

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135100.8 - phase: 0
(420 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
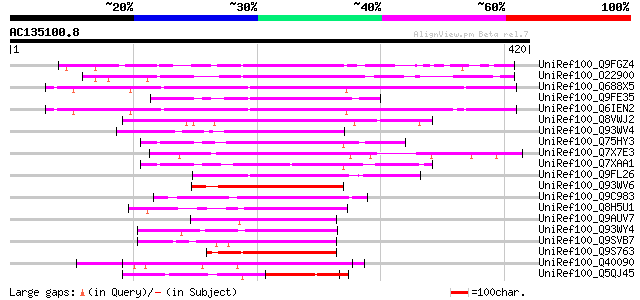
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FGZ4 Probable WRKY transcription factor 48 [Arabidop... 186 1e-45
UniRef100_O22900 Probable WRKY transcription factor 23 [Arabidop... 171 3e-41
UniRef100_Q688X5 WRKY transcription factor [Oryza sativa] 151 4e-35
UniRef100_Q9FE35 DNA-binding protein WRKY2-like [Oryza sativa] 150 7e-35
UniRef100_Q6IEN2 WRKY transcription factor 49 [Oryza sativa] 149 2e-34
UniRef100_Q8VWJ2 Probable WRKY transcription factor 28 [Arabidop... 146 9e-34
UniRef100_Q93WV4 Probable WRKY transcription factor 71 [Arabidop... 145 3e-33
UniRef100_Q75HY3 Hypothetical protein OSJNBb0035N21.13 [Oryza sa... 144 6e-33
UniRef100_Q7X7E3 WRKY3 [Oryza sativa] 137 6e-31
UniRef100_Q7XAA1 WRKY17 [Oryza sativa] 136 1e-30
UniRef100_Q9FL26 Probable WRKY transcription factor 8 [Arabidops... 128 3e-28
UniRef100_Q93WV6 Probable WRKY transcription factor 68 [Arabidop... 128 3e-28
UniRef100_Q9C983 Probable WRKY transcription factor 57 [Arabidop... 128 3e-28
UniRef100_Q8H5U1 Putative DNA-binding protein WRKY2 [Oryza sativa] 124 4e-27
UniRef100_Q9AUV7 Putative DNA binding protein [Oryza sativa] 123 1e-26
UniRef100_Q93WY4 Probable WRKY transcription factor 12 [Arabidop... 119 1e-25
UniRef100_Q9SVB7 Probable WRKY transcription factor 13 [Arabidop... 116 1e-24
UniRef100_Q9S763 Probable WRKY transcription factor 45 [Arabidop... 115 2e-24
UniRef100_Q40090 SPF1 protein [Ipomoea batatas] 115 3e-24
UniRef100_Q5QJ45 WRKY3 [Nicotiana attenuata] 114 5e-24
>UniRef100_Q9FGZ4 Probable WRKY transcription factor 48 [Arabidopsis thaliana]
Length = 399
Score = 186 bits (471), Expect = 1e-45
Identities = 141/380 (37%), Positives = 199/380 (52%), Gaps = 79/380 (20%)
Query: 40 SIFDM----IPPSQSDDQQSSSFAGYIDLLSTN----DFTSSSATLFDWFPTIDTDMSAP 91
SIFD P S +D S++ ++DLL + ++SS+ FD FP + + +
Sbjct: 69 SIFDTSSLPFPYSYFEDHSSNNPNSFLDLLRQDHQFASSSNSSSFSFDAFPLPNNNNN-- 126
Query: 92 PSTTQLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELE 151
T+ LP P +SEV+N T + S+S+SSSSN A +
Sbjct: 127 --TSFFTDLPLPQAESSEVVNTTPTSPN-STSVSSSSNEAA--------------NDNNS 169
Query: 152 GDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGY 211
G E KD++ G + Q +Q K QLK KKKN+ KK R AR F TKSD+D+LDDGY
Sbjct: 170 GKEVTVKDQEEG----DQQQEQKGTKPQLKAKKKNQ-KKAREARFAFLTKSDIDNLDDGY 224
Query: 212 RWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLR 271
RWRKYGQK VKNSP+PRSYYRCT C VKKR+ERS+ D SIV+T+YEG H H P+ R
Sbjct: 225 RWRKYGQKAVKNSPYPRSYYRCTTVGCGVKKRVERSSDDPSIVMTTYEGQHTHPFPMTPR 284
Query: 272 AANLGIMSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMSQSQQYQKFQDQNQNVLHHQR 331
++G+++ P + H TT+ S+ + ++Q HHQ
Sbjct: 285 -GHIGMLTSP------ILDHGATTASSSSFSIPQPRYLLTQ---------------HHQ- 321
Query: 332 QAVPSLLYNSDYNNSFNISPLSNIVNSADNLMIS---STSFGGFLQNQENYGGGNSSLSS 388
P +YN NNS ++ S+D ++ S+SF GF G + S +S
Sbjct: 322 ---PYNMYN---NNSLSMINR----RSSDGTFVNPGPSSSFPGF--------GYDMSQAS 363
Query: 389 SMMTSTNDMVRNNGLLQDMI 408
TST+ +R++GLLQD++
Sbjct: 364 ---TSTSSSIRDHGLLQDIL 380
>UniRef100_O22900 Probable WRKY transcription factor 23 [Arabidopsis thaliana]
Length = 337
Score = 171 bits (434), Expect = 3e-41
Identities = 128/364 (35%), Positives = 175/364 (47%), Gaps = 83/364 (22%)
Query: 60 GYIDLLSTN---DFTSSSATLF---DWFPTIDTDMSAPPSTTQLPPLPAPSPAASEV--- 110
G+++LLS+ DF + S F P T SA S++ + P+ S+V
Sbjct: 34 GFMELLSSQQHQDFATVSPHSFLLQTSQPQTQTQPSAKLSSSIIQAPPSEQLVTSKVESL 93
Query: 111 ------LNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHGI 164
+N PATP S SSISS+S+ A +K E+NE E E+
Sbjct: 94 CSDHLLINPPATPNS--SSISSASSEALNEEKPKTEDNEEEGGED--------------- 136
Query: 165 RLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNS 224
Q ++ KKQLK KK N+ K+ R ARV F TKS+VDHL+DGYRWRKYGQK VKNS
Sbjct: 137 -----QQEKSHTKKQLKAKKNNQ-KRQREARVAFMTKSEVDHLEDGYRWRKYGQKAVKNS 190
Query: 225 PFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAANLGIMSDPSGF 284
PFPRSYYRCT +C VKKR+ERS D S V+T+YEG H H+SP+ R + G SG
Sbjct: 191 PFPRSYYRCTTASCNVKKRVERSFRDPSTVVTTYEGQHTHISPLTSRPISTGGFFGSSGA 250
Query: 285 QEQVHQHHNTTSGVVGSSVLGNEFAMSQSQQYQKFQDQNQNVLHHQRQAVPSLLYNSDYN 344
+ +G G + G+ Q QQ ++ HHQ+Q
Sbjct: 251 ASSL------GNGCFGFPIDGSTLISPQFQQLVQY--------HHQQQ------------ 284
Query: 345 NSFNISPLSNIVNSADNLMISSTSFGGFLQNQENYGGGNSSLSSSMMTSTNDMVRNNGLL 404
LM +L + N G ++ + S + +V++NGLL
Sbjct: 285 --------------QQELMSCFGGVNEYLNSHANEYGDDNRVKKSRV-----LVKDNGLL 325
Query: 405 QDMI 408
QD++
Sbjct: 326 QDVV 329
>UniRef100_Q688X5 WRKY transcription factor [Oryza sativa]
Length = 419
Score = 151 bits (381), Expect = 4e-35
Identities = 123/399 (30%), Positives = 187/399 (46%), Gaps = 23/399 (5%)
Query: 30 FPFSPALPSCSIFDMIPPSQS-----DDQQSSSFAGYIDLLSTNDFTSSSATLFDWFPTI 84
FPF L S+F PP+ + QQ SF Y L ++ ++ F
Sbjct: 11 FPFHDEL--ASLFAERPPNGAMPGMLQQQQPWSFIDYHHHLMQESAPTTPPLDYEAFAGE 68
Query: 85 DTDMSAPPSTTQ----------LPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVA 134
D AP + P A + AA+ + P TP S+S S S+SS A GV
Sbjct: 69 FDDDVAPLEEVKRELVVDGVGLFPGGGASAAAAAAAVAGPMTPNSMSVS-STSSEACGVG 127
Query: 135 DKVMV-EENEHEHEEELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRP 193
EE+ + ++E EGD ++ ++ + + +D+ KK K K K + + +P
Sbjct: 128 GGAGGDEESAGKCKKEEEGDGGDDDGKEGSSTTKGDGDGEDKNKKGGKGKGKGEKRPRQP 187
Query: 194 ARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSI 253
R F TKS+VDHL+DGYRWRKYGQK VKNSPFPRSYYRCT C VKKR+ERS D+++
Sbjct: 188 -RFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTTQKCPVKKRVERSYQDAAV 246
Query: 254 VLTSYEGHHIHLSPVLLR--AANLGIMSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMS 311
V+T+YEG H H P LR A LG + + H + VG +
Sbjct: 247 VITTYEGKHTHPIPATLRGTAHLLGAAAAAHHHGGLQYHHPGHFAAAVGHRLPPQPHDAL 306
Query: 312 QSQQYQKFQDQNQNVLHHQRQAVPSLLYNSDYNNSFNISPLSNIVNSADNLMISSTSFGG 371
Q+ + + HQ Q + + ++ + + ++ +I+ST+ G
Sbjct: 307 GGGLLAPPHAQHLHAMQHQMQLAAAAAASGGSLHAAAMQQMPQPDHAGLVAIIASTT-GA 365
Query: 372 FLQNQENYGGGNSSLSSSMMTSTNDMVRNNGLLQDMIAP 410
G+++ +++ + + M ++ GLLQDM P
Sbjct: 366 STTPPPPPATGSAAAATTPLRMQHFMAQDYGLLQDMFIP 404
>UniRef100_Q9FE35 DNA-binding protein WRKY2-like [Oryza sativa]
Length = 379
Score = 150 bits (379), Expect = 7e-35
Identities = 87/186 (46%), Positives = 107/186 (56%), Gaps = 22/186 (11%)
Query: 115 ATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHGIRLVENQNDQD 174
ATP S +S SS A G ++ + + EEE ++KDE+ G +
Sbjct: 129 ATPNSSASFSSSDGEAEGGKSSRRCKKGQAKAEEE------DDKDEEDG----------E 172
Query: 175 QMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCT 234
KK KPKKK + ++ +P RV F TKS+VDHL+DGYRWRKYGQK VKNSP+PRSYYRCT
Sbjct: 173 NSKKPNKPKKKAEKRQRQP-RVAFLTKSEVDHLEDGYRWRKYGQKAVKNSPYPRSYYRCT 231
Query: 235 AGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAANLGIMSDPSGFQEQVHQHHNT 294
C VKKR+ERS D S V+T+YEG H H SP LR G+ G H HH
Sbjct: 232 TPKCGVKKRVERSYQDPSTVITTYEGQHTHHSPASLRGGGGGV-----GIVGGHHHHHLF 286
Query: 295 TSGVVG 300
GV G
Sbjct: 287 MPGVHG 292
>UniRef100_Q6IEN2 WRKY transcription factor 49 [Oryza sativa]
Length = 418
Score = 149 bits (375), Expect = 2e-34
Identities = 127/400 (31%), Positives = 187/400 (46%), Gaps = 26/400 (6%)
Query: 30 FPFSPALPSCSIFDMIPPSQS-----DDQQSSSFAGYIDLLSTNDFTSSSATLFDWFPTI 84
FPF L S+F PP+ + QQ SF Y L ++ ++ F
Sbjct: 11 FPFHDEL--ASLFAERPPNGAMPGMLQQQQPWSFIDYHHHLMQESAPTTPPLDYEAFAGE 68
Query: 85 DTDMSAPPSTTQ----------LPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVA 134
D AP + P A + AA+ + P TP S+S S S+SS A GV
Sbjct: 69 FDDDVAPLEEVKRELVVDGVGLFPGGGASAAAAAAAVAGPMTPNSMSVS-STSSEACGVG 127
Query: 135 DKVMV-EENEHEHEEELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRP 193
EE+ + ++E EGD ++ ++ + + +D+ KK K K K + + +P
Sbjct: 128 GGAGGDEESAGKCKKEEEGDGGDDDGKEGSSTTKGDGDGEDKNKKGGKGKGKGEKRPRQP 187
Query: 194 ARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSI 253
R F TKS+VDHL+DGYRWRKYGQK VKNSPFPRSYYRCT C VKKR+ERS D+++
Sbjct: 188 -RFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTTQKCPVKKRVERSYQDAAV 246
Query: 254 VLTSYEGHHIHLSPVLLR--AANLGIMSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMS 311
V+T+YEG H H P LR A LG + + H + VG +
Sbjct: 247 VITTYEGKHTHPIPATLRGTAHLLGAAAAAHHHGGLQYHHPGHFAAAVGHRLPPQPHDAL 306
Query: 312 QSQQYQKFQDQNQNVLHHQRQ-AVPSLLYNSDYNNSFNISPLSNIVNSADNLMISSTSFG 370
Q+ + + HQ Q A + S + + P + A +I+ST+ G
Sbjct: 307 GGGLLAPPHAQHLHAMQHQMQLAAAAASGGSLHAAAMQQMPQPDHAGLA--AIIASTT-G 363
Query: 371 GFLQNQENYGGGNSSLSSSMMTSTNDMVRNNGLLQDMIAP 410
G+++ +++ + + M ++ GLLQDM P
Sbjct: 364 ASTTPAPPPATGSAAAAATPLRMQHFMAQDYGLLQDMFIP 403
>UniRef100_Q8VWJ2 Probable WRKY transcription factor 28 [Arabidopsis thaliana]
Length = 318
Score = 146 bits (369), Expect = 9e-34
Identities = 98/276 (35%), Positives = 138/276 (49%), Gaps = 27/276 (9%)
Query: 92 PSTTQLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEE-------NEH 144
PST SPAA E L +S SSS +S+ ++ + + NE
Sbjct: 44 PSTYSFTDCLQSSPAAYESLLQKTFGLSPSSSEVFNSSIDQEPNRDVTNDVINGGACNET 103
Query: 145 EHE---EELEGDEAEEKDEDHGI--RLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFK 199
E EA+ ED G R E ++DQ+ K++ KK + KK R RV+F
Sbjct: 104 ETRVSPSNSSSSEADHPGEDSGKSRRKRELVGEEDQISKKVGKTKKTEVKKQREPRVSFM 163
Query: 200 TKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYE 259
TKS+VDHL+DGYRWRKYGQK VKNSP+PRSYYRCT C VKKR+ERS D ++V+T+YE
Sbjct: 164 TKSEVDHLEDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSFQDPTVVITTYE 223
Query: 260 GHHIHLSPVLLRAANLGI------MSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMSQS 313
G H H P LR ++ + P F + + T+G GS ++ QS
Sbjct: 224 GQHNHPIPTNLRGSSAAAAMFSADLMTPRSFAHDMFRTAAYTNG--GSVAAALDYGYGQS 281
Query: 314 QQYQKFQDQNQNVLHHQ-------RQAVPSLLYNSD 342
+ + + ++HQ R+ PS+ + +
Sbjct: 282 GYGSVNSNPSSHQVYHQGGEYELLREIFPSIFFKQE 317
>UniRef100_Q93WV4 Probable WRKY transcription factor 71 [Arabidopsis thaliana]
Length = 282
Score = 145 bits (365), Expect = 3e-33
Identities = 79/185 (42%), Positives = 108/185 (57%), Gaps = 17/185 (9%)
Query: 87 DMSAPPSTTQLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEH 146
D ++ + P +P + S +NNP ++ +S + SSS+ G +EN ++
Sbjct: 32 DYNSLEKVFKFSPYSSPFQSVSPSVNNPYLNLTSNSPVVSSSSNEGEP-----KENTNDK 86
Query: 147 EEELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDH 206
+++E +E + HG+ + K+ K KK KK R RV F TKS++DH
Sbjct: 87 SDQMEDNEGDL----HGVG--------ESSKQLTKQGKKKGEKKEREVRVAFMTKSEIDH 134
Query: 207 LDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLS 266
L+DGYRWRKYGQK VKNSP+PRSYYRCT C VKKR+ERS D SIV+T+YEG H H
Sbjct: 135 LEDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSFQDPSIVITTYEGKHNHPI 194
Query: 267 PVLLR 271
P LR
Sbjct: 195 PSTLR 199
>UniRef100_Q75HY3 Hypothetical protein OSJNBb0035N21.13 [Oryza sativa]
Length = 337
Score = 144 bits (362), Expect = 6e-33
Identities = 96/224 (42%), Positives = 120/224 (52%), Gaps = 30/224 (13%)
Query: 107 ASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHGIRL 166
A V PATP S S +SSSS AAG D ++ +EE EE+ +D G +
Sbjct: 103 AGGVAAAPATPNS--SVLSSSSEAAGGDDLRRCKKGRRPEDEE------EEEIDDEGSAV 154
Query: 167 VENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPF 226
Q K K K K KK R RV F TKS+VDHL+DGYRWRKYGQK VKNS +
Sbjct: 155 --------QSCKTNKMKNKKGAKKEREPRVAFMTKSEVDHLEDGYRWRKYGQKAVKNSSY 206
Query: 227 PRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPV---------LLRAANLGI 277
PRSYYRCTA C VKKR+ERS D S+V+T+YEG H H SPV L+ + G+
Sbjct: 207 PRSYYRCTAPRCGVKKRVERSEQDPSMVITTYEGQHTHPSPVSYHMHRQQGLMHVSARGV 266
Query: 278 MSDPSGFQEQVHQHHNTTSGVVG-SSVLGNEFAMSQSQQYQKFQ 320
M +G +Q ++G L M+ +QQ Q+ Q
Sbjct: 267 MPGAAG----AYQFGAPPPPLLGFDEALAARVRMTMNQQQQQQQ 306
>UniRef100_Q7X7E3 WRKY3 [Oryza sativa]
Length = 565
Score = 137 bits (345), Expect = 6e-31
Identities = 109/335 (32%), Positives = 148/335 (43%), Gaps = 67/335 (20%)
Query: 114 PATPVSVSSSISSSSNAAGVADK--VMVEENEHEHEEELEGDEAEEKDEDHGIRLVENQN 171
P TP + SS +SSS G E E + +E E K+E G E++
Sbjct: 261 PLTPNTTSSMSTSSSEGVGGGGGGGAGAGAGEEESPARCKKEEDENKEEGKG---EEDEG 317
Query: 172 DQDQMKKQLKPKKKNKN-KKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSY 230
+++ K K K K+ R R F TKS+VDHL+DGYRWRKYGQK VKNSP+PRSY
Sbjct: 318 HKNKKGSAAKGGKAGKGEKRARQPRFAFMTKSEVDHLEDGYRWRKYGQKAVKNSPYPRSY 377
Query: 231 YRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAAN--LGIMSDPSGFQEQV 288
YRCT C VKKR+ERS D ++V+T+YEG H H P LR + L + +
Sbjct: 378 YRCTTQKCPVKKRVERSYQDPAVVITTYEGKHTHPIPATLRGSTHLLAAHAQAAAAAAAA 437
Query: 289 HQ--HHNTTSGVVGSSVLGNEFAMSQSQQYQKFQDQNQNVLHHQRQAVPSLLYN---SDY 343
HQ HH+ G H A P L + + +
Sbjct: 438 HQLHHHHGHHG-------------------------------HHGMAPPLPLGSGAAAQF 466
Query: 344 NNSFNISPLSNIVNSADNLMISSTSFGGF---------LQNQENYGGGNSSLSSSMMT-- 392
S I LS+ + A T+ GG L + + GGG SS ++S +T
Sbjct: 467 GRSSGIDVLSSFLPRAAAAHHGMTTMGGAAATTTTSHGLNSAISGGGGVSSETTSAVTVA 526
Query: 393 ------------STNDMVRNNGLLQDMIAPMEMGG 415
+ M ++ GLLQDM+ P + G
Sbjct: 527 ASAQPSSPAALQMQHFMAQDLGLLQDMLLPSFIHG 561
>UniRef100_Q7XAA1 WRKY17 [Oryza sativa]
Length = 502
Score = 136 bits (343), Expect = 1e-30
Identities = 98/245 (40%), Positives = 122/245 (49%), Gaps = 40/245 (16%)
Query: 107 ASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHGIRL 166
A V PATP S S +SSSS AAG D ++ +EE EE+ +D G +
Sbjct: 103 AGGVAAAPATPNS--SVLSSSSEAAGGDDLRRCKKGRRPEDEE------EEEIDDEGSAV 154
Query: 167 VENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPF 226
N K K K KK R RV F TKS+VDHL+DGYRWRKYGQK VKNS +
Sbjct: 155 QSCTN---------KMKNKKGAKKEREPRVAFMTKSEVDHLEDGYRWRKYGQKAVKNSSY 205
Query: 227 PRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPV---------LLRAANLGI 277
P SYYRCTA C VKKR+ERS D S+V+T+YEG H H SPV L+ + G+
Sbjct: 206 P-SYYRCTAPRCGVKKRVERSEQDPSMVITTYEGQHTHPSPVSYHMHRQQGLMHVSARGV 264
Query: 278 MSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMSQSQQYQKFQDQNQNVLHHQRQAVPSL 337
M +G + G +LG + A++ Q Q + QR SL
Sbjct: 265 MPGAAGAYQ---------FGAPPPPLLGFDEALAARSQDTVLVSQEDRLSQQQR----SL 311
Query: 338 LYNSD 342
Y D
Sbjct: 312 YYRVD 316
>UniRef100_Q9FL26 Probable WRKY transcription factor 8 [Arabidopsis thaliana]
Length = 326
Score = 128 bits (322), Expect = 3e-28
Identities = 71/185 (38%), Positives = 103/185 (55%), Gaps = 10/185 (5%)
Query: 149 ELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLD 208
E + E+ + R V + + DQ +++ KK + KK P RV+F TK++VDHL+
Sbjct: 125 EADHHPGEDSGKIRKKREVRDGGEDDQRSQKVVKTKKKEEKKKEP-RVSFMTKTEVDHLE 183
Query: 209 DGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPV 268
DGYRWRKYGQK VKNSP+PRSYYRCT C VKKR+ERS D ++V+T+YE H H P
Sbjct: 184 DGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSYQDPTVVITTYESQHNHPIPT 243
Query: 269 LLRAANLGIMSDPSGFQEQVHQHHNTTSGVVGSSVLGNEFAMSQSQQYQ-KFQDQNQNVL 327
R A SG ++ ++S + ++ + S ++ + N N
Sbjct: 244 NRRTAMF------SG--TTASDYNPSSSPIFSDLIINTPRSFSNDDLFRVPYASVNVNPS 295
Query: 328 HHQRQ 332
+HQ+Q
Sbjct: 296 YHQQQ 300
>UniRef100_Q93WV6 Probable WRKY transcription factor 68 [Arabidopsis thaliana]
Length = 277
Score = 128 bits (322), Expect = 3e-28
Identities = 63/124 (50%), Positives = 84/124 (66%), Gaps = 10/124 (8%)
Query: 148 EELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLK-PKKKNKNKKLRPARVTFKTKSDVDH 206
E + GD+ EE+D + Q + KK+ K K K K + +V+F T+S+V H
Sbjct: 66 EAVNGDDEEEED---------GEEQQHKTKKRFKFTKMSRKQTKKKVPKVSFITRSEVLH 116
Query: 207 LDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLS 266
LDDGY+WRKYGQKPVK+SPFPR+YYRCT C+VKKR+ERS +D S V+T+YEG H H
Sbjct: 117 LDDGYKWRKYGQKPVKDSPFPRNYYRCTTTWCDVKKRVERSFSDPSSVITTYEGQHTHPR 176
Query: 267 PVLL 270
P+L+
Sbjct: 177 PLLI 180
>UniRef100_Q9C983 Probable WRKY transcription factor 57 [Arabidopsis thaliana]
Length = 287
Score = 128 bits (322), Expect = 3e-28
Identities = 74/173 (42%), Positives = 94/173 (53%), Gaps = 13/173 (7%)
Query: 117 PVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHGIRLVENQNDQDQM 176
P +V+SS SSS+ V V + +E+ E+ + + +
Sbjct: 68 PTTVTSSCSSSA-------AVSVAVTSTNNNPSATSSSSEDPAENSTASAEKTPPPETPV 120
Query: 177 KKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAG 236
K+ KK K++R R F TKSDVD+L+DGYRWRKYGQK VKNSPFPRSYYRCT
Sbjct: 121 KE-----KKKAQKRIRQPRFAFMTKSDVDNLEDGYRWRKYGQKAVKNSPFPRSYYRCTNS 175
Query: 237 NCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAANLGIMSDPSGFQEQVH 289
C VKKR+ERS+ D SIV+T+YEG H H + R L DP F H
Sbjct: 176 RCTVKKRVERSSDDPSIVITTYEGQHCHQTIGFPRGGIL-TAHDPHSFTSHHH 227
>UniRef100_Q8H5U1 Putative DNA-binding protein WRKY2 [Oryza sativa]
Length = 290
Score = 124 bits (312), Expect = 4e-27
Identities = 77/180 (42%), Positives = 93/180 (50%), Gaps = 36/180 (20%)
Query: 97 LPPLPAPSPAASEV---LNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGD 153
LPPL P A+ L+ P TP S+ ++SS + AD G
Sbjct: 43 LPPLDPPGELATPPPPPLDLPETPAGSSADGAASSCSTDDADG---------------GK 87
Query: 154 EAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRW 213
A E K L P KK + R R F TKS++DHL+DGYRW
Sbjct: 88 PAAASTE--------------AASKSLTPGKK----RARQPRFAFMTKSEIDHLEDGYRW 129
Query: 214 RKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVLLRAA 273
RKYGQK VKNSPFPRSYYRCT C VKKR+ERS+ D S+V+T+YEG H H + RAA
Sbjct: 130 RKYGQKAVKNSPFPRSYYRCTNSKCTVKKRVERSSDDPSVVITTYEGQHSHHTVTFPRAA 189
>UniRef100_Q9AUV7 Putative DNA binding protein [Oryza sativa]
Length = 314
Score = 123 bits (308), Expect = 1e-26
Identities = 61/133 (45%), Positives = 76/133 (56%), Gaps = 15/133 (11%)
Query: 147 EEELEGDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKK---------------KNKNKKL 191
EEE E +EA + D + + + +P K K+
Sbjct: 59 EEEEEEEEAPARSGDGAAAAASSSSSGEPAAPDKRPAAAEAAPAAAATATATAKKGQKRA 118
Query: 192 RPARVTFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADS 251
R R F TKS++DHL+DGYRWRKYGQK VKNSPFPRSYYRCT C VKKR+ERS+ D
Sbjct: 119 RQPRFAFMTKSEIDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTNSKCTVKKRVERSSDDP 178
Query: 252 SIVLTSYEGHHIH 264
S+V+T+YEG H H
Sbjct: 179 SVVITTYEGQHCH 191
>UniRef100_Q93WY4 Probable WRKY transcription factor 12 [Arabidopsis thaliana]
Length = 218
Score = 119 bits (299), Expect = 1e-25
Identities = 66/167 (39%), Positives = 99/167 (58%), Gaps = 13/167 (7%)
Query: 104 SPAASEVLNNPATPVSVSSSISSSSNAAGVADKV-----MVEENEHEHEEELEGDEAEEK 158
S ++S L++P+ P+ SSS +++ G ++ + + + ++ +E G
Sbjct: 44 SSSSSSSLSSPSFPIHNSSSTTTTHAPLGFSNNLQGGGPLGSKVVNDDQENFGGGT---N 100
Query: 159 DEDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQ 218
++ H + + MK ++K ++K LR R F+TKSDVD LDDGY+WRKYGQ
Sbjct: 101 NDAHSNSWWRSNSGSGDMKNKVKIRRK-----LREPRFCFQTKSDVDVLDDGYKWRKYGQ 155
Query: 219 KPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHL 265
K VKNS PRSYYRCT NC VKKR+ER + D +V+T+YEG H H+
Sbjct: 156 KVVKNSLHPRSYYRCTHNNCRVKKRVERLSEDCRMVITTYEGRHNHI 202
>UniRef100_Q9SVB7 Probable WRKY transcription factor 13 [Arabidopsis thaliana]
Length = 304
Score = 116 bits (291), Expect = 1e-24
Identities = 70/169 (41%), Positives = 95/169 (55%), Gaps = 15/169 (8%)
Query: 104 SPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDEAEEKDEDHG 163
S ++ L+NP + SS N G ++ V++N+ E GD D+D+G
Sbjct: 118 SISSQRPLSNPWAWSCQAGYGSSQKNNHG--SEIDVDDNDDE-----VGDGGGINDDDNG 170
Query: 164 IRL---VENQNDQDQ-----MKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRK 215
+++D+ + LK KK +K+R R FKT S+VD LDDGYRWRK
Sbjct: 171 RHHHHDTPSRHDKHNTASLGVVSSLKMKKLKTRRKVREPRFCFKTLSEVDVLDDGYRWRK 230
Query: 216 YGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIH 264
YGQK VKN+ PRSYYRCT C VKKR+ER A D +V+T+YEG H+H
Sbjct: 231 YGQKVVKNTQHPRSYYRCTQDKCRVKKRVERLADDPRMVITTYEGRHLH 279
>UniRef100_Q9S763 Probable WRKY transcription factor 45 [Arabidopsis thaliana]
Length = 147
Score = 115 bits (288), Expect = 2e-24
Identities = 55/105 (52%), Positives = 71/105 (67%), Gaps = 5/105 (4%)
Query: 160 EDHGIRLVENQNDQDQMKKQLKPKKKNKNKKLRPARVTFKTKSDVDHLDDGYRWRKYGQK 219
E HG+ + + Q + KP+ K K KK R AR F+T+S VD LDDGYRWRKYGQK
Sbjct: 22 EFHGV----DNSAQPTTSSEEKPRSKKK-KKEREARYAFQTRSQVDILDDGYRWRKYGQK 76
Query: 220 PVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIH 264
VKN+PFPRSYY+CT C VKK+++R D +V+T+Y+G H H
Sbjct: 77 AVKNNPFPRSYYKCTEEGCRVKKQVQRQWGDEGVVVTTYQGVHTH 121
>UniRef100_Q40090 SPF1 protein [Ipomoea batatas]
Length = 549
Score = 115 bits (287), Expect = 3e-24
Identities = 66/188 (35%), Positives = 99/188 (52%), Gaps = 2/188 (1%)
Query: 92 PSTTQLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELE 151
P +T+ S A++ + P S S SN G D V EN + E
Sbjct: 268 PQSTRRSSSSTASSASTLAAQSYNAPASDVPDQSYWSNGNGQMDSVATPENSSISVGDDE 327
Query: 152 GDEAEEKDEDHGIRLVENQNDQDQMKKQLKPK--KKNKNKKLRPARVTFKTKSDVDHLDD 209
+++ +K E G E++ D + K + + + ++ +R RV +T SD+D LDD
Sbjct: 328 FEQSSQKRESGGDEFDEDEPDAKRWKVENESEGVSAQGSRTVREPRVVVQTTSDIDILDD 387
Query: 210 GYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSPVL 269
GYRWRKYGQK VK +P PRSYY+CT+ C V+K +ER++ D V+T+YEG H H P
Sbjct: 388 GYRWRKYGQKVVKGNPNPRSYYKCTSQGCPVRKHVERASHDIRSVITTYEGKHNHDVPAA 447
Query: 270 LRAANLGI 277
+ + G+
Sbjct: 448 RGSGSHGL 455
Score = 85.1 bits (209), Expect = 3e-15
Identities = 74/271 (27%), Positives = 115/271 (42%), Gaps = 43/271 (15%)
Query: 55 SSSFAGYIDLLSTNDFTSSSATLFDWFPTIDTDMSAPPSTTQLPP--LPAPSPAAS---- 108
SS + + DLL+++ ++ S + + S P LPP LP SPA S
Sbjct: 23 SSFMSSFTDLLASDAYSGGSVSRGLGDRIAERTGSGVPKFKSLPPPSLPLSSPAVSPSSY 82
Query: 109 ----------EVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELEGDE---- 154
E+L++P +S S+ + S + A + + + +E+++ +E
Sbjct: 83 FAFPPGLSPSELLDSPVL-LSSSNILPSPTTGTFPAQTFNWKNDSNASQEDVKQEEKGYP 141
Query: 155 -----------------AEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNK-NKKLRPARV 196
++ KDE + ++ + QM Q ++ N + P
Sbjct: 142 DFSFQTNSASMTLNYEDSKRKDELNSLQSLPPVTTSTQMSSQNNGGSYSEYNNQCCPPSQ 201
Query: 197 TFKTKSDVDHLDDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLT 256
T + + DDGY WRKYGQK VK S PRSYY+CT NC KK++ER A D I
Sbjct: 202 TLREQR---RSDDGYNWRKYGQKQVKGSENPRSYYKCTHPNCPTKKKVER-ALDGQITEI 257
Query: 257 SYEGHHIHLSPVLLRAANLGIMSDPSGFQEQ 287
Y+G H H P R ++ S S Q
Sbjct: 258 VYKGAHNHPKPQSTRRSSSSTASSASTLAAQ 288
>UniRef100_Q5QJ45 WRKY3 [Nicotiana attenuata]
Length = 354
Score = 114 bits (285), Expect = 5e-24
Identities = 62/178 (34%), Positives = 96/178 (53%), Gaps = 10/178 (5%)
Query: 92 PSTTQLPPLPAPSPAASEVLNNPATPVSVSSSISSSSNAAGVADKVMVEENEHEHEEELE 151
P +T+ A S +A + N + S S NA + +++HEH +
Sbjct: 89 PQSTRRSSSTASSSSAVQPYNTQTNEIPDHQSYGS--NATPENSSISFGDDDHEHSSQKS 146
Query: 152 GDEAEEKDEDHGIRLVENQNDQDQMKKQLKPKKKNK--NKKLRPARVTFKTKSDVDHLDD 209
++ DE+ E + D + K++ + + + ++ +R RV +T SD+D LDD
Sbjct: 147 RSRGDDFDEE------EEEPDSKRWKRESESESLSAPGSRTVREPRVVVQTTSDIDILDD 200
Query: 210 GYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSP 267
GYRWRKYGQK VK +P PRSYY+CT+ C V+K +ER++ D V+T+YEG H H P
Sbjct: 201 GYRWRKYGQKVVKGNPNPRSYYKCTSPGCPVRKHVERASQDIRSVITTYEGKHNHDVP 258
Score = 74.7 bits (182), Expect = 5e-12
Identities = 35/67 (52%), Positives = 43/67 (63%), Gaps = 1/67 (1%)
Query: 208 DDGYRWRKYGQKPVKNSPFPRSYYRCTAGNCEVKKRIERSAADSSIVLTSYEGHHIHLSP 267
+DGY WRKYGQK VK S PRSYY+CT NC KK++ERS D I Y+G+H H P
Sbjct: 31 EDGYNWRKYGQKQVKGSENPRSYYKCTFPNCPTKKKVERS-LDGQITEIVYKGNHNHPKP 89
Query: 268 VLLRAAN 274
R ++
Sbjct: 90 QSTRRSS 96
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.309 0.126 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 727,060,832
Number of Sequences: 2790947
Number of extensions: 32789270
Number of successful extensions: 244886
Number of sequences better than 10.0: 2282
Number of HSP's better than 10.0 without gapping: 842
Number of HSP's successfully gapped in prelim test: 1576
Number of HSP's that attempted gapping in prelim test: 225245
Number of HSP's gapped (non-prelim): 10845
length of query: 420
length of database: 848,049,833
effective HSP length: 130
effective length of query: 290
effective length of database: 485,226,723
effective search space: 140715749670
effective search space used: 140715749670
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 76 (33.9 bits)
Medicago: description of AC135100.8


