I L K+ K +EIL + + L P L
Sbjct: 1 MAGEELLKLFEQNWSERPIFKKDKENLNGKSREKRGEKEILEERREEEALKNFPVSFLVE 60
Query: 49 RFYSEQNLSIIGS---VFSDSP-----SPNSVL----TSSKLRTIPSEREFGEFSTGTSS 96
R S++ + S +FS S SP SVL T KL+TI S +E F+
Sbjct: 61 RAMSDETMMTTSSKTSLFSSSSDDLFLSPRSVLPVKPTPMKLQTILSGKEVNAFTIAERE 120
Query: 97 ENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEED-KDSKLVS 155
+KE+ KKK+S R RK +S+S+LE++ELKGFMDLGFVFSE+D KDS LVS
Sbjct: 121 RLLSEKEEQRKKKKKKSNV---RTRKGKSMSDLEYEELKGFMDLGFVFSEDDHKDSDLVS 177
Query: 156 LIPGLQRLGRENDDA--EEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERK---NPLV 210
++PGLQRL +++D EE E+EE KI N + +PYLSEAWD R+ K P +
Sbjct: 178 ILPGLQRLVKKDDGVTKEEEEEEEEDKIGGNRAA--RPYLSEAWDHCGGRKGKKQITPEI 235
Query: 211 NWRV--PDKGSEIDMKDNLKFWAHAVASIAR 239
WRV P SE+D+KDNL+ WAHAVAS R
Sbjct: 236 KWRVPAPAAASEVDLKDNLRLWAHAVASTIR 266
>UniRef100_Q5Z943 Hypothetical protein P0659D09.34 [Oryza sativa]
Length = 283
Score = 111 bits (277), Expect = 2e-23
Identities = 60/143 (41%), Positives = 81/143 (55%), Gaps = 21/143 (14%)
Query: 117 RRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDE 176
RRR+R+ RS+SELEF+E+KG DLGF FSE+D D++L S++PGL+R + DDA E
Sbjct: 142 RRRQRRGRSMSELEFEEVKGLQDLGFTFSEDDVDAELASIVPGLRRRRSDEDDAREAPAA 201
Query: 177 EHKKIDENVLSD------------------NKPYLSEAWDVFDQRERKNPLVNWRVP--D 216
+E S +PYLSEAWD ++ E + L NWR+P
Sbjct: 202 AAASAEEEAASSRRIGSAPAGTSSSFSSAPRRPYLSEAWD-DEEEEMRRMLRNWRIPPAG 260
Query: 217 KGSEIDMKDNLKFWAHAVASIAR 239
G D+K+ L+ WAH VAS R
Sbjct: 261 DGDGADLKEQLRLWAHTVASAVR 283
>UniRef100_Q9ARP7 OSJNBa0010K01.19 protein [Oryza sativa]
Length = 383
Score = 65.5 bits (158), Expect = 1e-09
Identities = 44/130 (33%), Positives = 65/130 (49%), Gaps = 14/130 (10%)
Query: 121 RKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQ--------RLGRENDDAEE 172
+K +S S+LE E++GF DLGFVF +E+ L ++PGL+ +NDDA
Sbjct: 254 KKWKSSSDLESIEVQGFRDLGFVFDQEELRESLADVLPGLRGKPTPTGSGSASDNDDANT 313
Query: 173 GEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNP---LVNWRVPDKGSEIDMKDNLKF 229
+ V + +PYLSEAW + ++P + + D S +MKD L+
Sbjct: 314 ATTATG---SDAVAAVRRPYLSEAWYHVRRPAPRSPAAAAMRLQQADARSAAEMKDQLRM 370
Query: 230 WAHAVASIAR 239
WA AVA R
Sbjct: 371 WAQAVACNVR 380
>UniRef100_Q6K5H3 Hypothetical protein OJ1791_B03.25 [Oryza sativa]
Length = 167
Score = 52.0 bits (123), Expect = 1e-05
Identities = 36/101 (35%), Positives = 54/101 (52%), Gaps = 21/101 (20%)
Query: 114 QSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEG 173
+S RR RR+ RS+S LE K D GF FSE+D D++LVS++ GL+R + DDA +
Sbjct: 39 RSGRRWRRRRRSMS-LELK----LQDFGFTFSEDDVDAELVSIVSGLRRQRSDEDDARKA 93
Query: 174 ---------EDEEHKKI-------DENVLSDNKPYLSEAWD 198
E+ ++I + + +P+LSE WD
Sbjct: 94 PAFAVASAEEEAAGRRIGFAPTGASSSSSTLRRPHLSEVWD 134
>UniRef100_Q8VZ71 Hypothetical protein At2g31560 [Arabidopsis thaliana]
Length = 202
Score = 51.6 bits (122), Expect = 2e-05
Identities = 34/119 (28%), Positives = 58/119 (48%), Gaps = 14/119 (11%)
Query: 122 KSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKI 181
+++SL++ + +ELKG +DLGF FS D+ +L + +P L+ + + + + H K
Sbjct: 94 RAKSLTDDDLEELKGCLDLGFGFS-YDEIPELCNTLPALELCYSMSQKFLDDKQQNHHKS 152
Query: 182 DENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSE-IDMKDNLKFWAHAVASIAR 239
E S P + P+ NW++ G + D+K LK+WA VA R
Sbjct: 153 QEEDDSSPPPTTTA------------PIANWKISSPGDDPDDVKARLKYWAQTVACTVR 199
>UniRef100_Q94JW9 Hypothetical protein At2g43340; T1O24.8 [Arabidopsis thaliana]
Length = 189
Score = 48.1 bits (113), Expect = 2e-04
Identities = 51/194 (26%), Positives = 86/194 (44%), Gaps = 24/194 (12%)
Query: 56 LSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQS-- 113
LS S +D S + + TSS SE E E + G +K+ KKK +
Sbjct: 7 LSRCSSYEADVDSDSEISTSSFSSCSYSESEEEEINNGFGVVQS-KKQTKKLEKKKSNVL 65
Query: 114 -------QSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRE 166
+ ++++SL++ + +ELKG +DLGF F+ E+ +L + +P L+
Sbjct: 66 LEGYVVDSAVNDDLKRTKSLTDDDLEELKGCVDLGFGFNYEE-IPELCNTLPALELCYSM 124
Query: 167 NDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKG-SEIDMKD 225
+ + + H S + P + V D +P+ +W++ G + D+K
Sbjct: 125 SQKFIDQDHHHH--------SSSSP--EKKSSVLD--SPVSPIASWKISSPGDNPDDVKA 172
Query: 226 NLKFWAHAVASIAR 239
LKFWA AVA R
Sbjct: 173 RLKFWAQAVACTVR 186
>UniRef100_O22849 Hypothetical protein At2g43340 [Arabidopsis thaliana]
Length = 241
Score = 47.8 bits (112), Expect = 2e-04
Identities = 51/194 (26%), Positives = 86/194 (44%), Gaps = 24/194 (12%)
Query: 56 LSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQS-- 113
LS S +D S + + TSS SE E E + G +K+ KKK +
Sbjct: 7 LSRCSSYEADVDSDSEISTSSFSSCSYSESEEEEINNGFGVVQS-KKQTKKLEKKKSNVL 65
Query: 114 -------QSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRE 166
+ ++++SL++ + +ELKG +DLGF F+ E+ +L + +P L+
Sbjct: 66 LEGYVVDSAVNDDLKRTKSLTDDDLEELKGCVDLGFGFNYEE-IPELCNTLPALELCYSM 124
Query: 167 NDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKG-SEIDMKD 225
+ + + H S + P + V D +P+ +W++ G + D+K
Sbjct: 125 SQKFIDQDHHHH--------SSSSP--EKKSSVLD--SPVSPIASWKISSPGDNPDDVKA 172
Query: 226 NLKFWAHAVASIAR 239
LKFWA AVA R
Sbjct: 173 RLKFWAQAVACTMR 186
>UniRef100_Q9MA39 T20M3.14 protein [Arabidopsis thaliana]
Length = 189
Score = 47.4 bits (111), Expect = 3e-04
Identities = 38/152 (25%), Positives = 73/152 (48%), Gaps = 34/152 (22%)
Query: 88 GEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEE 147
G T +SS QK+D+ +S+SL++ + ++L+G +DLGF FS
Sbjct: 61 GYVETASSSSVDDQKDDLT---------------RSKSLTDDDLEDLRGCLDLGFGFS-Y 104
Query: 148 DKDSKLVSLIPGLQ---RLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRE 204
D+ +L + +P L+ + ++ D ++ + E +++ + P ++
Sbjct: 105 DEIPELCNTLPALELCYSMSQKFLDDKQNKSPETSSVED---CPSPPLVT---------- 151
Query: 205 RKNPLVNWRVPDKG-SEIDMKDNLKFWAHAVA 235
P+ NW++ G + D+K LK+WA AVA
Sbjct: 152 -ATPIANWKISSPGDNPDDVKARLKYWAQAVA 182
>UniRef100_Q86J01 Hypothetical protein [Dictyostelium discoideum]
Length = 923
Score = 40.0 bits (92), Expect = 0.051
Identities = 37/116 (31%), Positives = 59/116 (49%), Gaps = 5/116 (4%)
Query: 84 EREFGEFSTGTSSENHHQKEDVAYTKKKQ--SQSYRRRRRKSRSLSELEFKELKGF-MDL 140
E E GE SSE K+ V K++ +S + SE E +E +G +
Sbjct: 448 EEESGEEEEEESSEEEQVKQPVKKPIKQEPKEESEEEESEEEEEESEEEEEESEGEESEE 507
Query: 141 GFVFSEEDKDS-KLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSE 195
SEE+KD+ + V +P L++ E+++++E EDEE DE+ SD++ LSE
Sbjct: 508 EEEESEEEKDNGRKVFRVP-LRKPTTESEESDEDEDEEESDEDEDEESDSEENLSE 562
>UniRef100_UPI000049A3A5 UPI000049A3A5 UniRef100 entry
Length = 1657
Score = 39.7 bits (91), Expect = 0.066
Identities = 29/123 (23%), Positives = 61/123 (49%), Gaps = 8/123 (6%)
Query: 97 ENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSL 156
E ++K+++ ++K+ + R+ ++ + ELE ++ K + EE+++ KL
Sbjct: 793 EEENKKKEIEENERKKKEEEERQLKEEQKKKELEEQQRKEEEERERKLEEEERERKLEEE 852
Query: 157 IPGLQRLGRENDDAEEGEDEEHKKIDENVL-----SDNKPYLSEAWDVFDQRERKNPLVN 211
I +R+ E +E E+E +K+ E L +NK + E + D++E K+ + N
Sbjct: 853 I---KRIEEEERKLQEEEEENERKLQEEELKIKQEEENKRKVVENKPLKDKKEWKDLVEN 909
Query: 212 WRV 214
V
Sbjct: 910 EEV 912
>UniRef100_UPI00003379F7 UPI00003379F7 UniRef100 entry
Length = 2099
Score = 38.1 bits (87), Expect = 0.19
Identities = 27/131 (20%), Positives = 57/131 (42%), Gaps = 2/131 (1%)
Query: 76 SKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELK 135
SK + +E + E+ SS E+++ +++ R K + +K+ K
Sbjct: 928 SKPKEGSAESDDSEYEKSDSSAESGMSEELSESEETGKHRKTRAATKKDEEKQRSYKQKK 987
Query: 136 GFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSE 195
+ V + K+ + LQ+ G E+D+ ++G+D + +K +L D+K +E
Sbjct: 988 KRRRIK-VQESSSSNEKVFEMFQTLQKSGEEDDEQQDGDDRKGRKKIRKILKDDK-LRTE 1045
Query: 196 AWDVFDQRERK 206
D + E +
Sbjct: 1046 TRDALKEEEER 1056
>UniRef100_Q7RQ80 Subtilisin-like protease 2 [Plasmodium yoelii yoelii]
Length = 1225
Score = 38.1 bits (87), Expect = 0.19
Identities = 39/155 (25%), Positives = 74/155 (47%), Gaps = 30/155 (19%)
Query: 100 HQKEDVAYTKKKQ----SQSYRRRR-------RKSRSLSELEFKELKG-FMDLGFVFSEE 147
H+K+D+A KK++ QS RR R S L +++F EL + ++G S++
Sbjct: 76 HKKKDIADKKKRRRYGNQQSIENRRIAEENERRLSNQLDDIQFIELSNKYPNIGKQNSQQ 135
Query: 148 DKDSKLVSLIPGLQRLG--RENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQ--- 202
+K +K+ + R ++D +EGEDE+ D++++ K L E D+ ++
Sbjct: 136 NKVNKINNQNGASNSNDNIRNDEDEDEGEDEDE---DDDLIEGRKDNLEED-DLVEKNGA 191
Query: 203 ---------RERKNPLVNWRVPDKGSEIDMKDNLK 228
+E KN +N ++ + ++ DN K
Sbjct: 192 NLKRGNMHGQEEKNKNINTTPGNENNSKNVNDNKK 226
>UniRef100_P27476 Nuclear localization sequence binding protein [Saccharomyces
cerevisiae]
Length = 414
Score = 38.1 bits (87), Expect = 0.19
Identities = 32/114 (28%), Positives = 51/114 (44%), Gaps = 24/114 (21%)
Query: 64 SDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKS 123
S+S S +S + S+ T E + S+ SS + ++E+ TKK++S+ S
Sbjct: 64 SESSSSSSSDSESEAETKKEESKDSSSSSSDSSSDEEEEEEKEETKKEESKESSSSDSSS 123
Query: 124 RSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEE 177
S S+ E ++ EE D K R+++DAEE EDEE
Sbjct: 124 SSSSDSESEK------------EESNDKK------------RKSEDAEEEEDEE 153
>UniRef100_UPI00002EA5E5 UPI00002EA5E5 UniRef100 entry
Length = 260
Score = 37.7 bits (86), Expect = 0.25
Identities = 46/171 (26%), Positives = 68/171 (38%), Gaps = 25/171 (14%)
Query: 21 TNKNPSKVEEILPQDTNLLLVPTPNLDVRFYSEQNLSIIG----SVFSDSPSPNSVLTSS 76
+NK+ K + D + PTP R + E S S S P+S S+
Sbjct: 25 SNKSTPKEDHSKEDDRSADRTPTPVSRSRSHKESPRSSKSHSKTSRTSSERKPSSQTPST 84
Query: 77 KLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKG 136
+ R++P ER +G+S + +HQ + K SQ RR K RS S F +
Sbjct: 85 RTRSVPEERT----QSGSSFDEYHQS-----STKASSQGQRRNVPKQRSSSYTTFARAET 135
Query: 137 FMDLGFVFSEEDKDSKLV--------SLIPGLQRLGRENDDAEEGEDEEHK 179
F E ++ S +V S+ GLQ L D E++ HK
Sbjct: 136 TPSKRDQFPETEEMSSVVDTNELVSISVSSGLQSL----RDMFFDENDSHK 182
>UniRef100_Q8CFW7 RIKEN cDNA 5730509K17 gene [Mus musculus]
Length = 1633
Score = 37.7 bits (86), Expect = 0.25
Identities = 32/132 (24%), Positives = 57/132 (42%), Gaps = 3/132 (2%)
Query: 61 SVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRR 120
++ D P+P S R +E EFG + + + + +Y++ K S R+ +
Sbjct: 118 ALLRDVPTPRPRRLRSPSREKETETEFGTEPSTEVQDTQKEDDTKSYSRIKFRDSVRKIK 177
Query: 121 RKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEG--EDEEH 178
K + E + + F F E ++S+ S + G +R E+ + EEG E
Sbjct: 178 SKPQLPPGFPSAE-EAYNFFTFNFDPEPEESEEKSPVKGGERAHHEDQEGEEGTQAQERA 236
Query: 179 KKIDENVLSDNK 190
KK +E L + K
Sbjct: 237 KKTEEEELLNGK 248
>UniRef100_Q8BP57 Mus musculus 8 days embryo whole body cDNA, RIKEN full-length
enriched library, clone:5730509K17 product:hypothetical
C2 domain containing protein, full insert sequence [Mus
musculus]
Length = 541
Score = 37.7 bits (86), Expect = 0.25
Identities = 32/132 (24%), Positives = 57/132 (42%), Gaps = 3/132 (2%)
Query: 61 SVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRR 120
++ D P+P S R +E EFG + + + + +Y++ K S R+ +
Sbjct: 69 ALLRDVPTPRPRRLRSPSREKETETEFGTEPSTEVQDTQKEDDTKSYSRIKFRDSVRKIK 128
Query: 121 RKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEG--EDEEH 178
K + E + + F F E ++S+ S + G +R E+ + EEG E
Sbjct: 129 SKPQLPPGFPSAE-EAYNFFTFNFDPEPEESEEKSPVKGGERAHHEDQEGEEGTQAQERA 187
Query: 179 KKIDENVLSDNK 190
KK +E L + K
Sbjct: 188 KKTEEEELLNGK 199
>UniRef100_Q17384 Hum-3 [Caenorhabditis elegans]
Length = 846
Score = 37.7 bits (86), Expect = 0.25
Identities = 23/89 (25%), Positives = 44/89 (48%), Gaps = 8/89 (8%)
Query: 147 EDKDSKLVSLIPGLQRLGRENDDA-------EEGEDEEHKKIDENVLSDNKPYLSEAWDV 199
ED ++ S+I L +L END+ +E ED+E ++ +E + + + + W
Sbjct: 703 EDCVKRVDSIIADL-KLQLENDELAEVERARKEAEDKERREAEEKAAASRRKFCAAKWRK 761
Query: 200 FDQRERKNPLVNWRVPDKGSEIDMKDNLK 228
+R RK+ NWR + S + ++ +K
Sbjct: 762 NVRRPRKSTSCNWRCRSRSSPSEAEEEVK 790
>UniRef100_Q8T2Q4 Hypothetical protein [Dictyostelium discoideum]
Length = 595
Score = 37.7 bits (86), Expect = 0.25
Identities = 49/206 (23%), Positives = 86/206 (40%), Gaps = 37/206 (17%)
Query: 13 FWFETTILTNKN--PSKVEEILPQDTNLLLVPTPNLDVRFYSEQN----LSIIGSVFSDS 66
F + +T L N+N P V + + N L+P P + + N L I F
Sbjct: 233 FGYNSTSLINQNTIPPSVTHV---ELNEFLIPKPWIGFQSNPFPNSVTYLKINAYSFYQI 289
Query: 67 PSPNSVLT-------SSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRR- 118
P P +V T S K T PS T ++SEN +Q+ ++ K ++ +
Sbjct: 290 PVPPNVKTLIYSGNFSFKNSTFPSSSS-SSIPTSSTSENINQESIFSFPKTLKTLTLSSD 348
Query: 119 -----RRRKSRSLSELEF-------------KELKGFMDLGFVFSEEDKDSKLVSLIPGL 160
+ S+++L F K +K F+ G F++ K +L S+ P
Sbjct: 349 FNQPLKNLLPNSITKLTFGGAFNCQLENSIPKSIK-FLKFGNSFNKIIKIGELTSISPSS 407
Query: 161 QRLGRENDDAEEGEDEEHKKIDENVL 186
++ EN+D E ++ + +I+E+ L
Sbjct: 408 NQINNENNDNEINDNNNNNEIEESNL 433
>UniRef100_UPI0000499A95 UPI0000499A95 UniRef100 entry
Length = 351
Score = 37.4 bits (85), Expect = 0.33
Identities = 43/200 (21%), Positives = 85/200 (42%), Gaps = 28/200 (14%)
Query: 26 SKVEEILPQDTNLLLVPTPNLDVRFYSEQNLSIIGSVFSDSPSPNSVLTSSKLR--TIPS 83
S +E+ +P+ T +P F + S S+S S N + +++ T P+
Sbjct: 153 SPIEKTIPKTTTTNHPYSPTFQTPFSGIPQGDV--STISNSASNNHLTRDYRIKGNTPPT 210
Query: 84 EREFGEFSTGTSSENHHQKEDVAYTKKK----------QSQSYRR-------RRRKSRSL 126
E F E + ++E++AYTK K Q Q Y+ R++++S
Sbjct: 211 EGSFQEVIDQLRDQLKMKEEEIAYTKNKIMWYNDMLNIQDQRYKELENNLEYERKRTKSA 270
Query: 127 SELEFKELKGFMDLGFVFSE-EDKDSKLVSLIPG----LQRLGRENDDAEEGEDEEHKKI 181
+E E + K + V ++++++L +L+ +Q + N +E + +KI
Sbjct: 271 NENESQSKKELAKMSSVLKRYQEENTQLRNLLQDNDRKIQHIESYNSQLKEEIQTKEEKI 330
Query: 182 D--ENVLSDNKPYLSEAWDV 199
E L+D K + +D+
Sbjct: 331 KQLEETLNDRKMIIEGTFDL 350
>UniRef100_UPI00002E7E5C UPI00002E7E5C UniRef100 entry
Length = 209
Score = 37.4 bits (85), Expect = 0.33
Identities = 25/109 (22%), Positives = 50/109 (44%), Gaps = 2/109 (1%)
Query: 68 SPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLS 127
+PN T+S R +++ F E +++ + TK++ + + + K +
Sbjct: 98 TPNKTATNSN-RFEENKKSFSESENQIENDDFQVADTRKKTKERSKKEFEKDELKQKKPK 156
Query: 128 ELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDE 176
FK+ + F +E+ D L+SL L+R E++D +E ED+
Sbjct: 157 NPSFKKQNKYGKFS-EFDDEEDDISLLSLRKELEREDFEDEDEDEDEDD 204
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.311 0.130 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 412,735,875
Number of Sequences: 2790947
Number of extensions: 18341588
Number of successful extensions: 62401
Number of sequences better than 10.0: 245
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 225
Number of HSP's that attempted gapping in prelim test: 62120
Number of HSP's gapped (non-prelim): 430
length of query: 239
length of database: 848,049,833
effective HSP length: 124
effective length of query: 115
effective length of database: 501,972,405
effective search space: 57726826575
effective search space used: 57726826575
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 73 (32.7 bits)
Medicago: description of AC135160.3


