

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
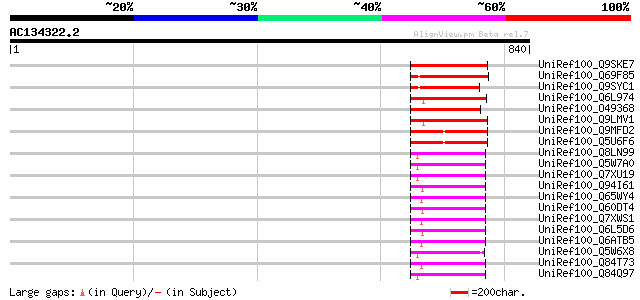
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SKE7 Putative retroelement pol polyprotein [Arabidop... 116 3e-24
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 114 1e-23
UniRef100_Q9SYC1 F11M15.5 protein [Arabidopsis thaliana] 107 1e-21
UniRef100_Q6L974 GAG-POL [Vitis vinifera] 100 2e-19
UniRef100_O49368 Hypothetical protein F10M6.140 [Arabidopsis tha... 99 5e-19
UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana] 99 5e-19
UniRef100_Q9MFD2 Orf764 protein [Beta vulgaris subsp. vulgaris] 97 3e-18
UniRef100_Q5U6F6 Orf764 protein [Beta vulgaris subsp. vulgaris] 97 3e-18
UniRef100_Q8LN99 Putative gag-pol [Oryza sativa] 96 6e-18
UniRef100_Q5W7A0 Putative polyprotein [Oryza sativa] 96 6e-18
UniRef100_Q7XU19 OSJNBa0091D06.19 protein [Oryza sativa] 96 6e-18
UniRef100_Q94I61 Putative retroelement [Oryza sativa] 96 6e-18
UniRef100_Q65WY4 Putative polyprotein [Oryza sativa] 95 1e-17
UniRef100_Q60DT4 Putative polyprotein [Oryza sativa] 94 1e-17
UniRef100_Q7XWS1 OSJNBa0091C12.11 protein [Oryza sativa] 94 1e-17
UniRef100_Q6L5D6 Putative polyprotein [Oryza sativa] 94 1e-17
UniRef100_Q6ATB5 Putative polyprotein [Oryza sativa] 94 2e-17
UniRef100_Q5W6X8 Putative polyprotein [Oryza sativa] 94 2e-17
UniRef100_Q84T73 Putative polyprotein [Oryza sativa] 94 2e-17
UniRef100_Q84Q97 Putative polyprotein [Oryza sativa] 94 2e-17
>UniRef100_Q9SKE7 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 889
Score = 116 bits (290), Expect = 3e-24
Identities = 61/126 (48%), Positives = 85/126 (67%), Gaps = 2/126 (1%)
Query: 650 LADFLVEMTAEK--GSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSS 707
LADF++++ G+ KW+L VDGSSN +GSG G+ L P G +IEQSL+ F +S
Sbjct: 622 LADFVIKLPLADLDGTNSNKKWLLHVDGSSNRQGSGVGIQLTSPTGEVIEQSLQLGFNAS 681
Query: 708 NNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQ 767
NN++EYEAL AG++LA + ++E+ A SDSQLV Q G ++ KD ++ YLE V L Q
Sbjct: 682 NNESEYEALIAGIKLAQEKGIREIHAYSDSQLVTSQFHGEYEAKDERMEAYLELVKTLAQ 741
Query: 768 KFVSFK 773
+F SFK
Sbjct: 742 QFESFK 747
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 114 bits (285), Expect = 1e-23
Identities = 58/128 (45%), Positives = 87/128 (67%), Gaps = 3/128 (2%)
Query: 649 SLADFLVEMTAEKGSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSN 708
+LADF E+T + +W L VDGSSN + SGAGV LEGP +++EQ+++F+FK+SN
Sbjct: 1309 ALADFTAELTPYL-TERTPRWTLYVDGSSNSRSSGAGVVLEGPGEIVVEQAMKFEFKTSN 1367
Query: 709 NQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQK 768
NQAEYEA+ AG+ LA+++ V + KSDS+LV Q+TG ++ ++ L Y V L +
Sbjct: 1368 NQAEYEAIIAGLHLAIELEVTNITCKSDSRLVVGQLTGEYEVRETLLQQYFHFVKNLLNR 1427
Query: 769 F--VSFKY 774
F +SF++
Sbjct: 1428 FKEISFQH 1435
>UniRef100_Q9SYC1 F11M15.5 protein [Arabidopsis thaliana]
Length = 1295
Score = 107 bits (267), Expect = 1e-21
Identities = 57/114 (50%), Positives = 82/114 (71%), Gaps = 4/114 (3%)
Query: 650 LADFLVEM---TAEKGSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKS 706
LADFL+E+ TAE + + W+L VDG+S+ +GSGAGV L+ P G ++EQS R F +
Sbjct: 902 LADFLIELPLATAES-DVPSQPWILHVDGASSKQGSGAGVRLKLPTGEVLEQSFRLAFNA 960
Query: 707 SNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLE 760
SNN+AEYE+L AG+RLA+ V V+ L+A DSQLV Q +G+++ KD ++ YL+
Sbjct: 961 SNNEAEYESLLAGLRLAIGVGVRALQAHCDSQLVASQYSGDYEAKDSRMEAYLD 1014
>UniRef100_Q6L974 GAG-POL [Vitis vinifera]
Length = 1027
Score = 100 bits (248), Expect = 2e-19
Identities = 51/127 (40%), Positives = 78/127 (61%), Gaps = 4/127 (3%)
Query: 650 LADFLVEMTAEKGSMEAAK----WVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFK 705
+ADF++E + + G E ++ W L VDG+S GSG G+ L+ P G +EQ++R F
Sbjct: 431 MADFVLEYSRKPGQHEGSRKKEWWTLRVDGASRSSGSGVGLLLQSPTGEHLEQAIRLGFS 490
Query: 706 SSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYL 765
+SNN+AEYEA+ +G+ LAL + V +L SDSQLV + V ++ KD ++ YL V
Sbjct: 491 ASNNEAEYEAILSGLDLALALSVSKLRIFSDSQLVVKHVQEEYEAKDARMARYLAKVRNT 550
Query: 766 TQKFVSF 772
Q+F +
Sbjct: 551 LQQFTEW 557
>UniRef100_O49368 Hypothetical protein F10M6.140 [Arabidopsis thaliana]
Length = 452
Score = 99.0 bits (245), Expect = 5e-19
Identities = 51/114 (44%), Positives = 77/114 (66%), Gaps = 1/114 (0%)
Query: 650 LADFLVEMTAEK-GSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSN 708
LADF+ E++ + + W+L DGSS+L+GS G+ L+ P ++EQSLR FK+SN
Sbjct: 189 LADFMTELSPDVIDVIPDENWILYSDGSSSLQGSSLGILLQSPTREILEQSLRLQFKASN 248
Query: 709 NQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESV 762
N+AEYEAL AG+RLA + K+++A SDSQLV + +G F+ K+ ++ +L V
Sbjct: 249 NEAEYEALLAGLRLAKGLGAKQIKAFSDSQLVVSRFSGEFEAKNERMRAHLSMV 302
>UniRef100_Q9LMV1 F5M15.26 [Arabidopsis thaliana]
Length = 1838
Score = 99.0 bits (245), Expect = 5e-19
Identities = 51/129 (39%), Positives = 80/129 (61%), Gaps = 5/129 (3%)
Query: 650 LADFLVEMTAEKGSMEAA-----KWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDF 704
LADFL+E+ + + +W L VDGSS+ +GSG G+ L P ++EQS R F
Sbjct: 1233 LADFLIELPLQSAERAVSGNRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRF 1292
Query: 705 KSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMY 764
++NN AEYE L AG+RLA + + + A +DSQL+ Q++G ++ K+ ++ YL+ V
Sbjct: 1293 VATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQL 1352
Query: 765 LTQKFVSFK 773
+T+ F +FK
Sbjct: 1353 MTKDFENFK 1361
>UniRef100_Q9MFD2 Orf764 protein [Beta vulgaris subsp. vulgaris]
Length = 764
Score = 96.7 bits (239), Expect = 3e-18
Identities = 46/125 (36%), Positives = 79/125 (62%), Gaps = 2/125 (1%)
Query: 649 SLADFLVEMTAEKGSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSN 708
+LADF+VE + E+ W L+VDGS+ GSGAG+ + P G + E +++F K+SN
Sbjct: 177 ALADFIVEASYEEEEEPVGVWKLAVDGSAAQTGSGAGIIMTSPEGNVFEYAIKF--KASN 234
Query: 709 NQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQK 768
N+AEYEA AG+++ + K++ ++DSQLV Q+ G ++ ++P + YL+ + L +
Sbjct: 235 NEAEYEAAIAGIKMCMAADAKKIRLQTDSQLVASQIRGEYEAREPAMQKYLQMIKDLAAQ 294
Query: 769 FVSFK 773
+S +
Sbjct: 295 LISLE 299
>UniRef100_Q5U6F6 Orf764 protein [Beta vulgaris subsp. vulgaris]
Length = 764
Score = 96.7 bits (239), Expect = 3e-18
Identities = 46/125 (36%), Positives = 79/125 (62%), Gaps = 2/125 (1%)
Query: 649 SLADFLVEMTAEKGSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSN 708
+LADF+VE + E+ W L+VDGS+ GSGAG+ + P G + E +++F K+SN
Sbjct: 177 ALADFIVEASYEEEEEPVGVWKLAVDGSAAQTGSGAGIIMTSPEGNVFEYAIKF--KASN 234
Query: 709 NQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQK 768
N+AEYEA AG+++ + K++ ++DSQLV Q+ G ++ ++P + YL+ + L +
Sbjct: 235 NEAEYEAAIAGIKMCMAADAKKIRLQTDSQLVASQIRGEYEAREPAMQKYLQMIKDLAAQ 294
Query: 769 FVSFK 773
+S +
Sbjct: 295 LISLE 299
>UniRef100_Q8LN99 Putative gag-pol [Oryza sativa]
Length = 1740
Score = 95.5 bits (236), Expect = 6e-18
Identities = 53/134 (39%), Positives = 78/134 (57%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMT----------AEKGSME---AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T A G + A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1135 ALADFVAEWTPAPEPVSIPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 1194
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
++ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV+ +Q DPQ+
Sbjct: 1195 LKYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVSKEYQCSDPQM 1254
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1255 DAYVRQVRRMERHF 1268
>UniRef100_Q5W7A0 Putative polyprotein [Oryza sativa]
Length = 1207
Score = 95.5 bits (236), Expect = 6e-18
Identities = 53/134 (39%), Positives = 78/134 (57%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMT----------AEKGSME---AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T A G + A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 592 ALADFVAEWTPAPEPVSIPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 651
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
++ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV+ +Q DPQ+
Sbjct: 652 LKYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVSKEYQCSDPQM 711
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 712 DAYVRQVRRMERHF 725
>UniRef100_Q7XU19 OSJNBa0091D06.19 protein [Oryza sativa]
Length = 1910
Score = 95.5 bits (236), Expect = 6e-18
Identities = 53/134 (39%), Positives = 78/134 (57%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMT----------AEKGSME---AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T A G + A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1340 ALADFVAEWTPAPEPVSIPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 1399
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
++ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV+ +Q DPQ+
Sbjct: 1400 LKYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVSKEYQCSDPQM 1459
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1460 DAYVRQVRRMERHF 1473
>UniRef100_Q94I61 Putative retroelement [Oryza sativa]
Length = 1541
Score = 95.5 bits (236), Expect = 6e-18
Identities = 52/134 (38%), Positives = 75/134 (55%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSMEA-------------AKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T + A A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 917 ALADFVAEWTPAPEPVSAPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 976
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV +Q DPQ+
Sbjct: 977 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLRVLGDSQLVVNQVCKEYQCSDPQM 1036
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1037 DAYVRQVRRMERHF 1050
>UniRef100_Q65WY4 Putative polyprotein [Oryza sativa]
Length = 2027
Score = 94.7 bits (234), Expect = 1e-17
Identities = 51/134 (38%), Positives = 77/134 (57%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSME-------------AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T ++ AA WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1403 ALADFVAEWTPAPETVSIPEANTDPTQLPHAAHWVMQFDGSLSLQGAGAGVTLTSPNGDV 1462
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A+ + ++ L DSQLV QV ++ DPQ+
Sbjct: 1463 LRYLVRLDFRATNNMAEYEGLLAGLRVAVGLGIRRLLVLGDSQLVVNQVCKEYRCSDPQM 1522
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1523 DAYVRQVRRMERHF 1536
>UniRef100_Q60DT4 Putative polyprotein [Oryza sativa]
Length = 1796
Score = 94.4 bits (233), Expect = 1e-17
Identities = 52/134 (38%), Positives = 75/134 (55%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSMEA-------------AKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T + A A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1172 ALADFVAEWTPAPEPVSAPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 1231
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV +Q DPQ+
Sbjct: 1232 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVRKEYQCSDPQM 1291
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1292 DAYVRQVRRMERHF 1305
>UniRef100_Q7XWS1 OSJNBa0091C12.11 protein [Oryza sativa]
Length = 1992
Score = 94.4 bits (233), Expect = 1e-17
Identities = 54/134 (40%), Positives = 77/134 (57%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTA--EKGSM-----------EAAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T E SM AA WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1368 ALADFVAEWTPAPETVSMPEASTNPSQLPHAAHWVMQFDGSLSLQGAGAGVTLTSPNGDV 1427
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV ++ DPQ+
Sbjct: 1428 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYRCSDPQM 1487
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1488 DAYVRQVRRMERHF 1501
>UniRef100_Q6L5D6 Putative polyprotein [Oryza sativa]
Length = 1514
Score = 94.4 bits (233), Expect = 1e-17
Identities = 52/134 (38%), Positives = 75/134 (55%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSMEA-------------AKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T + A A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 929 ALADFVAEWTPAPEPVSAPEASSGPTQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 988
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV +Q DPQ+
Sbjct: 989 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYQCSDPQM 1048
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1049 DAYVRQVRRMERHF 1062
>UniRef100_Q6ATB5 Putative polyprotein [Oryza sativa]
Length = 1989
Score = 94.0 bits (232), Expect = 2e-17
Identities = 51/134 (38%), Positives = 76/134 (56%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSME-------------AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T ++ AA WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1365 ALADFVAEWTPAPETVSIPEASADPSQLPHAAHWVMQFDGSLSLQGAGAGVTLTSPNGDV 1424
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV ++ DPQ+
Sbjct: 1425 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYRCSDPQM 1484
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1485 DAYVRQVRRMERHF 1498
>UniRef100_Q5W6X8 Putative polyprotein [Oryza sativa]
Length = 1688
Score = 94.0 bits (232), Expect = 2e-17
Identities = 54/133 (40%), Positives = 77/133 (57%), Gaps = 14/133 (10%)
Query: 649 SLADFLVEMT----------AEKGSME---AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T A G + A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1158 ALADFVAEWTPAPEPVSIPEASSGPSQLPHTAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 1217
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
++ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV+ +Q DPQ+
Sbjct: 1218 LKYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVSKEYQCSDPQM 1277
Query: 756 LMYLESVMYLTQK 768
Y+ YL K
Sbjct: 1278 DAYVRQA-YLADK 1289
>UniRef100_Q84T73 Putative polyprotein [Oryza sativa]
Length = 2026
Score = 94.0 bits (232), Expect = 2e-17
Identities = 51/134 (38%), Positives = 76/134 (56%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMTAEKGSME-------------AAKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T ++ AA WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1402 ALADFVAEWTPAPETVSIPEASADPSQLPHAAHWVMQFDGSLSLQGAGAGVTLTSPNGDV 1461
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV ++ DPQ+
Sbjct: 1462 LRYLVRLDFRATNNMAEYEGLLAGLRVAAGLGIRRLLVLGDSQLVVNQVCKEYRCSDPQM 1521
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1522 DAYVRQVRRMERHF 1535
>UniRef100_Q84Q97 Putative polyprotein [Oryza sativa]
Length = 1918
Score = 94.0 bits (232), Expect = 2e-17
Identities = 53/134 (39%), Positives = 76/134 (56%), Gaps = 13/134 (9%)
Query: 649 SLADFLVEMT----------AEKGSMEA---AKWVLSVDGSSNLKGSGAGVTLEGPYGVL 695
+LADF+ E T A G + A WV+ DGS +L+G+GAGVTL P G +
Sbjct: 1416 ALADFVAEWTPAPEPVSIPEASSGPSQLPHIAYWVMQFDGSLSLQGAGAGVTLTSPSGDV 1475
Query: 696 IEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQL 755
+ +R DF+++NN AEYE L AG+R+A + ++ L DSQLV QV +Q DPQ+
Sbjct: 1476 LRYLVRLDFRATNNMAEYEGLLAGLRVATGLGIRRLLVLGDSQLVVNQVCKEYQCSDPQM 1535
Query: 756 LMYLESVMYLTQKF 769
Y+ V + + F
Sbjct: 1536 DAYVRQVRRMERHF 1549
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,125,177,027
Number of Sequences: 2790947
Number of extensions: 37961483
Number of successful extensions: 144455
Number of sequences better than 10.0: 517
Number of HSP's better than 10.0 without gapping: 499
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 143796
Number of HSP's gapped (non-prelim): 607
length of query: 840
length of database: 848,049,833
effective HSP length: 136
effective length of query: 704
effective length of database: 468,481,041
effective search space: 329810652864
effective search space used: 329810652864
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 79 (35.0 bits)
Medicago: description of AC134322.2


