

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.24 + phase: 0
(456 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
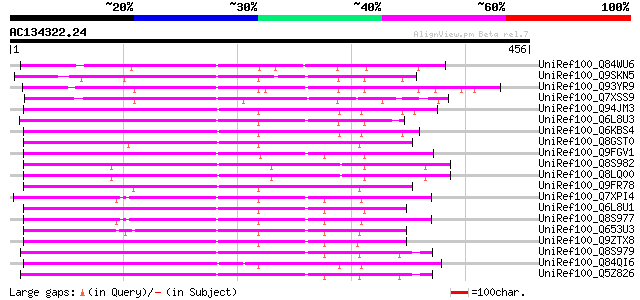
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q84WU6 Auxin response factor 17 [Arabidopsis thaliana] 262 1e-68
UniRef100_Q9SKN5 Auxin response factor 10 [Arabidopsis thaliana] 239 2e-61
UniRef100_Q93YR9 Auxin response factor 16 [Arabidopsis thaliana] 235 2e-60
UniRef100_Q7XSS9 OSJNBa0039K24.27 protein [Oryza sativa] 186 2e-45
UniRef100_Q94JM3 Auxin response factor 2 [Arabidopsis thaliana] 181 3e-44
UniRef100_Q6L8U3 Auxin response factor 1 [Cucumis sativus] 180 7e-44
UniRef100_Q6KBS4 Putative auxin response factor [Brassica napus] 179 1e-43
UniRef100_Q8GST0 Auxin response factor 1 [Oryza sativa] 178 3e-43
UniRef100_Q9FGV1 Auxin response factor 8 [Arabidopsis thaliana] 177 4e-43
UniRef100_Q8S982 Auxin response factor 2 [Oryza sativa] 177 6e-43
UniRef100_Q8LQ00 Auxin response factor 2 [Oryza sativa] 177 6e-43
UniRef100_Q9FR78 Putative auxin response factor 1 [Oryza sativa] 176 9e-43
UniRef100_Q7XPI4 OSJNBb0004A17.5 protein [Oryza sativa] 175 2e-42
UniRef100_Q6L8U1 Auxin response factor 3 [Cucumis sativus] 175 2e-42
UniRef100_Q8S977 Auxin response factor 8 [Oryza sativa] 175 3e-42
UniRef100_Q653U3 Putative auxin response factor [Oryza sativa] 175 3e-42
UniRef100_Q9ZTX8 Auxin response factor 6 [Arabidopsis thaliana] 174 5e-42
UniRef100_Q8S979 Auxin response factor 7a [Oryza sativa] 174 6e-42
UniRef100_Q84QI6 Auxin response factor-like protein [Mangifera i... 174 6e-42
UniRef100_Q5Z826 Putative auxin response factor 7a [Oryza sativa] 174 6e-42
>UniRef100_Q84WU6 Auxin response factor 17 [Arabidopsis thaliana]
Length = 585
Score = 262 bits (670), Expect = 1e-68
Identities = 159/439 (36%), Positives = 231/439 (52%), Gaps = 74/439 (16%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
V P IW CA A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++
Sbjct: 16 VDPTIWRACAGASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIIT 68
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDV 125
++ LLADP TDEVF L+L P+T +++D N+V +F K LT SD
Sbjct: 69 SIQLLADPVTDEVFAHLILQPMTQQQFTPTNYSRFGRFDGDVDDNNKVTTFAKILTPSDA 128
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
NN F +PRFCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L +
Sbjct: 129 NNGGGFSVPRFCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTG 187
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQ 228
W+ FV KKL+AGDSV+FM+ S ++F+G+RR + + ++
Sbjct: 188 WSKFVNSKKLIAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVA 247
Query: 229 KKDE---------------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDE 273
K+D+ +AV +A+ A + FE+V+YP W +FVV V+
Sbjct: 248 KEDDGSPKKTFRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVES 306
Query: 274 SMKIQWNPRMRVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQVNWDEFQVSQ 325
SM + W P RVK M+T+ SSRI +QG +S + + WR LQ+ WDE ++ Q
Sbjct: 307 SMSMYWTPGTRVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQITWDEPEILQ 366
Query: 326 IPRRVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSD 364
+RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 367 NVKRVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPD 426
Query: 365 KKETLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 427 PSPSMFSYSTFPAGMQGAR 445
>UniRef100_Q9SKN5 Auxin response factor 10 [Arabidopsis thaliana]
Length = 693
Score = 239 bits (609), Expect = 2e-61
Identities = 156/416 (37%), Positives = 217/416 (51%), Gaps = 76/416 (18%)
Query: 5 KQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFR--P 62
+Q + P++WH CA + V+IP L+S V+YF QGH E+A H + R P
Sbjct: 2 EQEKSLDPQLWHACAGSMVQIPSLNSTVFYFAQGHTEHA--------HAPPDFHAPRVPP 53
Query: 63 FTLCIVSAVDLLADPHTDEVFVKLLLTPIT-NDVHLEN-------PKEEVANLNDRNEVV 114
LC V +V LAD TDEVF K+ L P+ ND+ LEN P N N + +
Sbjct: 54 LILCRVVSVKFLADAETDEVFAKITLLPLPGNDLDLENDAVLGLTPPSSDGNGNGKEKPA 113
Query: 115 SFVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
SF KTLT+SD NN F +PR+CA+ +FP+LD AE Q + D+HGE KF H+ RG
Sbjct: 114 SFAKTLTQSDANNGGGFSVPRYCAETIFPRLDYSAEPPVQTVIAKDIHGETWKFRHIYRG 173
Query: 175 FPKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR----------------- 217
P+R++L + W++FV +KKL+AGDS++F++ +G + VGIRR
Sbjct: 174 TPRRHLL-TTGWSTFVNQKKLIAGDSIVFLRSESGDLCVGIRRAKRGGLGSNAGSDNPYP 232
Query: 218 -------------------------NTQFVAAAAEQKKDELEKAVMEALKLAEENKAFEI 252
N AAA + + E AV EA+ A +AFE+
Sbjct: 233 GFSGFLRDDESTTTTSKLMMMKRNGNNDGNAAATGRVRVE---AVAEAVARAACGQAFEV 289
Query: 253 VYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSN 309
VYYP+ +F V V +M+I+W MR KM +T+ SSRI + GT+S V
Sbjct: 290 VYYPRAST-PEFCVKAADVRSAMRIRWCSGMRFKMAFETEDSSRISWFMGTVSAVQVADP 348
Query: 310 L------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQ 357
+ WR+LQV WDE + Q +RV+PW VEL+S+ P +PF KK R Q
Sbjct: 349 IRWPNSPWRLLQVAWDEPDLLQNVKRVSPWLVELVSNMPTIHLSPFSPRKKIRIPQ 404
>UniRef100_Q93YR9 Auxin response factor 16 [Arabidopsis thaliana]
Length = 670
Score = 235 bits (600), Expect = 2e-60
Identities = 173/519 (33%), Positives = 245/519 (46%), Gaps = 107/519 (20%)
Query: 12 PEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAV 71
P++WH CA V++P ++S+V+YFPQGH ENA P LC V A+
Sbjct: 18 PQLWHACAGGMVRMPPMNSKVFYFPQGHAENAYDCVDFGN------LPIPPMVLCRVLAI 71
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN----DRNEVVSFVKTLTRSDVNN 127
+AD +DEVF KL L P+ +D ++++ + + N + + SF KTLT+SD NN
Sbjct: 72 KYMADAESDEVFAKLRLIPLKDDEYVDHEYGDGEDSNGFESNSEKTPSFAKTLTQSDANN 131
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +PR+CA+ +FP+LD AE Q + DVHG+V KF H+ RG P+R++L + W+
Sbjct: 132 GGGFSVPRYCAETIFPRLDYNAEPPVQTILAKDVHGDVWKFRHIYRGTPRRHLL-TTGWS 190
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--------NTQFVA---------------- 223
+FV +KKLVAGDS++FM+ G + VGIRR ++ A
Sbjct: 191 NFVNQKKLVAGDSIVFMRAENGDLCVGIRRAKRGGIGNGPEYSAGWNPIGGSCGYSSLLR 250
Query: 224 ------------AAAEQKKDELEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVV 271
+ A++K ++V+EA LA + FE+VYYP+ +F V
Sbjct: 251 EDESNSLRRSNCSLADRKGKVTAESVIEAATLAISGRPFEVVYYPRAST-SEFCVKALDA 309
Query: 272 DESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSNL------WRMLQVNWDEFQ 322
+M+I W MR KM +T+ SSRI + GT+S V+ + + WR+LQV WDE
Sbjct: 310 RAAMRIPWCSGMRFKMAFETEDSSRISWFMGTVSAVNVSDPIRWPNSPWRLLQVAWDEPD 369
Query: 323 VSQIPRRVNPWWVELISH-KPAP-TPF-PQTKKFRTTQ---------SSAQLSDKKETLL 370
+ Q +RVNPW VEL+S+ P P T F P KK R Q S S L+
Sbjct: 370 LLQNVKRVNPWLVELVSNVHPIPLTSFSPPRKKMRLPQHPDYNNLINSIPVPSFPSNPLI 429
Query: 371 NG-------DGFPVDIQRVRHDLVSISGPIHS---HIILNSSETKFP------------- 407
D PV +Q RH+ G S H LN P
Sbjct: 430 RSSPLSSVLDNVPVGLQGARHNAHQYYGLSSSDLHHYYLNRPPPPPPPSSLQLSPSLGLR 489
Query: 408 ---------------ATHNCNTKNDSDGSIMLFGKIIQP 431
T CN I+LFGK+I P
Sbjct: 490 NIDTKNEKGFCFLTMGTTPCNDTKSKKSHIVLFGKLILP 528
>UniRef100_Q7XSS9 OSJNBa0039K24.27 protein [Oryza sativa]
Length = 529
Score = 186 bits (471), Expect = 2e-45
Identities = 129/401 (32%), Positives = 197/401 (48%), Gaps = 50/401 (12%)
Query: 14 IWHTCATAAVKIPKLHSRVYYFPQGHLENA-SPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
+W CA +IP + ++V YFP+GH E +P + H F LC ++AVD
Sbjct: 28 VWLACAAPLSRIPVVGTQVSYFPEGHAEQCPAPLPDPLPSAHRFF-------LCTITAVD 80
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN----DRNEVVSFVKTLTRSDVNNA 128
L AD T E + + L P+ +D P A + E + K LT+SD NN
Sbjct: 81 LSADTTTGEPYATISLLPLRHDAPAPAPAPAPAAAELAEAESQEFRYYAKQLTQSDANNG 140
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR CAD++FP L+L+ + Q L + D+ G+ +F H+ RG P+R++L + W+
Sbjct: 141 GGFSVPRLCADHIFPALNLDDDPPVQSLTMGDLQGDSWEFRHIYRGTPRRHLL-TTGWSK 199
Query: 189 FVKRKKLVAGDSVIFM----KDSTGKIFVGIRRNTQFVAAAAEQKKDELE-KAVMEALKL 243
FV K+LVAGD+V+FM K+ VG+RR ++ +A + ++ + VMEA++L
Sbjct: 200 FVNAKQLVAGDTVVFMWCGAPAPERKLLVGVRRAARYSGESACNARGRVQPQEVMEAVRL 259
Query: 244 AEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVK---MKTDKSSRIPY-QG 299
A E AF + YYP+ +FVV VD+ + W M+V+ M+ + + R+ + G
Sbjct: 260 AAEQAAFRVTYYPRHGAG-EFVVPRVEVDKGLTTPWRCGMQVRAQVMEAEDTRRLAWLNG 318
Query: 300 TISIVSRTSNLWRMLQVNWDEFQVSQI--PRRVNPWWVELISHKPAPTPFPQTKKFRTTQ 357
T++ + R +WR L+V WD S R VNPW V+ + P P K
Sbjct: 319 TLTNL-RHQQIWRTLEVEWDASAASSSMKNRFVNPWQVQPVDF----PPLPMGLKISNNN 373
Query: 358 SSAQLSDKKETLLNGDGF-------------PVDIQRVRHD 385
SA + NGD P DIQ RH+
Sbjct: 374 ISA-------PVCNGDSLLVPPILMHPQPQPPADIQGARHN 407
>UniRef100_Q94JM3 Auxin response factor 2 [Arabidopsis thaliana]
Length = 859
Score = 181 bits (460), Expect = 3e-44
Identities = 124/381 (32%), Positives = 182/381 (47%), Gaps = 18/381 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKLLCRVINVD 120
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 121 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 180
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 181 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 239
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 240 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 299
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 300 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 359
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFR-----T 355
+ WR L+V WDE P RV+PW VE PA P P P+ K+ R +
Sbjct: 360 PTRWPKSKWRSLKVRWDETSSIPRPDRVSPWKVEPALAPPALSPVPMPRPKRPRSNIAPS 419
Query: 356 TQSSAQLSDKKETLLNGDGFP 376
+ S+ L+ + T N D P
Sbjct: 420 SPDSSMLTREGTTKANMDPLP 440
>UniRef100_Q6L8U3 Auxin response factor 1 [Cucumis sativus]
Length = 1081
Score = 180 bits (457), Expect = 7e-44
Identities = 115/350 (32%), Positives = 174/350 (48%), Gaps = 18/350 (5%)
Query: 9 HVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIV 68
++ E+WH CA V +P + S V YFPQGH E + S + T + +C++
Sbjct: 19 NINSELWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMNKETDFIPNYPNLPSKLICML 78
Query: 69 SAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
V L ADP TDEV+ ++ L P+ ++ R F KTLT SD +
Sbjct: 79 HNVTLHADPETDEVYAQMTLQPVNKYEKEALLASDIGLKQSRQPAEFFCKTLTASDTSTH 138
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR A+ +FP LD + +Q L D+H F H+ RG PKR++L + W+
Sbjct: 139 GGFSVPRRAAEKIFPPLDYSMQPPAQELVARDLHDNSWTFRHIYRGQPKRHLL-TTGWSV 197
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAE 245
FV K+L AGDSV+F++D ++ +GIRR Q +++ D + ++ A A
Sbjct: 198 FVSTKRLFAGDSVLFIRDEKSQLLLGIRRANRQQPALSSSVISSDSMHIGILASAAHAAA 257
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIPYQGTISI 303
N F I Y P+ +FV+ +++M Q + MR +M +T++S Y GTI+
Sbjct: 258 NNSPFTIFYNPRASP-SEFVIPLAKYNKAMYTQVSLGMRFRMMFETEESGVRRYMGTITG 316
Query: 304 VSRTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPF 347
+S ++ WR LQV WDE + P RV+ W VE P TPF
Sbjct: 317 ISDMDSVRWKNSQWRNLQVGWDESAAGERPNRVSIWEVE-----PVVTPF 361
>UniRef100_Q6KBS4 Putative auxin response factor [Brassica napus]
Length = 848
Score = 179 bits (455), Expect = 1e-43
Identities = 118/360 (32%), Positives = 175/360 (47%), Gaps = 13/360 (3%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ L LC V VD
Sbjct: 56 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQVEASTNQAAEQQMPLYDLPSKILCRVINVD 115
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P KE R +V SF KTLT SD + F
Sbjct: 116 LKAEADTDEVYAQITLLPEPVQDENSIEKEAPPPPPPRFQVHSFCKTLTASDTSTHGGFS 175
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 176 VLRRHADECLPPLDMSRQPPTQELVAKDLHASEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 234
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 235 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 294
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+KI ++ MR KM+ +++ + GTI + +
Sbjct: 295 FTVYYKPRTSPSEFIVPFDQYTESVKINYSIGMRFKMRFEGEEAPEQRFTGTIVGIEDSD 354
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQSSA 360
+ WR L+V WDE P RV+PW +E PA P P P+ K+ R+ +S+
Sbjct: 355 PTRWAKSKWRSLKVRWDETTSIPRPDRVSPWKIEPALSPPALSPVPMPRPKRPRSNLASS 414
>UniRef100_Q8GST0 Auxin response factor 1 [Oryza sativa]
Length = 836
Score = 178 bits (451), Expect = 3e-43
Identities = 119/359 (33%), Positives = 175/359 (48%), Gaps = 18/359 (5%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+W CA V +P++ +V+YFPQGH+E S++ + L + LC V V+
Sbjct: 24 ELWSACAGPLVTVPRVGEKVFYFPQGHIEQVEASTNQVGEQRMQLYNLPWKILCEVMNVE 83
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEE-----VANLNDRNEVVSFVKTLTRSDVNN 127
L A+P TDEV+ +L L P + EE A + R V SF KTLT SD +
Sbjct: 84 LKAEPDTDEVYAQLTLLPESKQQEDNGSTEEEVPSAPAAGHVRPRVHSFCKTLTASDTST 143
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F + R AD P LD+ + +Q L D+HG +F H+ RG P+R++L S W+
Sbjct: 144 HGGFSVLRRHADECLPPLDMSRQPPTQELVAKDLHGVEWRFRHIFRGQPRRHLLQ-SGWS 202
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAE 245
FV K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 203 VFVSAKRLVAGDAFIFLRGENGELRVGVRRAMRQQTNVPSSVISSHSMHLGVLATAWHAV 262
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
VYY +FVV + ES+K ++ MR KM+ + + T +IV
Sbjct: 263 NTGTMFTVYYKPRTSPAEFVVPYDRYMESLKQNYSIGMRFKMRFEGEEAPEQRFTGTIVG 322
Query: 306 R--------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVE-LISHKPA-PTPFPQTKKFR 354
+ WR L+V WDE P RV+PW +E +S P P P P+TK+ R
Sbjct: 323 MGDSDPAGWPESKWRSLKVRWDEASSIPRPERVSPWQIEPAVSPPPVNPLPVPRTKRLR 381
>UniRef100_Q9FGV1 Auxin response factor 8 [Arabidopsis thaliana]
Length = 811
Score = 177 bits (450), Expect = 4e-43
Identities = 118/374 (31%), Positives = 189/374 (49%), Gaps = 16/374 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P SRV YFPQGH E + +++ H S P +C + V
Sbjct: 22 ELWHACAGPLVSLPSSGSRVVYFPQGHSEQVAATTNKEVDGHIPNYPSLPPQLICQLHNV 81
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+T + E + + F KTLT SD + F
Sbjct: 82 TMHADVETDEVYAQMTLQPLTPEEQKETFVPIELGIPSKQPSNYFCKTLTASDTSTHGGF 141
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 142 SVPRRAAEKVFPPLDYTLQPPAQELIARDLHDVEWKFRHIFRGQPKRHLL-TTGWSVFVS 200
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNT--QFVAAAAEQKKDELEKAVMEALKLAE-ENK 248
K+LVAGDSVIF+++ ++F+GIR T Q + ++ D + ++ A A N
Sbjct: 201 AKRLVAGDSVIFIRNEKNQLFLGIRHATRPQTIVPSSVLSSDSMHIGLLAAAAHASATNS 260
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
F + ++P+ +FV+ + +++ +I R R+ +T++SS Y GTI+ +S
Sbjct: 261 CFTVFFHPRASQ-SEFVIQLSKYIKAVFHTRISVGMRFRMLFETEESSVRRYMGTITGIS 319
Query: 306 RTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQS 358
++ WR ++V WDE + RV+ W +E ++ P P+ FP K
Sbjct: 320 DLDSVRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSLFPLRLKRPWHAG 379
Query: 359 SAQLSDKKETLLNG 372
++ L D + L +G
Sbjct: 380 TSSLPDGRGDLGSG 393
>UniRef100_Q8S982 Auxin response factor 2 [Oryza sativa]
Length = 791
Score = 177 bits (449), Expect = 6e-43
Identities = 120/391 (30%), Positives = 182/391 (45%), Gaps = 18/391 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P++ V+YFPQGH+E S + + + L LC V V+
Sbjct: 7 ELWHACAGPLVTVPRVGDLVFYFPQGHIEQVEASMNQVADSQMRLYDLPSKLLCRVLNVE 66
Query: 73 LLADPHTDEVFVKLLL--TPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARS 130
L A+ TDEV+ +++L P N++ +E + R V SF KTLT SD +
Sbjct: 67 LKAEQDTDEVYAQVMLMPEPEQNEMAVEKTTPTSGPVQARPPVRSFCKTLTASDTSTHGG 126
Query: 131 FHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFV 190
F + R AD P LD+ +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 127 FSVLRRHADECLPPLDMTQSPPTQELVAKDLHSMDWRFRHIFRGQPRRHLLQ-SGWSVFV 185
Query: 191 KRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELEKAVMEALKLAEENK 248
K+LVAGD+ IF++ G++ VG+RR + ++ + V+ A K
Sbjct: 186 SSKRLVAGDAFIFLRGENGELRVGVRRAMRQLSNVPSSVISSQSMHLGVLATAWHAINTK 245
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVSRTS 308
+ VYY +F++ + ES+K ++ MR +M+ + P Q + +
Sbjct: 246 SMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFE-GEEAPEQRFTGTIIGSE 304
Query: 309 NL--------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQSS 359
NL WR L+V WDE P RV+PW +E S P P P + K+ R
Sbjct: 305 NLDPVWPESSWRSLKVRWDEPSTIPRPDRVSPWKIEPASSPPVNPLPLSRVKRPRPNAPP 364
Query: 360 AQLSD---KKETLLNGDGFPVDIQRVRHDLV 387
A KE D P QR ++ V
Sbjct: 365 ASPESPILTKEAATKVDTDPAQAQRSQNSTV 395
>UniRef100_Q8LQ00 Auxin response factor 2 [Oryza sativa]
Length = 826
Score = 177 bits (449), Expect = 6e-43
Identities = 120/391 (30%), Positives = 182/391 (45%), Gaps = 18/391 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P++ V+YFPQGH+E S + + + L LC V V+
Sbjct: 19 ELWHACAGPLVTVPRVGDLVFYFPQGHIEQVEASMNQVADSQMRLYDLPSKLLCRVLNVE 78
Query: 73 LLADPHTDEVFVKLLL--TPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARS 130
L A+ TDEV+ +++L P N++ +E + R V SF KTLT SD +
Sbjct: 79 LKAEQDTDEVYAQVMLMPEPEQNEMAVEKTTPTSGPVQARPPVRSFCKTLTASDTSTHGG 138
Query: 131 FHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFV 190
F + R AD P LD+ +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 139 FSVLRRHADECLPPLDMTQSPPTQELVAKDLHSMDWRFRHIFRGQPRRHLLQ-SGWSVFV 197
Query: 191 KRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELEKAVMEALKLAEENK 248
K+LVAGD+ IF++ G++ VG+RR + ++ + V+ A K
Sbjct: 198 SSKRLVAGDAFIFLRGENGELRVGVRRAMRQLSNVPSSVISSQSMHLGVLATAWHAINTK 257
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVSRTS 308
+ VYY +F++ + ES+K ++ MR +M+ + P Q + +
Sbjct: 258 SMFTVYYKPRTSPSEFIIPYDQYMESVKNNYSVGMRFRMRFE-GEEAPEQRFTGTIIGSE 316
Query: 309 NL--------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQSS 359
NL WR L+V WDE P RV+PW +E S P P P + K+ R
Sbjct: 317 NLDPVWPESSWRSLKVRWDEPSTIPRPDRVSPWKIEPASSPPVNPLPLSRVKRPRPNAPP 376
Query: 360 AQLSD---KKETLLNGDGFPVDIQRVRHDLV 387
A KE D P QR ++ V
Sbjct: 377 ASPESPILTKEAATKVDTDPAQAQRSQNSTV 407
>UniRef100_Q9FR78 Putative auxin response factor 1 [Oryza sativa]
Length = 857
Score = 176 bits (447), Expect = 9e-43
Identities = 118/359 (32%), Positives = 177/359 (48%), Gaps = 18/359 (5%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+W CA V +P++ + +YFPQGH+E S++ + L + LC V V+
Sbjct: 40 ELWSACAGPLVTVPRVGEKEFYFPQGHIEQVEASTNQVGEQRMQLYNLPWKILCEVMNVE 99
Query: 73 LLADPHTDEVFVKLLLTP-ITNDVHLENPKEEVANL----NDRNEVVSFVKTLTRSDVNN 127
L A+P TDEV+ +L L P + + +EEV + + R V SF KTLT SD +
Sbjct: 100 LKAEPDTDEVYAQLTLLPELKRQEDNGSTEEEVPSAPAAGHVRPRVHSFCKTLTASDTST 159
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F + R AD P LD+ + +Q L D+HG +F H+ RG P+R++L S W+
Sbjct: 160 HGGFSVLRRHADECLPPLDMSRQPPTQELVAKDLHGVEWRFRHIFRGQPRRHLLQ-SGWS 218
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAE 245
FV K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 219 VFVSAKRLVAGDAFIFLRGENGELRVGVRRAMRQQTNVPSSVISSHSMHLGVLATAWHAV 278
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
VYY +FVV + ES+K ++ MR KM+ + + T +IV
Sbjct: 279 NTGTMFTVYYKPRTSPAEFVVPYDRYMESLKRNYSIGMRFKMRFESEEAPEQRFTGTIVG 338
Query: 306 R--------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVE-LISHKPA-PTPFPQTKKFR 354
+ WR L+V WDE P RV+PW +E +S P P P P+TK+ R
Sbjct: 339 MGDSDPAGWPESKWRSLKVRWDEASSIPRPERVSPWQIEPAVSPPPVNPLPVPRTKRLR 397
>UniRef100_Q7XPI4 OSJNBb0004A17.5 protein [Oryza sativa]
Length = 818
Score = 175 bits (444), Expect = 2e-42
Identities = 125/385 (32%), Positives = 191/385 (49%), Gaps = 24/385 (6%)
Query: 4 QKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPF 63
+++ + E+WH CA V +P + +RV YFPQGH E + S++ H P
Sbjct: 22 EEEKKCLNSELWHACAGPLVCLPTVGTRVVYFPQGHSEQVAASTNKEVEGHIPNYPNLPA 81
Query: 64 TL-CIVSAVDLLADPHTDEVFVKLLLTPIT----NDVHLENPKEEVANLNDRNEVVSFVK 118
L C + V + AD TDEV+ ++ L P+ ND +L P E + + F K
Sbjct: 82 QLICQLHDVTMHADVETDEVYAQMTLQPLNPQEQNDAYL--PAE--MGIMSKQPTNYFCK 137
Query: 119 TLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKR 178
TLT SD + F +PR A+ VFP LD + +Q L D+H KF H+ RG PKR
Sbjct: 138 TLTASDTSTHGGFSVPRRAAERVFPPLDFTQQPPAQELIARDIHDIEWKFRHIFRGQPKR 197
Query: 179 NMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKA 236
++L + W+ FV K+LVAGDSV+F+ + ++ +GIRR Q V ++ D +
Sbjct: 198 HLL-TTGWSVFVSAKRLVAGDSVLFIWNEKNQLLLGIRRASRPQTVMPSSVLSSDSMHIG 256
Query: 237 VM-EALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKS 292
++ A A N F I Y P+ +FV+ + +++ +I R R+ +T++S
Sbjct: 257 LLAAAAHAAATNSRFTIFYNPRASP-SEFVIPLSKYIKAVFHTRISVGMRFRMLFETEES 315
Query: 293 SRIPYQGTISIVSRT------SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PT 345
S Y GTI+ VS S+ WR ++V WDE + P RV+ W +E ++ P P+
Sbjct: 316 SVRRYMGTITEVSDADPVRWPSSYWRSVKVGWDESTAGERPPRVSLWEIEPLTTFPMYPS 375
Query: 346 PFPQTKKFRTTQSSAQLSDKKETLL 370
FP K A L D L+
Sbjct: 376 LFPLRVKHPWYSGVASLHDDSNALM 400
>UniRef100_Q6L8U1 Auxin response factor 3 [Cucumis sativus]
Length = 916
Score = 175 bits (444), Expect = 2e-42
Identities = 115/350 (32%), Positives = 175/350 (49%), Gaps = 16/350 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P + SRV YFPQGH E + S++ H S P +C + V
Sbjct: 23 ELWHACAGPLVSLPAVGSRVVYFPQGHSEQVAISTNREVDAHIPSYPSLPPQLICQLHNV 82
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+T E + F KTLT SD + F
Sbjct: 83 TMHADIETDEVYAQMTLQPLTAQEQKEPYLPAELGAPSKQPTNYFCKTLTASDTSTHGGF 142
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 143 SVPRRAAEKVFPPLDFSQQPPAQELIARDLHDNEWKFRHIFRGQPKRHLL-TTGWSVFVS 201
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAEENK 248
K+LVAGDSVIF+ + ++ +GIRR Q V ++ D + ++ A A N
Sbjct: 202 AKRLVAGDSVIFIWNEKNQLLLGIRRANRPQTVMPSSVLSSDSMHLGLLAAAAHAAATNS 261
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
F I Y P+ +F++ +++ ++ R R+ +T++SS Y GTI+ +S
Sbjct: 262 RFTIFYNPRASP-SEFIIPLAKYVKAVYHTRVSVGMRFRMLFETEESSVRRYMGTITGIS 320
Query: 306 RTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFP 348
+ WR ++V WDE + RV+ W +E ++ P P+PFP
Sbjct: 321 DLDSTRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSPFP 370
>UniRef100_Q8S977 Auxin response factor 8 [Oryza sativa]
Length = 795
Score = 175 bits (443), Expect = 3e-42
Identities = 125/376 (33%), Positives = 187/376 (49%), Gaps = 24/376 (6%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTL-CIVSAV 71
E+WH CA V +P + +RV YFPQGH E + S++ H P L C + V
Sbjct: 8 ELWHACAGPLVCLPTVGTRVVYFPQGHSEQVAASTNKEVEGHIPNYPNLPAQLICQLHDV 67
Query: 72 DLLADPHTDEVFVKLLLTPIT----NDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNN 127
+ AD TDEV+ ++ L P+ ND +L P E + + F KTLT SD +
Sbjct: 68 TMHADVETDEVYAQMTLQPLNPQEQNDAYL--PAE--MGIMSKQPTNYFCKTLTASDTST 123
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+
Sbjct: 124 HGGFSVPRRAAERVFPPLDFTQQPPAQELIARDIHDIEWKFRHIFRGQPKRHLL-TTGWS 182
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLA 244
FV K+LVAGDSV+F+ + ++ +GIRR Q V ++ D + ++ A A
Sbjct: 183 VFVSAKRLVAGDSVLFIWNEKNQLLLGIRRASRPQTVMPSSVLSSDSMHIGLLAAAAHAA 242
Query: 245 EENKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTI 301
N F I Y P+ +FV+ + +++ +I R R+ +T++SS Y GTI
Sbjct: 243 ATNSRFTIFYNPRASP-SEFVIPLSKYIKAVFHTRISVGMRFRMLFETEESSVRRYMGTI 301
Query: 302 SIVSRT------SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFR 354
+ VS S+ WR ++V WDE + P RV+ W +E ++ P P+ FP K
Sbjct: 302 TEVSDADPVRWPSSYWRSVKVGWDESTAGERPPRVSLWEIEPLTTFPMYPSLFPLRVKHP 361
Query: 355 TTQSSAQLSDKKETLL 370
A L D L+
Sbjct: 362 WYSGVASLHDDSNALM 377
>UniRef100_Q653U3 Putative auxin response factor [Oryza sativa]
Length = 917
Score = 175 bits (443), Expect = 3e-42
Identities = 117/352 (33%), Positives = 185/352 (52%), Gaps = 20/352 (5%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P + SRV YFPQGH E + S++ + + P +C + V
Sbjct: 30 ELWHACAGPLVSLPAVGSRVVYFPQGHSEQVAASTNKEMESQIPNYPNLPPQLICQLHNV 89
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENP--KEEVANLNDRNEVVSFVKTLTRSDVNNAR 129
+ AD TDEV+ ++ L P++ L++P E+ + N + F KTLT SD +
Sbjct: 90 TMHADAETDEVYAQMTLQPLSPQ-ELKDPYLPAELGSAN-KQPTNYFCKTLTASDTSTHG 147
Query: 130 SFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSF 189
F +PR A+ VFP LD + +Q L D+HG KF H+ RG PKR++L + W+ F
Sbjct: 148 GFSVPRRAAEKVFPPLDFTQQPPAQELIAKDLHGNEWKFRHIFRGQPKRHLL-TTGWSVF 206
Query: 190 VKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAEE 246
V K+LVAGDSV+F+ + ++ +GIRR Q V ++ D + ++ A A
Sbjct: 207 VSAKRLVAGDSVLFIWNDNNQLLLGIRRANRPQTVMPSSVLSSDSMHIGLLAAAAHAAST 266
Query: 247 NKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISI 303
N F I Y P+ +FV+ + +++ +I R R+ +T++SS Y GTI+
Sbjct: 267 NSRFTIFYNPRASP-SEFVIPLSKYVKAVYHTRISVGMRFRMLFETEESSVRRYMGTITG 325
Query: 304 VSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFP 348
+S ++ WR ++V WDE + RV+ W +E ++ P P+PFP
Sbjct: 326 ISDLDAARWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSPFP 377
>UniRef100_Q9ZTX8 Auxin response factor 6 [Arabidopsis thaliana]
Length = 933
Score = 174 bits (441), Expect = 5e-42
Identities = 115/350 (32%), Positives = 177/350 (49%), Gaps = 16/350 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P + SRV YFPQGH E + S++ H S P +C + V
Sbjct: 23 ELWHACAGPLVSLPPVGSRVVYFPQGHSEQVAASTNKEVDAHIPNYPSLHPQLICQLHNV 82
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+ + + R F KTLT SD + F
Sbjct: 83 TMHADVETDEVYAQMTLQPLNAQEQKDPYLPAELGVPSRQPTNYFCKTLTASDTSTHGGF 142
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 143 SVPRRAAEKVFPPLDYSQQPPAQELMARDLHDNEWKFRHIFRGQPKRHLL-TTGWSVFVS 201
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAEENK 248
K+LVAGDSV+F+ + ++ +GIRR Q V ++ D + ++ A A N
Sbjct: 202 AKRLVAGDSVLFIWNDKNQLLLGIRRANRPQTVMPSSVLSSDSMHLGLLAAAAHAAATNS 261
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIV- 304
F I Y P+ +FV+ +++ ++ R R+ +T++SS Y GTI+ +
Sbjct: 262 RFTIFYNPRASP-SEFVIPLAKYVKAVYHTRVSVGMRFRMLFETEESSVRRYMGTITGIC 320
Query: 305 ----SRTSNL-WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFP 348
+R +N WR ++V WDE + RV+ W +E ++ P P+PFP
Sbjct: 321 DLDPTRWANSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSPFP 370
>UniRef100_Q8S979 Auxin response factor 7a [Oryza sativa]
Length = 1123
Score = 174 bits (440), Expect = 6e-42
Identities = 123/377 (32%), Positives = 183/377 (47%), Gaps = 23/377 (6%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTL-CIV 68
+ E+WH CA V +P S V YFPQGH E + S H P L C++
Sbjct: 9 INSELWHACAGPLVSLPPAGSLVVYFPQGHSEQVAASMQKDVDAHVPSYPNLPSKLICLL 68
Query: 69 SAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
V L ADP TDEV+ ++ L P+T+ E+A R + F KTLT SD +
Sbjct: 69 HNVTLHADPETDEVYAQMTLQPVTSYGKEALQLSELALKQARPQTEFFCKTLTASDTSTH 128
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR A+ +FP LD + +Q L D+H V F H+ RG PKR++L + W+
Sbjct: 129 GGFSVPRRAAEKIFPPLDFSMQPPAQELQARDLHDNVWTFRHIYRGQPKRHLL-TTGWSL 187
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR-NTQFV-AAAAEQKKDELEKAVM-EALKLAE 245
FV K+L AGDSVIF++D ++ +GIRR N Q +++ D + ++ A A
Sbjct: 188 FVSGKRLFAGDSVIFVRDEKQQLLLGIRRANRQPTNISSSVLSSDSMHIGILAAAAHAAA 247
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTIS 302
N F I Y P+ +FV+ +++ +I R R+ +T++ Y GTI+
Sbjct: 248 NNSPFTIFYNPRASP-TEFVIPFAKYQKAVYGNQISLGMRFRMMFETEELGTRRYMGTIT 306
Query: 303 IVSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHK--PAPTPFPQTKKFR 354
+S ++ WR LQV WDE + RV+ W +E ++ P PF K+ R
Sbjct: 307 GISDLDPVRWKNSQWRNLQVGWDESAAGERRNRVSIWEIEPVAAPFFICPPPFFGAKRPR 366
Query: 355 TTQSSAQLSDKKETLLN 371
QL D+ + N
Sbjct: 367 ------QLDDESSEMEN 377
>UniRef100_Q84QI6 Auxin response factor-like protein [Mangifera indica]
Length = 840
Score = 174 bits (440), Expect = 6e-42
Identities = 119/377 (31%), Positives = 174/377 (45%), Gaps = 12/377 (3%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RVYYFPQGH+E S++ + R LC V V
Sbjct: 37 ELWHACAGPLVTVPRQGERVYYFPQGHIEQVEASTNQFADQQMPIYDLRSKILCRVINVQ 96
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A P TDEVF ++ L P N KE L R V SF KTLT SD + F
Sbjct: 97 LKAKPDTDEVFAQITLLPEPNQDENAVEKEPPPPLLPRFHVHSFCKTLTASDTSTHGGFS 156
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R A+ P+LD+ + +Q L D+HG +F H+ RG P+R++L S W+ FV
Sbjct: 157 VLRRHAEECLPELDMSQQPPTQDLVAKDLHGNEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 215
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ ++ VG+RR Q ++ + V+ A
Sbjct: 216 KRLVAGDAFIFLR-CEKELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAVSTGTM 274
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ M +M+ +++ Y GTI +
Sbjct: 275 FTVYYKPRISPAEFIVPFDQYMESVKSNYSIGMGFEMRFEGEEAPEQRYTGTIVGIEDAD 334
Query: 308 -----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPFPQTKKFRTTQSSAQL 362
+ WR L+V WDE P RV+PW +E A P P ++ R +
Sbjct: 335 PQRWPDSKWRCLKVRWDETSTVPRPERVSPWKIEPALAPLALNPLPLSRPKRPRSNMVPS 394
Query: 363 SDKKETLLNGDGFPVDI 379
S L F V++
Sbjct: 395 SPDSSVLTREGSFKVNV 411
>UniRef100_Q5Z826 Putative auxin response factor 7a [Oryza sativa]
Length = 991
Score = 174 bits (440), Expect = 6e-42
Identities = 123/377 (32%), Positives = 183/377 (47%), Gaps = 23/377 (6%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTL-CIV 68
+ E+WH CA V +P S V YFPQGH E + S H P L C++
Sbjct: 47 INSELWHACAGPLVSLPPAGSLVVYFPQGHSEQVAASMQKDVDAHVPSYPNLPSKLICLL 106
Query: 69 SAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
V L ADP TDEV+ ++ L P+T+ E+A R + F KTLT SD +
Sbjct: 107 HNVTLHADPETDEVYAQMTLQPVTSYGKEALQLSELALKQARPQTEFFCKTLTASDTSTH 166
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR A+ +FP LD + +Q L D+H V F H+ RG PKR++L + W+
Sbjct: 167 GGFSVPRRAAEKIFPPLDFSMQPPAQELQARDLHDNVWTFRHIYRGQPKRHLL-TTGWSL 225
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR-NTQFV-AAAAEQKKDELEKAVM-EALKLAE 245
FV K+L AGDSVIF++D ++ +GIRR N Q +++ D + ++ A A
Sbjct: 226 FVSGKRLFAGDSVIFVRDEKQQLLLGIRRANRQPTNISSSVLSSDSMHIGILAAAAHAAA 285
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTIS 302
N F I Y P+ +FV+ +++ +I R R+ +T++ Y GTI+
Sbjct: 286 NNSPFTIFYNPRASP-TEFVIPFAKYQKAVYGNQISLGMRFRMMFETEELGTRRYMGTIT 344
Query: 303 IVSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHK--PAPTPFPQTKKFR 354
+S ++ WR LQV WDE + RV+ W +E ++ P PF K+ R
Sbjct: 345 GISDLDPVRWKNSQWRNLQVGWDESAAGERRNRVSIWEIEPVAAPFFICPPPFFGAKRPR 404
Query: 355 TTQSSAQLSDKKETLLN 371
QL D+ + N
Sbjct: 405 ------QLDDESSEMEN 415
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.133 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 764,623,245
Number of Sequences: 2790947
Number of extensions: 31388948
Number of successful extensions: 83645
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 72
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 83242
Number of HSP's gapped (non-prelim): 158
length of query: 456
length of database: 848,049,833
effective HSP length: 131
effective length of query: 325
effective length of database: 482,435,776
effective search space: 156791627200
effective search space used: 156791627200
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 76 (33.9 bits)
Medicago: description of AC134322.24


